Article
|
Open Access
Featured
-
-
Article
| Open Access Characterizing chromatin landscape from aggregate and single-cell genomic assays using flexible duration modeling
Characterizing chromatin landscape from aggregate and single-cell genomic assays using flexible duration modeling
Most currently available statistical tools for the analysis of ATAC-seq data were repurposed from tools developed for other functional genomics data (e.g. ChIP-seq). Here, Gabitto et al develop ChromA, a Bayesian statistical approach for the analysis of both bulk and single-cell ATAC-seq data.
- Mariano I. Gabitto
- , Anders Rasmussen
- & Richard Bonneau
-
Article
| Open Access High-throughput identification of synthetic riboswitches by barcode-free amplicon-sequencing in human cells
High-throughput identification of synthetic riboswitches by barcode-free amplicon-sequencing in human cells
Riboswitches can mediate ligand-dependent RNA cleavage and splicing to control gene expression. Here the authors present a method to functionally screen large libraries and identify functional variants.
- Benjamin Strobel
- , Maike Spöring
- & Sebastian Kreuz
-
Article
| Open Access Bridging non-overlapping reads illuminates high-order epistasis between distal protein sites in a GPCR
Bridging non-overlapping reads illuminates high-order epistasis between distal protein sites in a GPCR
Epistasis effects among amino acids at distal sites within binding pockets can have important impacts on protein fitness landscapes. Here the authors present BRIDGE, which matches non-overlapping sequence reads with their cognate DNA templates.
- Justin I. Yoo
- , Patrick S. Daugherty
- & Michelle A. O’Malley
-
Article
| Open Access Chromatin mapping and single-cell immune profiling define the temporal dynamics of ibrutinib response in CLL
Chromatin mapping and single-cell immune profiling define the temporal dynamics of ibrutinib response in CLL
Ibrutinib, a Bruton tyrosine kinase inhibitor, provides effective treatment for chronic lymphocytic leukemia (CLL). Here, the authors describe time-dependent molecular changes to malignant cells and to the immune system in patients undergoing ibrutinib therapy, with can be used for therapy monitoring.
- André F. Rendeiro
- , Thomas Krausgruber
- & Christoph Bock
-
Article
| Open Access Genome-wide rare variant analysis for thousands of phenotypes in over 70,000 exomes from two cohorts
Genome-wide rare variant analysis for thousands of phenotypes in over 70,000 exomes from two cohorts
Population-based association analyses of rare genetic variants with complex traits are limited by the availability of data from sufficiently large cohorts. Here, Cirulli et al. report gene-based collapsing analysis of exomes from 49,960 participants of the UK Biobank and 21,866 participants of the Healthy Nevada Project over a total of 4377 traits.
- Elizabeth T. Cirulli
- , Simon White
- & Nicole L. Washington
-
Article
| Open Access Transcriptomic and open chromatin atlas of high-resolution anatomical regions in the rhesus macaque brain
Transcriptomic and open chromatin atlas of high-resolution anatomical regions in the rhesus macaque brain
Non-human primates share many features with humans and are an important animal model in neuroscience. Here, the authors present a comprehensive transcriptomic and open chromatin atlas of the rhesus macaque brain.
- Senlin Yin
- , Keying Lu
- & Yang Yu
-
Article
| Open Access Non-invasive characterization of human bone marrow stimulation and reconstitution by cell-free messenger RNA sequencing
Non-invasive characterization of human bone marrow stimulation and reconstitution by cell-free messenger RNA sequencing
Circulating cell-free mRNA holds great promise as a non-invasive diagnostic biomarker. Here the authors show that cell-free mRNA captures transcripts from the bone marrow and can be used to non-invasively monitor dynamic changes in bone marrow physiology.
- Arkaitz Ibarra
- , Jiali Zhuang
- & Michael Nerenberg
-
Article
| Open Access Mutational signatures in tumours induced by high and low energy radiation in Trp53 deficient mice
Mutational signatures in tumours induced by high and low energy radiation in Trp53 deficient mice
Mutational signatures induced by ionising radiation remain largely unexplored. Here in TP53 mutant mice, the authors characterise the genomic landscape of tumours induced by high- and low-energy radiation.
- Yun Rose Li
- , Kyle D. Halliwill
- & Allan Balmain
-
Article
| Open Access Mikania micrantha genome provides insights into the molecular mechanism of rapid growth
Mikania micrantha genome provides insights into the molecular mechanism of rapid growth
Mikania micrantha is an extremely fast-growing invasive plant species that can cause serious damage to natural ecosystems. Here, the authors assemble its chromosome-scale reference genome and explore possible mechanisms that contribute to its rapid growth.
