Abstract
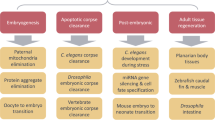
Autophagy involves the sequestration of a portion of the cytosolic contents in an enclosed double-membrane autophagosomal structure and its subsequent delivery to lysosomes for degradation. Autophagy activity functions in multiple biological processes during Caenorhabditis elegans development. The basal level of autophagy in embryos removes aggregate-prone proteins, paternal mitochondria and spermatid-specific membranous organelles (MOs). Autophagy also contributes to the efficient removal of embryonic apoptotic cell corpses by promoting phagosome maturation. During larval development, autophagy modulates miRNA-mediated gene silencing by selectively degrading AIN-1, a component of miRNA-induced silencing complex, and thus participates in the specification of multiple cell fates controlled by miRNAs. During development of the hermaphrodite germline, autophagy acts coordinately with the core apoptotic machinery to execute genotoxic stress-induced germline cell death and also cell death when caspase activity is partially compromised. Autophagy is also involved in the utilization of lipid droplets in the aging process in adult animals. Studies in C. elegans provide valuable insights into the physiological functions of autophagy in the development of multicellular organisms.
Similar content being viewed by others
Introduction
Macroautophagy (herein referred to as autophagy) is an evolutionarily conserved lysosome-mediated degradation process. It involves the formation of a double-membrane structure, called the autophagosome, and its subsequent maturation and fusion with lysosomes to form degradative autolysosomes, in which the sequestrated cytosolic contents and the autophagosomal inner membrane are degraded1,2. Autophagy is induced in response to various metabolic stresses, and engulfs a portion of cytoplasm for degradation, thereby acting as a cell survival mechanism. Autophagy also functions as a quality control system by selectively removing misfolded and aggregate-prone proteins, and damaged or superfluous organelles. Dysfunction of autophagy has been linked to the development of a broad range of human diseases, including tumorigenesis, neurodegeneration and immune diseases3. This review focuses on the role of autophagy during C. elegans development.
Conservation of the autophagic machinery in C. elegans
The establishment of Saccharomyces cerevisiae as a genetic model to study autophagy has laid the groundwork for our understanding of the molecular mechanism of autophagy4,5,6. Under nutrient-deficient conditions, yeast cells lacking vacuolar proteinases accumulate spherical membrane structures in the vacuole (the yeast lysosome counterpart). These vesicles, also known as autophagic bodies, are derived from fusion of autophagosomes with the vacuole, and consist of the enclosed autophagosomal inner membrane and its sequestrated cytosolic contents4. The transport of vacuolar enzymes including the precursor form of aminopeptidase I (prApeI) and α-mannosidase under vegetative growth conditions involves their selective packaging into autophagosome-like vesicles, called Cvt vesicles, and subsequent fusion of Cvt vesicles with the vacuole7. Genetic screens have identified approximately 18 ATG (autophagy related genes) that are essential for the formation of autophagosomes and Cvt vesicles2,5,6. These Atg proteins form distinct complexes involved in different steps of autophagosome biogenesis. The Atg1/Atg13 complex and the class III phosphatidylinositol 3-kinase Vps34 complex are required for the induction and nucleation of the crescent-shaped double membrane known as the isolation membrane or phagophore. Expansion of isolation membranes into autophagosomes requires the two ubiquitin-like conjugation systems. The ubiquitin-like protein, Atg8, through the actions of the E1-like enzyme Atg7 and the E2-like conjugating enzyme Atg3, is conjugated to phosphatidylethanolamine (PE), while the ubiquitin-like protein Atg12 is conjugated to Atg5 via the sequential reactions of Atg7 and the E2-like enzyme Atg10. The Atg12-Atg5 conjugate further interacts with the self-oligomerized protein Atg16 to form a multimeric complex, which possesses an E3-like ligase activity for Atg8-PE conjugation and also targets Atg8-PE to the preautophagosomal structure (PAS), from which autophagosomes are originated2. The multispanning membrane protein Atg9 traffics between mobile Atg9-positive vesicles in the cytoplasm and the PAS to provide a membrane source for autophagosomes8,9. The Atg2/Atg18 complex and the Atg1 complex are required for retrieval of Atg9 from the PAS8,10. How these Atg proteins act coordinately to regulate autophagosome formation remains largely unknown.
Study of autophagy in higher eukaryotes is greatly facilitated by the conservation of Atg genes and their roles in autophagosome formation. The autophagic process, however, exhibits fundamental differences in higher eukaryotes, including integration of various signaling pathways during development and also the maturation of nascent autophagosomes before fusion with lysosomes11. Autophagic removal of a variety of protein substrates during embryogenesis establishes C. elegans as a model suitable for genetic screens for essential autophagy genes12,13. Worm homologs of yeast Atg genes, including Atg1-10, 12, 16 and Atg18, and genes that have no yeast counterparts or that are distantly related homologs of Atg genes, including epg-1, -3, -4, -5, -6, -8 and -9, have been identified from genetic screens14,15. epg-1 and epg-8 encode highly divergent Atg13 and Atg14 homologs, respectively16,17. epg-9 encodes the mammalian Atg101 homolog, and EPG-9 forms a complex with UNC-51/Atg1 and EPG-1/Atg1318. epg-3 and epg-4, encoding transmembrane proteins, and epg-6, encoding a WD40 repeat PtdIn(3)P-binding protein, are involved in progression of omegasomes/isolation membranes into autophagosomes14,15. The novel protein epg-5 is essential for the formation of degradative autolysosomes14,19. Thus, the more elaborate autophagic machinery in higher eukaryotes involves highly conserved counterparts of yeast Atg proteins, highly divergent components and also metazoan-specific factors. The presence of multiple homologs of the same yeast Atg genes endows another layer of complexity on the autophagic machinery. The two Atg4 homologs and Atg16 homologs have partially redundant functions in, but also contribute differentially to, the autophagic degradation of protein aggregates20,21. A detailed summary of the autophagy pathway in C. elegans is described in the recent review of Lu et al.11.
Embryogenesis: autophagy as a picky eater
Degradation of protein aggregates
A variety of protein aggregates are degraded by autophagy during embryogenesis P granules, specialized protein-RNA aggregates, are maternally loaded and dispersed throughout the cytoplasm of a newly fertilized embryo, but during a series of asymmetric cell divisions they become localized exclusively in the germline blastomeres P1, P2, P3, and P4 and finally in the two germ precursor cells, Z2 and Z3, derived from P422. The P granule components, PGL-1 and PGL-3, which are partitioned into somatic cells during early asymmetric cell divisions, are degraded by autophagy (Figure 1)12. PGL-1 and PGL-3 form numerous aggregates in somatic cells in autophagy mutants. Autophagic degradation of PGL-1 and PGL-3 and their accumulation into aggregates in somatic cells require the self-oligomerizing protein SEPA-112. In early-stage embryos, SEPA-1 forms aggregates independent of PGL-1 and PGL-3, and is itself removed by autophagy, resulting in the absence of SEPA-1 in late-stage embryos12. In autophagy mutants, SEPA-1 completely colocalizes with PGL-1 and PGL-3 forming aggregates termed PGL granules that persist throughout embryogenesis12.
