Abstract
Background:
Some childhood acute lymphoblastic leukaemias (ALL) can be traced back to a prenatal origin, where a virus infection could be involved in the first pre-leukaemic clone development. The DNA virome of 95 children who later developed ALL was characterised from neonatal blood spots (NBS) using unbiased next-generation sequencing (NGS) and compared with the virome of 95 non-ALL controls.
Methods:
DNA was individually extracted from the ALL-patients and controls, pooled, randomly amplified and sequenced using the Illumina MiSeq Sequencing System.
Results:
Virus-like sequences identified in both groups mapped to human endogenous retroviruses and propionibacterium phage, considered a part of the normal microbial flora. Potential pathogens human herpesvirus type 6 (HHV-6) and parvovirus B19 were also identified, but only few samples in both ALL and controls tested positive by PCR follow-up.
Conclusions:
Unbiased NGS was employed to search for DNA from potential infectious agents in neonatal samples of children who later developed ALL. Although several viral candidates were identified in the NBS samples, further investigation by PCR suggested that these viruses did not have a major role in ALL development.
Similar content being viewed by others
Main
The etiology of childhood leukaemia is unclear in 95% of all cases. The development of some acute lymphoblastic leukaemias (ALL) can be traced back to a prenatal origin, starting in utero with a pre-leukaemic clone, followed by the acquisition of postnatal genomic losses or gains originated from the prenatal cytogenetic aberration. The evidence of an in utero first event is documented by molecular studies from neonatal blood spots (NBS), cord blood, twin studies and space-time clustering data (Gustafsson and Carstensen, 2000; Greaves et al, 2003; Greaves 2005, 2006a, b, 2009; Bateman et al, 2010). Many leukaemic clones found at time of diagnosis in ALL can also be detected in NBS, such as ETV6-RUNX1 fusion, hyper diploid ALLs, infant MLL-rearrangements, Phil+ALLs and rare forms of infant T-cell ALLs (Wiemels et al, 2002; Maia et al, 2004; Eguchi-Ishimae et al, 2008; Bateman et al, 2010; Cazzaniga et al, 2011; Mansur et al, 2011, 2015; Alpar et al, 2014). Concerning the MLL gene fusions, there is evidence of a transplacental chemical carcinogenesis (Greaves, 2005). However, the cause of cytogenic aberrations in the major part of leukaemias still remains unknown.
Several human tumour DNA viruses are known to be involved in the development of a malignant clone (Lindblom et al, 2005; Weitzman and Ornelles, 2005; Hudnall et al, 2008; Chang and Moore, 2012; Chabay and Preciado, 2013). These viruses can persist in the lymphoid cells and suppress double-stranded DNA-break repair, also known as DNA damage response (DDR). As the DDR protects the genome from the accumulation of deleterious mutations, down-regulation is associated with an increased risk of clonal development. This suggests that a virus infection may generate the aberrant clones of lymphocytes that precede development of ALL (Boggs, 1999; Hollingworth and Grand, 2015; Ornelles et al, 2015).
Although epidemiological evidence proposes that ALL may be initiated by in utero infection with a common pathogen, the identification of such a pathogen has remained elusive (Gustafsson and Carstensen, 1999; Wiemels et al, 1999; Canfield et al, 2004; Isa et al, 2004). Unlike viral-specific methods, unbiased next-generation sequencing (NGS) can provide a holistic picture of the virome of ALL patients, thus facilitating the identification of viral candidates that might lead to ALL development. The use of unbiased NGS in the study of the human virome has been widespread during the last decade, aiding in the detection of known and unknown viruses from both isolated cases and from major disease outbreaks (Cox-Foster et al, 2007; Palacios et al, 2008; Towner et al, 2008; Nakamura et al, 2009; Schowalter et al, 2010; de Vries et al, 2011; Yozwiak et al, 2012).
The aim of this study is to identify viral candidates that could be involved in the first step of leukaemogenesis by investigating whether in utero infections of DNA viruses are more common in newborns who later developed ALL. To accomplish this, unbiased NGS was used to characterise and compare the DNA virome at the time of birth from ALL and non-ALL children.
