Abstract
Background:
MicroRNA-7 (miR-7) has been reported to be a tumour suppressor gene. However, whether it has a role in the growth of non-small-cell lung cancer (NSCLC) and what is its target involved in the tumour growth is still under investigation.
Methods:
NSCLC tissue sample, NSCLC cell lines and tissue microarray were investigated in this study. Total RNA, miRNA and protein were used for RT-PCR and western blot analysis. Immunohistochemistry was performed in tissues microarray. Cell culture and intervention experiments were performed in vitro and in vivo. Bioinformatics prediction, western blot and luciferase assay were identified the target of miR-7.
Results:
In this study, we found that the expression of miR-7 was significantly downregulated not only in NSCLC cell lines, but also in human NSCLC tissues compared with the matched adjacent tissues. Restoration of its expression through miR-7 mimics in A549 and H1299 NSCLC cells inhibited cell proliferation, colony formation, and cell-cycle progression in vitro. More importantly, the tumorigenicity in nude mice was reduced after administration of miR-7 in vivo. In advance, through bioinformatic analysis, luciferase assay and western blot, we identified a novel target of miR-7, PA28gamma (a proteasome activator) to be enrolled in the regulation with tumour. PA28gamma mRNA and protein levels are markedly upregulated in NSCLC cell lines and tumour samples, exhibiting a strong inverse relation with that of miR-7. In addition, knockdown of PA28gamma induced similar effects as overexpression of miR-7 in NSCLC cells. Furthermore, miR-7 overexpression or silencing of PA28gamma reduced the cyclinD1 expression at mRNA and protein level in NSCLC cell lines.
Conclusion:
All these findings strongly imply that the overexpression of PA28gamma resulted from miR-7 downexpression in NSCLC has an important role in promoting cancer cell progress and consequently results in NSCLC growth. Thus, strategies targeting PA28gamma and/or miR-7 may become promising molecular therapies in NSCLC treatment.
Similar content being viewed by others
Main
Non-small-cell lung cancer (NSCLC) remains the leading cause of death from cancer in the world and with poor prognosis in most cases(Cataldo et al, 2011; Siegel et al, 2012). The consistent poor 5-year survival underscores the need for further exploration for its molecular mechanisms (O'Mahony et al, 2005; Owonikoko et al, 2010). Most studies nowadays are focused on the molecular network of protein-coding genes and pathways, such as MAPK (Lee et al, 2005), p53 (Lee et al, 2007; Sinthupibulyakit et al, 2010) and Wnt (Klaus and Birchmeier, 2008; DiMeo et al, 2009) signalling, and attempts have been made to identify important genes and related pathways that contribute to the tumorigenesis of NSCLC. However, knowledge of genomic aberrations associated with non-coding genes, such as microRNAs (miRNAs) and their contributions to NSCLC, are relatively limited, and the identification of oncogenic miRNAs that exert key effects in NSCLC cells may offer new therapeutic targets.
MiRNAs are an abundant group of short, non-coding RNAs with the potential to target and silence multiple genes across diverse biological processes, including cell differentiation, proliferation, growth, mobility, and apoptosis (Wiemer, 2007; Cushing et al, 2011; Sato et al, 2011; Valastyan and Weinberg, 2011). Much exciting evidence has assigned an important regulatory role for miRNAs in several human diseases, including cancer (Suzuki et al, 2009; Liu and Tang, 2011; Png et al, 2011; Zhang et al, 2012). MicroRNAs are directly involved in cancer initiation and progression by regulating the expression of important cancer-related genes and thereby functioning as tumour suppressors or oncogenes (Duan et al, 2010; Bader et al, 2011; Mazar et al, 2011; Kurashige et al, 2012). Recently, miR-7 was identified as a tumour suppressor in several human cancers (Kefas et al, 2008; Reddy et al, 2008; Jiang et al, 2010; Saydam et al, 2011). Differential expression of miR-7 has been reported between neoplastic and normal tissues in brain cancer including glioblastoma and schwannoma (Kefas et al, 2008; Saydam et al, 2011). Only a few reports were focused on lung cancer and indicated a potential role in tumorigenesis of lung cancer (Webster et al, 2009; Chou et al, 2010; Rai et al, 2011; Xiong et al, 2011). However, the results in these researches are contradictory, moreover, the identified targets by which miR-7 exerts its function are still limited, necessitating studies for further elucidation.
To this end, we screened for the expression of miR-7 in 41 pairs of primary NSCLC tissues along with their matched adjacent normal tissues, and explored its function in NSCLC. We found that miR-7 is downregulated both in NSCLC cell lines and tumour tissues, and we provide the first evidence that PA28gamma (a proteasome activator), a novel target of miR-7, is inversely upregulated in NSCLC tissues as well as in NSCLC cell lines. In addition, both overexpression of miR-7 and silencing of PA28gamma downregulated cyclin D1 expression, and therefore result in the inhibition of NSCLC cells growth. Taken together, our data indicate that miR-7 is a candidate tumour suppressor miRNA and functions through the negative regulation of PA28gamma expression in NSCLC. These findings also suggest that low-level miR-7 has a potentially important role on the initiation and/or progression of NSCLC, likely via upregulation of PA28gamma.
Materials and methods
Tissue samples, mice, and cell lines
Forty-one pairs of primary NSCLC tissues and matched adjacent tissues (adenocarcinoma vs squamous cell carcinoma=28 : 13) were obtained from patients in Zhongshan Hospital, Fudan University from 2008 to 2013 with informed consent and agreement. This study was carried out according to the World Medical Association Declaration of Helsinki and approved by the Medical Ethics Committee of Zhongshan Hospital, Fudan University (Approval ID: Y2012-133). Written informed consents were obtained from participants involved in our study. All tissue samples were from untreated patients undergoing surgery and were snap frozen in liquid nitrogen and then stored at −80 °C. Grossly normal and abnormal areas of sectioned patients’ samples were identified by the pathologists and histologically classified and graded according to WHO guidelines and pTNM (UICC) pathological staging criteria. The percentage of tumour is over 90% in squamous cell carcinoma tissues and over 70% in adenocarcinoma tissues (Supplementary Figure 1). For all the samples, clinicopathologic information is summarised in Supplementary Table 1. According to the previous reports, tumour volume >4.5 cm was defined as larger tumour volume group (Strong et al, 2007; Park et al, 2011).
Nude mice, aged 4–6 weeks, were purchased from Shanghai Experimental Center, Chinese Academy of Science. All animal experiments were carried out in compliance with the Guide for the Care and Use of Laboratory Animals published by the US National Institutes of Health (NIH publication no. 85-23, revised 1996) and the Guidelines of Shanghai Medical Laboratory Animal Care and Use Committee (Approval ID: 20120302-023).