The hierarchical recruitment of receptor and scaffold proteins triggers selective degradation of protein aggregates. In degradation of PGL granules, methylated PGL-1 and PGL-3 are recruited into SEPA-1 aggregates, which associate with the scaffold protein EPG-2. The scaffold protein EPG-7 mediates the degradation of AIN-1 and also of SQST-1. The cargo/receptor/scaffold complex triggers the formation of surrounding autophagosomal membranes. EPG-2 and EPG-7 directly interact with multiple ATG proteins, including LGG-1/Atg8. SEPA-1, AIN-1 and SQST-1 also interact with LGG-1.
Like p62/sequestosome-1 (SQSTM1) in mammalian cells and flies, the C. elegans homolog of p62, SQST-1 (SeQueSTosome-related protein), is also degraded by autophagy14. In autophagy mutant embryos, SQST-1 is present at dramatically increased levels and accumulates into a large number of aggregates that are distinct from PGL granules14. In addition to PGL granules and SQST-1, other proteins, including SEPA-1 family members and W07G4.5, are also selectively removed by autophagy during embryogenesis12,23. Why many protein aggregates are selectively degraded by autophagy during embryogenesis? The development of C. elegans embryos relies on the degradation of maternally-loaded factors rather than external nutrients. Autophagic degradation of P granules and other protein substrates may provide a source of nutrients for embryogenesis. Some protein factors may function only at a specific embryonic stage and their persistence would be detrimental to embryonic development.
The scaffold protein confers cargo selectivity and degradation efficiency A family of proteins that simultaneously associate with the cargo and Atg8/LC3 has been suggested to act as receptors for tethering protein aggregates to the autophagic machinery24. p62, which contains a self-oligomerization PB1 domain, an LC3-interacting LIR motif and a UBA domain, acts as a receptor for autophagic degradation of ubiquitinated protein aggregates24. SEPA-1 exhibits many features similar to p62, including self-oligomerization and direct interactions with the cargo PGL-3 and LGG-1/Atg8, and thus acts as a receptor for degradation of PGL-1 and PGL-312. However, the colocalization of PGL granules with LGG-1/Atg8 puncta also requires the scaffold protein EPG-214. EPG-2 forms aggregates and is removed by autophagy, and both of these processes are independent of pgl-1, pgl-3 or sepa-114. EPG-2 directly interacts with SEPA-1 and also with multiple Atg proteins25. Therefore, degradation of PGL-1 and PGL-3 involves the hierarchical recruitment of receptor and scaffold proteins.
EPG-7 acts as a scaffold protein in degradation of SQST-1 during embryogenesis (Figure 1)23. EPG-7 self-oligomerizes and is itself degraded by autophagy in a SQST-1-independent manner. EPG-7 directly interacts with SQST-1 and also associates with the autophagy proteins: LGG-1, LGG-3/Atg12, ATG-18 and ATG-923. Like the role of EPG-2 in degradation of PGL granules, EPG-7 links SQST-1 aggregates with LGG-1 puncta23. Thus, autophagic degradation of different types of protein aggregates is mediated by distinct scaffold proteins. The cargo/receptor/scaffold protein complex may act as a PAS-like structure, providing a platform for recruiting core Atg proteins to protein aggregates, which then trigger the assembly of surrounding autophagosomal membranes.
The utilization of a scaffold protein appears to be a general mechanism for promoting the delivery of the cargo/receptor complex to the autophagosome. In the Cvt pathway, Atg11 interacts with the receptor protein Atg19 to transport the prApe1 complex to the PAS26,27. Atg11 also interacts with multiple Atg proteins, including Atg1 and Atg9, which acts at the top of the hierarchical recruitment of Atg proteins to the PAS28,29,30. Yeast Atg11 also interacts with the mitochondrial outer membrane-spanning protein Atg32 and the peroxisome-associated protein PpAtg30, which act as receptors for removal of excess or damaged mitochondria (mitophagy) and peroxisomes (pexophagy), respectively31,32,33. The phosphatidylinositol 3-phosphate-binding protein ALFY scaffolds a complex between p62-positive aggregates and the Atg5/Atg12/Atg16L complex and LC3 to increase degradation efficiency34. EPG-7 displays a sequence similarity to yeast Atg11 and mammalian FIP200, and all three proteins contain an Atg11 motif at the C-terminus23. In mammalian cells, FIP200 is a component of the ULK1 complex and is essential for autophagy induction35. Whether FIP200 also has a function independent of the ULK1 complex in promoting cargo degradation efficiency remains unknown.
Association of the cargo/receptor complex with the scaffold protein is regulated by post-translational modification How is the hierarchical recruitment of cargo, receptor and scaffold proteins regulated? These proteins may be expressed in a sequential temporal pattern; alternatively, their interactions may be controlled at the post-translational level. In the degradation of PGL-1 and PGL-3, SEPA-1 is zygotically synthesized and displays a dynamic expression pattern that overlaps with EPG-225. A recent study demonstrated that the interaction between the cargo (PGL-1 and PGL-3) and the receptor (SEPA-1) and also the interaction of the cargo/receptor complex (PGL granule) with the scaffold protein (EPG-2) are controlled by post-translational arginine methylation25. Loss of function of epg-11, which encodes the C. elegans arginine methyltransferase PRMT1 homolog, causes accumulation of PGL granules25. EPG-11 directly binds to PGL-1 and PGL-3 and methylates their RGG domains25. Arginine methylation of the RGG domains of PGL-1 and PGL-3 may prevent the formation of PGL-1 and PGL-3 aggregates, resulting in the diffuse localization of PGL-1 and PGL-3 and their strict dependence on SEPA-1 for formation of aggregates (Figure 1). The interaction between PGL-3 and SEPA-1 is greatly enhanced in epg-11 mutant embryos25. The altered composition or organization of PGL granules in epg-11 mutants reduces the binding affinity of SEPA-1 with EPG-2, and thus the autophagic degradation efficiency of PGL granules is greatly impaired. Ectopic accumulation of PGL-1/PGL-3 in somatic cells impairs the degradation of SEPA-1, but has no effect on the autophagic removal of EPG-2 and other autophagy substrates25. Therefore, in addition to conferring cargo selectivity and promoting degradation efficiency, the tightly controlled association of the cargo/receptor complex with the scaffold protein also prevents ectopic accumulation of a specific type of cargo/receptor complex from saturating the autophagic flux and thus impairing degradation of other selective cargoes.
The interaction among cargo, receptor and scaffold proteins is also modulated by post-translational modification in other systems. When mutant huntingtin protein is acetylated, it exhibits preferential interaction with p62, which promotes its trafficking into autophagosomes36. Phosphorylation of p62 at serine 403 in the UBA domain increases its binding affinity with polyubiquitinated chains37. Phosphorylation of PpAtg30 and Atg32 facilitates their interactions with Atg11 and triggers pexophagy and mitophagy in yeast31,38. The hierarchical recruitment of the receptor and scaffold proteins provides multiple regulatory layers for cellular or developmental modulation of degradation efficiency.