- Bo Liu
- , Jian Yan
- & Fanghao Wan
-
Article
| Open Access CaSpER identifies and visualizes CNV events by integrative analysis of single-cell or bulk RNA-sequencing data
CaSpER identifies and visualizes CNV events by integrative analysis of single-cell or bulk RNA-sequencing data
RNA-sequencing is mostly used to assess gene expression; however, it can also give information about genetic variants. Here, the authors present CaSpER, a statistical framework that utilises RNA-sequencing reads to identify and visualise CNV events by integrating transcriptome-wide expression and allelic shift profiles.
- Akdes Serin Harmanci
- , Arif O. Harmanci
- & Xiaobo Zhou
-
Article
| Open Access Identification of recurrent FHL2-GLI2 oncogenic fusion in sclerosing stromal tumors of the ovary
Identification of recurrent FHL2-GLI2 oncogenic fusion in sclerosing stromal tumors of the ovary
Little is known about the genetics of sclerosing stromal tumor of the ovary, a rare type of sex cord-stromal tumor. Here, the authors use sequencing strategies to show that in a cohort of 26 tumor samples 65% carry a FHL2-GLI2 fusion gene and demonstrate in vitro that the fusion gene has oncogenic properties.
- Sarah H. Kim
- , Arnaud Da Cruz Paula
- & Britta Weigelt
-
Article
| Open Access A 5700 year-old human genome and oral microbiome from chewed birch pitch
A 5700 year-old human genome and oral microbiome from chewed birch pitch
Birch pitch is thought to have been used in prehistoric times as hafting material or antiseptic and tooth imprints suggest that it was chewed. Here, the authors report a 5,700 year-old piece of chewed birch pitch from Denmark from which they successfully recovered a complete ancient human genome and oral microbiome DNA.
- Theis Z. T. Jensen
- , Jonas Niemann
- & Hannes Schroeder
-
Article
| Open Access Single-nuclei RNA-seq on human retinal tissue provides improved transcriptome profiling
Single-nuclei RNA-seq on human retinal tissue provides improved transcriptome profiling
The retina is a heterogeneous tissue composed of multiple cell types. Via single-nuclei RNA sequencing on human neural retinal tissue, the authors characterise the transcriptome profile for individual cell types in the human retina.
- Qingnan Liang
- , Rachayata Dharmat
- & Rui Chen
-
Article
| Open Access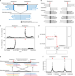 LinkedSV for detection of mosaic structural variants from linked-read exome and genome sequencing data
LinkedSV for detection of mosaic structural variants from linked-read exome and genome sequencing data
Compared to single nucleotide variants and short indels, structural variants (SVs) are often more challenging to detect using high-throughput sequencing based methods. Here, the authors develop LinkedSV, a computational tool for SV detection using linked-read exome and genome sequencing data.
- Li Fang
- , Charlly Kao
- & Kai Wang
-
Article
| Open Access Trio deep-sequencing does not reveal unexpected off-target and on-target mutations in Cas9-edited rhesus monkeys
Trio deep-sequencing does not reveal unexpected off-target and on-target mutations in Cas9-edited rhesus monkeys
The off-target effects and on-target mutations of CRISPR editing in higher non-human primates are complex. Here the authors perform whole genome trio sequencing of rhesus monkeys and see no unexpected mutations.
- Xin Luo
- , Yaoxi He
- & Bing Su
-
Article
| Open Access The art of using t-SNE for single-cell transcriptomics
The art of using t-SNE for single-cell transcriptomics
t-SNE is widely used for dimensionality reduction and visualization of high-dimensional single-cell data. Here, the authors introduce a protocol to help avoid common shortcomings of t-SNE, for example, enabling preservation of the global structure of the data.
- Dmitry Kobak
- & Philipp Berens
-
Article
| Open Access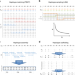 Accurate, scalable and integrative haplotype estimation
Accurate, scalable and integrative haplotype estimation
Haplotype information inferred by phasing is useful in genetic and genomic analysis. Here, the authors develop SHAPEIT4, a phasing method that exhibits sub-linear running time, provides accurate haplotypes and enables integration of external phasing information.