Non-small-cell lung cancer cell lines (A549, H1299, H1355, and H460), human bronchial epithelial (HBE) cell line and 293T cells were purchased by the Institute of Biochemistry and Cell Biology of Chinese Academy of Science, China. A549, H1299, H1355, H460, and 293T cells were grown in DMEM supplemented with 10% foetal bovine serum (FBS), 2 μ M glutamine, 100 IU ml−1 penicillin, and 100 μg ml−1 streptomycin sulphate except HBE cells were cultured in RPMI-1640 cell culture medium supplemented with 10% FBS.
RNA extraction
Total RNA of tissues and cultured cells was extracted with Trizol reagent (Invitrogen, Carlsbad, CA, USA) according to the manufacturer’s protocol. RNAs were stored at −80 °C before quantitative reverse transcription polymerase chain reaction (qRT–PCR) analysis.
Quantitative RT-PCR for miRNA
For mature miRNA expression analysis, ∼10 ng of total RNA was converted to cDNA using the ABI miRNA reverse transcription kit (Applied Biosystems, Foster City, CA, USA) and miR-7-specific primers (Applied Biosystems). After reverse transcription, quantitative PCR was performed in triplicate for each sample on the ABI 7500 thermocycler (Applied Biosystems) according to the manufacturer’s protocol. U6 was used as a normalisation control for all tumour and adjacent normal lung tissues and cell line samples.
Quantitative RT-PCR for mRNA expression
Synthesis of cDNA was performed with 0.5–1 μg of total RNA per sample with the primerScript RT reagent Kit (TaKaRa, Dalian, China) according to the manufacturer’s manual. Quantitative RT–PCR was performed in triplicate for each paired tissue or cell sample by the FastStart Universal SYBR Green Master kit (Roche, Mannheim, Germany) according to the manufacturer’s instructions. Primer oligonucleotides were designed by the Primer Premier 5.0 software, and synthesised by Sangon Biotech (Shanghai, China). GAPDH was used as a housekeeping gene for normalisation. The relative gene expression was determined in tumour tissue relative to the first matched normal tissue or HBE cells. The sequences of oligonucleotide primers in this study are summarised in Supplementary Table 2.
miRNAs, small interfering RNAs, and transfection
The human miR-7 duplex mimic, negative control miRNA duplex (miR-NC), and miR-7 inhibitor were designed and synthesised by Ribobio Inc. (Guangzhou, Guangdong, China). miR-7 inhibitor is chemically modified, single-stranded nucleic acids designed to specifically bind to and inhibit endogenous miR-7 molecules. The siRNA pool against PA28gamma mRNA (PA28gamma-siRNA) and siRNA NC were designed and synthesised by Thermo Scientific Dharmacon (Chicago, IL, USA) (Lykke-Andersen et al, 2008). The sequences of the miRNA duplex and siRNA are presented in Supplementary Table 3. Confluent cells (30–50%) were transfected with small interfering RNAs (50 nM) or miRNAs (50 nM) by Lipofectamine 2000 (Invitrogen) according to the manufacturer’s protocol. The transfection efficiencies were detected by fluorescence microscope and FACS (supplementary Figure 2). The expression efficiencies were measured by real-time PCR (Supplementary Figure 3). Total RNA was extracted 24 h after transfection, and total cell proteins were extracted 48 or 72 h after transfection.
Cell proliferation assay
Cell proliferation was determined by the Cell Counting Kit-8 assay (Dojindo, Kumamoto, Japan), a redox assay similar to 3-(4, 5-dimethylthiazol-2-yl)-2,5-diphenyltetrazolium bromide, according to the manufacturer’s protocol and our previously studies (Cai et al, 2011; Liu et al, 2011). Cell proliferation assay was carried out with six replicates.
Colony formation assay
Colony formation assays were performed as described previously (Liu et al, 2011). Briefly, transfected cells (500–1000) were re-plated into 6 cm tissue culture dishes. After 10–15 days, visible colonies were fixed and stained with crystal violet solution (Sigma, St. Louis, MO, USA), and the colony numbers were quantified. The results are expressed as a percent (colony number/original cells). This assay was carried out in triplicate.
Cell-cycle analysis by DNA content
For cell-cycle analysis, cells were transfected with control, miR-7, or PA28gamma-siRNA. Three days later, cells were pre-fixed in cold 70% ethanol, washed, and re-suspended in phosphate-buffered saline containing 50 μg ml−1 propidium iodide and 50 μg ml−1 RNAase A, and DNA content was detected by flow cytometry on a FACScan (BD Biosciences, Franklin Lakes, NJ, USA). Analysis of the cell cycle was performed by the CellFit software.
miRNA target prediction
Putative miR-7 target genes were identified by the combined use of miRBase (http://microrna.sanger.ac.uk), TargetScan (http://genes.mit.edu/targetscan), and PicTar (http://pictar.bio.nyu.edu) target prediction programs (Lewis et al, 2003; Krek et al, 2005; Griffiths-Jones et al, 2008; Zheng et al, 2012).
Vector constructs
To construct the wild-type pGL3-con-PA28gamma-UTR-WT plasmid, a wild-type 3′-UTR fragment of the human PA28gamma mRNA (Genbank accession no. NM_005789) containing the putative miR-7 binding sequence (bases 1413–1419) was amplified by PCR and cloned into the XbaI/FseI site of the pGL3-control vector (Promega, Madison, WI, USA), which is downstream of the luciferase reporter gene. A mutant of the single miR-7 binding site in the 3′-UTR of PA28gamma (5′-GUCUUCCC-3′ to 5′-GUCCGAGC-3′) was made by the Site-Directed Mutagenesis Kit (SBS Genetech, Beijing, China). The mutagenesis primers and the nucleotide sequences of all the PCR cloning primers are listed in Supplementary Table 2.
Luciferase assay
HEK293T cells were transfected with pGL3-con-PA28gamma-3′-UTR-Wt, pGL3-con-PA28gamma-3′-UTR-Mut, a control Renilla luciferase pRL-TK vector (Promega), and miR-7 mimic or negative control (miR-NC) using Lipofectamine 2000 reagent (Invitrogen) according to the manufacturer’s protocol. Luminescence was assayed 48 h later using the Dual-Luciferase Reporter Assay System (Promega) according to the manufacturer’s instructions. Results were normalised to the Renilla luminescence from the same vector and are shown as the ratio between the various treatments and cells transfected with mutant vector.
Tumorigenicity assay in nude mice
Nude mice were used for xenograft studies. miR-7- and miR-NC-transfected H1299 cells (3 × 106) were injected subcutaneously into either side of the posterior flank of the same female nude mouse. After 10 days, the tumour volume was measured with a vernier caliper at weekly intervals. Tumour volume was calculated by the following formula: volume=W2 × L/2, where W is the short diameter and L is the long diameter.