Clearance of paternal mitochondria and MOs
In animals, offspring inherit the mitochondrial genome (mtDNA) only from the female parent, known as maternal inheritance. In Drosophila melanogaster, two developmental barriers act prior to fertilization to prevent paternal mtDNA from being transmitted to the next generation39. mtDNA is eliminated by the mitochondrial endonuclease G during sperm elongation, and is also removed from spermatids into waste bags by a cellular remodeling process during the spermatid individualization stage39. In mice, sperm mtDNA is also pre-eliminated before fertilization40. If a sperm containing mtDNA enters the zygote, the mtDNA is not eliminated and becomes unevenly distributed in blastomeres during cleavage, persisting in only a small fraction of embryonic cells40. Unlike in flies and mice, in which maternal inheritance of mtDNA is a passive process, autophagy selectively degrades paternal mitochondria and mtRNA in C. elegans41,42,43. In two- and four-cell stage embryos, sperm-derived mitochondria are randomly partitioned into blastomeres, but they are eliminated before the 64-cell embryonic stage. In autophagy mutant zygotes, paternal mitochondria and mtDNA persist in late embryos and even in larvae. Paternal mitochondria are associated with LGG-1-labeled autophagosomes in the early embryo (Figure 2)41,42. Ubiquitination has been shown to act as a trigger for selective autophagic degradation of various autophagy substrates24. How C. elegans sperm mitochondria, which are not ubiquitinated, are recognized and degraded by the autophagic machinery has yet to be elucidated. In mice, sperm mitochondria are labeled by p62, LC3 and ubiquitin immediately after fertilization, but are not engulfed by autophagosomes40. p62 and LC3 eventually disengage from sperm mitochondria40.
Autophagic degradation of paternal mitochondria and MOs after fertilization. Schematic of the C. elegans spermatogenesis process. The two secondary spermatocytes may or may not remain attached by a cytoplasmic bridge. The cytosolic components are differentially segregated into developing spermatids and the residual body. Spermatids inherit mitochondria, Golgi-derived fibrous body-membranous organelles (FB-MOs) and a haploid nucleus, while the residual body contains all ribosomes, nearly all actin and myosin and most of the tubulin. Once spermatids bud from the residual body, the FB-MO complex is reorganized and FBs disappear. Sperm mitochondria and MOs are degraded by autophagy after fertilization. Residual bodies are engulfed and degraded by the gonadal sheath cells.
C. elegans spermatids also contain membranous organelles (MOs), which are specialized vesicular structures. MOs are also engulfed by LGG-1-positive autophagosomes (Figure 2)41,42. Unlike mitochondria, MOs in newly fertilized embryos are ubiquitinated, which probably acts as the signal to trigger their selective recognition and degradation by autophagy41,42. The 19S regulatory subunit of the proteasome also surrounds MOs, suggesting that proteasome activity may contribute to MO degradation42.
As spermatids terminally differentiate into spermatozoa in mammals, the bulk cytoplasmic contents are segregated into the residual bodies, which are detached from the spermatids and eliminated44. It has been suggested that disposal of the residual body is initiated by autophagy and then completed by phagocytosis by Sertoli cells45. During C. elegans spermatogenesis, residual bodies are generated during the second meiotic division of spermatocytes when spermatids are generated (Figure 2)46. All ribosomes, nearly all the actin and myosin, and most of the tubulin and many internal membranes are partitioned into the residual body47,48. Autophagy mutants show no evident accumulation of residual bodies49. Residual bodies, which expose the “eat-me” signal phosphatidylserine (PS) on the surface, are recognized, engulfed and degraded by gonadal epithelial sheath cells (Figure 2), a process that uses the same set of genes underlying apoptotic cell removal49. Efficient removal of residual bodies regulates the number of spermatids and also the effective transfer of spermatids during male mating49.
Removal of apoptotic cell corpses during embryogenesis
Phagocytosis involves the engulfment of extracellular constituents in a single-membrane phagosome, which then undergoes extensive membrane remodeling, including sequential recruitment of PtdIns(3)P and RAB small GTPases, and finally fuses with lysosomes for degradation. Autophagy proteins have been shown to participate in phagosome-mediated degradation. LC3 is recruited to phagosomes containing apoptotic cells, macropinosomes and entotic vacuoles (which form when one live epithelial cell engulfs another), facilitating their fusion with lysosomes, and consequently the degradation of phagocytosed cargoes50,51,52. Recruitment of LC3 depends on the PtdIn(3)P-generating complex (Beclin1 and Vps34) and the two conjugation systems (Atg5 and Atg7), but not the ULK-1-ATG13-FIP200 complex50,51. In the clearance of apoptotic cell corpses during cavitation in mouse embryoid bodies and chick retinal development, autophagy contributes to this process by generating the cellular ATP that is required to expose the “eat-me” signal PS on the apoptotic cell surface53,54.
During C. elegans embryonic development, 113 somatic cells undergo programmed cell death55. The neighboring cells recognize the PS exposed on the apoptotic cell surface, and then envelop and internalize the cell corpse to form a phagosome56,57. Two partially redundant signaling pathways, with ced-2, ced-5 and ced-12 in one pathway and ttr-52, ced-1, ced-7, ced-6 and dyn-1 in the other, mediate recognition and phagocytosis of cell corpses56,57. The nascent cell corpse-containing phagosomes undergo stepwise maturation processes, involving the dynamic coating of PdtIns(3)P and sequential recruitment of RAB proteins, including RAB-5, RAB-7, UNC-108/RAB2 and RAB-14, before fusion with lysosomes to generate phagolysosomes for degradation56,57,58,59. The class II PI/PtdIns(3)-kinase PIKI-1 and the class III PtdIns 3-kinase VPS-34 act coordinately to regulate PtdIns(3)P generation and accumulation on phagosomes60,61. Loss of function of autophagy genes acting at distinct steps of the autophagy pathway, including genes encoding components of the UNC-51/EPG-1 Atg1 complex, the VPS-34/BEC-1/EPG-8 complex and the two conjugation systems, atg-9 and epg-5, causes an increased number of cell corpses in C. elegans embryos62,63. In autophagy mutant embryos, the frequency of cell death is normal but the duration of cell corpses is prolonged compared to wild type63. Loss of function of autophagy genes has no effect on PS exposure on the dying cell, nor on the internalization of cell corpses62,63. Phagosome maturation, however, is impaired by defective autophagy. PtdIns(3)P level on phagosomes is reduced and phagosomal association of RAB-5 and RAB-7 is decreased63. Autophagy genes act in the same pathway with VPS-34 but in parallel to PIKI-1 to regulate phagosomal PtdIns(3)P levels63. The involvement of autophagy genes acting at all steps of the autophagy pathway in cell corpse removal argues that the autophagy process itself, rather than additional functions of autophagy genes, is involved. Autophagy may modulate the formation of different VPS-34 complexes with distinct functions in the endocytic and autophagic pathways. Alternatively, PtdIns(3)P-containing autophagic structures may directly fuse with phagosomes. Taken together, the existing evidence shows that autophagy activity is essential for selective removal of protein aggregates, sperm mitochondria, MOs and also apoptotic cell corpses during embryogenesis.