Materials and methods
Patients
Children who later developed ALL were randomly identified from the Nordic Society of Paediatric Haematology and Oncology register and linked to the Swedish Medical Birth register to gain access to the personal codes of mothers and children necessary for NBS identification. Controls, matching for age and birthplace were also collected for further comparisons. In total, NBS from 95 children diagnosed with ALL between 1992–2006 and 95 non-ALL children were analysed. The patient population included 39 girls and 56 boys with a median age at time of diagnosis of 5 years (5 months–17 years). Eighty-five patients were diagnosed with a B-cell leukaemia, 9 children with a T-cell-origin and one patient was diagnosed with an undifferentiated lineage. Cytogenetic characteristics at diagnosis corresponded to 25 children with hyperdiploidy, 2 patients with a hypodiploidy, 8 children were diagnosed with t(12;21), 2 with t(1;14), 2 children with t(4;11) (MLL-rearrangements) and 3 children were positive for Philadelphia chromosomes t(9;22). One child was positive for both MLL and Philadelphia chromosomes. Overall, 20 children were diagnosed with other cytogenetic changes, 20 children were diagnosed with no chromosomal changes and no cytogenetic analysis was performed in 39 diagnosed very early. This study was approved by The Regional Ethical Review Board, Stockholm, Sweden.
Sample collection
NBS were collected at 2–5 days of age for screening for several inborn metabolic diseases. They corresponded to four droplets of capillary blood containing ∼3 × 104 nucleated cells/spot blotted into a filter paper (Guthrie card), which are later dried and stored at 4 °C. The median storage time for the NBS was 9.9 years (2.6–22.8).
DNA isolation
DNA was extracted using the minimal essential medium (Barbi et al, 1996; Yamagishi et al, 2006) after punching four uniform discs of 3 mm in diameter from one of four blood spots. Extracted DNA was individually tested for the human albumin gene (ALB) using TagMan Real-Time quantitative PCR (Laurendeau et al, 1999) to ensure the availability of DNA for subsequent DNA amplification and sequencing.
Unbiased NGS
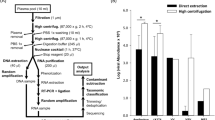
Extracted DNA from the ALL patients and controls was independently pooled. DNA from both pools was randomly amplified with the illustra GenomiPhi V2 DNA Amplification Kit and sequenced using the Illumina MiSeq Sequencing System at Science for Life Laboratory, Stockholm, Sweden. A computational pipeline estimated then the composition of the viral community from resulting sequencing data. The pipeline trimmed adapters and low-quality bases and discarded human sequences by a mapping approach. Multiple assembly strategies were performed on the remaining reads. Taxonomical annotation of contigs and unassembled reads was performed with BLASTN and BLASTX (Altschul et al, 1990) with E value 10−3 to frozen versions of the NCBI databases nt and nr, respectively, applying the lowest common ancestor algorithm in MEGAN5 (Huson et al, 2007). Moreover, further analyses were conducted in sequences that could not be classified with MEGAN. In here, (i) contigs and merged paired-end reads longer than 400 bp were selected, (ii) sequences were translated to proteins in all six open-reading frames, and (iii) virus-like motifs were searched using hmmscan from the HMMER3 suite (Eddy, 2011) against the Pfam-A v28.0 (Finn et al, 2014) and vFam-A 2014 (Skewes-Cox et al, 2014) protein family databases. Viral candidates identified by the pipeline with potential associations to ALL were later investigated using agent-specific PCR.
Results
All samples were positive for the ALB gene after DNA extraction, ensuring thus the presence of genetic material and excluding PCR inhibitors. The MiSeq Sequencing System generated approximately seven million paired-end reads from each pool. About 25% of the sequences were kept after quality filtering and 95% of remaining reads mapped to the human genome. After filtering out human sequences, 5,112 and 3,259 reads kept from ALL and controls, respectively, which were later assembled into contigs. Sequencing results are summarised in Table 1. Unassembled reads and contigs were then assigned a taxonomical classification as described in the ‘Material and methods’ section.
Analysis of the libraries yielded two candidates potentially involved in leukemogenesis. They corresponded to sequence hits detected only in the ALL library, assigned to HHV-6 and human parvovirus B19. Viral assignments of unassembled reads and contigs with BLASTN and BLASTX, and visualisation of virome datasets are shown in Table 2 and Figure 1, respectively. To further investigate a putative role of HHV-6 and parvovirus B19 in ALL development, the presence of the viruses in the original samples was determined by real-time PCR.
Visualisation of virome datasets by MEGAN. Taxonomical annotation of contigs and unassembled reads with BLASTN and BLASTX with E value 10−3 to frozen versions of the NCBI databases nt and nr, respectively, applying the lowest common ancestor algorithm in MEGAN5 (Huson et al, 2007). (A) ALL BLASTN, (B) ALL BLASTX, (C) non-ALL BLASTN, (D) non-ALL BLASTX.