Western blot analysis
Proteins extracted from cells or tumour tissues were immunoblotted with different antibodies following a published protocol (Xiong et al, 2009). The primary antibodies used were PA28gamma (1 : 3000 dilution), Cyclin D1 (1 : 2000 dilution) and GAPDH (1 : 5000 dilution) (Cell Signaling, Danvers, MA, USA).
Tissue microarray and immunohistochemistry
An independent tissue microarray including NSCLC, small-cell lung cancer (SCLC), and their adjacent normal tissues were obtained from OUTDO Biotech (Shanghai, China). The clinicopathologic information for the tissues from the array is summarised in Supplementary Table 4. Immunohistochemistry was performed using a two-step protocol as previously described (Gao et al, 2007). Primary rabbit anti-PA28gamma antibody (1 : 60 dilution) was incubated overnight at 4 °C, and HRP-goat anti-rabbit IgG (Cell Signaling; 1 : 100 dilution) was then incubated for 30 min at room temperature. Then, slides were incubated in fresh DAB solution. The reaction was stopped by washing in running water, and slides were counterstained with haematoxylin.
Evaluation of immunohistochemical staining
Immunohistochemical staining was assessed by two independent pathologists with no knowledge of the patient characteristics, and any discrepancies were resolved by consensus. The mean±s.e.m. percentage value of two cores was considered representative of one tumour. PA28gamma staining was mainly located in the nucleus, and the intensity of PA28gamma staining was evaluated using the following criteria: 0=no yellow; 1=weak yellow; 2=yellow or orange; 3=brown or dark brown. Only yellow, orange, and brown staining were considered a positive result.
Statistical analysis
Data were expressed as mean±s.e.m. of two or three independent experiments, and the statistical analyses were performed by PRISM 5.0 software (GraphPad Software Inc., San Diego, CA, USA). The level of miR-7 between tumour and adjacent tumour tissues was calculated using paired t-test analysis. The ANOVA analysis was used for the multi-groups comparison, whereas Student’s t-test was applied for the comparison between two groups. The correlation was analysed by Pearson correlation analysis. P-values <0.05 were considered statistically significant.
Results
miR-7 is downregulated in NSCLC tissues
In an attempt to explore the expression and significance of miR-7 in NSCLC carcinogenesis, we first analysed its expression by real-time qRT–PCR in human NSCLC cell lines including A549, H1299, H1355, and H460, in which it was greatly downregulated in each compared with the control normal HBE cells (Figure 1A). In addition, miR-7 was further examined in 41 cases of NSCLC tissue. As shown in Figure 1B, the expression of miR-7 significantly reduced in NSCLC tissues relative to their matched adjacent tissues (P=0.0031). The average of miR-7 expression levels was 3.00 in normal tissues and 1.20 in tumour tissues, respectively. MiR-7 level reduced 2.5 times in NSCLC tissues.
Expression of miR-7 is significantly downregulated in primary NSCLC cell lines (A) and NSCLC tissues (B). Mature miR-7 expression in primary NSCLC tissues, matched adjacent lung tissues, and lung cancer cell lines were analysed by real-time PCR and normalised to the endogenous control U6. miR-7 expression was determined in tumour tissue relative to the first patient-matched adjacent normal tissue and relative expression of miR-7 in NSCLC cell lines was also normalised to that in HBE cells. Each sample was analysed in triplicate (**P<0.01, ***P<0.001).
These data suggest that miR-7 is significantly downregulated in NSCLC tissues and cell lines, especially in squamous cell carcinoma and adenocarcinoma. Such reduced expression of miR-7 may correlate with the tumorigenicity of NSCLC.
In addition, the expression of miR-7 in H460 was lower than that in high-invasive A549 and H1299 cells (Supplementary Figure 4A). However, there was no significant difference between low-invasive cell line 95C and high-invasive cell line 95D as shown in Supplementary Figure 4B. Therefore, its expression may be associated with the subtype of NSCLC cancer.
miR-7 inhibits tumour cell growth both in vitro and in vivo
Given the low expression of miR-7 in NSCLC tissues and cell lines, we further evaluated the biological roles of miR-7 in NSCLC cell lines. miR-7 mimic and the control (miR-NC) were transfected into the NSCLC cell lines A549 and H1299 cells separately. As shown in Figure 2A, miR-7 markedly suppressed cell proliferation of both A549 and H1299 cells compared with the negative control. In addition, miR-7 induced cell-cycle arrest at the G1 phase (Figure 2B). Colony formation rate was also reduced after overexpression of miR-7 (Figure 2C).
Ectopic expression of miR-7 inhibits tumour growth in vitro and in vivo. (A) A549 and H1299 cells were transfected with miR-7 mimics and miR-NC. At 24, 48, and 72 h, cell proliferation was assessed by CCK-8 assay. (B) A549 and H1299 cells were transfected with miR-7, or miR-NC in six-well plates. Seventy-two hours later, cells were harvested, and cell cycle was analysed by FACS. (C) Transfected A549 and H1299 cells were replated into 6 cm dishes. Ten to fifteen days later, visible colonies were fixed and stained with crystal violet, and the colony formation ratio was calculated with the following formula: (visible colony numbers/original number of cells) × 100%. Experiments were repeated at least three times. (D) The effect of miR-7 on tumorigenicity was measured in the nude mice xenograft model. miR-7- and miR-NC-transfected H1299 cells (3 × 106) were injected subcutaneously into either side of the posterior flank of the same female nude mouse (n=6). Tumour volume is shown as mean±s.e.m. All data are representative of three or two independent experiments (*P<0.05; **P<0.01; ***P<0.001).
We next explored the role of miR-7 in NSCLC tumour growth in vivo. H1299 cells were transfected with either miR-7 or miR-NC. Six hours later, they were implanted subcutaneously into either posterior flank of the same nude mice (n=6). Six weeks after injection, only 16.7% of mice had palpable tumours observed in the flanks injected with the miR-7-transfected H1299 cells. In contrast, tumours appeared in the opposite sites injected with miR-NC in 100% of mice. Mice were killed when the tumour volume reached 100 mm3. At day 17, all the mice in the control group but none in miR-7-transfected group have reached the critical volume (Figure 2D). The growth curve of the miR-7-transfected group was significantly suppressed compared with the control group. These results show that miR-7 can inhibit the growth of NSCLC cells both in vitro and in vivo.
