This week, peering inside the proton, identifying research misconduct and making sense of mystery genes.
In this episode:
00:45 Probing the inside of protons
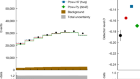
Researchers have made the first attempt to identify the 3D structure within a proton. Research paper: Burkert et al.
06:55 The pitfalls of research misconduct
Being aware of the factors that lead to bad decisions can help researchers avoid scientific misconduct. Comment: Nine pitfalls of research misconduct; Nature Special: Lab Health
13:29 Research Highlights
How pain helps flesh-eating bacteria proliferate, and the carbon emissions of vacations. Research Highlight: Flesh-eating bacteria thrive on pain; Research Highlight: Holiday trips spell trouble for the future of life on Earth
15:06 What are the functions of mystery genes?
How a new method might help identify what bacterial genes do. Research paper: Price et al.; ‘Fitness Browser’ interactive database
21:52 News Chat
Wikipedia’s most cited science papers, and what taking down dams might mean for local ecology. News: Wikipedia’s top-cited scholarly articles — revealed; News: Europe is demolishing its dams to restore ecosystems
Never miss an episode: Subscribe to the Nature Podcast on iTunes or your favourite podcast app. Head here for the Nature Podcast RSS feed.
Transcript
Nature Podcast
Introduction
This is a transcript of the 17th May 2018 edition of the weekly Nature Podcast. Audio files for the current show and archive episodes can be accessed from the Nature Podcast index page , which also contains details on how to subscribe to the Nature Podcast for FREE, and has troubleshooting top-tips. Send us your feedback to podcast@nature.com.
[Jingle]
Interviewer: Benjamin Thompson
Welcome back to the Nature Podcast. This week on the show, we’re probing protons and learning how to maintain a healthy lab environment.
Interviewer: Shamini Bundell
Plus, we’ll be finding out how to make sense of mystery microbe genes. This is the Nature Podcast for the 17th May 2018. I’m Shamini Bundell.
Interviewer: Benjamin Thompson
And I’m Benjamin Thompson.
[Jingle]
Interviewer: Benjamin Thompson
First up this week, reporter Lizzie Gibney is peering into the heart of matter.
Interviewer: Lizzie Gibney
Here at Nature, we talk a fair amount about dark matter. But what about boring old ordinary matter? It may account for just 15% of material in the Universe, but it's a pretty important part. After all, it makes up everything we can see, from stars to us. And even though it’s visible, it still hides plenty of mysteries. The bulk of the matter in the Universe is made up of protons. But what makes up the proton? That's where things get a little fuzzy. Protons are far too small to see under a microscope – around 100,000 times smaller than an atom. So instead, physicists study protons by pinging high-energy electrons off them. These experiments show that each proton must consist of more fundamental particles: three quarks, which are held together by the strong nuclear force. But scientists haven’t known much about how the quarks are arranged in 3D or anything about the proton's mechanical properties. Only now are physicists developing techniques that allow them to probe inside the proton: the particle that's crucial to anything being here at all. To hear how, I spoke to physicist Latifa Elouadrhiri, who explained how techniques to study the proton have evolved.
Interviewee: Latifa Elouadrhiri
Prior to the 90s, the only thing we could study is one-dimensional structure of the proton. And in the 90s there were developments of new formalism that enabled us to connect electromagnetic processes to do three-dimensional structure of the proton. Let me just make simple analogy – so what we have we been doing prior to the 90s and 00s, is like we want to study the heart, and we are studying it through electrography, which is the process of just recording electrical activity of the heart that give us one-dimensional structure that tells us lots about the heart, but not everything. Now with the heart, we have the medical 3D imaging technology that now allow the doctors to learn more in non-invasive manner, the structure of the heart. And this is what we want to do with the new generation of experiments.
Interviewer: Lizzie Gibney
And how do you go about doing those experiments then?
Interviewee: Latifa Elouadrhiri
We are firing high-energy electrons at our protons with very precise measurements. But in order to understand the structure, we want to be able to understand the energy and the momentum that is transferred to the quark so that you acquire both the developments in the theory to interpret results, but also developments in the technology to perform the measurements.
Interviewer: Lizzie Gibney
So, new kinds of theory can connect how electrons bounce of the protons with what’s actually going on inside the proton, things like the forces on the quarks?
Interviewee: Latifa Elouadrhiri
Exactly, and it’s only able to do this interpretation if from the experiment, we have measured all the necessary observables.