Besides HHV-6 and human parvovirus 6, assignments to human endogenous retroviruses (HERV) and propionibacterium phage (PP) were detected in both libraries. HERVs are considered remnants of ancient retroviral infections, which results in around 8% of the human genome with high similarity to HERV (Belshaw et al, 2004; Nelson et al, 2004). Although HERV sequences should be depleted during human filtering step in the bioinformatics pipeline, the genomic variability of these regions with respect to the human genome reference sequence used in the mapping approach might facilitate HERVs displaying after host depletion. HERV assignments using BLASTN and BLASTX in ALL and controls are shown in Table 3. In particular, 15 sequences were assigned to HERV with BLASTX in ALL whereas 4 and 1 sequences mapped with BLASTX and BLASTN in controls, respectively. Although HERVs are generally considered to be harmless, replicatively active HERVs have been associated with carcinogenesis (Brodsky et al, 1993a, 1993b; Yin et al, 1997; Depil et al, 2002). Hence, the HERV strains found by NGS in ALL were further assessed by PCR. The same HERV composition was found in both ALL and controls, suggesting that there is no association between HERV and ALL in these samples. The other viral finding, PP is considered part of the normal human microbial flora and a likely contaminant from skin contact. One sequence was assigned to PP with BLASTX in ALL whereas 2 and 3 sequences were also assigned with BLASTX and BLASTN, respectively, in controls. No follow-up PCR was performed for PP, as it was detected in both groups and is very unlikely to be associated to disease.
Regarding additional analyses carried out in sequences not classified with MEGAN, one viral-like motif hit was found in ALL. The hit displayed was a 24 amino acid stretch located in one of the contigs mapping to a nucleopolyhedrovirus protein (vFam_71, E value 2.2e−6). Nucleopolyhedrovirus is a genus of the family Baculoviridae, primarily pathogenic for insects and not capable of replicating in vertebrate cells (Summers, 1975). Owing to the lack of pathogenicity of nucleopolyhedrovirus shown in humans together with the difficulty of oligonucleotide design because of the limited sequence information obtained by NGS, we decided not to follow this finding up using PCR.
HHV-6 and parvovirus B19 determinations by qPCR
Extracted DNA from each ALL and control samples previously analysed by NGS was subjected to Real-Time qualitative PCR for HHV-6 and parvovirus B19 to explore their possible association with the development of ALL.
HHV-6 can be chromosomally integrated (ciHHV-6) into the human genome (Pellett et al, 2012) and transmitted to the descendent after germ-line virus infection (Daibata et al, 1998). Then, the determination of an in utero HHV-6 infection by qPCR might be disturbed by the presence of vertically transmitted ciHHV-6 unless a clear difference between the number of positive PCR reactions in ALL and controls was observed. HHV-6 was only detected in 2 out 94 ALL patients (one patient sample with insufficient material) and 3 out of 95 controls by PCR (Collot et al, 2002). Interestingly, the fact that HHV-6 was found in the control group by qPCR but not by NGS could be attributed to the very low copy numbers of HHV-6 viral genomes in the samples (free or integrated viral particles) and/or high-host content masking the presence of viruses during unbiased NGS. Regarding parvovirus B19, viral determinations were carried out as in (Broliden et al, 1998). The finding of parvovirus was confirmed only in 1 out 94 patient samples and in none of the 95 controls. According to results obtained, HHV-6 and parvovirus B19 in utero infection did not have a major role in ALL development.
Discussion
The role of viral infections during pregnancy in the development of childhood ALL has been investigated in several studies but no specific virus has been defined (Smith, 1997; Eden, 2010). Our group has previously examined the presence of adenoviruses, herpesviruses (cytomegalovirus, Epstein–Barr virus and HHV-6), polyomaviruses and parvovirus B19 by PCR from NBS in children who later in life developed ALL. So far, only adenovirus was identified in the NBS (Priftakis et al, 2003; Isa et al, 2004; Gustafsson et al, 2006, 2007, 2012; Gustafsson and Bogdanovic, 2007) where an increased frequency of adenoviral DNA in ALL versus non-ALL controls was observed. However, this trend could not be replicated in an extended study (Gustafsson et al, 2012).