PA28gamma emerges as a novel target for miR-7
To explore the underlying molecular mechanism responsible for lung cancer growth suppression caused by miR-7, we performed bioinformatic analyses to search for miR-7 target genes. Three prediction algorithms (PicTar, Target Scan, and miRanda) were used to identify the putative target genes, and 49 candidate genes were commonly predicted to be the possible targets of miR-7 (Figure 3A). Among these potential target genes, oncogenes or anti-apoptosis genes involved in the regulation of tumour growth were selected for analysis due to the fact that miRNA-mediated gene repression results in downregulation of target genes. We also found that PA28gamma was a potential target of miR-7. Bioinformatics analysis revealed that the binding sites of the 3′-UTR sequence of PA28gamma and miR-7 are highly conserved in human, chimpanzee, monkey, mouse, cow, horse, hedgehog, and elephant (Figure 3B). The predicted binding site of miR-7 to the 3′UTR of PA28gamma is at the bases from 1413 to 1419 (Figure 3C).
miR-7 post-transcriptionally downregulates PA28gamma expression by directly targeting its 3′-UTR. (A) Putative target genes were predicted by PicTar, Target Scan, and miRanda. Each circle represents one prediction algorithm with the number of predicted genes. The number listed in overlapping areas of the circles was simultaneously predicted by different algorithms. (B) The potential binding sequences within the 3′-UTR of PA28gamma for miR-7 is conserved in human (Hsa), chimpanzee (Ptr), rhesus (Mml), mouse (Mmu), rat, hedgehog (Eeu), shrew (Sar), horse (Eca), cow (Eta), and elephant (Dno). Seed sequences are highlighted. (C) The PA28gamma 3′-UTR and corresponding fragments were inserted into the region immediately downstream of the luciferase gene in the pGL3-con vector. The sequences of the predicted miR-7 binding sites within the PA28gamma 3′-UTR including the wild-type UTR or mutant UTR segments (dotted lines) are shown. (D) After co-transfection of miR-7 mimic or miR-NC with the above reporter plasmids and pRL-TK vector in HEK 293T cells, relative luciferase activity was analysed by Dual-Luciferase Reporter Assay. (E) A549 and H1299 cells were transfected with miR-7, miR-NC, PA28gamma-siRNA, and miR-7 inhibitor for 48 h. PA28gamma protein levels were examined by western blot using GAPDH as a loading control (one of three similar blots is shown). (F) H1299 cells were transfected with PA28gamma-siRNA, and/or miR-7 inhibitor for 72 h. PA28gamma protein levels were examined by western blot using GAPDH as a loading control (one of three similar blots is shown). These experiments were performed in triplicate and results are shown as the mean±s.e.m. (***P<0.001).
In order to determine whether PA28gamma is directly regulated by miR-7 through binding to its 3′-UTR, the 3′-UTR fragment, including the potential binding site (wild-type or mutant) (Figure 3C), was constructed and cloned into the pGL3-con vector immediately downstream of the luciferase reporter. For luciferase activity assays, HEK 293T cells were co-transfected with the PA28gamma-3′UTR-reporter plasmid and miR-7 mimic or negative control (miR-NC). Interestingly, the relative luciferase activity of the reporter containing the wild-type 3′-UTR was remarkably decreased when co-transfected with miR-7. In contrast, the luciferase activity of the mutant reporter was unchanged by simultaneous transfection of miR-7 mimic or negative control (Figure 3D).
miRNA-mediated gene repression mostly results in decreased protein levels of their target genes. To further assess whether miR-7 had a functional role in the downregulation of endogenous PA28gamma expression, A549 cells and H1299 cells were transfected with miR-7 mimic for 48 h and then assessed by western blot. As shown in Figure 3E, PA28gamma was sensitive to miR-7, as a significant decrease was detected in miR-7-transfected A549 cells and H1299 cells. On the contrary, miR-7 inhibitor partially rescued the inhibition of PA28gamma expression by miR-7. Moreover, inhibition of miR-7 upregulated the PA28gamma expression and partially rescued the effects of the PA28gamma knock down in H1299 cells (Figure 3F). These results indicate that miR-7 suppresses PA28gamma expression through directly targeting its 3′-UTR, and PA28gamma is a novel target of miR-7.
PA28gamma is significantly upregulated in NSCLC
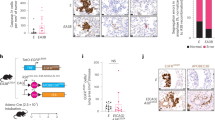
As PA28gamma is emerging as a novel target of miR-7, we further examined its expression in NSCLC by immunohistochemistry in an independent tissue microarray including 50 pairs of NSCLC samples and 42 cases of SCLC samples. The relative expression level of PA28gamma was scored by two independent pathologists. A total of 70% (35 out of 50) of NSCLC cancer tissues showed a net gain of immunoreactivity for nuclear PA28gamma compared with the matched normal lung tissues (Figure 4A and Table 1, P<0.001); SCLC cancer tissues also showed decreased reactivity when compared with NSCLC cancer tissues (Figure 4A and Table 1, P=0.003).
PA28gamma is overexpressed in NSCLC tissues. (A) Immunohistochemical staining of PA28gamma in an independent lung cancer tissue microarray including the matched adjacent normal tissues. (SCC=squamous cell carcinoma; ADC=adenocarcinoma; SCLC=small-cell lung carcinoma. The tissue pairs were isolated from the same patient). The staining intensities were scored, and statistical analysis was performed according to the scoring guidelines (Table 1). The protein level of PA28gamma in representative resected fresh NSCLC tissues along with their matched non-tumour tissues (B) and NSCLC cell lines (C) was analysed by western blot analysis and normalised to GAPDH, and further normalised to the value in the first normal specimen or HBE cells. Two or three independent experiments were performed, and representative data are shown (N=non-tumour tissue; T=tumour tissue).
To further validate the expression of PA28gamma in human NSCLC, we also analysed 41 pairs of fresh resected tumour samples in our study by western blot assay. Similar to the results of immunohistochemistry in the independent tissue microarray, we found that protein levels of PA28gamma in NSCLC tumour samples were significantly higher compared with that in matched adjacent normal tissues (Figure 4B). In addition, we also analysed the expression of PA28gamma in NSCLC cell lines by western blot assay. Similar results were observed in A549, H1299 cells compared with the normal HBE cell line (Figure 4C), in spite of the unclear significant difference between H1355 and HBE cells. This data indicated that PA28gamma is overexpressed in NSCLC and may become a novel biomarker for diagnosis of NSCLC.
Inverse relationship between PA28gamma and miR-7 is detected in NSCLC tissues
On the basis of the overexpression of PA28gamma in NSCLC cancer tissues and the direct negative regulation of miR-7 on PA28gamma, we hypothesised that there may be an inverse relationship between miR-7 and PA28gamma expression in NSCLC. Thus, we analysed the mRNA level of PA28gamma in 41 cases of human NSCLC tumour samples and their matched adjacent normal tissues by qRT–PCR analysis. Interestingly, we found that mRNA levels of PA28gamma in NSCLC tumour samples were significantly higher compared with their matched adjacent normal tissues (P<0.0001) (Figure 5A). In contrast, expression of miR-7 was significantly (P=0.0015) reduced in NSCLC tissues compared with their matched controls (Figure 1B). Pearson’s correlation analysis revealed a significant inverse relationship between the levels of PA28gamma and miR-7 in human NSCLC tissue samples (r=−0.4013; P=0.0002; Figure 5C). In addition, we further analysed their expression in three different NSCLC cell lines: A549, H1299, and H1355. In all of these, PA28gamma expression was upregulated (Figure 5B), whereas miR-7 expression was also downregulated compared with the normal HBE cell line (Figure 1A). Taken together, these results strongly implicate a role of miR-7 in the negative regulation of PA28gamma expression.