Post-embryonic cell fate specification: autophagy and miRNA-mediated gene silencing
miRNAs, ∼22 nucleotide noncoding RNAs, post-transcriptionally repress gene expression by forming non-perfect complementary interactions with the 3′ untranslated region of target mRNAs to inhibit their translation and/or promote their degradation64. miRNA-mediated gene silencing is mediated by the miRNA-induced silencing complex, which contains Argonaute (Ago), miRNA and a member of the GW182 family of proteins65. In C. elegans, miRNAs post-transcriptionally regulate the expression of heterochronic genes to specify the temporal fate of a row of lateral hypodermal seam cells, which undergo stage-specific developmental programs66. The let-7 family miRNAs, including mir-84, mir-48 and mir-241, inhibit the expression of hbl-1 to control succession from the L2 to L3 larval stage, while let-7 miRNA downregulates the expression of lin-41 in specifying the L4-adult switch67,68,69. Loss of function of the miRNA pathway, including mutations in dcr-1 (Dicer homolog), alg-1 and alg-2 (the Argonaute homologs), and ain-1 and ain-2 (the GW182 homologs), leads to a retarded heterochronic defect, in which the L2-stage-specific division is reiterated at the L3 stage and terminal differentiation at the late L4 stage is delayed, manifested as an increased number of seam cells and incomplete formation of cuticular alae structures70,71. Autophagy mutants exhibit neither precocious nor retarded heterochronic phenotypes72. However, loss-of-autophagy mutations partially suppress the retarded heterochronic defects associated with dcr-1, alg-1 or ain-1 mutations72. Levels of reporters containing the 3′ untranslated region from hbl-1 and lin-41, two genetically verified targets of let-7 family miRNAs, are downregulated in autophagy mutants in a let-7-dependent manner, while elevation of autophagy activity by inactivation of Tor signaling or starvation increases their expression72.
Autophagy also regulates other developmental processes controlled by miRNAs. The let-7 family miRNAs negatively regulate let-60/RAS, which is essential for vulval cell fate specification73. Vulval precursor cell fate is sensitive to the dosage of let-60 activity. Loss of function of let-60 causes a vulvaless phenotype, while elevated let-60 activity results in generation of supernumerary vulva-like structures, known as multivulva74. Autophagy mutants show normal vulval development. Loss of autophagy activity, however, ameliorates the multivulva phenotype associated with activating mutations in let-60 and this suppression effect is dependent on the miRNA pathway72. The miRNA lsy-6 controls the left-right asymmetry of the two ASE taste neurons ASE left (ASEL) and ASE right by repressing the Nkx-type homeodomain gene cog-1 in ASEL75. lsy-6(null) mutants show a loss of the ASEL fate with a concomitant adoption of the ASE right-specific fate. This defect has incomplete penetrance in lsy-6 hypomorphic mutants. Loss of autophagy activity has no effect on ASEL fate specification, but suppresses the defect in lsy-6(hypo) mutants72. In contrast, elevated autophagy activity exacerbates the ASEL specification defect in lsy-6(hypo) mutants72. Thus, autophagy participates in diverse biological processes regulated by various miRNAs. Loss of autophagy activity enhances miRNA-mediated gene silencing.
Autophagy modulates miRNA-induced gene silencing by degrading components of miRNA-induced silencing complex. Compared to wild-type animals, AIN-1 is present at increased levels and accumulates into a large number of aggregates that colocalize with SQST-1 in autophagy mutants72. EPG-7, the scaffold protein for removal of SQST-1, mediates the degradation of AIN-1 (Figure 1)72. Degradation of components of the RNA-induced silencing complex (RISC) by autophagy has recently emerged as a mechanism for modulating small interfering RNA (siRNA)- or miRNA-mediated gene silencing. In mammalian cells, DICER and AGO2 are targeted for autophagic degradation by the receptor NDP5276. In Arabidopsis, ARGONAUTE1 is degraded by autophagy, a process triggered by the viral suppressor of RNA silencing protein P0 and also by impaired production of miRNAs77.
To live: autophagy, lipid metabolism and aging
During C. elegans post-embryonic development, autophagy functions as a survival mechanism when the animal experiences metabolic stresses such as starvation. In the absence of food, newly hatched larvae fail to initiate larval development and arrest at the L1 diapause, remaining viable for about 1-2 weeks. Inhibition of autophagy greatly shortens the survival of worms during L1 starvation14,15,16,17,18,20,21,78. When early larvae experience harsh environmental conditions, including high population density, limited food supply or high temperature, they adopt a specialized third larval stage, called the dauer diapause, for long-term survival. Loss of autophagy activity significantly reduces dauer formation79. At the adult stage, autophagy activity slows down the aging process13,21. Autophagy activity is also required for the extended life-span associated with germline removal, TOR inhibition, dietary restriction, reduced insulin/IGF-1 signaling and reduced mitochondrial respiration13. Recent studies demonstrated that autophagy acts concomitantly with lipolysis to modulate the aging process.
Lipid metabolism and autophagy
Lipid droplets (LDs) contain a hydrophobic lipid core enclosed by a phospholipid monolayer that is decorated by membrane-associated proteins80. Excess cellular nutrients can be converted into triglyceride and stored in LDs. Under starvation conditions, lipases are recruited to the surface of LDs and break down triglyceride into free fatty acids that are utilized by mitochondria to generate energy through β-oxidation81. In hepatocytes, starvation induces engulfment of LDs by autophagosomes, which are subsequently delivered to lysosomes for the hydrolysis of LD triglyceride, a process termed lipophagy82. Inhibition of autophagy increases the triglyceride level and the number and size of LDs, and decreases β-oxidation in hepatocytes82. The livers of Atg7 knockout mice also display increased triglyceride storage in LDs compared to wild-type animals under starvation conditions82. How the LDs are selectively recognized and degraded by the autophagic machinery has yet to be determined. In contrast to hepatocytes, autophagy is required in adipocytes for normal adipogenesis83,84. In adipocyte-specific Atg7 knockout mice, the white adipose tissue exhibits characteristics of brown adipocytes, containing multilocular LDs in contrast with the single large unilocular LD in white adipose cells, and numerous mitochondria, and displays increased levels of fatty acid β-oxidation83,84.
In C. elegans, autophagy also facilitates lipid storage in the intestine85. Compared to wild-type animals, autophagy mutants, including bec-1 and unc-51 mutants, have fewer LDs and lower triglyceride levels in the intestinal cells85. The C. elegans intestine performs multiple functions, including digestion of food, absorption of processed material and nutrients, and lipid biogenesis. Autophagy may facilitate lipid storage by directly providing raw materials generated from the recycling of misfolded proteins and damaged or superfluous organelles, or it may act to stimulate lipogenesis indirectly.