Interviewer: Lizzie Gibney
So, what do you actually do then? You fire your electron at the proton, and what happens?
Interviewee: Latifa Elouadrhiri
So, we fire a high-energy electron because by increasing the energy, the wavelength is smaller so we can see deeper into the object. We measure the light that is emitted by the quark, together with the scattered electron, then we also measure the proton. So, we leave the proton intact, we measure it, we measure the produced photon, the light, and the scattered electron. We need to detect all these particles in the final state in order to use the formalism and understand the structure, and that was, this is what was not possible in earlier experiments with electron scattering.
Interviewer: Lizzie Gibney
And what is it then, what did you see, what did you discover about the structure inside the proton?
Interviewee: Latifa Elouadrhiri
So, this is our, the first measurement of the pressure distribution experienced by the quark inside the proton. So, what we found is that there is this extreme outward pressure, but if there was only this pressure in the centre, the proton would explode. But there is another pressure that is going in the other direction, that balances this pressure at the centre that makes the proton stable.
Interviewer: Lizzie Gibney
Which is something we’re very grateful for.
Interviewee: Latifa Elouadrhiri
Yes!
Interviewer: Lizzie Gibney
And can you give me an idea of the scale of the forces that we are talking about, or the pressure at the heart of the proton?
Interviewee: Latifa Elouadrhiri
So, the pressure that we measured is 10 to 35 pascal. This is 10 times larger than, for example, the pressure inside the neutron star.
Interviewer: Lizzie Gibney
Wow, and that’s pretty much the densest matter that we know of, right?
Interviewee: Latifa Elouadrhiri
Exactly.
Interviewer: Lizzie Gibney
And does that match with what theorists predicted?
Interviewee: Latifa Elouadrhiri
That matches some of them. There was a model that were theoretical prediction before this experiment, but this is the first observation, yeah.
Interviewer: Lizzie Gibney
And are you going to be able to use this technique to find out anything else about what’s going on inside a proton?
Interviewee: Latifa Elouadrhiri
This measurement now is just the beginning of a new field of research. So, the paper we published is related to the pressure distributions inside the proton, but next will be to calculate forces, and then move on and understand the 3D imaging of the proton, the spatial distribution of the quark inside the proton, and also the motion of the quark inside the proton.
Interviewer: Lizzie Gibney
Gosh, so the proton might not be so much of a mystery anymore.
Interviewee: Latifa Elouadrhiri
That’s the beginning of solving the mysteries.
Interviewer: Benjamin Thomspon
That was Latifa Elouadrhiri who’s based at the Jefferson Lab in the United States, speaking with reporter Lizzie Gibney. You can find her paper over at nature.com/nature.
Interviewer: Shamini Bundell
This week, Nature is publishing a special on maintaining a healthy, happy and productive lab. A good environment promotes good research, but sometimes things can go wrong. Research misconduct not only degrades the reliability of research findings, but in the most extreme cases can end a career. Tina Gunsalus is Director of the National Center for Professional and Research Ethics, and she’s co-authored a Comment piece outlining the pitfalls of misconduct that can plague departments. She’s also come up with a mnemonic, to help researchers identify the causes. Reporter Geoff Marsh gave her a call.
Interviewer: Geoff Marsh
You’ve worked with troubled departments as part of your job. Could you share some anecdotes of research misconduct that you see coming up time and time again?
Interviewee: Tina Gunsalus
I think what happens is that people with power get cross, they want results, they feel pressured themselves, where are the data, where are the data, why aren’t you doing this right? And the person with no power doesn’t know how to respond effectively, and doesn’t know how to account for themselves, so they start to pretty the data up and make it look like what’s expected, as opposed to hewing to the actual results that the actual research provided. So, I think that that’s a place that things really go wrong - when people are so afraid and don’t have the tools to interact, that they start cutting corners.
Interviewer: Geoff Marsh
Right, so in an effort to help researchers spot the most common pitfalls associated with misconduct, you’ve created a suitably alarming mnemonic which is TRAGEDIES. Could you break that down for us?
Interviewee: Tina Gunsalus
So, it starts with temptation, rationalisation, ambition, group and authority pressure, entitlement, deception, incrementalism, embarrassment and stupid systems. Most of them you’ll see are feelings. Temptation is a feeling. Rationalisation is something entirely internal, you know, and it’s always possible to rationalise any scummy thing you want to do. Entitlement is a feeling. Embarrassment is a feeling. Stupid systems is something that we all exist within, all the time, and if you don’t know how to deal with the mixed messages and mixed incentives, and sometimes perverse incentives in an environment, you can really easily end up in a career tragedy.