Expression of PA28gamma correlates inversely with miR-7 levels in NSCLC tissues. PA28gamma expression was examined by qRT–PCR in 41 NSCLC tissues, their matched non-tumour tissues (A), NSCLC cell lines and their control HBE cells (B). PA28gamma expression was normalised to GAPDH mRNA, and the relative gene expression of PA28gamma were determined in tumour tissue relative to the first matched normal tissue sample or normal HBE cell line. Pearson’s correlation analysis was performed for PA28gamma and miR-7 in human NSCLC tissue samples (C). Each sample was analysed in triplicate.
Specific knockdown of PA28gamma by siRNA elicits similar phenotypes of growth arrest
To directly address miR-7 specificity and involvement of PA28gamma in miR-7-regulated inhibition of proliferation and tumour suppression, we treated A549 and H1299 cells with a pool of siRNAs specific to PA28gamma mRNA. Western blot analysis confirmed specific knockdown of PA28gamma by siRNA (Figure 6A). Further, PA28gamma-siRNA-transfected A549 and H1299 cells showed inhibited proliferation (Figure 6B), cell-cycle arrest at G1 phase (Figure 6D) and decreased colony formation (Figure 6E) compared with siRNA-NC-transfected cells. On the contrary, inhibition of miR-7 promoted cell proliferation (Figure 6C), progression of cell cycle (Figure 6D), and colony formation (Figure 6E) in H1299 cells and partially rescued the effects of PA28gamma inhibition (Figure 6C–E).
PA28gamma mediates miR-7-induced inhibition of NSCLC cell growth. (A) A549 and H1299 cells were transfected with negative control siRNA (siRNA-NC) or a PA28gamma siRNA pool. Forty-eight hours later, PA28gamma expression was examined by western blot. (B) A549 and H1299 cells were transfected with siRNA-NC or PA28gamma siRNA in 96-well plates. Cell proliferation was assessed by CCK-8 assay at 24, 48, and 72 h. (C) H1299 cells were transfected with negative control siRNA (SiRNA-NC), PA28gamma siRNA, and/or miR-7 inhibitor in 96-well plates. Cell proliferation was assessed by CCK-8 assay at 72 h. (D) H1299 cells were transfected with siRNA-NC, PA28gamma siRNA, and/or miR-7 inhibitor in six-well plates. Seventy-two hours later, cells were harvested, and cell cycle was analysed by FACS. (E) One thousand transfected H1299 cells were plated to 6 cm dishes to perform the colony formation assays. Ten to fifteen days later, visible colonies were fixed and stained with crystal violet. Three independent experiments were performed, and representative data are shown (*P<0.05; **P<0.01; ***P<0.001).
These data suggest that PA28gamma, which is downregulated by miR-7, positively regulates cell growth of NSCLC.
miR-7 represses cell-cycle protein cyclin D1 expression through downregulation of PA28gamma
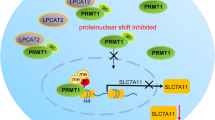
PA28gamma, is an activator of 20S proteasome, and may be involved in cell growth and cell-cycle progression (Masson et al, 2001; Kanai et al, 2011). Cyclin D1 is proposed to serve as an important switch in the regulation of continued cell-cycle progression and is often overexpressed in a variety of cancers (Marchetti et al, 1998; Lamb et al, 2003). To determine if PA28gamma repression by miR-7 affects cyclin D1 expression, we transfected A549 and H1299 cells with miR-7 or PA28gamma siRNA and evaluated cyclinD1 expression levels. As shown in Figure 7A and B, analysis of mRNA expression by real-time PCR indicated a profound reduction in cyclin D1 levels after miR-7 compared with control treatments. Western blot analysis further confirmed that repression of miR-7 inhibited cyclin D1 protein expression both in A549 and H1299 cells (Figure 7C and D). These effects were similar with direct knockdown of PA28gamma by siRNA (Figure 7A–D). These data suggest that part of the anti-cancer effect of miR-7 may occur through inhibition of the PA28gamma and subsequent reduction of cyclin D1 expression. Therefore, it induces NSCLC tumour cell growth arrest (Figure 7E).
miR-7 represses cyclin D1 expression through reducing PA28gamma level. A549 and H1299 cells were transfected with miR-7, PA28gamma siRNA and their controls at 50 nM for 24 or 72 h. The mRNA expression levels of cyclin D1 in A549 (A) and H1299 cells (B) were measured by real-time PCR and standardized to GAPDH. Protein levels of cyclin D1 in A549 (C) and H1299 cells (D) were determined by western blot with GAPDH used as a loading control. (**P<0.01; ***P<0.001). (E) Hypothetical model of miR-7 suppressive function in NSCLC.
Discussion
In this study, we reported that miR-7 is downregulated in NSCLC tumour tissue. It is assumed that the downexpressed miRNAs in cancers may function as tumour-suppressors. Hence, we inferred that miR-7 might be a negative regulator in NSCLC. As a result, restoration of miR-7 inhibited cell proliferation, colony formation, cell-cycle progression in vitro, and the tumorigenicity in vivo, which indicated the importance of miR-7 in tumorigenesis of NSCLC. Previous studies have also shown that miR-7 expression was significantly decreased in recurrent lung cancers and emerged as a potential tumour suppressor (Webster et al, 2009; Duncavage et al, 2010; Xiong et al, 2011). However, a recent report demonstrated that miR-7 promoted, rather than inhibited, cell growth in lung cancer cell line CL1-50 (Chou et al, 2010). In contract, Webster et al (2009) and Wang et al (2013) showed that miR-7 functioned as a tumour suppressor by directly targeting epidermal growth factor receptor (EGFR). Obviously, the interaction between miR-7 and EGFR is complicated. Similar to Webster’s report, our present study showed that miR-7 inhibited rather than promoted the lung tumorigenesis in A549 cells. Therefore, the role of miR-7 may be different in various cancer types and diverse biological backgrounds of cell lines. The exact role of miR-7 in lung cancer development remained contradictive. On the basis of our previous work (Xiong et al, 2011) and results in this study, we provide further evidence that miR-7 is a functional regulator in NSCLC and support the hypothesis that miR-7 serves as a tumour suppressor in NSCLC.