Lipid metabolism, autophagy and aging in C. elegans
Autophagy activity is induced and is required for longevity in germline-deficient glp-1 mutants and insulin signaling-compromised daf-2 mutants79,86. The transcription factor PHA-4/FOXA is required for the induction of autophagy and the extended lifespan in glp-1 mutants86. The longevity of glp-1 and daf-2 mutant animals is also linked to an increased expression of the triacylglycerol lipase, LIPL-4, which is upregulated in a daf-16-dependent manner (Figure 3)86,87. Autophagy and LIPL-4 are interdependently regulated in glp-1 mutants. Autophagy is required to maintain high lipase activity, while LIPL-4 is required for autophagy induction86. The effect of LIPL-4 appears to be mediated by ω-3/6 polyunsaturated fatty acids (PUFAs), the production of which is induced by overexpression of lipl-488. Inactivation of lipl-4 also suppresses the longevity of daf-2 mutants87. Autophagy induction in daf-2 mutants is independent of daf-1689. Thus, LIPL-4, activated by DAF-16, acts coordinately with autophagy to mediate the longevity of daf-2 mutants.
Autophagy, lipolysis and aging. The expression of LIPL-4 is activated by DAF-16. Autophagy and LIPL-4 are interdependently regulated. In response to nutrient availability, HLH-30 acts antagonistically to MXL-3 to regulate the expression of lysosomal lipases, including LIPL-1 and LIPL-3. Enhanced lipid metabolism extends lifespan of animals.
Autophagy is also required for utilization of lipid stores for lifespan extension in C. elegans. In addition to LIPL-4, other lipases in C. elegans, including LIPL-1, -2, -3, and -5, are upregulated under starvation conditions90. LIPL-1, -2, and -3, but not LIPL-4, localize to lysosomal structures in the intestine90. LIPL-1 and LIPL-3 are required for lipid breakdown through lipophagy in worms under starvation conditions (Figure 3)90.
MXL-3 and HLH-30/TFEB in lipid metabolism
The basic-helix–loop–helix transcription factor MXL-3 (Max-like 3) represses expression of the lysosomal lipases lipl-1, -2, -3 and -5, but not lipl-4, in well-fed animals (Figure 3)90. As an acute response to nutrient starvation, MXL-3 is downregulated and consequently the expression of lipl-1 and -3 is induced to mobilize lipids as an energy supply90. mxl-3 mutants are long lived, a phenotype that is independent of daf-16, pha-4 or rheb-190. mxl-3 mutants exhibit increased lipolysis activity, but show no enhanced autophagy activity, suggesting that increased lysosomal lipolysis extends the worm lifespan90.
The helix-loop-helix transcription factor TFEB is a master regulator for expression of genes essential for the autophagy-lysosomal pathway91. TFEB activity is regulated by serine phosphorylation mediated by the extracellular signal-regulated kinase ERK2, which controls the nuclear localization of TFEB91. HLH-30, the worm homolog of mammalian TFEB, acts antagonistically to MXL-3 to regulate lysosomal lipases in response to nutrient availability90,92. Under starvation conditions, HLH-30 is transcriptionally activated and translocates into the nucleus where it activates the expression of lipl genes and autophagy genes (Figure 3)92. hlh-30 mutants show impaired autophagy induction and lipase expression, and consequently the utilization of LDs is slowed down90. hlh-30 mutants show a modest reduction in lifespan, while overexpression of HLH-30 extends the lifespan90,92. hlh-30 is required for the lifespan extension in animals with germline removal, TOR inhibition, reduced insulin/IGF-1 signaling and mitochondrial dysfunction92. These long-lived animals also exhibit increased HLH-30 nuclear localization and elevated autophagy activity92. Loss of hlh-30 function also suppresses the longevity phenotype of mxl-3 mutants90. Therefore, HLH-30 mediates lifespan extension in multiple longevity pathways by autophagy induction and/or increased lysosomal lipolysis.
The role of TFEB in regulation of lipid metabolism by autophagy is evolutionarily conserved93. Expression of TFEB is induced by starvation through an autoregulatory loop, which then exerts global transcriptional control on genes involved in lipid metabolism93. Liver-specific depletion of TFEB impairs, while overexpression of TFEB facilitates, lipid degradation93. Autophagy is required for lipid degradation regulated by TFEB93.
To die: autophagy contributes to cell death
Autophagy has been shown to act as a mechanism for executing cell death in the context of impaired functions of apoptosis factors in mammalian cells94. For example, various apoptotic stimuli such as etoposide and staurosporine induce non-apoptotic death of Bax/Bak knockout fibroblasts in an autophagy-dependent manner95. During Drosophila development, autophagy and caspases exhibit context-specific roles in physiological cell death in different types of tissues96. In metamorphic cell death of the salivary gland, caspase activation occurs in parallel with autophagy, and their coordinated action leads to the complete removal of salivary glands97,98. Autophagy acts upstream of caspase activation to promote cell death in the female germline99, while autophagy per se functions as a cell death mechanism in degeneration of the midgut100. During C. elegans development, 131 somatic cells undergo programmed cell death. The death program is initiated by transcriptional activation of the BH3-only protein EGL-1, which binds to the Bcl-2-like anti-apoptotic protein CED-9, and subsequently leads to activation of the pro-apoptotic proteins CED-4 and CED-3, which are the C. elegans orthologs of mammalian Apaf1 and caspases, respectively101. In the gonad of adult hermaphrodites, a large number of germline cells at the meiotic pachytene stage of oogenesis also undergo programmed cell death102. Physiological germ cell death is executed via a cascade of apoptotic factors, composed of CED-9, CED-4 and CED-3, but independent of EGL-1103. Genotoxic stress such as DNA damage also induces meiotic pachytene-stage germ cell death that is triggered by the activation of CEP-1, the worm homolog of mammalian p53, which transcriptionally activates egl-1 and ced-13 (which encodes another BH3 domain-only protein)102. Autophagy activity is dispensable for physiological germline cell death104. However, compared to wild-type animals, the number of germline cell corpses is lower in autophagy mutants after γ-ray treatment104. The engulfment and degradation of germ cell corpses, which is mediated by gonadal sheath cells, are not altered in autophagy mutants104. Autophagy appears to function downstream of EGL-1 and CED-13 or in parallel to the core cell death machinery to contribute to genotoxic stress-induced germline cell death104.
In C. elegans, when caspase activity is partially compromised, cells that normally undergo programmed cell death have the ability to escape death, even after activation of the apoptotic program105,106. This reversion of fate is enhanced by mutations in engulfment genes. The six Pn.aap cells (posterior daughters of the anterior daughters of the anterior daughters of P blast cells) generated from P1, P2, P9 to P12 in the ventral cord normally die, but some of them survive in mutants with partial loss of ced-3 activity105,107. Autophagy itself does not trigger cell death in the ventral cord. However, inhibiting autophagy increases the survival of Pn.aap cells generated from P1, P2, P9 to P12 when ced-3 activity is partially compromised104. Autophagy activity also contributes to physiological germline cell death in mutants with partially reduced caspase activity104.