Interviewer: Geoff Marsh
And who do you think that the separate elements of the TRAGEDIES mnemonic kind of relate to? Because it sounds almost like it’s mainly focused at maybe the newer scientists who are feeling pressure from above, you know the people at the lower end of the power dynamic. Is that who you’re sort of talking to here?
Interviewee: Tina Gunsalus
Actually, I think it’s pretty universal. I mean, one of the things I did for several decades of my career was, I was the Research Integrity Officer, and I did, led internal investigations for my university. And that included things that weren’t about research, sometimes they were about human subjects, sometimes they were financial, sometimes they were policy violations. And I spent a lot of time investigating very highly-educated, very smart people who had ended up in a situation where they were facing some sort of reputational disaster, shambles. I never met anyone who said, yeah, you know, that was the day I woke up and decided to you know, put my career at risk, potentially go to jail, embarrass my family, lose my job. Organisations are shaped by their leaders, and they’re shaped by the human beings within them, and human beings are susceptible to these pitfalls.
Interviewer: Geoff Marsh
Yeah, and so, individually these can actually be quite small occurrences. They kind of ironically don’t tend to look like actual tragedies themselves. The point is that they, if you spot them early, hopefully you’ll avoid an actual tragedy.
Interviewee: Tina Gunsalus
Sure, I mean I think people don’t know what they don’t know about how really smart individuals get into trouble. And so, if you learn about the TRAGEDIES, and learn to be aware of them in yourself, and how each one of them feels individually, and how they interact, you have more tools for building other professional skills for having the career that you want to have.
Interviewer: Geoff Marsh
In terms of learning, you mention in the article that training alone, formal training, won’t necessarily be as effective for changing attitudes as the informal curriculum, how we see the other people around us working.
Interviewee: Tina Gunsalus
Well, I think that it takes attention both to intentional professional development where people teach practical professional skills, that include how to have disputes and how to raise questions, and it includes attention to the informal environment and the informal curriculum. What happens when someone makes a mistake here? How do we treat them? Is it educational or is it shaming and blaming? So, there’s a whole series of things and they interlock. There’s not one magic bullet, it’s being aware of and taking responsibility for our professional conduct and our professional environments.
Interviewer: Geoff Marsh
What advice would you give to a young PhD student, who did spot some ethically dubious behaviour going on from above?
Interviewee: Tina Gunsalus
Well, I think there are two parts to it, and the most cited paper I ever wrote is called ‘How to blow the whistle and still have a career afterwards’, and the first six steps are all about how to assure that you’re actually in a situation that requires that, because it’s very easy at the bottom of the power curve when you are working so hard, to get tunnel vision and not see the bigger picture. And then, and only then, do you follow the steps for actually blowing the whistle, and there are effective ways to do it. So, there are things that work and things that don’t work, and you have to know that. You don’t know what you don’t know when you start out, and that’s what intentional professional development is about, and that’s why learning about the TRAGEDIES is important.
Interviewer: Geoff Marsh
How do you think research departments should be sort of assessing their own conduct health?
Interviewee: Tina Gunsalus
Well I think there are two very time-effective ways to do it. One of them is the Survey of Organizational Research Climate, the SOuRCe, which can be done in under fifteen minutes. We also have an informal, very quick self-assessment called the Academic Unit Diagnostic Tool available on our website that just gives you a place to start having conversations.
Interviewer: Shamini Bundell
That was Tina Gunsalus talking with Geoff Marsh. You can read more about TRAGEDIES and how to avoid them in her co-authored Comment over at nature.com/news, along with the rest of the special issue, filled with tips and advice for keeping lab culture healthy.
Interviewer: Benjamin Thompson
Still to come in the show, we’ll be taking a look at the most cited science on Wikipedia. That’s coming up in the News Chat. Right now though, Noah Baker’s here, and he’s bought the Research Highlights with him.
[Jingle]
Interviewer: Noah Baker
Flesh-eating bacteria have an excruciating trick up their sleeves. Streptococcus pyogenes destroys skin and soft tissue. But before these symptoms even show up, the infection is extremely painful. Now, researchers have found out why. The bacteria actually release a toxin, which cause pain-sensing neurons in the brain to fire. This sends signals to the immune system to hold back its response. When researchers blocked the neuron signals in mice, the immune system could fight off the infection. The hope is that this could help treat this gruesome disease. Find that paper in Cell.