- Olivier Delaneau
- , Jean-François Zagury
- & Emmanouil T. Dermitzakis
-
Article
| Open Access GraphTyper2 enables population-scale genotyping of structural variation using pangenome graphs
GraphTyper2 enables population-scale genotyping of structural variation using pangenome graphs
Structural variants may be omitted in sequence analysis despite their importance in genome variation and phenotypic impact. Here the authors present GraphTyper2, which uses pangenome graphs to genotype structural variants using short-reads and can be applied in large-scale sequencing studies.
- Hannes P. Eggertsson
- , Snaedis Kristmundsdottir
- & Pall Melsted
-
Article
| Open Access Assembly of chromosome-scale contigs by efficiently resolving repetitive sequences with long reads
Assembly of chromosome-scale contigs by efficiently resolving repetitive sequences with long reads
Repetitive sequences in complex eukaryote genomes can cause fragmented assemblies with incomplete gene sequences and unanchored or mispositioned contigs. Here, the authors report HERA, a method to improve genome assemblies by efficiently resolving repeats using single-molecule sequencing data.
- Huilong Du
- & Chengzhi Liang
-
Article
| Open Access Poly(A) inclusive RNA isoform sequencing (PAIso−seq) reveals wide-spread non-adenosine residues within RNA poly(A) tails
Poly(A) inclusive RNA isoform sequencing (PAIso−seq) reveals wide-spread non-adenosine residues within RNA poly(A) tails
The poly(A) tails on mRNA are vital for their function but it is difficult to map full-length sequences of mRNA isoforms with the entire poly(A) tails. Here the authors develop PAIso−seq which can measure isoform specific poly(A) tail length and base composition at single-cell sensitivity.
- Yusheng Liu
- , Hu Nie
- & Falong Lu
-
Article
| Open Access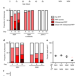 A fln-2 mutation affects lethal pathology and lifespan in C. elegans
A fln-2 mutation affects lethal pathology and lifespan in C. elegans
C. elegans is a commonly used model organism in the study of ageing, and differences in genetic background can result in varying strain longevity. Here the authors demonstrate that a background mutation in fln-2 affects life-limiting pharyngeal infection and that in the mutant background the beneficial effect of sir-2.1 over-expression is suppressed.
- Yuan Zhao
- , Hongyuan Wang
- & David Gems
-
Article
| Open Access Full-length transcriptome reconstruction reveals a large diversity of RNA and protein isoforms in rat hippocampus
Full-length transcriptome reconstruction reveals a large diversity of RNA and protein isoforms in rat hippocampus
It is challenging to characterize diverse transcript isoforms by short-read sequencing. Here the authors report full-length transcriptomes in rat hippocampus by hybrid-sequencing, predict isoform-specific translational status, and reconstruct open reading frames validated by mass spectrometry.
- Xi Wang
- , Xintian You
- & Wei Chen
-
Article
| Open Access CUTseq is a versatile method for preparing multiplexed DNA sequencing libraries from low-input samples
CUTseq is a versatile method for preparing multiplexed DNA sequencing libraries from low-input samples
Genomics DNA library preparation from formalin-fixed paraffin-embedded tissues is challenging. Here the authors describe CUTseq that uses restriction enzymes and in vitro amplification to barcode samples for reduced representation genome sequencing.
- Xiaolu Zhang
- , Silvano Garnerone
- & Nicola Crosetto
-
Article
| Open Access De novo and recessive forms of congenital heart disease have distinct genetic and phenotypic landscapes
De novo and recessive forms of congenital heart disease have distinct genetic and phenotypic landscapes
Large whole-exome sequencing studies have suggested that the genetic architecture of syndromic congenital heart disease (CHD) is different from sporadic forms. Here, Watkins et al. estimate the relative contribution of damaging recessive and de novo genotypes to CHD in 2391 trios and find them to be associated with different gene functions.
- W. Scott Watkins
- , E. Javier Hernandez
- & Martin Tristani-Firouzi
-
Article
| Open Access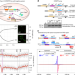 Detection of spacer precursors formed in vivo during primed CRISPR adaptation
Detection of spacer precursors formed in vivo during primed CRISPR adaptation
Primed adaptation in the CRISPR-Cas system helps recognition of previously encountered sequence elements and promotes the formation of new memories. Here the authors characterized spacer precursors of type I-E and type I-F CRISPR-Cas system using in vivo models.
