Abstract
Hereditary spastic paraplegias (HSP) are neurodegenerative disorders characterized by lower limb spasticity and weakness that can be complicated by other neurological or non-neurological signs. Despite a high genetic heterogeneity (>60 causative genes), 40–70% of the families remain without a molecular diagnosis. Analysis of one of the pioneer cohorts of 193 HSP families generated in the early 1990s in Portugal highlighted that SPAST and SPG11 are the most frequent diagnoses. We have now explored 98 unsolved families from this series using custom next generation sequencing panels analyzing up to 70 candidate HSP genes. We identified the likely disease-causing variant in 20 of the 98 families with KIF5A being the most frequently mutated gene. We also found 52 variants of unknown significance (VUS) in 38% of the cases. These new diagnoses resulted in 42% of solved cases in the full Portuguese cohort (81/193). Segregation of the variants was not always compatible with the presumed inheritance, indicating that the analysis of all HSP genes regardless of the inheritance mode can help to explain some cases. Our results show that there is still a large set of unknown genes responsible for HSP and most likely novel mechanisms or inheritance modes leading to the disease to be uncovered, but this will require international collaborative efforts, particularly for the analysis of VUS.
Similar content being viewed by others
Introduction
Hereditary spastic paraplegias (HSP) are a clinically and genetically heterogeneous group of rare neurodegenerative disorders. HSP have a variable age at onset and are mainly characterized by progressive lower limb spasticity and weakness. These hallmarks are caused by degeneration of the motor axons that mostly affects the distal ends of the long central nervous system tracts.1 Their estimated prevalence ranges from 1–10/100 000 depending on the geographic localization2 and is 4.1/100 000 in Portugal.3
These disorders are historically divided on clinical grounds into pure and complex forms, according to the absence or presence of additional neurological and extraneurological features, such as neuropathy, cognitive impairment, cerebellar ataxia and epilepsy. Consequently, ancillary tests such as brain/spinal cord magnetic resonance imaging, electroneuromyography and others can show various abnormal patterns. Genetically, all classical modes of inheritance have been described, with 79 SPG-associated loci and more than 60 identified genes, a number that is constantly increasing.4, 5, 6, 7, 8, 9, 10 Moreover, at least 11 additional genes have also been associated with spastic paraplegia, some of them very recently.11, 12 Although a considerable number of genes have already been identified, most of the families worldwide have not been extensively screened so far and remain without a molecular diagnosis, as was the case of 68.9% families in Portugal, before this study. Indeed, among the HSP a considerable number of genes and variants are apparently orphan, affecting single families.
Identification of the responsible genes has a great impact on patients and their families as it reveals the cause of the disease, and may also be important to better understand how these variants arise (founder effects and susceptibility chromosomes, among others) and how they lead to specific phenotypes. This study aimed to provide a molecular diagnosis for the Portuguese families identified through a population-based survey3 using targeted next generation sequencing (NGS) to screen these patients for the presence of variants in 70 genes already associated with HSP or candidate genes, covering all classical modes of inheritance.
Materials and methods
Patients
In Portugal, 193 families with hereditary spastic paraplegia were identified in a national, population-based survey.3, 13 After exclusion of the most common genes in most of the families and isolated cases (ATL1, SPAST and REEP1 in families with dominant inheritance, and SPG11, ZFYVE26 and CYP7B1 in the recessive ones), there were still 68.9% (133/193) of the families without a molecular diagnosis, 62% (98/159) if we consider only the families with DNA available for testing. Taking that into consideration, 98 unsolved families (ie, with DNA available) were selected to be screened for variants in known genes using one patient per family.
Among these 98 Portuguese families, 38 showed a probable recessive inheritance, 44 a dominant inheritance and 16 were isolated cases. Pure and complex forms were present and age at onset in the whole cohort ranged from less than 1 year to 65 years. Written informed consent for genetic testing was obtained from all tested individuals or their respective legal guardian and the genetic analyses were approved by the ethical committees of ICBAS (Portugal) and of the Paris-Necker Hospital (France).
