Article
|
Open Access
Featured
-
-
Article
| Open Access A genome-wide scan statistic framework for whole-genome sequence data analysis
A genome-wide scan statistic framework for whole-genome sequence data analysis
Whole-genome sequencing data reveals a large number of variants for testing their associations with phenotypic traits and diseases. Here, the authors develop WGScan, a statistical method for detecting the existence and estimating the locations of the association signal at genome-wide scale.
- Zihuai He
- , Bin Xu
- & Iuliana Ionita-Laza
-
Article
| Open Access DNA assembly for nanopore data storage readout
DNA assembly for nanopore data storage readout
A major bottleneck in using DNA as a data storage medium is the slowness of sequencing. Here the authors decode 1.67 megabytes of information using a portable nanopore platform with an assembly strategy for increased throughput.
- Randolph Lopez
- , Yuan-Jyue Chen
- & Luis Ceze
-
Article
| Open Access Quantitative proteomics and single-nucleus transcriptomics of the sinus node elucidates the foundation of cardiac pacemaking
Quantitative proteomics and single-nucleus transcriptomics of the sinus node elucidates the foundation of cardiac pacemaking
The sinus node generates rhythmic heartbeat but the molecular basis of pacemaking is still under debate. Here, the authors combine quantitative proteomics and single-nucleus transcriptomics to characterize the molecular composition of the sinus node and provide insights into the underpinnings of pacemaking.
- Nora Linscheid
- , Sunil Jit R. J. Logantha
- & Alicia Lundby
-
Article
| Open Access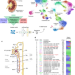 A single-nucleus RNA-sequencing pipeline to decipher the molecular anatomy and pathophysiology of human kidneys
A single-nucleus RNA-sequencing pipeline to decipher the molecular anatomy and pathophysiology of human kidneys
Single-cell studies in solid tissues remain challenging and have benefited from the development of single-nuclei RNA sequencing strategies. Here Lake et al. apply single-nucleus RNA sequencing to human kidney tissues to provide a comprehensive molecular and cellular atlas of the human kidney, with potential implications for the understanding of kidney physiology and disease.
- Blue B. Lake
- , Song Chen
- & Sanjay Jain
-
Article
| Open Access Common and distinct transcriptional signatures of mammalian embryonic lethality
Common and distinct transcriptional signatures of mammalian embryonic lethality
The transcriptional signature of embryonic lethality has not been defined. Here, the authors, as part of the Deciphering the Mechanisms of Developmental Disorders programme, define genes causing murine embryonic lethality around E9.5 and identify developmental delay transcriptional signatures.
- John E. Collins
- , Richard J. White
- & Elisabeth M. Busch-Nentwich
-
Article
| Open Access HIV-1 DNA sequence diversity and evolution during acute subtype C infection
HIV-1 DNA sequence diversity and evolution during acute subtype C infection
The dynamics of HIV-1 DNA sequences early after HIV-1 transmission remains poorly characterized. Here, the authors perform a longitudinal evaluation of HIV-1 DNA sequences in subtype C-infected individuals during acute infection, providing a landscape of the nature and evolution of the very early viral genome.
- Guinevere Q. Lee
- , Kavidha Reddy
- & Mathias Lichterfeld
-
Article
| Open Access Simulating multiple faceted variability in single cell RNA sequencing
Simulating multiple faceted variability in single cell RNA sequencing
Simulated single cell RNA sequencing data is useful for method development and comparison. Here, the authors developed SymSim, a simulator that explicitly models the main factors of variation in single cell data.
- Xiuwei Zhang
- , Chenling Xu
- & Nir Yosef
-
Article
| Open Access Comprehensive analysis of coding variants highlights genetic complexity in developmental and epileptic encephalopathy
Comprehensive analysis of coding variants highlights genetic complexity in developmental and epileptic encephalopathy
Many causative genes are known for epileptic or developmental and epileptic encephalopathies (EE/DEE) yet a genetic diagnosis cannot be made for many patients. Here, the authors analyse whole exome sequencing data from a Japanese case−control cohort to identify common, rare and ultra-rare coding variants associated with EE/DEE.