- Anna A. Shiriaeva
- , Ekaterina Savitskaya
- & Ekaterina Semenova
-
Article
| Open Access Massively parallel RNA device engineering in mammalian cells with RNA-Seq
Massively parallel RNA device engineering in mammalian cells with RNA-Seq
Synthetic RNA-based devices can dynamically control a wide range of processes. Here the authors develop a quantitative and high-throughput mammalian cell-based RNA-seq assay to efficiently engineer ribozyme switches.
- Joy S. Xiang
- , Matias Kaplan
- & Christina D. Smolke
-
Article
| Open Access Linked-read sequencing of gametes allows efficient genome-wide analysis of meiotic recombination
Linked-read sequencing of gametes allows efficient genome-wide analysis of meiotic recombination
Meiotic crossovers (COs) generate genetic variation and ensure proper chromosome segregation. Here, the authors develop a method for identifying COs at kilobase resolution in pooled recombinants using linked-read sequencing data, and apply it to investigate genome-wide CO landscapes of Arabidopsis thaliana.
- Hequan Sun
- , Beth A. Rowan
- & Korbinian Schneeberger
-
Article
| Open Access Single-cell profiling guided combinatorial immunotherapy for fast-evolving CDK4/6 inhibitor-resistant HER2-positive breast cancer
Single-cell profiling guided combinatorial immunotherapy for fast-evolving CDK4/6 inhibitor-resistant HER2-positive breast cancer
The benefit of combined CDK4/6 and anti-HER2 therapy in breast cancer is limited due to acquired resistance. Here, the authors perform single-cell analysis and show an immature myeloid cell population to infiltrate resistant tumors, and that combined cabozantinib and checkpoint therapy overcome this resistance with a sustained efficacy.
- Qingfei Wang
- , Ian H. Guldner
- & Siyuan Zhang
-
Article
| Open Access Tuberculous meningitis in children is characterized by compartmentalized immune responses and neural excitotoxicity
Tuberculous meningitis in children is characterized by compartmentalized immune responses and neural excitotoxicity
Tuberculosis meningitis (TBM) is a severe form of TB with limited treatment options. Here, the authors perform RNA sequencing on whole blood and on ventricular and lumbar cerebrospinal fluid (CSF) samples from pediatric patients treated for TBM to characterize the immune response and tissue damage.
- Ursula K. Rohlwink
- , Anthony Figaji
- & Rachel P. J. Lai
-
Article
| Open Access Mitogenic and progenitor gene programmes in single pilocytic astrocytoma cells
Mitogenic and progenitor gene programmes in single pilocytic astrocytoma cells
Pilocytic astrocytoma is a low-grade pediatric glioma, characterized by a single BRAF rearrangement. Here, Reitman and colleagues use single-cell RNA sequencing to reveal molecular hallmarks of the disease that might be targeted therapeutically.
- Zachary J. Reitman
- , Brenton R. Paolella
- & Rameen Beroukhim
-
Article
| Open Access The pause-initiation limit restricts transcription activation in human cells
The pause-initiation limit restricts transcription activation in human cells
Gene activation requires an increase of successful initiation events. Here, by employing a genome-wide kinetic analysis of transcription, the authors showed that gene activation generally requires a decrease in RNA Polymerase II (Pol II) promoter-proximal pausing while transcription of enhancer elements is not limited by Pol II pausing.
- Saskia Gressel
- , Björn Schwalb
- & Patrick Cramer
-
Article
| Open Access Saturation mutagenesis of twenty disease-associated regulatory elements at single base-pair resolution
Saturation mutagenesis of twenty disease-associated regulatory elements at single base-pair resolution
Interpreting genetic variation in the noncoding genome remains challenging, with functional effects difficult to predict. Here, the authors perform saturation mutagenesis combined with massively parallel reporter assays for 20 disease-associated regulatory elements, quantifying the effects of over 30,000 variants.
- Martin Kircher
- , Chenling Xiong
- & Nadav Ahituv
-
Article
| Open Access A comprehensive examination of Nanopore native RNA sequencing for characterization of complex transcriptomes
A comprehensive examination of Nanopore native RNA sequencing for characterization of complex transcriptomes
The Oxford Nanopore system is capable of direct RNA sequencing but complex transcriptome analysis still remains to be thoroughly investigated. Here the authors perform sequencing of native RNA as well as cDNA to characterise the strengths and weaknesses of the system.