Methods used in previous studies for pathogen identification enabled us to detect only sequences from the chosen target of different viruses defined by the primers. In this study, unbiased NGS was applied to search for viral candidates in NBS from newborns who later developed ALL. We opted to investigate only the DNA virome because of the nature of the starting material. The available samples consisted of extracted DNA from NBS used in previous studies, where we did not expect sufficient amounts of intact RNA. This technology facilitated the screening of all potential pathogens simultaneously and was capable of inspecting the virome in NBS, which contained low numbers of free viral particles, bacteriophages, intracellular viruses and proviruses together with a large number of white blood cells. Among the commonly detected viruses, the presence of HERV in both patient and control group is likely explained by the detection of integrated retroviral genomes in the human genome, favoured by the high host background in the samples. Similarly, finding of bacteriophage sequences in both groups corresponds to a part of microbial flora in human material and as such was not considered as a potential pathogen.
HHV-6 and parvovirus B19, both ubiquitous pathogens (Cohen and Buckley, 1988; Lamont et al, 2011), were the only viruses identified in ALL group and, therefore, further investigated. HHV-6 is a double stranded DNA virus with observed oncogenic potential in vitro (Razzaque, 1990). A unique feature among the human herpes viruses is the integration of whole HHV-6 genome into one of human chromosomes and germ-line transmission (Pellett et al, 2012), being detected in 1% of the general population (Pellett et al, 2012). The frequency of ciHHV-6 in children with ALL has been reported to be similar to that of the general population (Gravel et al, 2013). The detection of HHV-6 in both patients and controls by PCR could be associated with either active infection or detection of integrated viral genomes, which complicates the assessment of the role of viral infection in leukaemias. However, and according to negative reports obtained in our previous studies (Bogdanovic et al, 2004), the fact that HHV-6 was detected in both groups and in few patients suggests that it does not have a major role in ALL development.
The role of parvovirus B19 in the pathogenesis of leukaemias has been speculated about, but not been broadly examined. In children with ALL, parvovirus infection is associated with serious complications such as cytopenia (Lindblom et al, 2005). Approximately 30–50% of pregnant women are seronegative for parvovirus B19 and vertical transmission is possible following maternal infection in pregnancy (Lamont et al, 2011). In this study parvovirus B19 was identified in one patient and none of the controls and together with the negative results in our previous study, no evidence is provided to support a putative association between parvovirus B19 and ALL. It has been shown that parvovirus B19 DNA remains detectable in human tissues after past infection (Söderlund-Venermo et al, 2002), but it is difficult to conclude whether any of the studied children actually were infected in utero based on the present findings.
All HERV strains detected by NGS in ALL were also found in controls. Although none of the strains tested seemed to be predominant in patients with ALL, the involvement of these mobile genetic elements cannot be fully discarded. Further studies are needed to investigate whether polymorphisms, recombination or transposition to oncogenes or tumour suppressor genes could possibly lead to genetic instability involved in the causation of ALL.
Torque teno virus (TTV), a single-stranded DNA virus, has been frequently detected in human samples (Okamoto, 1999, 2009) with prevalence in serum of healthy individuals of up to 90%. Moreover, TTV has been also detected during pregnancy in serum from mothers of children who later developed leukaemias (Bzhalava et al, 2012), and its transmission during pregnancy has been previously suggested in several studies (Gerner et al, 2000; Schröter et al, 2000). In our study no TTV-like sequences were identified in NBS from ALL children or controls indicating that primary infection with TTV may occur postnatally.
An important limitation of this study was the small amounts of starting DNA and high proportion of human DNA relative to viruses in the blood spots. Both factors limit the sensitivity of NGS for the detection of viruses. Despite these limitations, our approach was capable of detecting parvovirus (5.5 kb) in a pool of 95 samples containing high host background. After individual qPCR inspection only one sample was positive (Ct>27). Nonetheless, caution is needed when interpreting the significance of the negative findings.
In conclusion, unbiased NGS was employed for search for potential DNA infectious agents in neonatal samples of children who later developed ALL. Here, NGS was capable of identifying viral candidates in NBS from ALL-patients characterised by containing high host background. However, further investigation by PCR suggested that viruses reported did not have a major role in ALL development.
Change history
11 October 2016
This paper was modified 12 months after initial publication to switch to Creative Commons licence terms, as noted at publication
References
Alpar D, Wren D, Ermini L, Mansur MB, van Delft FW, Bateman CM, Titley I, Kearney L, Szczepanski T, de Castro DG, Ford A M, Potter NE, Greaves M (2014) Clonal origins of ETV6-RUNX1(+) acute lymphoblastic leukemia: studies in monozygotic twins. Leukemia 29: 1–34.
Altschul SF, Gish W, Miller W, Myers EW, Lipman DJ (1990) Basic local alignment search tool. J Mol Biol 215: 403–410.
Barbi M, Binda S, Primache V, Novelli C (1996) Cytomegalovirus in peripheral blood leukocytes of infants with congenital or postnatal infection. Pediatr Infect Dis J 15: 898–903.