- Atsushi Takata
- , Mitsuko Nakashima
- & Naomichi Matsumoto
-
Article
| Open Access qDSB-Seq is a general method for genome-wide quantification of DNA double-strand breaks using sequencing
qDSB-Seq is a general method for genome-wide quantification of DNA double-strand breaks using sequencing
Measuring relative frequencies of DNA double-strand breaks between loci does not provide the full physiological relevance of those breaks. Here Rowicka and colleagues present qDSB-Seq method which uses spike-in double-strand breaks induced by a restriction enzyme to accurately quantify DNA damage.
- Yingjie Zhu
- , Anna Biernacka
- & Maga Rowicka
-
Article
| Open Access Complete deconvolution of cellular mixtures based on linearity of transcriptional signatures
Complete deconvolution of cellular mixtures based on linearity of transcriptional signatures
Complete gene expression deconvolution remains a challenging problem. Here, the authors provide a solution based on the recognition that expression levels of cell type specific genes are mutually linear across mixtures and mutually linear gene clusters correspond to cell type-specific signatures.
- Konstantin Zaitsev
- , Monika Bambouskova
- & Maxim N. Artyomov
-
Article
| Open Access Hydro-Seq enables contamination-free high-throughput single-cell RNA-sequencing for circulating tumor cells
Hydro-Seq enables contamination-free high-throughput single-cell RNA-sequencing for circulating tumor cells
Transcriptome analysis of circulating tumor cells (CTCs) provides insights into monitoring target therapeutics and underlying tumor metastasis. Here the authors present Hydro-Seq, a contamination-free high-throughput hydrodynamic scRNA-seq barcoding technique for rare CTCs.
- Yu-Heng Cheng
- , Yu-Chih Chen
- & Euisik Yoon
-
Article
| Open Access Accurate high throughput alignment via line sweep-based seed processing
Accurate high throughput alignment via line sweep-based seed processing
Alignment is an important stage in the analysis of sequencing data. Here, the authors present a fast and accurate alignment approach suitable for long and short reads, and introduce two line sweep-based techniques which can replace the often-used chaining approach.
- Markus Schmidt
- , Klaus Heese
- & Arne Kutzner
-
Article
| Open Access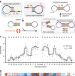 High-resolution specificity profiling and off-target prediction for site-specific DNA recombinases
High-resolution specificity profiling and off-target prediction for site-specific DNA recombinases
The development of site-specific recombinases as genome editing tools is limited by the difficulty of altering their DNA sequence specificity. Here the authors present Rec-seq, a method for identifying specificity determinants and off-target substrates of recombinases in an unbiased manner.
- Jeffrey L. Bessen
- , Lena K. Afeyan
- & David R. Liu
-
Article
| Open Access Single-cell trajectories reconstruction, exploration and mapping of omics data with STREAM
Single-cell trajectories reconstruction, exploration and mapping of omics data with STREAM
The increasing accessibility of single cell omics technologies beyond transcriptomics demands parallel advances in analysis. Here, the authors introduce STREAM, a pipeline for reconstruction and visualization of differentiation trajectories from both single-cell RNA-seq and ATAC-seq data.
- Huidong Chen
- , Luca Albergante
- & Luca Pinello
-
Article
| Open Access Multi-platform discovery of haplotype-resolved structural variation in human genomes
Multi-platform discovery of haplotype-resolved structural variation in human genomes
Structural variants (SVs) in human genomes contribute diversity and diseases. Here, the authors use a multi-platform strategy to generate haplotype-resolved SVs for three human parent–child trios.
- Mark J. P. Chaisson
- , Ashley D. Sanders
- & Charles Lee
-
Article
| Open Access Decoupling tRNA promoter and processing activities enables specific Pol-II Cas9 guide RNA expression
Decoupling tRNA promoter and processing activities enables specific Pol-II Cas9 guide RNA expression
The utility of CRISPR-based technologies could be enhanced with the ability to control the spatial and temporal expression of gRNAs. Here the authors design a tRNA scaffold for highly specific gRNA production from a Pol II promoter.
- David J. H. F. Knapp
- , Yale S. Michaels
- & Tudor A. Fulga
-
Article
| Open Access Identification of human D lactate dehydrogenase deficiency
Identification of human D lactate dehydrogenase deficiency
D-lactic acidosis typically occurs in the context of short bowel syndrome; excess D-lactate is produced by intestinal bacteria. Here, the authors identify two point mutations in the human lactate dehydrogenase D (LDHD) gene that cause enzymatic loss of function and are associated with elevated plasma D-lactate.