Concluding remarks
The wealth of knowledge about its developmental processes makes C. elegans an excellent model system to investigate the functions of autophagy under normal physiological conditions and also when the animal experiences various stresses. In future, the powerful genetic tools that are available to study C. elegans may help us to reveal the molecular mechanisms underlying the selective autophagic removal of protein aggregates, paternal mitochondria, MOs and lipid droplets. Investigating how various developmental signaling and environmental cues are integrated into the autophagic machinery will also be of great importance in understanding autophagy regulation during C. elegans development. Ultimately, these studies will provide insights into how autophagy dysfunction leads to the development of a broad range of human diseases, such as metabolic disorders and neurodegeneration.
References
Xie ZP, Klionsky DJ . Autophagosome formation: core machinery and adaptations. Nat Cell Biol 2007; 9:1102–1109.
Nakatogawa H, Suzuki K, Kamada Y, Ohsumi Y . Dynamics and diversity in autophagy mechanisms: lessons from yeast. Nat Rev Mol Cell Biol 2009; 10:458–467.
Levine B, Kroemer G . Autophagy in the pathogenesis of disease. Cell 2008; 132:27–42.
Takeshige K, Baba M, Tsuboi S, Noda T, Ohsumi Y . Autophagy in yeast demonstrated with proteinase-deficient mutants and conditions for its induction. J Cell Biol 1992; 119:301–311.
Tsukada M, Ohsumi Y . Isolation and characterization of autophagy-defective mutants of Saccharomyces cerevisiae. FEBS Lett 1993; 333:169–174.
Harding TM, Morano KA, Scott SV, Klionsky DJ . Isolation and characterization of yeast mutants in the cytoplasm to vacuole protein targeting pathway. J Cell Biol 1995; 131:591–602.
Kim J, Klionsky DJ . Autophagy, cytoplasm-to-vacuole targeting pathway, and pexophagy in yeast and mammalian cells. Annu Rev Biochem 2000; 69:303–342.
Mari M, Griffith J, Rieter E, Krishnappa L, Klionsky DJ, Reggiori F . An Atg9-containing compartment that functions in the early steps of autophagosome biogenesis. J Cell Biol 2010; 190:1005–1022.
Yamamoto H, Kakuta S, Watanabe TM, et al. Atg9 vesicles are an important membrane source during early steps of autophagosome formation. J Cell Biol 2012; 198:219–233.
Reggiori F, Tucker KA, Stromhaug PE, Klionsky DJ . The Atg1-Atg13 complex regulates Atg9 and Atg23 retrieval transport from the pre-autophagosomal structure. Dev Cell 2004; 6:79–90.
Lu Q, Wu F, Zhang H . Aggrephagy: lessons from C. elegans. Biochem J 2013; 452:381–390.
Zhang YX, Yan LB, Zhou Z, et al. SEPA-1 mediates the specific recognition and degradation of P granule components by autophagy in C. elegans. Cell 2009; 136:308–321.
Kovács AL, Zhang H . Role of autophagy in Caenorhabditis elegans. FEBS Lett 2010; 584:1335–1341.
Tian Y, Li ZP, Hu WQ, et al. C. elegans screen identifies autophagy genes specific to multicellular organisms. Cell 2010; 141:1042–1055.
Lu Q, Yang PG, Huang XX, et al. The WD40 repeat PtdIns(3)P-binding protein EPG-6 regulates progression of omegasomes to autophagosomes. Dev Cell 2011; 21:343–357.
Tian E, Wang FX, Han JH, Zhang H . epg-1 functions in autophagy-regulated processes and may encode a highly divergent Atg13 homolog in C. elegans. Autophagy 2009; 5:608–615.
Yang PG, Zhang H . The coiled-coil domain protein EPG-8 plays an essential role in the autophagy pathway in C. elegans. Autophagy 2011; 7:159–165.
Liang QQ, Yang PG, Tian E, Han JH, Zhang H . The C. elegans ATG101 homolog EPG-9 directly interacts with EPG-1/Atg13 and is essential for autophagy. Autophagy 2012; 8:1426–1433.
Zhao HY, Zhao YG, Wang XW, et al. Mice deficient in Epg5 exhibit selective neuronal vulnerability to degeneration. J Cell Biol 2013; 200:731–741.
Wu F, Li YP, Wang F, Noda NN, Zhang H . Differential function of the two Atg4 homologues in the aggrephagy pathway in Caenorhabditis elegans. J Biol Chem 2012; 287:29457–29467.
Zhang H, Wu F, Wang XW, Du HW, Wang XC, Zhang H . The two C. elegans Atg16 homologs have partially redundant functions in the basal autophagy pathway. Autophagy 2013; 9: (in press).
Strome S . Specification of the germ line. WormBook 2005 Jul 28:1–10.
Lin L, Yang PG, Huang XX, Zhang H, Lu Q, Zhang H . The scaffold protein EPG-7 links cargo-receptor complexes with the autophagic assembly machinery. J Cell Biol 2013; 201:113–129.
Johansen T, Lamark T . Selective autophagy mediated by autophagic adapter proteins. Autophagy 2011; 7:279–296.
Li SH, Yang PG, Tian E, Zhang H . Arginine methylation modulates autophagic degradation of PGL granules in C. elegans. Mol Cell 2013; 52:421–433.
Shintani T, Huang WP, Stromhaug PE, Klionsky DJ . Mechanism of cargo selection in the cytoplasm-to-vacuole targeting pathway. Dev Cell 2002; 3:825–837.
Nair U, Klionsky DJ . Molecular mechanisms and regulation of specific and nonspecific autophagy pathways in yeast. J Biol Chem 2005; 280:41785–41788.
He C, Song H, Yorimitsu T, et al. Recruitment of Atg9 to the preautophagosomal structure by Atg11 is essential for selective autophagy in budding yeast. J Cell Biol 2006; 175:925–935.
Yorimitsu T, Klionsky DJ . Atg11 links cargo to the vesicle-forming machinery in the cytoplasm to vacuole targeting pathway. Mol Biol Cell 2005; 16:1593–1605.
Suzuki K, Kubota Y, Sekito T, Ohsumi Y . Hierarchy of Atg proteins in pre-autophagosomal structure organization. Genes Cells 2007; 12:209–218.
Farré JC, Manjithaya R, Mathewson RD, Subramani S . PpAtg30 tags peroxisomes for turnover by selective autophagy. Dev Cell 2008; 14:365–376.
Okamoto K, Kondo-Okamoto N, Ohsumi Y . Mitochondria-anchored receptor Atg32 mediates degradation of mitochondria via selective autophagy. Dev Cell 2009; 17:87–97.
Kanki T, Wang K, Cao Y, Baba M, Klionsky DJ . Atg32 is a mitochondrial protein that confers selectivity during mitophagy. Dev Cell 2009; 17:98–109.
Filimonenko M, Isakson P, Finley KD, et al. The selective macroautophagic degradation of aggregated proteins requires the PI3P-binding protein Alfy. Mol Cell 2010; 38:265–279.
Hara T, Takamura A, Kishi C, et al. FIP200, a ULK-interacting protein, is required for autophagosome formation in mammalian cells. J Cell Biol 2008; 181:497–510.
Jeong H, Then F, Melia TJ, et al. Acetylation targets mutant huntingtin to autophagosomes for degradation. Cell 2009; 137:60–72.