[Jingle]
Interviewer: Noah Baker
Holidays are heating up the planet. Researchers used an in-depth model to unpick the impact of tourism, and found that worldwide, it’s responsible for 8% of greenhouse gas emissions. That’s quadruple what scientists had previously thought. Plus, this carbon footprint is increasing over time. According to the researchers, American tourists have the biggest footprint, but the impact varies from destination to destination. A holiday in the Maldives has the biggest carbon cost, which puts the country in a tricky situation. Tourism is the largest contributor to the Maldivian economy, but as a low-lying island nation, it’s also one of the countries most threatened by sea level rise. Find out more in Nature Climate Change.
[Jingle]
Interviewer: Shamini Bundell
Sequencing the first human genome took scientists around the world 13 years. Today, that time has been reduced to days, if not hours. But while science may be able to tell you the sequences of genes in your body, in many cases it can’t tell you what the proteins that the genes code for actually do. The same is true for bacteria – you can sequence a bacterial genome, but how do you work out what all those genes are for? I rang up Adam Deutschbauer, to find out how his research is trying to keep up with advances in gene sequencing.
Interviewee: Adam Deutschbauer
The ability to sequence genomes and genes is just extraordinary. It’s really, really easy, right, so we have just in databases, we have millions and millions of different gene sequences, and virtually none of those genes have been studied experimentally.
Interviewer: Shamini Bundell
Why is it hard to figure out what all these genes are doing?
Interviewee: Adam Deutschbauer
The reason why it’s hard to figure out the function of a gene, is that there’s actually not that much data beyond genome sequence in microbiology. And so, the best that we can do, because we have the sequences of so many genes, is there’s basically servers that will try to predict the functions or annotate the functions of genes using automated approaches.
Interviewer: Shamini Bundell
And how are these, like, databases making these predictions about function, just based on a gene sequence?
Interviewee: Adam Deutschbauer
All of the existing servers do it based on the sequence similarity, the something that they have in their database, right? So, they will say, well your, you know, your protein that you sequenced is 40% identical to some characterised protein family. The issue is that we know that even for a closely related genes and proteins that have a high sequence similarity, they can perform very different functions within the physiology of a different bacterium.
Interviewer: Shamini Bundell
And there are people out there doing studies on particular proteins, and obviously it takes a lot of time and effort. And as you’ve said, we’re sequencing more and more bacterial genomes every day, we have more and more data and we can’t keep up with the, sort of identifying the protein functions, or annotating the protein functions. So that’s where you and your colleagues have sort of taken a slightly different approach to solving this problem.
Interviewee: Adam Deutschbauer
The approach that we take is large-scale genetics. So, I’m trained with a genticist, right, Genetics 101 is that you make mutations in an organism, and then you ask what are the consequences of making the mutation. So, the consequences, what you can measure, and the organism that we call the phenotype. And, that’s basically exactly what we did in our story. We made and measured lots and lots of phenotypes of mutants in bacteria across many, many conditions.
Interviewer: Shamini Bundell
And how difficult was it to do something like that at scale, because I can imagine you could take a particular gene, introduce a mutation, and you look at the outcome for that bacterium, but how do you do that for multiple genes?
Interviewee: Adam Deutschbauer
We actually make a large population of mutants in a given bacterium, and we mix together say 100,000 different mutations of an organism, and we mix them all together in the tube and we let them compete against each other under different growth conditions. And so, as they’re competing in different growth conditions, we can measure which genes when knocked out were the winners, and which ones were the losers, across many, many conditions.
Interviewer: Shamini Bundell
So, if you’re growing bacteria in conditions where they need to, for example, get carbon from a particular food source, or be able to resist an antibiotic, you can then find out based on which bacteria do well, which mutations are beneficial or harmful to those processes.
Interviewee: Adam Deutschbauer
So, if you take a population of 100,000 mutants, and many of those mutants are in say, a gene involved in eating sucrose, right, as a carbon substrate. If I grow that population, that population of mutants, and I let them compete against each other in a condition where the only food is sucrose, then those mutants won’t grow.
Interviewer: Shamini Bundell
And then that’s how you know that that gene has something to do with sucrose metabolism. But then you didn’t want to target specific genes for mutations, you actually created a whole load of mutations, tested them all, and then just sort of looked at what came out.