- Glen R. Monroe
- , Albertien M. van Eerde
- & Judith J. Jans
-
Article
| Open Access A study of the extraordinarily strong and tough silk produced by bagworms
A study of the extraordinarily strong and tough silk produced by bagworms
Spider silk is widely studied for its structural properties; however, other creatures produce silk that could be of interest. Here, the authors study the properties and structure of Bagworm silk and report it as being extraordinarily strong and tough compared to other known silks.
- Taiyo Yoshioka
- , Takuya Tsubota
- & Tsunenori Kameda
-
Article
| Open Access Diagnosis of fusion genes using targeted RNA sequencing
Diagnosis of fusion genes using targeted RNA sequencing
Rapid and accurate detection of fusion genes is important in cancer diagnostics. Here, the authors demonstrate that targeted RNA sequencing provides fast, sensitive and quantitative gene fusion detection and overcomes the limitations of approaches currently in clinical use.
- Erin E. Heyer
- , Ira W. Deveson
- & James Blackburn
-
Article
| Open Access Chiral DNA sequences as commutable controls for clinical genomics
Chiral DNA sequences as commutable controls for clinical genomics
Any DNA sequence can be represented by a chiral partner sequence – an exact copy arranged in reverse nucleotide order. Here, the authors show that chiral DNA sequence pairs share important properties and show the utility of synthetic chiral sequences (sequins) as controls for clinical genomics.
- Ira W. Deveson
- , Bindu Swapna Madala
- & Tim R. Mercer
-
Article
| Open Access Whole-genome resequencing of 472 Vitis accessions for grapevine diversity and demographic history analyses
Whole-genome resequencing of 472 Vitis accessions for grapevine diversity and demographic history analyses
Despite the importance of grapevine cultivation in human history and the economic values of cultivar improvement, large-scale genomic variation data are lacking. Here the authors resequence 472 Vitis accessions and use the identified genetic variations for domestication history, demography, and GWAS analyses.
- Zhenchang Liang
- , Shengchang Duan
- & Yang Dong
-
Article
| Open Access The use of technical replication for detection of low-level somatic mutations in next-generation sequencing
The use of technical replication for detection of low-level somatic mutations in next-generation sequencing
Somatic mutations of low allele frequencies are often difficult to detect. Here, the authors develop RePlow, a computational method that leverages technical replication for detecting low-level somatic mutations using next-generation sequencing.
- Junho Kim
- , Dachan Kim
- & Sangwoo Kim
-
Article
| Open Access Single gametophyte sequencing reveals that crossover events differ between sexes in maize
Single gametophyte sequencing reveals that crossover events differ between sexes in maize
Meiotic crossover (CO) landscape differs inter- and intra-species, as well as between sexes. Here, the authors show that male meiosis produces more COs than female in maize and detect CO maturation inefficiency in some genetic backgrounds, which may help to improve breeding efficiency.
- Cheng Luo
- , Xiang Li
- & Jianbing Yan
-
Article
| Open Access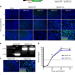 Non-lytic clearance of influenza B virus from infected cells preserves epithelial barrier function
Non-lytic clearance of influenza B virus from infected cells preserves epithelial barrier function
Infection of a cell with influenza B virus (IBV) often results in cell death and the role of surviving cells in pathogenesis is unclear. Here, Dumm et al. generate a recombinant IBV that activates a host-cell reporter to permanently label infected cells, and show that surviving cells are important to preserve epithelial barrier function.
- Rebekah E. Dumm
- , Jessica K. Fiege
- & Nicholas S. Heaton
-
Article
| Open Access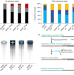 Direct RNA sequencing on nanopore arrays redefines the transcriptional complexity of a viral pathogen
Direct RNA sequencing on nanopore arrays redefines the transcriptional complexity of a viral pathogen
Here, Depledge et al. use nanopore arrays for direct RNA sequencing to profile the HSV-1 transcriptome in productively infected cells. Sequencing of individual RNAs reveals a highly complex viral transcriptome including mRNAs encoding new viral fusion proteins derived by read-through transcription.
- Daniel P. Depledge
- , Kalanghad Puthankalam Srinivas
- & Angus C. Wilson
-
Article
| Open Access Dual RNA-seq identifies human mucosal immunity protein Mucin-13 as a hallmark of Plasmodium exoerythrocytic infection
Dual RNA-seq identifies human mucosal immunity protein Mucin-13 as a hallmark of Plasmodium exoerythrocytic infection
Host-parasite interactions during the exoerythrocytic stage of Plasmodium infection remains poorly understood. Using dual RNA-Seq, the authors show that human mucosal immunity protein mucin-13 is upregulated during Plasmodium hepatic-stage infection and marks infected cells independent of tested Plasmodium species.