HSP panels
Two sequential custom NGS panels were developed at ICM, France, to search for variants in HSP-related genes (Supplementary Table S1). This strategy was favored to exome sequencing for ethical concerns regarding the risk of secondary or incidental findings in HSP diagnosis in agreement with our local ethics committees and for the cost effectiveness given the number of genes involved in HSP that could carry a mutation. The first panel covered 34 genes among which 31 were known to be responsible for dominant and recessive HSP forms and the other three (ALS2, SACS and SETX) were implicated in overlapping phenotypes.14, 15, 16 The 34 genes corresponded to 531 regions with a total length of 109 768 base pairs. In order to increase achieving a molecular diagnosis, a second panel was later designed that covered 70 genes. This added 30 newly identified genes5 to the previous panel as well as three genes (FBXO7, GJA1 and SAMHD1) causing overlapping phenotypes17, 18, 19 and three candidate genes that encode partners of known HSP proteins (AP5B1, AP5M1 and AP5S1). This panel covered a total length of 210 363 bases corresponding to 1001 regions. These two panels targeted exonic regions and at least 20 intronic bases at the exon-intron boundaries.
Genetic analysis was performed using a customized Roche/Nimblegen capture followed by NGS in the MiSeq apparatus (Illumina, San Diego, CA, USA). The procedure consisted of four major steps: (1) library preparation, where we used the Illumina TruSeq DNA LT Sample Preparation kit v2-Set A (Illumina) according to the manufacturer’s protocol for the first panel and the Kapa HTP Library Prep Kit Illumina (Roche, Basel, Switzerland) with its corresponding protocol for the second panel; (2) double capture using the Roche NimbleGen SeqCap EZ Reagent Kit and SeqCap EZ Library (Roche) using the SeqCap EZ Library SR User’s Guide protocol; (3) massive parallel sequencing using the Illumina MiSeq Benchtop Sequencer; and (4) data analysis using the CLC Bio-Genomics Workbench 6.5.1 software (https://www.qiagenbioinformatics.com/). After the alignment with the Homo sapiens (hg19) reference sequence, data were filtered taking into consideration that the variants had to: (i) be present in the target regions, (ii) cause a change at the protein level (nonsense, amino acid change or a splicing effect), (iii) be present in the local database with a frequency below 15% and, (iv) if known in the databases, its minor allele frequency had to be below 1% for autosomal recessive inheritance and 0.2% for autosomal dominant transmission. All filtered variants were further analyzed using Alamut v.2.9.0 software (Interactive Biosoftware, La Rochelle, France) for functional effect prediction with SpliceSiteFinder, MaxEntScan, NNSPLICE, GeneSplicer, Human Splicing finder, Polyphen-2, SIFT, MutationTaster, Align GVGD and UMD-Predictor.
Twelve patients per experiment were sequenced using the first panel, while 24 were analyzed simultaneously using the second panel. The first 25 Portuguese patients tested were sequenced using the first panel and the remaining 73 patients were screened with the second panel.
In order to detect rearrangements, we analyzed the coverage of all the regions with an in-house algorithm based on the analysis of the coverage. In the genes where a suggestive alteration in dosage was detected, multiplex ligation-dependent probe amplification or quantitative real-time PCR was applied to confirm the deletion/duplication.
All the variants that were not excluded after filtering and segregation analysis (whenever possible) were submitted to ClinVar (http://www.ncbi.nlm.nih.gov/clinvar/). ClinVar accession numbers are SCV000574442–SCV000574511.
Sanger sequencing
We confirmed the presence of the variants identified by NGS checking, by Sanger sequencing, the region where the variant was present, including additional family members when DNA was available. Primers were designed using Primer3Plus software (http://www.bioinformatics.nl/cgi-bin/primer3plus/primer3plus.cgi). The PCR amplification was done using the DreamTaq DNA Polymerase Kit (ThermoFisher Scientific, Waltham, MA, USA) or when necessary, AccuPrime GC-Rich DNA polymerase (Invitrogen, Carlsbad, CA, MA, USA), followed by sequencing by GATC BioTech using BigDye chemistry in an ABI3730 sequencer (Applied Biosystems, Foster City, CA, USA). Sequence analysis was performed using SeqScape 2.6 software (Applied Biosystems).
Multiplex ligation-dependent probe amplification
In order to confirm the presence of large gene rearrangements in the SPG11 gene we performed multiplex ligation-dependent probe amplification using the SALSA MLPA kit P306 (MRC Holland, Amsterdam, Netherlands) according to manufacturer’s instructions and analyzed the resulting fragments on an ABI 3130xl Genetic Analyzer using 500-LIZ (Applied Biosystems), as a size standard, and GeneMarker v1.90 (SoftGenetics, State College, PA, USA).