- Charlotte Soneson
- , Yao Yao
- & Shobbir Hussain
-
Article
| Open Access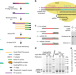 Single-cell CAS-seq reveals a class of short PIWI-interacting RNAs in human oocytes
Single-cell CAS-seq reveals a class of short PIWI-interacting RNAs in human oocytes
PIWI-interacting RNAs (piRNAs) are ~25–33 nt small RNAs expressed in animal germ cells. Here, the authors develop a single-cell small RNA sequencing method and report that a class of ~20-nt piRNAs lacking 3′ end 2′-O-methylation are associated with PIWIL3 protein and predominantly expressed in human and monkey oocytes.
- Qiyuan Yang
- , Ronghong Li
- & Ligang Wu
-
Article
| Open Access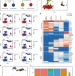 Single cell analysis of human foetal liver captures the transcriptional profile of hepatobiliary hybrid progenitors
Single cell analysis of human foetal liver captures the transcriptional profile of hepatobiliary hybrid progenitors
The liver parenchyma consists of several cell types, but the origin of this tissue in humans is unclear. Here, the authors perform single cell RNA sequencing of human fetal and adult liver to identify a hepatobiliary hybrid progenitor population of cells, which have a similar gene signature to mouse oval cells.
- Joe M. Segal
- , Deniz Kent
- & S. Tamir Rashid
-
Article
| Open Access FDA-ARGOS is a database with public quality-controlled reference genomes for diagnostic use and regulatory science
FDA-ARGOS is a database with public quality-controlled reference genomes for diagnostic use and regulatory science
To be able to use infectious disease next generation sequencing as a diagnostic tool, appropriate reference datasets are required. Here, Sichtig et al. describe FDA-ARGOS, a reference database for high-quality microbial reference genomes, and demonstrate its utility on the example of two use cases.
- Heike Sichtig
- , Timothy Minogue
- & Uwe Scherf
-
Article
| Open Access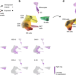 Predicting bacterial infection outcomes using single cell RNA-sequencing analysis of human immune cells
Predicting bacterial infection outcomes using single cell RNA-sequencing analysis of human immune cells
Complex interactions between different host immune cell types can determine the outcome of pathogen infections. Here, Avraham and colleagues present a deconvolution algorithm that uses single-cell RNA and bulk RNA sequencing measurements of pathogen-infected cells to predict disease risk outcomes.
- Noa Bossel Ben-Moshe
- , Shelly Hen-Avivi
- & Roi Avraham
-
Article
| Open Access NASC-seq monitors RNA synthesis in single cells
NASC-seq monitors RNA synthesis in single cells
Sequencing of newly synthesised RNA can reveal the transcriptional dynamics in a population of cells. Here the authors develop NASC-seq to bring this sensitivity and temporal resolution to single-cell analysis.
- Gert-Jan Hendriks
- , Lisa A. Jung
- & Rickard Sandberg
-
Article
| Open Access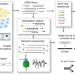 High-throughput targeted long-read single cell sequencing reveals the clonal and transcriptional landscape of lymphocytes
High-throughput targeted long-read single cell sequencing reveals the clonal and transcriptional landscape of lymphocytes
Single cell RNA sequencing generates short reads from one end of a template, providing incomplete transcript coverage and limiting identification of diverse sequences such as antigen receptors. Here the authors combine long read nanopore sequencing with short read profiling of barcoded libraries to generate full-length antigen receptor sequences.
- Mandeep Singh
- , Ghamdan Al-Eryani
- & Alexander Swarbrick
-
Article
| Open Access AMPA receptor GluA2 subunit defects are a cause of neurodevelopmental disorders
AMPA receptor GluA2 subunit defects are a cause of neurodevelopmental disorders
Genetic variants in ionotropic glutamate receptors have been implicated in neurodevelopmental disorders. Here, the authors report heterozygous de novo mutations in the GRIA2 gene in 28 individuals with intellectual disability and neurodevelopmental abnormalities associated with reduced Ca2+ transport and AMPAR currents.”
- Vincenzo Salpietro
- , Christine L. Dixon
- & Henry Houlden
-
Article
| Open Access A genome-wide scan statistic framework for whole-genome sequence data analysis
A genome-wide scan statistic framework for whole-genome sequence data analysis
Whole-genome sequencing data reveals a large number of variants for testing their associations with phenotypic traits and diseases. Here, the authors develop WGScan, a statistical method for detecting the existence and estimating the locations of the association signal at genome-wide scale.