- Gregory M. LaMonte
- , Pamela Orjuela-Sanchez
- & Elizabeth A. Winzeler
-
Article
| Open Access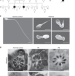 Loss-of-function mutations in QRICH2 cause male infertility with multiple morphological abnormalities of the sperm flagella
Loss-of-function mutations in QRICH2 cause male infertility with multiple morphological abnormalities of the sperm flagella
Multiple morphological abnormalities of the sperm flagella (MMAF) is a cause of male infertility. Here the authors identify homozygous nonsense mutations of the glutamine rich 2 (QRICH2) gene in two MMAF patients from 2 consanguineous families and show using QRICH2 knockout mice that the protein is required for sperm flagellar formation and motility.
- Ying Shen
- , Feng Zhang
- & Wenming Xu
-
Article
| Open Access C1 CAGE detects transcription start sites and enhancer activity at single-cell resolution
C1 CAGE detects transcription start sites and enhancer activity at single-cell resolution
Single-cell transcriptomic profiling allows the exploration of cellular heterogeneity but commonly focuses on the 3′-end of the transcript. Here the authors introduce C1 CAGE, which detects the 5′ transcript end in a multiplexed microfluidic system.
- Tsukasa Kouno
- , Jonathan Moody
- & Jay W. Shin
-
Article
| Open Access Biological relevance of computationally predicted pathogenicity of noncoding variants
Biological relevance of computationally predicted pathogenicity of noncoding variants
Researchers can make use of a variety of computational tools to prioritize genetic variants and predict their pathogenicity. Here, the authors evaluate the performance of six of these tools in three typical biological tasks and find generally low concordance of predictions and experimental confirmation.
- Li Liu
- , Maxwell D. Sanderford
- & Sudhir Kumar
-
Article
| Open Access Chromosome-level assembly of the water buffalo genome surpasses human and goat genomes in sequence contiguity
Chromosome-level assembly of the water buffalo genome surpasses human and goat genomes in sequence contiguity
Despite technological advances, chromosome-level assemblies of mammalian genomes are still rare. Here, the authors use PacBio, Chicago and Hi-C approaches to generate a highly contiguous and partially-phased genome assembly for the water buffalo, Bubalus bubalis
- Wai Yee Low
- , Rick Tearle
- & John L. Williams
-
Article
| Open Access Single-cell microRNA-mRNA co-sequencing reveals non-genetic heterogeneity and mechanisms of microRNA regulation
Single-cell microRNA-mRNA co-sequencing reveals non-genetic heterogeneity and mechanisms of microRNA regulation
Single-cell RNA sequencing allows characterizing cell-to-cell heterogeneity at transcriptome scale. Here, the authors present an approach that enables microRNA and mRNA sequencing in the same single cell, providing insights into the origins of non-genetic cellular variability and mechanisms of miRNA regulation.
- Nayi Wang
- , Ji Zheng
- & Jun Lu
-
Article
| Open Access Genome-wide profiling of adenine base editor specificity by EndoV-seq
Genome-wide profiling of adenine base editor specificity by EndoV-seq
Adenine base editors are an important contribution to the genome editing toolbox. Here the authors present EndoV-seq, an endonuclease-based assay for evaluating genomewide off-target effects of base editing.
- Puping Liang
- , Xiaowei Xie
- & Zhou Songyang
-
Article
| Open Access Microbiome characterization by high-throughput transfer RNA sequencing and modification analysis
Microbiome characterization by high-throughput transfer RNA sequencing and modification analysis
The authors present a molecular approach and a software suite for high-throughput sequencing and characterization of microbial tRNA transcripts, which provide unique physiological insights into complex environmental microbes.
- Michael H. Schwartz
- , Haipeng Wang
- & A. Murat Eren
-
Article
| Open Access Chromatin conformation analysis of primary patient tissue using a low input Hi-C method
Chromatin conformation analysis of primary patient tissue using a low input Hi-C method
Chromatin conformation studies are limited by the large amounts of starting material required to perform current protocols. Here the authors present Low-C, a Hi-C method for low amounts of input material and produce Low-C maps from primary B-cells of a diffuse large B-cell lymphoma patient, demonstrating the suitability of Low-C to analyse rare cell populations.