Results
Study design
A total of 98 index cases of Portuguese families without molecular diagnosis were screened for variants using either the first or the second custom sequencing panel. The mean coverage varied from 126 to 801 reads: 419 reads per base with 94% of the regions with a coverage ≥30 × with the first panel and 362 reads with 97% of the regions with at least 30 × of coverage with the second panel. All the variants considered relevant (rare, predicted deleterious, expected to have an impact given the suspected mode of inheritance) were confirmed by Sanger sequencing and segregation analysis was performed in all the available family members. The regions with less than 30-fold coverage were screened by Sanger sequencing in autosomal recessive cases in which only one heterozygous potentially disease-causing variant was identified. Relevant insufficiently covered exons in patients with a compatible phenotype were also sequenced by Sanger (ie, exon 1 of SPG7 in patients with cerebellar atrophy and/or optic atrophy). An in-house index of coverage was calculated for each exon and for each patient to search for genomic rearrangements.
Genetic characterization
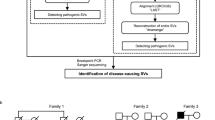
A molecular diagnosis of HSP was confirmed in 20.4% (20/98) of the cases. The coverage analysis was responsible for the identification of one large homozygous deletion encompassing exons 12 to 14 in SPG11 that was confirmed by multiplex ligation-dependent probe amplification (Table 1, Supplementary Figure S1) and was considered to affect spatacsin function. Additionally, after filtering of the NGS data, we found 139 nucleotide variants present in 74 families with minor allele frequency below 1.5% that potentially affected protein function according to at least one prediction software. These variants were divided in: (i) Disease-causing variants (17 variants in 14 families), (ii) Likely disease-causing variants (6 variants in 5 families), (iii) variants of unknown significance – VUS (52 variants in 37 families) and (iv) Excluded variants (64 variants in 45 families) by several criteria detailed in Figure 1. In the remaining 24 families, no disease-causing variants were found after filtering.
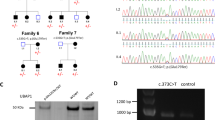
We considered 24 (of the 140) variants to be affecting protein function in 20 families (Table 1, Figure 2 and Supplementary Figure S1). Most (n=18) were classified as disease-causing variants because they were already reported as disease-causing variants, caused an early stop codon (nonsense and frameshift) or were large gene rearrangements. The other ones (n=6) were considered as likely disease-causing because we did not perform functional studies to prove it. All of them had low frequency in databases and a predicted deleterious effect (Table 1) and additionally, in 16 of these families, we could demonstrate their segregation with the disease (Figure 2 and Supplementary Figure S1), which included all the families with likely disease-causing variants.
Pedigree with segregation analysis, electropherogram and conservation of the seven novel missense variants. A black circles and squares indicate affected individuals with HSP; an asterisk in a symbol indicates family members with available DNA. Electropherogram with the position of the missense variant boxed. Conservation of the region of the altered amino acid (boxed).
Among the 52 VUS (Table 2), three are illustrative of the difficulties we faced in the interpretation of their biological relevance (Supplementary Figure S2): a homozygous missense variant in a gene causing a recessive form where family members were not available for segregation analysis (family SR26); a variant in a gene causing a dominant form, of interest only if we consider incomplete penetrance (family SR49) and a variant in the homozygous state in a gene causing a dominant form that segregated with the disease (family S88). In addition, in families SR65 and S86 we found compound heterozygous variants in two different genes known to interact at the protein level (Supplementary Figure S2). In both cases, the variants were found in two different members or interactors of the adaptor protein complex 5, but neither their cosegregation nor their effect at the cellular level could be analyzed, which would represent a prerequisite to prove these potential cases of digenism.
In this study, we report 10 new disease-causing variants among the 24 variants affecting protein function, of which seven are new missense variants (Figure 2), one is a nonsense variant, one a frameshift variant and the last is the deletion of three consecutive exons in SPG11 (Table 1).
The yield of mutational results varied according to inheritance mode (Figure 3). We found the disease-causing variants in nine of the 38 recessive families (23.7%) and in six of the 44 families with dominant transmission (13.6%). Of note, one family with a homozygous SPG7 variant was initially considered as dominant (Family S26, Supplementary Figure S1) because of a suspected family history of the disease in the father, who however was not examined and not tested. Five of the 16 isolated cases were also explained at the molecular level (31.3%), including one confirmed de novo variant in KIF1A (Family SR98, Figure 2). Segregation analysis could not be performed in three families, but all of them had variants already described as disease-causing in the literature.