Bateman CM, Colman SM, Chaplin T, Young BD, Eden TO, Bhakta M, Gratias EJ, Van Wering ER, Cazzaniga G, Harrison CJ, Hain R, Ancliff P, Ford AM, Kearney L, Greaves M (2010) Acquisition of genome-wide copy number alterations in monozygotic twins with acute lymphoblastic leukemia. Blood 115: 3553–3558.
Belshaw R, Pereira V, Katzourakis A, Talbot G, Paces J, Burt A, Tristem M (2004) Long-term reinfection of the human genome by endogenous retroviruses. Proc Natl Acad Sci USA 101: 4894–4899.
Bogdanovic G, Jernberg AG, Priftakis P, Grillner L, Gustafsson B (2004) Human herpes virus 6 or Epstein–Barr virus were not detected in Guthrie cards from children who later developed leukaemia. Br J Cancer 91: 913–915.
Boggs S (1999) The hematopoietic microenvironment: phylogeny and ontogeny of the hematopoietic microenvironment. Hematology 4 (1): 31–44.
Brodsky I, Foley B, Gillespie D (1993a) Expression of human endogenous retrovirus (HERV-K) in chronic myeloid leukemia. Leuk Lymphoma 11 (Suppl 1): 119–123.
Brodsky I, Foley B, Haines D, Johnston J, Cuddy K, Gillespie D (1993b) Expression of HERV-K proviruses in human leukocytes. Blood 81: 2369–2374.
Broliden K, Tolfvenstam T, Ohlsson S, Henter JI (1998) Persistent B19 parvovirus infection in pediatric malignancies. Med Pediatr Oncol 31: 66–72.
Bzhalava D, Ekström J, Lysholm F, Hultin E, Faust H, Persson B, Lehtinen M, de Villiers EM, Dillner J (2012) Phylogenetically diverse TT virus viremia among pregnant women. Virology 432: 427–434.
Canfield KN, Spector LG, Robison LL, Lazovich D, Roesler M, Olshan AF, Smith FO, Heerema NA, Barnard DR, Blair CK, Ross JA (2004) Childhood and maternal infections and risk of acute leukaemia in children with Down syndrome: a report from the Children’s Oncology Group. Br J Cancer 91: 1866–1872.
Cazzaniga G, Van Delft FW, Lo Nigro L, Ford AM, Score J, Iacobucci I, Mirabile E, Taj M, Colman SM, Biondi A, Greaves M (2011) Developmental origins and impact of BCR-ABL1 fusion and IKZF1 deletions in monozygotic twins with Ph+acute lymphoblastic leukemia. Blood 118: 5559–5564.
Chabay PA, Preciado M V. (2013) EBV primary infection in childhood and its relation to B-cell lymphoma development: a mini-review from a developing region. Int J Cancer 133: 1286–1292.
Chang Y, Moore PS (2012) Merkel cell carcinoma: a virus-induced human cancer. Annu Rev Pathol 7: 123–144.
Cohen BJ, Buckley MM (1988) The prevalence of antibody to human parvovirus B19 in England and Wales. J Med Microbiol 25: 151–153.
Collot S, Petit B, Bordessoule D, Alain S, Touati M, Denis F, Ranger-Rogez S (2002) Real-time PCR for quantification of human herpesvirus 6 DNA from lymph nodes and saliva. J Clin Microbiol 40: 2445–2451.
Cox-Foster DL, Conlan S, Holmes EC, Palacios G, Evans JD, Moran NA, Quan P-L, Briese T, Hornig M, Geiser DM, Martinson V, VanEngelsdorp D, Kalkstein AL, Drysdale A, Hui J, Zhai J, Cui L, Hutchison SK, Simons JF, Egholm M, Pettis JS, Lipkin WI (2007) A metagenomic survey of microbes in honey bee colony collapse disorder. Science 318: 283–287.
Daibata M, Taguchi T, Sawada T, Taguchi H, Miyoshi I (1998) Chromosomal transmission of human herpesvirus 6 DNA in acute lymphoblastic leukaemia. Lancet 352: 543–544.
Depil S, Roche C, Dussart P, Prin L (2002) Expression of a human endogenous retrovirus, HERV-K, in the blood cells of leukemia patients. Leukemia 16: 254–259.
Eddy SR (2011) Accelerated profile HMM searches. PLoS Comput Biol 7: e1002195.
Eden T (2010) Aetiology of childhood leukaemia. Cancer Treat Rev 36: 286–297.