Matsumoto G, Wada K, Okuno M, Kurosawa M, Nukina N . Serine 403 phosphorylation of p62/SQSTM1 regulates selective autophagic clearance of ubiquitinated proteins. Mol Cell 2011, 44:279–289.
Aoki Y, Kanki T, Hirota Y, et al. Phosphorylation of Serine 114 on Atg32 mediates mitophagy. Mol Biol Cell 2011; 22:3206–3217.
DeLuca SZ, O'Farrell PH . Barriers to male transmission of mitochondrial DNA in sperm development. Dev Cell 2012; 22:660–668.
Luo SM, Ge ZJ, Wang ZW, et al. Unique insights into maternal mitochondrial inheritance in mice. Proc Natl Acad Sci USA 2013; 110:13038–13043.
Sato M, Sato K . Degradation of paternal mitochondria by fertilization-triggered autophagy in C. elegans embryos. Science 2011, 334:1141–1144.
Rawi SA, Louvet-Vallée S, Djeddi A, et al. Postfertilization autophagy of sperm organelles prevents paternal mitochondrial DNA transmission. Science 2011; 334:1144–1147.
Zhou QH, Li HM, Xue D . Elimination of paternal mitochondria through the lysosomal degradation pathway in C. elegans. Cell Res 2011; 21:1662–1669.
Breucker H, Schafer E, Holstein AF . Morphogenesis and fate of the residual body in human spermiogenesis. Cell Tissue Res 1985; 240:303–309.
Chemes H . The phagocytic function of Sertoli cells: a morphological, biochemical, and endocrinological study of lysosomes and acid phosphatase localization in the rat testis. Endocrinology 1986; 119:1673–1681.
L'Hernault SW . Spermatogenesis. WormBook 2006 Feb 20:1–14.
Ward S . Asymmetric localization of gene products during the development of C. elegans spermatozoa. In Gametogenesis and the Early Embryo 44th Symposium of the Society for Developmental Biology. 1986; 55–75.
Ward S, Hogan E, Nelson GA . The initiation of spermiogenesis in the nematode Caenorhabditis elegans. Dev Biol 1983; 98:70–79
Huang J, Wang HB, Chen YY, Wang XC, Zhang H . Residual body removal during spermatogenesis in C. elegans requires genes that mediate cell corpse clearance. Development 2012; 139:4613–4622.
Sanjuan MA, Dillon CP, Tait SW, et al. Toll-like receptor signalling in macrophages links the autophagy pathway to phagocytosis. Nature 2007; 450:1253–1257.
Martinez J, Almendinger J, Oberst A, et al. Microtubule-associated protein 1 light chain 3 alpha (LC3)-associated phagocytosis is required for the efficient clearance of dead cells. Proc Natl Acad Sci USA 2011; 108:17396–17401.
Florey O, Kim SE, Sandoval CP, Haynes CM, Overholtzer M . Autophagy machinery mediates macroendocytic processing and entotic cell death by targeting single membranes. Nat Cell Biol 2011; 13:1335–1343.
Qu XP, Zou ZJ, Sun QH, et al. Autophagy gene-dependent clearance of apoptotic cells during embryonic development. Cell 2007; 128:931–946.
Mellén MA, de la Rosa EJ, Boya P . The autophagic machinery is necessary for removal of cell corpses from the developing retinal neuroepithelium. Cell Death Differ 2008; 15:1279–1290.
Sulston J E, Schierenberg E, White JG, Thomson JN . The embryonic cell lineage of the nematode Caenorhabditis elegans. Dev Biol 1983; 100:64–119.
Zhou Z, Yu X . Phagosome maturation during the removal of apoptotic cells: receptors lead the way. Trends Cell Biol 2008; 18:474–485.
Kinchen JM, Ravichandran KS . Phagosome maturation: going through the acid test. Nat Rev Mol Cell Biol 2008; 9:781–795.
Lu Q, Zhang Y, Hu T, Guo P, Li W, Wang X. C. elegans Rab GTPase 2 is required for the degradation of apoptotic cells. Development 2008; 135:1069–1080.
Guo P, Hu T, Zhang J, Jiang S, Wang X . Sequential action of Caenorhabditis elegans Rab GTPases regulates phagolysosome formation during apoptotic cell degradation. Proc Natl Acad Sci USA 2010; 107:18016–18021.
Kinchen JM, Doukoumetzidis K, Almendinger J, et al. A pathway for phagosome maturation during engulfment of apoptotic cells. Nat Cell Biol 2008; 10:556–566.
Lu N, Shen Q, Mahoney TR, Neukomm LJ, Wang Y, Zhou Z . Two PI 3-kinases and one PI 3-phosphatase together establish the cyclic waves of phagosomal PtdIns(3)P critical for the degradation of apoptotic cells. PLoS Biol 2012; 10:e1001245.
Huang SY, Jia KL, Wang Y, Zhou Z, Levine B . Autophagy genes function in apoptotic cell corpse clearance during C. elegans embryonic development. Autophagy 2013; 9:138–149.
Cheng SY, Wu YW, Lu Q, Yan JC, Zhang H, Wang XC . Autophagy genes coordinate with the class II PI3 kinase PIKI-1 to regulate maturation of apoptotic cell-containing phagosomes in C. elegans. Autophagy 2013; 9: (in press).
Carthew RW, Sontheimer EJ . Origins and mechanisms of miRNAs and siRNAs. Cell 2009; 136:642–655.
Ding L, Han M . GW182 family proteins are crucial for microRNA-mediated gene silencing. Trends Cell Biol 2007; 17:411–416.
Ambros V . MicroRNAs and developmental timing. Curr Opin Genet Dev 2011; 21:511–517.
Slack FJ, Basson M, Liu Z, Ambros V, Horvitz HR, Ruvkun G . The lin-41 RBCC gene acts in the C. elegans heterochronic pathway between the let-7 regulatory RNA and the LIN-29 transcription factor. Mol Cell 2000; 5:659–669.
Abbott AL, Alvarez-Saavedra E, Miska EA, et al. The let-7 MicroRNA family members mir-48, mir-84 and mir-241 function together to regulate developmental timing in Caenorhabditis elegans. Dev Cell 2005; 3:403–414.
Li M, Jones-Rhoades MW, Lau NC, Bartel DP, Rougvie AE . Regulatory mutations of mir-48, a C. elegans let-7 family microRNA, cause developmental timing defects. Dev Cell 2005; 9:415–422.
Ding L, Spencer A, Morita K, Han M . The developmental timing regulator AIN-1 interacts with miRISCs and may target the argonaute protein ALG-1 to cytoplasmic P bodies in C. elegans. Mol Cell 2005; 19:437–447.
Zhang L, Ding L, Cheung TH, et al. Systematic identification of C. elegans miRISC proteins, miRNAs, and mRNA targets by their interactions with GW182 proteins AIN-1 and AIN-2. Mol Cell 2007; 28:598–613.
Zhang PP, Zhang H . Autophagy modulates miRNA-mediated gene silencing and selectively degrades AIN-1/GW182 in C. elegans. EMBO Rep 2013; 14:568–576.