Clinical characterization
The phenotype of the patients was mostly similar to that of previously reported series. A detailed phenotypic description of the families is presented in Table 3. For example, in the two families carrying SPG11 variants, the clinical picture included early age at onset and a complex phenotype with cognitive impairment in both families and neuropathy in one of them. Magnetic resonance imaging was not available to check for the presence of a thin corpus callosum. In another family with a homozygous ZFYVE26 variant and an SPG11-like phenotype, thin corpus callosum was overt. Similarly to published cases, a pure form of the disease was observed in (i) the case with a de novo SPAST variant (SR4, Supplementary Figure S1), (ii) the patient presenting a variant in REEP1 (SR99), (iii) the two families with CYP7B1 variants (families S93 and SR97, Supplementary Figure S1) and the patient with an NIPA1 variant (S61, Supplementary Figure S1). On the contrary, in our KIF5A families (S41, S31, S27, S58, Figure 2 and Supplementary Figure S1), pure and complex forms were found, in agreement with Goizet and colleagues.28 We also identified three families with SPG7 variants (Supplementary Figure S1) with late-onset HSP and a complex phenotype: families SR75 and S26 also presenting ataxia and the sporadic case CI2 that has minor cerebellar atrophy at brain magnetic resonance imaging which confirms that the involvement of the cerebellum is part of the SPG7 core phenotype. In family SR84 (Supplementary Figure S1), presenting two known variants in the GBA2 gene, one affecting the catalytic site and one destabilizing the protein by disturbance of the charge balance,33 patients had an onset in the expected range (infancy or childhood) and a complex phenotype with cerebellar signs and ataxic gait (Table 3).
Interestingly, we confirmed the higher occurrence of cases with heterozygous vs homozygous/compound heterozygous (SPG30) variants in KIF1A. One patient in the SR98 family (Figure 2) carried a novel heterozygous de novo KIF1A variant located in the motor domain and presented a complex phenotype with mental retardation and neuropathy and an early age at onset, in the first year of life.
On the other hand, the clinical features of some of our patients extended the phenotypic spectrum of some clinico-genetic entities. This was the case of the family with the SACS variants (SR6, Figure 2 and Table 3), in which patients presented with spasticity and peripheral neuropathy but without ataxia, as in another single case recently reported in an Italian family.15 In the family with a C19orf12 variant (family SR88, Supplementary Figure S1), the phenotype was similar to SPG43 but also presenting cerebellar ataxia, mental retardation and dementia. Interestingly, the same variant was described in patients with neurodegeneration with brain iron accumulation type 4 (NBIA4),20 extending the spectrum of phenotypes associated with this variant. Brain imaging data were not available for this patient, so we could not check for the presence of brain iron accumulation. Lastly, patients of the S75 family (Figure 2) showing ERLIN2 variants had an early onset, at 8 and 19 years old, however with a pure phenotype, contrary to the complex phenotype previously described.34
Discussion
Genetic diagnostic yield
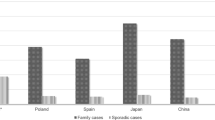
The identification of the genetic cause for HSP in only 20.4% of this cohort shows that there is still a large set of genes responsible for spastic paraplegia to be uncovered or that novel inheritance modes, as in the case of KIF1A,35 should be taken into account. Although most of these families had been previously screened for SPG3, SPG4 and SPG31 in the families with dominant forms36 and for SPG11, SPG15 and SPG5 in the families with recessive transmission (unpublished data), we expected a higher frequency of diagnosed families since we were testing almost all the known genetic causes of HSP, including all the genes identified by Novarino and colleagues.37 This frequency is very similar to that found in a previous study covering far fewer genes in patients where only SPAST (SPG4) variants were excluded.38 On the other hand, our results are also very similar and comparable to a study where exome sequencing identified disease-causing variants in eight of 48 HSP families (16.6%) and potentially disease-causing variants in another eight families where previous screening for the most likely genes had been performed.39 Taking into account the full cohort, we have found the responsible gene in 42% of the Portuguese families (81/193), or in 51% if we only take into account the families with available DNA (81/159), a frequency similar to a Greek study where only 16 HSP genes were screened.40 The most frequently mutated genes are (Figure 4): SPAST (SPG4) with a frequency of 18% (28/159) and SPG11 with a frequency of 13% (20/159). Interestingly, we found a high frequency of KIF5A variants, present in four out of 44 families with dominant inheritance (9.1%) and in 4 of the 159 families (2.5%) becoming the third most frequently mutated gene in dominant forms. This high occurrence was also found in the Greek population where it was the second cause of disease in families with dominant transmission40 and in the study by Warrenburg et al where it was found as the most frequently mutated gene (after exclusion of the most likely genes).39 Most of the other genes tested in our cohort have accounted for single families until now, as is the case of REEP2 (SPG72)41 and in the case of a family that was negative in this screening but after exome analysis proved to segregate ALDH18A1 (SPG9) variants.7 In line with our results showing that most of the new HSP genes are rarely involved, the yield of positive results obtained with the two panels, the second containing 30 additional HSP genes, was very similar. We cannot exclude that a variant in the 30 additional genes added to the second panel may explain some of the 17 cases without diagnosis (of the 25) tested with the first panel. This is however very unlikely given (1) the low relative frequencies of these genes,5, 37 (2) their involvement in recessive forms while 10 of 17 of our patients were associated with dominant inheritance of the disease, (3) and because seven of these cases processed under exome sequencing more recently do not carry mutations in the genes present in the second panel (unpublished data).