Eguchi-Ishimae M, Eguchi M, Kempski H, Greaves M (2008) NOTCH1 mutation can be an early, prenatal genetic event in T-ALL. Blood 111: 376–378.
Finn RD, Bateman A, Clements J, Coggill P, Eberhardt RY, Eddy SR, Heger A, Hetherington K, Holm L, Mistry J, Sonnhammer ELL, Tate J, Punta M (2014) Pfam: The protein families database. Nucleic Acids Res 42: D222–D230.
Gerner P, Oettinger R, Gerner W, Falbrede J, Wirth S (2000) Mother-to-infant transmission of TT virus: prevalence, extent and mechanism of vertical transmission. Pediatr Infect Dis J 19: 1074–1077.
Gravel A, Sinnett D, Flamand L (2013) Frequency of chromosomally-integrated human herpesvirus 6 in children with acute lymphoblastic leukemia. PLoS One 8: e84322.
Greaves M (2005) In utero origins of childhood leukaemia. Early Hum Dev 81: 123–129.
Greaves M (2006a) Infection, immune responses and the aetiology of childhood leukaemia. Nat Rev Cancer 6: 193–203.
Greaves M (2009) Darwin and evolutionary tales in leukemia. The Ham–Wasserman Lecture. Hematology Am Soc Hematol Educ Program 3–12; doi:10.1182/asheducation-2009.1.3.
Greaves MF (2006b) Cord blood donor cell leukemia in recipients. Leukemia 20: 1633–1634.
Greaves MF, Maia AT, Wiemels JL, Ford AM (2003) Leukemia in twins: lessons in natural history. Blood 102: 2321–2333.
Gustafsson B, Bogdanovic G (2007) Specific viruses were not detected in Guthrie cards from children who later developed leukemia. Pediatr Hematol Oncol 24: 607–613.
Gustafsson B, Carstensen J (1999) Evidence of space-time clustering of childhood acute lymphoblastic leukaemia in Sweden. Br J Cancer 79: 655–657.
Gustafsson B, Carstensen J (2000) Space-time clustering of childhood lymphatic leukaemias and non-Hodgkin’s lymphomas in Sweden. Eur J Epidemiol 16: 1111–1116.
Gustafsson B, Honkaniemi E, Goh S, Giraud G, Forestier E, von Dobeln U, Allander T, Dalianis T, Bogdanovic G (2012) KI, WU, and Merkel cell polyomavirus DNA was not detected in guthrie cards of children who later developed acute lymphoblastic leukemia. J Pediatr Hematol Oncol 34: 364–367.
Gustafsson B, Huang W, Bogdanovic G, Gauffin F, Nordgren A, Talekar G, Ornelles D A, Gooding LR (2007) Adenovirus DNA is detected at increased frequency in Guthrie cards from children who develop acute lymphoblastic leukaemia. Br J Cancer 97: 992–994.
Gustafsson B, Jernberg AG, Priftakis P, Bogdanovic G (2006) No CMV DNA in Guthrie cards from children who later developed ALL. Pediatr Hematol Oncol 23: 199–205.
Hollingworth R, Grand RJ (2015) Modulation of DNA damage and repair pathways by human tumour viruses. Viruses 7: 2542–2591.
Hudnall SD, Chen T, Amr S, Young KH, Henry K (2008) Detection of human herpesvirus DNA in Kikuchi–Fujimoto disease and reactive lymphoid hyperplasia. Int J Clin Exp Pathol 1: 362–368.
Huson DH, Auch AF, Qi J, Schuster SC (2007) MEGAN analysis of metagenomic data. Genome Res 17: 377–386.
Isa A, Priftakis P, Broliden K, Gustafsson B (2004) Human parvovirus B19 DNA is not detected in guthrie cards from children who have developed acute lymphoblastic leukemia. Pediatr Blood Cancer 42: 357–360.
Lamont RF, Sobel JD, Vaisbuch E, Kusanovic JP, Mazaki-Tovi S, Kim SK, Uldbjerg N, Romero R (2011) Parvovirus B19 infection in human pregnancy. BJOG 118: 175–186.
Laurendeau I, Bahuau M, Vodovar N, Larramendy C, Olivi M, Bieche I, Vidaud M, Vidaud D (1999) TaqMan PCR-based gene dosage assay for predictive testing in individuals from a cancer family with INK4 locus haploinsufficiency. Clin Chem 45: 982–986.
Lindblom A, Isa A, Norbeck O, Wolf S, Johansson B, Broliden K, Tolfvenstam T (2005) Slow clearance of human parvovirus B19 viremia following acute infection. Clin Infect Dis 41: 1201–1203.