Previously known targets of miR-7 include messages for signalling proteins including EGFR, murine leukaemia viral oncogene homologue 1RAF1 (v-raf-1), insulin receptor substrate 1 (IRS1), IRS2, activated CDC42 kinase 1 (Ack1), B-cell lymphoma 2(Bcl-2), and phosphoinositide-3-kinase, regulatory subunit 3 (PIK3R3) (Kefas et al, 2008; Reddy et al, 2008; Jiang et al, 2010; Saydam et al, 2011; Xiong et al, 2011; Xu et al, 2013). Among them, the interaction between miR-7 and ACK1, as well as their inverse expressions in human tumour tissues, has been well demonstrated (Saydam et al, 2011). We have also paid attention to the anti-apoptosis molecule Bcl-2, although miR-7 can directly target it and reduce its expression in NSCLC cell lines (Xiong et al, 2011), the inversed expression between miR-7 and Bcl-2 was not significant in human NSCLC samples (data not shown). Obviously, the molecular mechanism through which miR-7 functions as a tumour suppressor still needs to be explored. On the basis of bioinformatics analysis, we further predicted a number of additional miR-7 targets. PA28gamma was identified to be a novel target gene of miR-7 in this study.
PA28gamma, also called REGgamma, Ki antigen or PSME3, is an REG family member that binds and activates the 20S proteasome (Masson et al, 2001; Kanai et al, 2011). Although the exact physiological role of PA28gamma is still unclear, it has been suggested to be involved in cell growth and cell-cycle progression. PA28gamma knockout mice show growth retardation, and the embryonic fibroblasts from these animals displayed cell-cycle arrest at G1 phase (Murata et al, 1999; Barton et al, 2004). In recent years, PA28gamma has been shown to downregulate the steroid co-activating factor-3 (SRC-3), thereby resulting in cell growth (Masson et al, 2009; Zannini et al, 2009). Furthermore, three important cell-cycle regulators, p21, p16, and p19, are degraded in a PA28gamma-dependent and ubiquitin-independent manner (Chen et al, 2007; Wu et al, 2011). Thus, PA28gamma is involved the regulation of cell proliferation. However, the exact role of PA28gamma in tumours is unknown. Recently Sanchez et al (2013) reported that miR-7 triggers the cell cycle in G1/S transition by targeting multiple genes including PSME3, which further supports that PA28gamma (also named PSME3) is a functional target gene of miR-7.
In this study, we further analysed the expressions of miR-7 and PA28gamma in human samples. As a new target of miR-7, PA28gamma is inversely upregulated in both NSCLC cell lines and tumour tissues. Thus, as a post-transcriptional control, loss of miR-7 expression may contribute to PA28gamma overexpression in tumorigenesis. In addition, restoration of miR-7 and inhibition of PA28gamma repressed the expression of cyclin D1 at both mRNA and protein even though the detained mechanism is not known yet and need to be further explored.
This study has some limitations. We are mainly focused on the interaction of miR-7 and PA28gamma. Besides PA28gamma, some other important functional molecules have been identified in the previous studies, such as EGFR, v-raf-1, IRS1, IRS2, Ack1, Bcl-2, and PIK3R3 (Kefas et al, 2008; Reddy et al, 2008; Jiang et al, 2010; Saydam et al, 2011; Xiong et al, 2011; Xu et al, 2013). Indeed, how this complicated network regulates tumorigenesis remains unexplored. In addition, the function of PA28gamma, as well as its inverse expression with miR-7, is based on NSCLC. Our initial observations require more studies in different cancers.
In summary, the present study strongly imply that the overexpression of PA28gamma resulted from miR-7 downexpression in NSCLC has an important role in promoting cancer cell progress, and consequently results in NSCLC growth. Therefore, the present study enriches the knowledge about the interactions between miR-7 and the targets through demonstrating that miR-7 can not only promote cell apoptosis by targeting EGFR as reported, but also induce cell-cycle arrest through interaction with PA28gamma in regulating NSCLC development (Figure 7E). Thus, strategies targeting PA28gamma and/or miR-7 may be the promising molecular therapy in NSCLC treatment.
Change history
21 January 2014
This paper was modified 12 months after initial publication to switch to Creative Commons licence terms, as noted at publication
References
Bader AG, Brown D, Winkler M (2011) The promise of microRNA replacement therapy. Cancer Res 70: 7027–7030.
Barton LF, Runnels HA, Schell TD, Cho Y, Gibbons R, Tevethia SS, Deepe Jr GS, Monaco JJ (2004) Immune defects in 28-kDa proteasome activator gamma-deficient mice. J Immunol 172: 3948–3954.
Cai Y, Xiong S, Zheng Y, Luo F, Jiang P, Chu Y (2011) Trichosanthin enhances anti-tumour immune response in a murine Lewis lung cancer model by boosting the interaction between TSLC1 and CRTAM. Cell Mol Immunol 8: 359–367.
Cataldo VD, Gibbons DL, Perez-Soler R, Quintas-Cardama A (2011) Treatment of non-small-cell lung cancer with erlotinib or gefitinib. N Engl J Med 364: 947–955.
Chen X, Barton LF, Chi Y, Clurman BE, Roberts JM (2007) Ubiquitin-independent degradation of cell-cycle inhibitors by the REGgamma proteasome. Mol Cell 26: 843–852.
Chou YT, Lin HH, Lien YC, Wang YH, Hong CF, Kao YR, Lin SC, Chang YC, Lin SY, Chen SJ, Chen HC, Yeh SD, Wu CW (2010) EGFR promotes lung tumorigenesis by activating miR-7 through a Ras/ERK/Myc pathway that targets the Ets2 transcriptional repressor ERF. Cancer Res 70: 8822–8831.
Cushing L, Kuang PP, Qian J, Shao F, Wu J, Little F, Thannickal VJ, Cardoso WV, Lu J (2011) miR-29 is a major regulator of genes associated with pulmonary fibrosis. Am J Respir Cell Mol Biol 45: 287–294.
DiMeo TA, Anderson K, Phadke P, Fan C, Perou CM, Naber S, Kuperwasser C (2009) A novel lung metastasis signature links Wnt signaling with cancer cell self-renewal and epithelial-mesenchymal transition in basal-like breast cancer. Cancer Res 69: 5364–5373.
Duan W, Gao L, Wu X, Wang L, Nana-Sinkam SP, Otterson GA, Villalona-Calero MA (2010) MicroRNA-34a is an important component of PRIMA-1-induced apoptotic network in human lung cancer cells. Int J Cancer 127: 313–320.
Duncavage E, Goodgame B, Sezhiyan A, Govindan R, Pfeifer J (2010) Use of microRNA expression levels to predict outcomes in resected stage I non-small cell lung cancer. J Thorac Oncol 5: 1755–1763.
Gao Q, Qiu SJ, Fan J, Zhou J, Wang XY, Xiao YS, Xu Y, Li YW, Tang ZY (2007) Intratumoral balance of regulatory and cytotoxic T cells is associated with prognosis of hepatocellular carcinoma after resection. J Clin Oncol 25: 2586–2593.
Griffiths-Jones S, Saini HK, van Dongen S, Enright AJ (2008) miRBase: tools for microRNA genomics. Nucleic Acids Res 36: D154–D158.