Distribution of the disease-causing variants found in 81 families from the full cohort of Portuguese cases.# (a) Recessive families, (b) Dominant families, (c) Isolated cases. *Case described in this paper of a family classified with dominant inheritance but with a homozygous SPG7 variant. # Include mutations found in previously screened genes.7, 36, 41
Our relatively high percentage of diagnosis among the isolated cases (5 out of 16) is also an interesting finding that shows how important it is to test these patients that frequently are not considered in genetic analyses due to the absence of additional family members to confirm diagnosis.
This study also demonstrates that a panel strategy is also capable of detecting large rearrangements, as proved with the identification of a large deletion, although at the homozygous state, encompassing exons 12–14 in SPG11 in family SR17. This was possible by the analysis of the coverage, where we compared the mean coverage of all patients tested with each other taking into account the average coverage of each one.
Towards an unbiased approach
Our results also highlight the importance of testing all families for the same set of genes, regardless the presumed inheritance mode. This is important as sometimes the transmission mode is unclear in the pedigree because of its size or censured consanguinity, and because there is an increasing number of HSP genes associated with different modes of transmission (eg, BICD2, KIF1C and KIF1A), sometimes associated with a different disease presentation (eg, SPG7). We have some examples in our cohort of families with an apparent recessive mode of transmission (including sporadic cases) in whom we found a variant in a gene associated with a dominant form. This can happen when the disease is caused by a ‘de novo’ variant, as in families SR4 (Supplementary Figure S1) and SR98 (Figure 2), and also in late-onset families in which the disease could have been missed in the older generations. The reverse is also true since we report one case with a homozygous variant in SPG7, usually associated with a recessive form, found in a family classified as dominant (S26, Supplementary Figure S1). This could likely be explained by censured consanguinity or by the presence of older individuals reported as affected but with a different neurological condition. Family SR98, with a variant in the motor domain of KIF1A, is also an example of a gene that can have recessive or dominant inheritance depending on the localization of the variant. It was recently shown by Lee and colleagues that de novo heterozygous variants in the motor domain of KIF1A can be disease-causing35 and that this type of variants are more common than the homozygous ones, which has been later confirmed in several studies39, 42, 43 including our own. In family SR99 (Figure 2), we found a new variant (p.Ala27Pro) in REEP1 that, despite being present in the unaffected father, is likely the disease-causing variant because incomplete penetrance was already described for variants in this gene and because the same amino acid was affected in another HSP case.44
Need for international collaborative efforts
Our results also highlight the difficulty of reaching a conclusion as to the causative nature of the variants in 37% of the cases. Large screening studies, like ours, result in a large quantity of data that has to be interpreted, a task that is not always easy even with all the available software. This is especially difficult in HSP since the disease can have different inheritance modes. An example of the difficulty in interpretation without functional data is illustrated by family SR6 (Figure 2), where we found two new variants, within six bases of distance, that segregate together, and we were not able to demonstrate which one was deleterious, although the predictions on their effect on protein function is stronger in one of them.