- Zihuai He
- , Bin Xu
- & Iuliana Ionita-Laza
-
Article
| Open Access DNA assembly for nanopore data storage readout
DNA assembly for nanopore data storage readout
A major bottleneck in using DNA as a data storage medium is the slowness of sequencing. Here the authors decode 1.67 megabytes of information using a portable nanopore platform with an assembly strategy for increased throughput.
- Randolph Lopez
- , Yuan-Jyue Chen
- & Luis Ceze
-
Article
| Open Access Quantitative proteomics and single-nucleus transcriptomics of the sinus node elucidates the foundation of cardiac pacemaking
Quantitative proteomics and single-nucleus transcriptomics of the sinus node elucidates the foundation of cardiac pacemaking
The sinus node generates rhythmic heartbeat but the molecular basis of pacemaking is still under debate. Here, the authors combine quantitative proteomics and single-nucleus transcriptomics to characterize the molecular composition of the sinus node and provide insights into the underpinnings of pacemaking.
- Nora Linscheid
- , Sunil Jit R. J. Logantha
- & Alicia Lundby
-
Article
| Open Access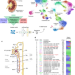 A single-nucleus RNA-sequencing pipeline to decipher the molecular anatomy and pathophysiology of human kidneys
A single-nucleus RNA-sequencing pipeline to decipher the molecular anatomy and pathophysiology of human kidneys
Single-cell studies in solid tissues remain challenging and have benefited from the development of single-nuclei RNA sequencing strategies. Here Lake et al. apply single-nucleus RNA sequencing to human kidney tissues to provide a comprehensive molecular and cellular atlas of the human kidney, with potential implications for the understanding of kidney physiology and disease.
- Blue B. Lake
- , Song Chen
- & Sanjay Jain
-
Article
| Open Access Common and distinct transcriptional signatures of mammalian embryonic lethality
Common and distinct transcriptional signatures of mammalian embryonic lethality
The transcriptional signature of embryonic lethality has not been defined. Here, the authors, as part of the Deciphering the Mechanisms of Developmental Disorders programme, define genes causing murine embryonic lethality around E9.5 and identify developmental delay transcriptional signatures.
- John E. Collins
- , Richard J. White
- & Elisabeth M. Busch-Nentwich
-
Article
| Open Access HIV-1 DNA sequence diversity and evolution during acute subtype C infection
HIV-1 DNA sequence diversity and evolution during acute subtype C infection
The dynamics of HIV-1 DNA sequences early after HIV-1 transmission remains poorly characterized. Here, the authors perform a longitudinal evaluation of HIV-1 DNA sequences in subtype C-infected individuals during acute infection, providing a landscape of the nature and evolution of the very early viral genome.
- Guinevere Q. Lee
- , Kavidha Reddy
- & Mathias Lichterfeld
-
Article
| Open Access Simulating multiple faceted variability in single cell RNA sequencing
Simulating multiple faceted variability in single cell RNA sequencing
Simulated single cell RNA sequencing data is useful for method development and comparison. Here, the authors developed SymSim, a simulator that explicitly models the main factors of variation in single cell data.
- Xiuwei Zhang
- , Chenling Xu
- & Nir Yosef
-
Article
| Open Access Comprehensive analysis of coding variants highlights genetic complexity in developmental and epileptic encephalopathy
Comprehensive analysis of coding variants highlights genetic complexity in developmental and epileptic encephalopathy
Many causative genes are known for epileptic or developmental and epileptic encephalopathies (EE/DEE) yet a genetic diagnosis cannot be made for many patients. Here, the authors analyse whole exome sequencing data from a Japanese case−control cohort to identify common, rare and ultra-rare coding variants associated with EE/DEE.
- Atsushi Takata
- , Mitsuko Nakashima
- & Naomichi Matsumoto
-
Article
| Open Access qDSB-Seq is a general method for genome-wide quantification of DNA double-strand breaks using sequencing
qDSB-Seq is a general method for genome-wide quantification of DNA double-strand breaks using sequencing
Measuring relative frequencies of DNA double-strand breaks between loci does not provide the full physiological relevance of those breaks. Here Rowicka and colleagues present qDSB-Seq method which uses spike-in double-strand breaks induced by a restriction enzyme to accurately quantify DNA damage.
- Yingjie Zhu
- , Anna Biernacka
- & Maga Rowicka
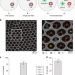
 Inference and effects of barcode multiplets in droplet-based single-cell assays
Inference and effects of barcode multiplets in droplet-based single-cell assays