Jiang L, Liu X, Chen Z, Jin Y, Heidbreder CE, Kolokythas A, Wang A, Dai Y, Zhou X (2010) MicroRNA-7 targets IGF1R (insulin-like growth factor 1 receptor) in tongue squamous cell carcinoma cells. Biochem J 432: 199–205.
Kanai K, Aramata S, Katakami S, Yasuda K, Kataoka K (2011) Proteasome activator PA28{gamma} stimulates degradation of GSK3-phosphorylated insulin transcription activator MAFA. J Mol Endocrinol 47: 119–127.
Kefas B, Godlewski J, Comeau L, Li Y, Abounader R, Hawkinson M, Lee J, Fine H, Chiocca EA, Lawler S, Purow B (2008) microRNA-7 inhibits the epidermal growth factor receptor and the Akt pathway and is down-regulated in glioblastoma. Cancer Res 68: 3566–3572.
Klaus A, Birchmeier W (2008) Wnt signalling and its impact on development and cancer. Nat Rev Cancer 8: 387–398.
Krek A, Grun D, Poy MN, Wolf R, Rosenberg L, Epstein EJ, MacMenamin P, da Piedade I, Gunsalus KC, Stoffel M, Rajewsky N (2005) Combinatorial microRNA target predictions. Nat Genet 37: 495–500.
Kurashige J, Watanabe M, Iwatsuki M, Kinoshita K, Saito S, Hiyoshi Y, Kamohara H, Baba Y, Mimori K, Baba H (2012) Overexpression of microRNA-223 regulates the ubiquitin ligase FBXW7 in oesophageal squamous cell carcinoma. Br J Cancer 106: 182–188.
Lamb J, Ramaswamy S, Ford HL, Contreras B, Martinez RV, Kittrell FS, Zahnow CA, Patterson N, Golub TR, Ewen ME (2003) A mechanism of cyclin D1 action encoded in the patterns of gene expression in human cancer. Cell 114: 323–334.
Lee HY, Oh SH, Suh YA, Baek JH, Papadimitrakopoulou V, Huang S, Hong WK (2005) Response of non-small cell lung cancer cells to the inhibitors of phosphatidylinositol 3-kinase/Akt- and MAPK kinase 4/c-Jun NH2-terminal kinase pathways: an effective therapeutic strategy for lung cancer. Clin Cancer Res 11: 6065–6074.
Lee YC, Wu CT, Kuo SW, Tseng YT, Chang YL (2007) Significance of extranodal extension of regional lymph nodes in surgically resected non-small cell lung cancer. Chest 131: 993–999.
Lewis BP, Shih IH, Jones-Rhoades MW, Bartel DP, Burge CB (2003) Prediction of mammalian microRNA targets. Cell 115: 787–798.
Liu C, Tang DG (2011) MicroRNA regulation of cancer stem cells. Cancer Res 71: 5950–5954.
Liu N, Zheng Y, Zhu Y, Xiong S, Chu Y (2011) Selective impairment of CD4+CD25+Foxp3+ regulatory T cells by paclitaxel is explained by Bcl-2/Bax mediated apoptosis. Int Immunopharmacol 11: 212–219.
Liu X, Sempere LF, Ouyang H, Memoli VA, Andrew AS, Luo Y, Demidenko E, Korc M, Shi W, Preis M, Dragnev KH, Li H, Direnzo J, Bak M, Freemantle SJ, Kauppinen S, Dmitrovsky E (2011) MicroRNA-31 functions as an oncogenic microRNA in mouse and human lung cancer cells by repressing specific tumor suppressors. J Clin Invest 120: 1298–1309.
Lykke-Andersen K, Gilchrist MJ, Grabarek JB, Das P, Miska E, Zernicka-Goetz M (2008) Maternal Argonaute 2 is essential for early mouse development at the maternal-zygotic transition. Mol Biol Cell 19: 4383–4392.
Marchetti A, Doglioni C, Barbareschi M, Buttitta F, Pellegrini S, Gaeta P, La Rocca R, Merlo G, Chella A, Angeletti CA, Dalla Palma P, Bevilacqua G (1998) Cyclin D1 and retinoblastoma susceptibility gene alterations in non-small cell lung cancer. Int J Cancer 75: 187–192.
Masson P, Andersson O, Petersen UM, Young P (2001) Identification and characterization of a Drosophila nuclear proteasome regulator. A homolog of human 11 S REGgamma (PA28gamma). J Biol Chem 276: 1383–1390.
Masson P, Lundin D, Soderbom F, Young P (2009) Characterization of a REG/PA28 proteasome activator homolog in Dictyostelium discoideum indicates that the ubiquitin- and ATP-independent REGgamma proteasome is an ancient nuclear protease. Eukaryot Cell 8: 844–851.
Mazar J, Khaitan D, DeBlasio D, Zhong C, Govindarajan SS, Kopanathi S, Zhang S, Ray A, Perera RJ (2011) Epigenetic regulation of microRNA genes and the role of miR-34b in cell invasion and motility in human melanoma. PLoS One 6: e24922.
Murata S, Kawahara H, Tohma S, Yamamoto K, Kasahara M, Nabeshima Y, Tanaka K, Chiba T (1999) Growth retardation in mice lacking the proteasome activator PA28gamma. J Biol Chem 274: 38211–38215.
O'Mahony D, Kummar S, Gutierrez ME (2005) Non-small-cell lung cancer vaccine therapy: a concise review. J Clin Oncol 23: 9022–9028.
Owonikoko TK, Ramalingam SS, Belani CP (2010) Maintenance therapy for advanced non-small cell lung cancer: current status, controversies, and emerging consensus. Clin Cancer Res 16: 2496–2504.
Park KJ, Kano H, Parry PV, Niranjan A, Flickinger JC, Lunsford LD, Kondziolka D (2011) Long-term outcomes after gamma knife stereotactic radiosurgery for nonfunctional pituitary adenomas. Neurosurgery 69: 1188–1199.
Png KJ, Halberg N, Yoshida M, Tavazoie SF (2011) A microRNA regulon that mediates endothelial recruitment and metastasis by cancer cells. Nature 481: 190–194.
Rai K, Takigawa N, Ito S, Kashihara H, Ichihara E, Yasuda T, Shimizu K, Tanimoto M, Kiura K (2011) Liposomal delivery of MicroRNA-7-expressing plasmid overcomes epidermal growth factor receptor tyrosine kinase inhibitor-resistance in lung cancer cells. Mol Cancer Ther 10: 1720–1727.
Reddy SD, Ohshiro K, Rayala SK, Kumar R (2008) MicroRNA-7, a homeobox D10 target, inhibits p21-activated kinase 1 and regulates its functions. Cancer Res 68: 8195–8200.