It is highly likely that in our 37 families with VUS, some of these VUS could be variants with new inheritance forms for already known genes. This could be the case in family SR88, where we found a KIF5A variant present at the homozygous state in patients in two generations of a family with no reported consanguinity (Supplementary Figure S2). This might represent the first autosomal recessive case due to KIF5A variants, but this cannot be ascertained in the absence of functional evidence or additional cases segregating in autosomal recessive pedigrees. We also found some variants in candidate genes selected by function (interactors) for which we were not able to conclude as to their causative effect due to the insufficient number of family members to validate their co-transmission (families SR65 and S86, Supplementary Figure S2), and the absence of cell lines to demonstrate the disruption of the AP5 complex by biochemical experiments.45 We observed a variant in a gene associated with dominant transmission, BSCL2, that is in a conserved region, is predicted to affect function by five software and is in the same domain as the two published variants (with reports of incomplete penetrance) but, since it is present in the non-affected mother and there are no other known carriers, we were not able to conclude as to the causative effect of this variant (case SR49, Supplementary Figure S2). Absence of family members to confirm segregation of a homozygous variant in the WDR48 was also an issue in family SR26 (Supplementary Figure S2). This family presents a complex phenotype with motor neuropathy that is in accordance with the reported clinical features,37 although with a later age at onset (20 and 22 years old vs 1-year-old), and only functional studies will allow to prove its causative role. Therefore, the availability of relatives to test segregation still remains crucial to conclude on a causative variant, and diagnosis in HSP should not rely only on the clinical/genealogical presentation because the clinical phenotype and full mutational spectrum associated with each gene are continuously expanding.5
In conclusion, this comprehensive study of a large cohort of 98 families of homogeneous origin (only Portuguese families) allowed us to conclude on the frequency of all the less common HSP genes. Also, it highlights the fact that although a high number of genes have been already identified in this condition, a large percentage of cases remain without molecular diagnosis, some of them due to the absence of extensive screening. Gene panel strategy is a cost-effective way to screen HSP families and probably a small panel like ours with 34 genes could give a diagnosis to almost half of the families if used as a first approach. The use of an extensive panel of genes like our second panel, including the more recent and less frequently mutated genes such as VCP39, 46 would only slightly increase the number of diagnosis. This suggests that performing exome sequencing in the families without diagnosis in the most frequent genes (34) is probably a good strategy in HSP.
References
Fink JK : Hereditary spastic paraplegia: clinico-pathologic features and emerging molecular mechanisms. Acta Neuropathol 2013; 126: 307–328.
Ruano L, Melo C, Silva MC, Coutinho P : The global epidemiology of hereditary ataxia and spastic paraplegia: a systematic review of prevalence studies. Neuroepidemiology 2014; 42: 174–183.
Coutinho P, Ruano L, Loureiro JL et al: Hereditary ataxia and spastic paraplegia in Portugal: a population-based prevalence study. JAMA Neurol 2013; 70: 746–755.
Lo Giudice T, Lombardi F, Santorelli FM, Kawarai T, Orlacchio A : Hereditary spastic paraplegia: clinical-genetic characteristics and evolving molecular mechanisms. Exp Neurol 2014; 261: 518–539.
Tesson C, Koht J, Stevanin G : Delving into the complexity of hereditary spastic paraplegias: how unexpected phenotypes and inheritance modes are revolutionizing their nosology. Hum Genet 2015; 134: 511–538.
Rinaldi C, Schmidt T, Situ AJ et al: Mutation in CPT1C Associated With Pure Autosomal Dominant Spastic Paraplegia. JAMA Neurol 2015; 72: 561–570.
Coutelier M, Goizet C, Durr A et al: Alteration of ornithine metabolism leads to dominant and recessive hereditary spastic paraplegia. Brain 2015; 138: 2191–2205.
Gan-Or Z, Bouslam N, Birouk N et al: Mutations in CAPN1 cause autosomal-recessive hereditary spastic paraplegia. Am J Hum Genet 2016; 98: 1038–1046.
Yang Y, Liu W, Fang Z et al: A newly identified missense mutation in FARS2 causes autosomal-recessive spastic paraplegia. Hum Mutat 2016; 37: 165–169.
Estrada-Cuzcano A, Martin S, Chamova T et al: Loss-of-function mutations in the ATP13A2/PARK9 gene cause complicated hereditary spastic paraplegia (SPG78). Brain 2017; 140: 287–305.
Josifova DJ, Monroe GR, Tessadori F et al: Heterozygous KIDINS220/ARMS nonsense variants cause spastic paraplegia, intellectual disability, nystagmus, and obesity. Hum Mol Genet 2016; 25: 2158–2167.
Ozes B, Karagoz N, Schule R et al: Pla2g6 mutations associated with a continuous clinical spectrum from neuroaxonal dystrophy to hereditary spastic paraplegia. Clin Genet 2017; e-pub ahead of print 13 March 2017; doi:10.1111/cge.13008.
Silva MC, Coutinho P, Pinheiro CD, Neves JM, Serrano P : Hereditary ataxias and spastic paraplegias: methodological aspects of a prevalence study in Portugal. J Clin Epidemiol 1997; 50: 1377–1384.
Wakil SM, Ramzan K, Abuthuraya R et al: Infantile-onset ascending hereditary spastic paraplegia with bulbar involvement due to the novel ALS2 mutation c.2761C>T. Gene 2014; 536: 217–220.