Johnson SM, Grosshans H, Shingara J, et al. RAS is regulated by the let-7 microRNA family. Cell 2005; 120:635–647.
Sternberg PW, Han M . Genetics of RAS signaling in C. elegans. Trends Genet 1998; 14:466–472.
Johnston RJ, Hobert O . A microRNA controlling left/right neuronal asymmetry in Caenorhabditis elegans. Nature 2003; 426:845–849.
Gibbings D, Mostowy S, Jay F, Schwab Y, Cossar P, Voinnet O . Selective autophagy degrades DICER and AGO2 and regulates miRNA activity. Nat Cell Biol 2012; 14:1314–1321.
Derrien B, Baumbergera N, Schepetilnikova M, et al. Degradation of the antiviral component ARGONAUTE1 by the autophagy pathway. Proc Natl Acad Sci USA 2012; 109:15942–15946.
Kang C, You YJ, Avery L . Dual roles of autophagy in the survival of Caenorhabditis elegans during starvation. Genes Dev 2007; 21:2161–2171.
Meléndez A, Tallóczy Z, Seaman M, Eskelinen EL, Hall DH, Levine B . Autophagy genes are essential for dauer development and life-span extension in C. elegans. Science 2003; 301:1387–1391.
Farese RV Jr, Walther TC . Lipid droplets finally get a little R-E-S-P-E-C-T. Cell 2009; 139:855–860.
Ducharme NA, Bickel PE . Lipid droplets in lipogenesis and lipolysis. Endocrinology 2008; 149:942–949.
Singh R, Kaushik S, Wang YJ, et al. Autophagy regulates lipid metabolism. Nature 2009; 458:1131–1135.
Singh R, Xiang YQ, Wang YJ, et al. Autophagy regulates adipose mass and differentiation in mice. J Clin Invest 2009; 119:3329–3339.
Zhang Y, Goldman S, Baerga R, Zhao Y, Komatsu M, Jin SK . Adipose-specific deletion of autophagy-related gene 7 (atg7) in mice reveals a role in adipogenesis. Proc Natl Acad Sci USA 2009; 106:19860–19865.
Lapierre LR, Silvestrini MJ, Nuñez L, et al. Autophagy genes are required for normal lipid levels in C. elegans. Autophagy 2013; 9:278–286.
Lapierre LR, Gelino S, Meléndez A, Hansen M . Autophagy and lipid metabolism coordinately modulate life span in germline-less C. elegans. Curr Biol 2011; 21:1507–1514.
Wang MC, O'Rourke EJ, Ruvkun G . Fat metabolism links germline stem cells and longevity in C. elegans. Science 2008; 322:957–960.
O'Rourke EJ, Kuballa P, Xavier R, Ruvkun G . ω-6 Polyunsaturated fatty acids extend life span through the activation of autophagy. Genes Dev 2013; 27:429–440.
Hansen M, Chandra A, Mitic LL, Onken B, Driscoll M, Kenyon C . A role for autophagy in the extension of lifespan by dietary restriction in C. elegans. PLoS Genet 2008; 4:e24.
O'Rourke EJ, Ruvkun G . MXL-3 and HLH-30 transcriptionally link lipolysis and autophagy to nutrient availability. Nat Cell Biol 2013; 15:668–676.
Settembre C, Di Malta C, Polito VA, et al. TFEB links autophagy to lysosomal biogenesis. Science 2011; 332:1429–1433.
Lapierre LR, De Magalhaes Filho CD, McQuary PR, et al. The TFEB orthologue HLH-30 regulates autophagy and modulates longevity in Caenorhabditis elegans. Nat Commun 2013; 4:2267.
Settembre C, De Cegli R, Mansueto G, et al. TFEB controls cellular lipid metabolism through a starvation-induced autoregulatory loop. Nat Cell Biol 2013; 15:647–658.
Yuan J, Kroemer G . Alternative cell death mechanisms in development and beyond. Genes Dev 2010; 24:2592–2602.
Shimizu S, Kanaseki T, Mizushima N, et al. Role of Bcl-2 family proteins in a non-apoptotic programmed cell death dependent on autophagy genes. Nat Cell Biol 2004; 6:1221–1228.
Ryoo HD, Baehrecke EH . Distinct death mechanisms in Drosophila development. Curr Opin Cell Biol 2010; 22:889–895.
Martin DN, Baehrecke EH . Caspases function in autophagic programmed cell death in Drosophila. Development 2004; 131:275–284.
Berry DL, Baehrecke EH . Growth arrest and autophagy are required for salivary gland cell degradation in Drosophila. Cell 2007; 131:1137–1148.
Nezis IP, Lamark T, Velentzas AD, et al. Cell death during Drosophila melanogaster early oogenesis is mediated through autophagy. Autophagy 2009; 5:298–302.
Denton D, Shravage B, Simin R, et al. Autophagy, not apoptosis, is essential for midgut cell death in Drosophila. Curr Biol 2009; 19:1741–1746.
Conradt B, Xue D . Programmed cell death. WormBook 2005; Oct 6:1–13.
Gartner A, Boag PR, Blackwell TK . Germline survival and apoptosis. WormBook 2008; Sep 4:1–20.
Gumienny TL, Lambie E, Hartwieg E, Horvitz HR, Hengartner MO . Genetic control of programmed cell death in the Caenorhabditis elegans hermaphrodite germline. Development 1999; 126:1011–1022.
Wang HB, Lu Q, Cheng SY, Wang XC, Zhang H . Autophagy activity contributes to programmed cell death in Caenorhabditis elegans. Autophagy 2013; 9: (in press).
Reddien PW, Cameron S, Horvitz HR . Phagocytosis promotes programmed cell death in C. elegans. Nature 2001; 412:198–202.
Hoeppner DJ, Hengartner MO, Schnabel R . Engulfment genes cooperate with ced-3 to promote cell death in Caenorhabditis elegans. Nature 2001; 412:202–206.
Sulston JE, Horvitz HR . Post-embryonic cell lineages of the nematode, Caenorhabditis elegans. Dev Biol 1977; 56:110–156.
Acknowledgements
We are grateful to Dr Isabel Hanson for editing the work. The work in Dr Hong Zhang's lab was supported by the National Basic Research Program of China (2013CB910100, 2011CB910100), the National Natural Science Foundation of China (31225018) and an International Early Career Scientist grant from the Howard Hughes Medical Institute.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Yang, P., Zhang, H. You are what you eat: multifaceted functions of autophagy during C. elegans development. Cell Res 24, 80–91 (2014). https://doi.org/10.1038/cr.2013.154
Published:
Issue Date:
DOI: https://doi.org/10.1038/cr.2013.154
Keywords
This article is cited by
-
Autophagy is required for spermatogonial differentiation in the Drosophila testis
Biologia Futura (2022)
-
Regulatory role of Wdr24 in autophagy activity during zebrafish embryogenesis
Molecular & Cellular Toxicology (2019)
-
Transcriptomic analysis of the autophagy machinery in crustaceans
BMC Genomics (2016)
-
Autophagy and regulation of cilia function and assembly
Cell Death & Differentiation (2015)
-
Current aging research in China
Protein & Cell (2015)