Maia AT, Tussiwand R, Cazzaniga G, Rebulla P, Colman S, Biondi A, Greaves M (2004) Identification of preleukemic precursors of hyperdiploid acute lymphoblastic leukemia in cord blood. Genes Chromosomes Cancer 40: 38–43.
Mansur MB, van Delft FW, Colman SM, Furness CL, Gibson J, Emerenciano M, Kempski H, Clappier E, Cave H, Soulier J, Pombo-de-Oliveira MS, Greaves M, Ford AM (2015) Distinctive genotypes in infants with T-cell acute lymphoblastic leukaemia. Br J Haematol 171: 574–584.
Mansur MB, Ford A M, van Delft FW, Gonzalez D, Emerenciano M, Maia RC, Greaves M, Pombo-de-Oliveira MS (2011) Occurrence of identical NOTCH1 mutation in non-twinned sisters with T-cell acute lymphoblastic leukemia. Leuk Off J Leuk Soc Am Leuk Res Fund, UK 25: 1368–1370.
Nakamura S, Yang CS, Sakon N, Ueda M, Tougan T, Yamashita A, Goto N, Takahashi K, Yasunaga T, Ikuta K, Mizutani T, Okamoto Y, Tagami M, Morita R, Maeda N, Kawai J, Hayashizaki Y, Nagai Y, Horii T, Iida T, Nakaya T (2009) Direct metagenomic detection of viral pathogens in nasal and fecal specimens using an unbiased high-throughput sequencing approach. PLoS One 4: e4219.
Nelson PN, Hooley P, Roden D, Ejtehadi HD, Rylance P, Warren P, Martin J, Murray PG (2004) Human endogenous retroviruses: transposable elements with potential? Clin Exp Immunol 138: 1–9.
Okamoto H (1999) Molecular virology of TT virus. Nihon Rinsho 57: 1239–1249.
Okamoto H (2009) History of discoveries and pathogenicity of TT viruses. Curr Top Microbiol Immunol 331: 1–20.
Ornelles DA, Gooding LR, Garnett-Benson C (2015) Neonatal infection with species C adenoviruses confirmed in viable cord blood lymphocytes. PLoS One 10: e0119256.
Palacios G, Druce J, Du L, Tran T, Birch C, Briese T, Conlan S, Quan P-L, Hui J, Marshall J, Simons JF, Egholm M, Paddock CD, Shieh W-J, Goldsmith CS, Zaki SR, Catton M, Lipkin WI (2008) A new arenavirus in a cluster of fatal transplant-associated diseases. N Engl J Med 358: 991–998.
Pellett PE, Ablashi DV, Ambros PF, Agut H, Caserta MT, Descamps V, Flamand L, Gautheret-Dejean A, Hall CB, Kamble RT, Kuehl U, Lassner D, Lautenschlager I, Loomis KS, Luppi M, Lusso P, Medveczky PG, Montoya JG, Mori Y, Ogata M, Pritchett JC, Rogez S, Seto E, Ward KN, Yoshikawa T, Razonable RR (2012) Chromosomally integrated human herpesvirus 6: questions and answers. Rev Med Virol 22: 144–155.
Priftakis P, Dalianis T, Carstensen J, Samuelsson U, Lewensohn-Fuchs I, Bogdanovic G, Winiarski J, Gustafsson B (2003) Human polyomavirus DNA is not detected in Guthrie cards (dried blood spots) from children who developed acute lymphoblastic leukemia. Med Pediatr Oncol 40: 219–223.
Razzaque A (1990) Oncogenic potential of human herpesvirus-6 DNA. Oncogene 5: 1365–1370.
Schowalter RM, Pastrana DV, Pumphrey KA, Moyer AL, Buck CB (2010) Merkel cell polyomavirus and two previously unknown polyomaviruses are chronically shed from human skin. Cell Host Microbe 7: 509–515.
Schröter M, Polywka S, Zöllner B, Schäfer P, Laufs R, Feucht HH (2000) Detection of TT virus DNA and GB virus type C/hepatitis G virus RNA in serum and breast milk: determination of mother-to-child transmission. J Clin Microbiol 38: 745–747.
Skewes-Cox P, Sharpton TJ, Pollard KS, DeRisi JL (2014) Profile hidden Markov models for the detection of viruses within metagenomic sequence data. PLoS One 9: e105067.
Smith M (1997) Considerations on a possible viral etiology for B-precursor acute lymphoblastic leukemia of childhood. J Immunother 20: 89–100.