- Noelia Díaz
- , Kai Kruse
- & Juan M. Vaquerizas
-
Article
| Open Access Expansive microbial metabolic versatility and biodiversity in dynamic Guaymas Basin hydrothermal sediments
Expansive microbial metabolic versatility and biodiversity in dynamic Guaymas Basin hydrothermal sediments
The diversity and function of microbes inhabiting hydrothermal areas is a topic of active interest in marine microbiology. Here, the authors assemble genomes from Guaymas Basin hydrothermal sediments and describe the metabolic roles of the bacterial community, which includes five new bacterial candidate phyla
- Nina Dombrowski
- , Andreas P. Teske
- & Brett J. Baker
-
Article
| Open Access Single cell RNA-sequencing identifies a metabolic aspect of apoptosis in Rbf mutant
Single cell RNA-sequencing identifies a metabolic aspect of apoptosis in Rbf mutant
The function of the Retinoblastoma (Rb) protein is regulated by its cellular environment. Here, the authors perform single cell RNA-sequencing during Drosophila eye development and identify the impact of an Rbf mutation, which sensitises specific cells to apoptosis by changing metabolism.
- Majd M. Ariss
- , Abul B. M. M. K. Islam
- & Maxim V. Frolov
-
Article
| Open Access Ancient Fennoscandian genomes reveal origin and spread of Siberian ancestry in Europe
Ancient Fennoscandian genomes reveal origin and spread of Siberian ancestry in Europe
Populations from North-eastern Europe, in particular those speaking Uralic languages, carry additional ancestry in similarity with modern East Asian populations. Here, the authors analyse ancient genomic data from 11 individuals from Finland and Northwest Russia, and identify genomic signals of migrations from Siberia that began at least 3500 years ago.
- Thiseas C. Lamnidis
- , Kerttu Majander
- & Stephan Schiffels
-
Article
| Open Access Defining human cardiac transcription factor hierarchies using integrated single-cell heterogeneity analysis
Defining human cardiac transcription factor hierarchies using integrated single-cell heterogeneity analysis
Human induced pluripotent stem cell derived cardiomyocytes are a powerful model for cardiogenesis and disease in vitro. Here the authors comprehensively map cardiac differentiation using multiple modalities, including single-cell RNA seq and CyTOF, in cells with a gain or loss of function in key cardiac transcription factors.
- Jared M. Churko
- , Priyanka Garg
- & Joseph C. Wu
-
Article
| Open Access A chromosome-scale assembly of the sorghum genome using nanopore sequencing and optical mapping
A chromosome-scale assembly of the sorghum genome using nanopore sequencing and optical mapping
Assembly of large, repeat-rich eukaryotic genomes remains challenging. Here, the authors use BioNano Genomics DLS optical mapping and single-molecule nanopore sequencing to generate a chromosome-scale assembly of a new Sorghum bicolor accession and identify variation compared to the publicly available S. bicolor genome.
- Stéphane Deschamps
- , Yun Zhang
- & Haining Lin
-
Article
| Open Access Cohort-wide deep whole genome sequencing and the allelic architecture of complex traits
Cohort-wide deep whole genome sequencing and the allelic architecture of complex traits
Rare genetic variants can contribute to complex traits but this contribution is not well understood. Here, the authors analyse deep whole genome sequencing data across 1457 individuals from an isolated Greek population and find association of rare variant burdens with cardiometabolic traits.
- Arthur Gilly
- , Daniel Suveges
- & Eleftheria Zeggini
-
Article
| Open Access Joint single-cell DNA accessibility and protein epitope profiling reveals environmental regulation of epigenomic heterogeneity
Joint single-cell DNA accessibility and protein epitope profiling reveals environmental regulation of epigenomic heterogeneity
Cellular heterogeneity in cancer is complex and difficult to study. Here, the authors introduce Protein-indexed Assay of Transposase Accessible Chromatin (Pi-ATAC), which combines single cell chromatin and proteomic profiling to provide deep insight into the tumor microenvironment, and reveal the role of hypoxia in shaping the regulome of a subset of breast cancer cells in vivo.