Sanchez N, Gallagher M, Lao N, Gallagher C, Clarke C, Doolan P, Aherne S, Blanco A, Meleady P, Clynes M, Barron N (2013) MiR-7 triggers cell cycle arrest at the G1/S transition by targeting multiple genes including Skp2 and Psme3. PLoS One 8: e65671.
Sato F, Tsuchiya S, Meltzer SJ, Shimizu K (2011) MicroRNAs and epigenetics. Febs J 278: 1598–1609.
Saydam O, Senol O, Wurdinger T, Mizrak A, Ozdener GB, Stemmer-Rachamimov AO, Yi M, Stephens RM, Krichevsky AM, Saydam N, Brenner GJ, Breakefield XO (2011) miRNA-7 attenuation in Schwannoma tumors stimulates growth by upregulating three oncogenic signaling pathways. Cancer Res 71: 852–861.
Siegel R, Naishadham D, Jemal A (2012) Cancer statistics, 2012. CA Cancer J Clin 62: 10–29.
Sinthupibulyakit C, Ittarat W, Clair St WH, Clair St DK (2010) p53 Protects lung cancer cells against metabolic stress. Int J Oncol 37: 1575–1581.
Strong VE, D'Angelica M, Tang L, Prete F, Gonen M, Coit D, Touijer KA, Fong Y, Brennan MF (2007) Laparoscopic adrenalectomy for isolated adrenal metastasis. Ann Surg Oncol 14: 3392–3400.
Suzuki HI, Yamagata K, Sugimoto K, Iwamoto T, Kato S, Miyazono K (2009) Modulation of microRNA processing by p53. Nature 460: 529–533.
Valastyan S, Weinberg RA (2011) Roles for microRNAs in the regulation of cell adhesion molecules. J Cell Sci 124: 999–1006.
Wang W, Dai LX, Zhang S, Yang Y, Yan N, Fan P, Dai L, Tian HW, Cheng L, Zhang XM, Li C, Zhang JF, Xu F, Shi G, Chen XL, Du T, Li YM, Wei YQ, Deng HX (2013) Regulation of epidermal growth factor receptor signaling by plasmid-based microRNA-7 inhibits human malignant gliomas growth and metastasis in vivo. Neoplasma 60: 274–283.
Webster RJ, Giles KM, Price KJ, Zhang PM, Mattick JS, Leedman PJ (2009) Regulation of epidermal growth factor receptor signaling in human cancer cells by microRNA-7. J Biol Chem 284: 5731–5741.
Wiemer EA (2007) The role of microRNAs in cancer: no small matter. Eur J Cancer 43: 1529–1544.
Wu Y, Wang L, Zhou P, Wang G, Zeng Y, Wang Y, Liu J, Zhang B, Liu S, Luo H, Li X (2011) Regulation of REGgamma cellular distribution and function by SUMO modification. Cell Res 21: 807–816.
Xiong S, Zheng Y, Jiang P, Liu R, Liu X, Chu Y (2011) MicroRNA-7 inhibits the growth of human non-small cell lung cancer A549 cells through targeting BCL-2. Int J Biol Sci 7: 805–814.
Xiong SD, Yu K, Liu XH, Yin LH, Kirschenbaum A, Yao S, Narla G, DiFeo A, Wu JB, Yuan Y, Ho SM, Lam YW, Levine AC (2009) Ribosome-inactivating proteins isolated from dietary bitter melon induce apoptosis and inhibit histone deacetylase-1 selectively in premalignant and malignant prostate cancer cells. Int J Cancer 125: 774–782.
Xu L, Wen Z, Zhou Y, Liu Z, Li Q, Fei G, Luo J, Ren T (2013) MicroRNA-7-regulated TLR9 signaling-enhanced growth and metastatic potential of human lung cancer cells by altering the phosphoinositide-3-kinase, regulatory subunit 3/Akt pathway. Mol Biol Cell 24: 42–55.
Zannini L, Buscemi G, Fontanella E, Lisanti S, Delia D (2009) REGgamma/PA28gamma proteasome activator interacts with PML and Chk2 and affects PML nuclear bodies number. Cell Cycle 8: 2399–2407.
Zhang J, Luo N, Luo Y, Peng Z, Zhang T, Li S (2012) microRNA-150 inhibits human CD133-positive liver cancer stem cells through negative regulation of the transcription factor c-Myb. Int J Oncol 40: 747–756.
Zheng Y, Xiong S, Jiang P, Liu R, Liu X, Qian J, Zheng X, Chu Y (2012) Glucocorticoids inhibit lipopolysaccharide-mediated inflammatory response by downregulating microRNA-155: a novel anti-inflammation mechanism. Free Radic Biol Med 52: 1307–1317.
Acknowledgements
We thank Dr Yi Sun (Division of Radiation and Cancer Biology, Department of Radiation Oncology, University of Michigan, USA), Dr Lijun Jia and Dr Rui He (Department of Immunology; Fudan University; Shanghai, China) for helpful scientific discussion. This work was supported by National Science Foundation of China (81272259, 81072408, and 81273215), the National 973 project 2011CB910 404) in China, the Science and Technology Commission of Shanghai Municipality (10JC 1401100) in China, the Major Research Plan of the National Natural Science Foundation of China (91229110), the China Postdoctoral Science Foundation (213M530001), Specialized Research Fund for the Doctoral Program of Higher Education (20120071110046), and the Graduates Creation Foundation of Fudan University (EZF101321).
Author information
Authors and Affiliations
Corresponding authors
Additional information
This work is published under the standard license to publish agreement. After 12 months the work will become freely available and the license terms will switch to a Creative Commons Attribution-NonCommercial-Share Alike 3.0 Unported License.
Supplementary Information accompanies this paper on British Journal of Cancer website
Rights and permissions
From twelve months after its original publication, this work is licensed under the Creative Commons Attribution-NonCommercial-Share Alike 3.0 Unported License. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-sa/3.0/
About this article
Cite this article
Xiong, S., Zheng, Y., Jiang, P. et al. PA28gamma emerges as a novel functional target of tumour suppressor microRNA-7 in non-small-cell lung cancer. Br J Cancer 110, 353–362 (2014). https://doi.org/10.1038/bjc.2013.728
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/bjc.2013.728
Keywords
This article is cited by
-
Reciprocal REGγ-mTORC1 regulation promotes glycolytic metabolism in hepatocellular carcinoma
Oncogene (2021)
-
REGγ deficiency suppresses tumor progression via stabilizing CK1ε in renal cell carcinoma
Cell Death & Disease (2018)
-
Comprehensive circular RNA profiling reveals the regulatory role of the circRNA-100338/miR-141-3p pathway in hepatitis B-related hepatocellular carcinoma
Scientific Reports (2017)
-
Breviscapine suppresses the growth of non-small cell lung cancer by enhancing microRNA-7 expression
Journal of Biosciences (2017)
-
Proteasome dysregulation in human cancer: implications for clinical therapies
Cancer and Metastasis Reviews (2017)