Söderlund-Venermo M, Hokynar K, Nieminen J, Rautakorpi H, Hedman K (2002) Persistence of human parvovirus B19 in human tissues. Pathol Biol (Paris) 50: 307–316.
Summers M (1975) Baculoviruses for insect pest control: safety considerations. Am Soc Microbiol, pp 186.
Towner JS, Sealy TK, Khristova ML, Albariño CG, Conlan S, Reeder SA, Quan PL, Lipkin WI, Downing R, Tappero JW, Okware S, Lutwama J, Bakamutumaho B, Kayiwa J, Comer JA, Rollin PE, Ksiazek TG, Nichol ST (2008) Newly discovered Ebola virus associated with hemorrhagic fever outbreak in Uganda. PLoS Pathog 4: e1000212.
de Vries M, Deijs M, Canuti M, van Schaik BDC, Faria NR, van de Garde MDB, Jachimowski LCM, Jebbink MF, Jakobs M, Luyf ACM, Coenjaerts FEJ, Claas ECJ, Molenkamp R, Koekkoek SM, Lammens C, Leus F, Goossens H, Ieven M, Baas F, van der Hoek L (2011) A sensitive assay for virus discovery in respiratory clinical samples. PLoS One 6: e16118.
Weitzman MD, Ornelles DA (2005) Inactivating intracellular antiviral responses during adenovirus infection. Oncogene 24: 7686–7696.
Wiemels JL, Cazzaniga G, Daniotti M, Eden OB, Addison GM, Masera G, Saha V, Biondi A, Greaves MF (1999) Prenatal origin of acute lymphoblastic leukaemia in children. Lancet 354: 1499–1503.
Wiemels JL, Xiao Z, Buffler PA, Maia AT, Ma X, Dicks BM, Smith MT, Zhang L, Feusner J, Wiencke J, Pritchard-Jones K, Kempski H, Greaves M (2002) In utero origin of t(8;21) AML1-ETO translocations in childhood acute myeloid leukemia. Blood 99: 3801–3805.
Yamagishi Y, Miyagawa H, Wada K, Matsumoto S, Arahori H, Tamura A, Taniguchi H, Kanekiyo T, Sashihara J, Yoda T, Kitagawa M, Ozono K (2006) CMV DNA detection in dried blood spots for diagnosing congenital CMV infection in Japan. J Med Virol 78: 923–925.
Yin H, Medstrand P, Andersson ML, Borg A, Olsson H, Blomberg J (1997) Transcription of human endogenous retroviral sequences related to mouse mammary tumor virus in human breast and placenta: similar pattern in most malignant and nonmalignant breast tissues. AIDS Res Hum Retroviruses 13: 507–516.
Yozwiak NL, Skewes-Cox P, Stenglein MD, Balmaseda A, Harris E, DeRisi JL (2012) Virus identification in unknown tropical febrile illness cases using deep sequencing. PLoS Negl Trop Dis 6: e1485.
Acknowledgements
Sources of support were given by the Swedish Childhood Cancer Foundation and ALF for this study.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
This work is published under the standard license to publish agreement. After 12 months the work will become freely available and the license terms will switch to a Creative Commons Attribution-NonCommercial-Share Alike 4.0 Unported License.
Author contributions
Britt Gustafsson designed and financed the original study including collecting Guthrie cards, clinical data from the entire leukaemia group and collected Guthrie cards for the healthy control from the Swedish Birth Registry. She was mainly responsible for writing the manuscript. Gordana Bogdanovic extracted DNA from the original study, using the MEM method, reassured the DNA availability in the Guthrie cards. She was responsible for storing the material. She co-designed this study, using the latest technique NGS, together with Tobias Allander and Annelie Bjerkner. She generated and analysed the experimental data and co-wrote the paper. Christian Pou, Mauricio Barrientos-Somarribas and Björn Andersson generated and analysed next-generation sequencing experimental data, agent specific PCRs and co-wrote the manuscript. All authors critically reviewed and approved the final draft of the manuscript.
Rights and permissions
From twelve months after its original publication, this work is licensed under the Creative Commons Attribution-NonCommercial-Share Alike 4.0 Unported License. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-sa/4.0/
About this article
Cite this article
Bogdanovic, G., Pou, C., Barrientos-Somarribas, M. et al. Virome characterisation from Guthrie cards in children who later developed acute lymphoblastic leukaemia. Br J Cancer 115, 1008–1014 (2016). https://doi.org/10.1038/bjc.2016.261
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/bjc.2016.261