Gregianin E, Vazza G, Scaramel E et al: A novel SACS mutation results in non-ataxic spastic paraplegia and peripheral neuropathy. Eur J Neurol 2013; 20: 1486–1491.
Strong MJ, Gordon PH : Primary lateral sclerosis, hereditary spastic paraplegia and amyotrophic lateral sclerosis: discrete entities or spectrum? Amyotroph Lateral Scler Other Motor Neuron Disord 2005; 6: 8–16.
Nisipeanu P, Kuritzky A, Korczyn AD : Familial levodopa-responsive parkinsonian-pyramidal syndrome. Mov Disord 1994; 9: 673–675.
Furuta N, Ikeda M, Hirayanagi K, Fujita Y, Amanuma M, Okamoto K : A novel GJA1 mutation in oculodentodigital dysplasia with progressive spastic paraplegia and sensory deficits. Intern Med 2012; 51: 93–98.
Dale RC, Gornall H, Singh-Grewal D, Alcausin M, Rice GI, Crow YJ : Familial Aicardi-Goutieres syndrome due to SAMHD1 mutations is associated with chronic arthropathy and contractures. Am J Med Genet A 2010; 152A: 938–942.
Hartig MB, Iuso A, Haack T et al: Absence of an orphan mitochondrial protein, c19orf12, causes a distinct clinical subtype of neurodegeneration with brain iron accumulation. Am J Hum Genet 2011; 89: 543–550.
Goizet C, Boukhris A, Durr A et al: CYP7B1 mutations in pure and complex forms of hereditary spastic paraplegia type 5. Brain 2009; 132: 1589–1600.
Hammer MB, Eleuch-Fayache G, Schottlaender LV et al: Mutations in GBA2 cause autosomal-recessive cerebellar ataxia with spasticity. Am J Hum Genet 2013; 92: 245–251.
Martin E, Schule R, Smets K et al: Loss of function of glucocerebrosidase GBA2 is responsible for motor neuron defects in hereditary spastic paraplegia. Am J Hum Genet 2013; 92: 238–244.
McDermott CJ, Dayaratne RK, Tomkins J et al: Paraplegin gene analysis in hereditary spastic paraparesis (HSP) pedigrees in northeast England. Neurology 2001; 56: 467–471.
Stevanin G, Santorelli FM, Azzedine H et al: Mutations in SPG11, encoding spatacsin, are a major cause of spastic paraplegia with thin corpus callosum. Nat Genet 2007; 39: 366–372.
Goizet C, Boukhris A, Maltete D et al: SPG15 is the second most common cause of hereditary spastic paraplegia with thin corpus callosum. Neurology 2009; 73: 1111–1119.
Tessa A, Silvestri G : de Leva MF et al: A novel KIF5A/SPG10 mutation in spastic paraplegia associated with axonal neuropathy. J Neurol 2008; 255: 1090–1092.
Goizet C, Boukhris A, Mundwiller E et al: Complicated forms of autosomal dominant hereditary spastic paraplegia are frequent in SPG10. Hum Mutat 2009; 30: E376–E385.
Chen S, Song C, Guo H et al: Distinct novel mutations affecting the same base in the NIPA1 gene cause autosomal dominant hereditary spastic paraplegia in two Chinese families. Hum Mutat 2005; 25: 135–141.
Tang B, Zhao G, Xia K et al: Three novel mutations of the spastin gene in Chinese patients with hereditary spastic paraplegia. Arch Neurol 2004; 61: 49–55.
Brugman F, Scheffer H, Wokke JH et al: Paraplegin mutations in sporadic adult-onset upper motor neuron syndromes. Neurology 2008; 71: 1500–1505.
Sanchez-Ferrero E, Coto E, Beetz C et al: SPG7 mutational screening in spastic paraplegia patients supports a dominant effect for some mutations and a pathogenic role for p.A510V. Clin Genet 2013; 83: 257–262.
Charoenwattanasatien R, Pengthaisong S, Breen I et al: Bacterial beta-glucosidase reveals the structural and functional basis of genetic defects in human glucocerebrosidase 2 (GBA2). ACS Chem Biol 2016; 11: 1891–1900.
Alazami AM, Adly N, Al Dhalaan H, Alkuraya FS : A nullimorphic ERLIN2 mutation defines a complicated hereditary spastic paraplegia locus (SPG18). Neurogenetics 2011; 12: 333–336.
Lee JR, Srour M, Kim D et al: De novo mutations in the motor domain of KIF1A cause cognitive impairment, spastic paraparesis, axonal neuropathy, and cerebellar atrophy. Hum Mutat 2015; 36: 69–78.