- Xingqi Chen
- , Ulrike M. Litzenburger
- & Howard Y. Chang
-
Article
| Open Access Cardiomyocyte gene programs encoding morphological and functional signatures in cardiac hypertrophy and failure
Cardiomyocyte gene programs encoding morphological and functional signatures in cardiac hypertrophy and failure
The mechanisms underlying the transition from cardiac hypertrophy to heart failure following pressure overload are incompletely understood. Here the authors identify the gene programs encoding the morphological and functional characteristics of cardiomyocytes during the transition from early hypertrophy to heart failure via single-cell transcriptomics, establishing a key role for p53 signalling.
- Seitaro Nomura
- , Masahiro Satoh
- & Issei Komuro
-
Article
| Open Access Single cell RNA sequencing of human liver reveals distinct intrahepatic macrophage populations
Single cell RNA sequencing of human liver reveals distinct intrahepatic macrophage populations
The development of single cell RNA sequencing technologies has been instrumental in advancing our understanding of tissue biology. Here, MacParland et al. performed single cell RNA sequencing of human liver samples, and identify distinct populations of intrahepatic macrophages that may play specific roles in liver disease.
- Sonya A. MacParland
- , Jeff C. Liu
- & Ian D. McGilvray
-
Article
| Open Access Germline pathogenic variants of 11 breast cancer genes in 7,051 Japanese patients and 11,241 controls
Germline pathogenic variants of 11 breast cancer genes in 7,051 Japanese patients and 11,241 controls
Association between variants in 11 different genes and breast cancer risk has been established and sequencing of these genes is recommended to provide personalized diagnosis, therapy, and surveillance for the high-risk patients and their relatives. Here the authors analyse the frequency of germline pathogenic mutations in these genes specifically in a Japanese population.
- Yukihide Momozawa
- , Yusuke Iwasaki
- & Michiaki Kubo
-
Article
| Open Access Functional equivalence of genome sequencing analysis pipelines enables harmonized variant calling across human genetics projects
Functional equivalence of genome sequencing analysis pipelines enables harmonized variant calling across human genetics projects
Sharing of whole genome sequencing (WGS) data improves study scale and power, but data from different groups are often incompatible. Here, US genome centers and NIH programs define WGS data processing standards and a flexible validation method, facilitating collaboration in human genetics research.
- Allison A. Regier
- , Yossi Farjoun
- & Ira M. Hall
-
Article
| Open Access Mapping protein selectivity landscapes using multi-target selective screening and next-generation sequencing of combinatorial libraries
Mapping protein selectivity landscapes using multi-target selective screening and next-generation sequencing of combinatorial libraries
Characterizing the binding selectivity landscape of interacting proteins is crucial in protein engineering. Here the authors use multi-target selective library screening and in silico next-generation sequencing to map the binding landscape of proteins and produce improved proteases inhibitors.
- Si Naftaly
- , Itay Cohen
- & Niv Papo
-
Article
| Open Access Robust single-cell DNA methylome profiling with snmC-seq2
Robust single-cell DNA methylome profiling with snmC-seq2
Single-cell DNA methylome profiling allows the study of epigenomic heterogeneity in tissues but has been impeded by library quality. Here the authors demonstrate snmC-seq2 which improves mapping, throughput and library complexity.
- Chongyuan Luo
- , Angeline Rivkin
- & Joseph R. Ecker
-
Article
| Open Access High-throughput chromatin accessibility profiling at single-cell resolution
High-throughput chromatin accessibility profiling at single-cell resolution
Single-cell chromatin accessibility is a promising means to identify regulatory programs in mixtures of cells. Here the authors describe µ-ATAC-seq, a low-cost method that can generate thousands of accessibility profiles per day.
- Anja Mezger
- , Sandy Klemm
- & William Greenleaf
-
Article
| Open Access Endocrine lineage biases arise in temporally distinct endocrine progenitors during pancreatic morphogenesis
Endocrine lineage biases arise in temporally distinct endocrine progenitors during pancreatic morphogenesis
Endocrine progenitors form early in pancreatic development but the diversity of this cell population is unclear. Here, the authors use single cell RNA sequencing of the mouse pancreas at e14.5 and e16.5 to show that endocrine progenitors are temporally distinct and those formed later are more likely to become beta cells
- Marissa A. Scavuzzo
- , Matthew C. Hill
- & Malgorzata Borowiak

 AMPA receptor GluA2 subunit defects are a cause of neurodevelopmental disorders
AMPA receptor GluA2 subunit defects are a cause of neurodevelopmental disorders