Interviewee: Adam Deutschbauer
Yeah, we tried to develop an approach that was scalable enough, and cheap enough, that we can measure at least genetically, sort of like measure everything for all genes.
Interviewer: Shamini Bundell
So, your experiment obviously couldn’t measure everything, but you got quite a big dataset. How many genes were you then able to predict or guess functions for?
Interviewee: Adam Deutschbauer
You know, we created lots and lots of genetic data across 32 different bacteria, and we have phenotypes for thousands and thousands of genes. The number of genes though that we can make really specific inferences about in their function, is probably in the order of like 600 or so.
Interviewer: Shamini Bundell
And so, if you’ve only got, well 600 is still quite a lot, but those are the ones you’re more confident in, but are the other data, this massive dataset, is that still going to be useful for you or other researchers?
Interviewee: Adam Deutschbauer
We hope so. My colleague Morgan Price created an interactive website for the comparative analysis of all of this data.
Interviewer: Shamini Bundell
And that anyone could go to, and find information about their specific protein that they’re looking in, or their particular species, or a particular pathway?
Interviewee: Adam Deutschbauer
Yes, so you can go to our site now and you can enter a sequence of your favourite protein, and basically do a search against all of our data, and find if we have data for any relatives of your favourite protein. You know, I think long term, right, because we can only mine the data so much ourselves, my hope is that ultimately there’s going to be multiple large databases of experimental data across many, many bacteria, that can be used to augment the existing annotation servers.
Interviewer: Shamini Bundell
That was Adam Deutschbauer of the Lawrence Berkeley National Laboratory. You can find his paper online in the usual place: nature.com/nature.
Interviewer: Benjamin Thompson
Right then listeners, it’s time for this week’s News Chat, and I’m joined here in the studio by Nisha Gaind, one of the News Editors here at Nature. Hi Nisha.
Interviewee: Nisha Gaind
Hi Ben.
Interviewer: Benjamin Thompson
Thanks for joining us. Two stories today, and our first one, we’re going to be delving into some, well, some metrics from Wikipedia, and I don’t think I need to explain to listeners what Wikipedia is, but what’s the story about?
Interviewee: Nisha Gaind
This is a really interesting story, a little data story that shows what are the most cited journal articles on Wikipedia.
Interviewer: Benjamin Thompson
Okay well who’s been doing this data diving then, and what specifically have they been looking at?
Interviewee: Nisha Gaind
So, the Wikimedia Foundation, which runs Wikipedia, released loads of its citation data, and a data scientist called Matt Miller looked at the data to see what the top-cited articles which had DOIs, which I’m sure lots of listeners will know are identifiers for scientific articles, looked at what the top ten most-cited DOI articles are.
Interviewer: Benjamin Thompson
Alright then Nisha, well let’s not muck about – what’s the number one article then?
Interviewee: Nisha Gaind
The most-cited article on the English language version of Wikipedia is a collection of human and mouse genes, and in fact there are lots of DNA sequence collections and gene collections throughout the top ten most-cited DOI journal articles.
Interviewer: Benjamin Thompson
Hmm, and how many citations does the number one spot have then?
Interviewee: Nisha Gaind
So, across all of English Wikipedia it’s got 4,700 citations, and what’s interesting about that is that that’s a lot more than it has even in the scientific literature.
Interviewer: Benjamin Thompson
Well listeners, back in 2014 we had a Feature article that looked at the top 100 citations in academic papers, and that was according to the Web of Science. Nisha, many of the top papers in that study were methods papers, which seem a lot more sparse in the Wikipedia citation list.
Interviewee: Nisha Gaind
Yeah, so what’s interesting about this, and maybe expected, is that lots of the most-cited articles are collections, reference collections, and really interestingly one of the papers which is the third most-cited paper, is a collection of the nomenclature of lunar craters.
Interviewer: Benjamin Thompson
Oh really? So space is always in there, everyone loves space.
Interviewee: Nisha Gaind
It’s always in there. Published in 1971, has thousands of citations on Wikipedia, but just 16 in the scientific literature.
Interviewer: Benjamin Thompson
Okay, and I guess that makes sense then, because I suppose each of these craters presumably has its own Wikipedia page, and each of them referenced this paper.
Interviewee: Nisha Gaind
Exactly, so lots of individual craters will have a page, and the same goes for genes, and they all tend to reference reliable collections that have this information.
Interviewer: Benjamin Thompson
So that’s the top ten of sort of English language Wikipedia then, but of course Wikipedia is a lot more than that. Have people been looking at sort of different languages as well?