Loureiro JL, Brandao E, Ruano L et al: Autosomal dominant spastic paraplegias: a review of 89 families resulting from a portuguese survey. JAMA Neurol 2013; 70: 481–487.
Novarino G, Fenstermaker AG, Zaki MS et al: Exome sequencing links corticospinal motor neuron disease to common neurodegenerative disorders. Science 2014; 343: 506–511.
Kumar KR, Blair NF, Vandebona H et al: Targeted next generation sequencing in SPAST-negative hereditary spastic paraplegia. J Neurol 2013; 260: 2516–2522.
van de Warrenburg BP, Schouten MI, de Bot ST et al: Clinical exome sequencing for cerebellar ataxia and spastic paraplegia uncovers novel gene-disease associations and unanticipated rare disorders. Eur J Hum Genet 2016; 24: 1460–1466.
Lynch DS, Koutsis G, Tucci A et al: Hereditary spastic paraplegia in Greece: characterisation of a previously unexplored population using next-generation sequencing. Eur J Hum Genet 2016; 24: 857–863.
Esteves T, Durr A, Mundwiller E et al: Loss of association of REEP2 with membranes leads to hereditary spastic paraplegia. Am J Hum Genet 2014; 94: 268–277.
Ylikallio E, Kim D, Isohanni P et al: Dominant transmission of de novo KIF1A motor domain variant underlying pure spastic paraplegia. Eur J Hum Genet 2015; 23: 1427–1430.
Citterio A, Arnoldi A, Panzeri E et al: Variants in KIF1A gene in dominant and sporadic forms of hereditary spastic paraparesis. J Neurol 2015; 262: 2684–2690.
Beetz C, Schule R, Deconinck T et al: REEP1 mutation spectrum and genotype/phenotype correlation in hereditary spastic paraplegia type 31. Brain 2008; 131: 1078–1086.
Hirst J, Edgar JR, Esteves T et al: Loss of AP-5 results in accumulation of aberrant endolysosomes: defining a new type of lysosomal storage disease. Hum Mol Genet 2015; 24: 4984–4996.
de Bot ST, Schelhaas HJ, Kamsteeg EJ, van de Warrenburg BP : Hereditary spastic paraplegia caused by a mutation in the VCP gene. Brain 2012; 135: e223, author reply e224.
Acknowledgements
We would like to thank the participating patients and families and also the clinicians as well as Federica Barrecca, Emeline Mundwiller and Delphine Bouteiller for their help in the screening. This study was funded in part by the FCT (Fundação para a Ciência e Tecnologia, grant FCT-ANR/BEX-GMG/0008/2013, to IA) and by the French National Agency for Research (SPATAX-QUEST, to GS), the European Union (Omics call of the 7th framework program, NEUROMICS, to AB; the E-rare program, NEUROLIPID and PREPARE, to GS), the VERUM Foundation (to AB and GS) and benefited from the program ‘Investissements d’Avenir’ ANR-10-IAIHU-06 (to the ICM). SM is the recipient of a fellowship (SFRH/BD/87189/2012) from the FCT supported by POPH/FSE funding.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Supplementary Information accompanies this paper on European Journal of Human Genetics website
Rights and permissions
This work is licensed under a Creative Commons Attribution-NonCommercial-NoDerivs 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-nd/4.0/
About this article
Cite this article
Morais, S., Raymond, L., Mairey, M. et al. Massive sequencing of 70 genes reveals a myriad of missing genes or mechanisms to be uncovered in hereditary spastic paraplegias. Eur J Hum Genet 25, 1217–1228 (2017). https://doi.org/10.1038/ejhg.2017.124
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ejhg.2017.124
This article is cited by
-
Hereditary spastic paraparesis type 46 (SPG46): new GBA2 variants in a large Italian case series and review of the literature
neurogenetics (2024)
-
Hereditary spastic paraparesis type 18 (SPG18): new ERLIN2 variants in a series of Italian patients, shedding light upon genetic and phenotypic variability
Neurological Sciences (2024)
-
Power of NGS-based tests in HSP diagnosis: analysis of massively parallel sequencing in clinical practice
neurogenetics (2023)
-
Clinical phenotyping and genetic diagnosis of a large cohort of Sudanese families with hereditary spinocerebellar degenerations
European Journal of Human Genetics (2023)
-
Reduced penetrance of an eastern French mutation in ATL1 autosomal-dominant inheritance (SPG3A): extended phenotypic spectrum coupled with brain 18F-FDG PET
neurogenetics (2022)