Interviewee: Nisha Gaind
Yeah, so the initial release from the Wikimedia Foundation contained information about citations across all of its language editions, and there are nearly 300 of them. So, someone else has looked at the top DOI citations for the whole of Wikipedia, and it’s got a really different paper in the number one slot, which has been cited almost 3,000,000 times. So, this article is a climate paper that looks at how the climate varies around the globe, and the reason that it’s been cited so many times is because it’s cited on lots of articles that are created by a bot, which is an automated computer programme that just creates articles and automatically cites this article.
Interviewer: Benjamin Thompson
Well that’s obviously kind of a heck of a data dive then, but to what end though? What value does this data have?
Interviewee: Nisha Gaind
Citations are really important, whether in Wikipedia or in the scientific literature, because they enable people to trust the information. And what’s really interesting about this study is that we can actually get at this citation data, and do these sorts of comparisons, which is actually quite difficult to do if you’re comparing it to the scientific literature because lots of that information is behind paywall services.
Interviewer: Benjamin Thompson
Alright then Nisha, well let’s move on to our second story then this week, and this is about dams and their effects on ecology. Often, I guess when we think about dams being built, we think about how this might negatively affect sort of wildlife in the area, but this new story perhaps is a little bit different.
Interviewee: Nisha Gaind
Yeah, so this story is about dam removal rather than dam building, which we don’t often talk about much. And the story is about a very large dam in Spain, which is about to be removed, and it’s going to be the biggest dam removal project in the European Union.
Interviewer: Benjamin Thompson
Well why are scientists interested in this then?
Interviewee: Nisha Gaind
So, dams and dam removal is quite important for the ecology of an ecosystem. And the removal of this large dam in western Spain is being hailed as a milestone by ecologists for river restoration efforts. It could help to revive some of the rivers ecosystems.
Interviewer: Benjamin Thompson
And are there any examples where sort of a dam removal has made a difference in the past?
Interviewee: Nisha Gaind
Yeah, absolutely. In the United States, more than a thousand barriers have been removed in recent decades, and they’ve generally had very positive effects on local ecosystems. For example, they’ve improved the habitats of species including certain types of salmon.
Interviewer: Benjamin Thompson
And is this endeavour in Spain part of a sort of wider trend then?
Interviewee: Nisha Gaind
Yeah, it looks as though dam removal is actually on the up around the European Union, after a piece of legislation was introduced in 2000. Now this legislation requires member states to improve ecological protection of rivers and lakes, and because there are hundreds of thousands of dams, some small, some large, many are no longer used, but their presence threatens the habitat of fish and wildlife.
Interviewer: Benjamin Thompson
I guess that some, many perhaps, of these dams were put up without necessarily thinking about local ecosystems. If this is a work in progress, how can we make sure that nothing bad happens?
Interviewee: Nisha Gaind
These projects do absolutely have to be monitored for negative effects as well. Dams are often huge structures, and their removal could damage the surrounding environment. They might allow invasive species to come into freshwater ecosystems, or they might move toxic sediment. So, experts are saying that even though dams might have been built with little regard for the impacts they might have on the ecosystem, we shouldn’t be making the same mistake when they’re then being removed.
Interviewer: Benjamin Thompson
Well finally from me then this week Nisha, what happens next?
Interviewee: Nisha Gaind
So, there are a number of small dams that are scheduled for removal later this year, in countries including the Netherlands, Denmark and Spain. But next year, French scientists are planning to monitor the removal of two massive hydropower dams, one of them 35 metres tall and the other 15 metres tall.
Interviewer: Benjamin Thompson
Oh my goodness, right, well, watch this space then – we’ll see how that goes next year. Nisha, thanks for joining us, and listeners for all the latest science news head over to nature.com/news.
[Jingle]
Interviewer: Shamini Bundell
That’s it for this week’s show. Don’t forget to follow the podcast on Twitter, we’re @NaturePodcast. And if you’d like to get in touch, you can do so on email: podcast@nature.com. I’m Shamini Bundell.
Interviewer: Benjamin Thompson
And I’m Benjamin Thompson, thanks for listening.
[Jingle]

 Nine pitfalls of research misconduct
Nine pitfalls of research misconduct
 Europe is demolishing its dams to restore ecosystems
Europe is demolishing its dams to restore ecosystems
 Wikipedia’s top-cited scholarly articles — revealed
Wikipedia’s top-cited scholarly articles — revealed