Featured
-
-
Article
| Open Access Loss of the RNA-binding protein TACO1 causes late-onset mitochondrial dysfunction in mice
Loss of the RNA-binding protein TACO1 causes late-onset mitochondrial dysfunction in mice
Mutations in the translational activator of cytochrome c oxidase subunit I (TACO1) causes cytochrome c oxidase deficiency and Leigh Syndrome in patients. Here, the authors characterize mice with a mutation that causes lack of TACO1 expression, identifying a mouse model that could be useful for preclinical trials.
- Tara R. Richman
- , Henrik Spåhr
- & Aleksandra Filipovska
-
Article
| Open Access Pre-40S ribosome biogenesis factor Tsr1 is an inactive structural mimic of translational GTPases
Pre-40S ribosome biogenesis factor Tsr1 is an inactive structural mimic of translational GTPases
Tsr1 is an essential ribosome biogenesis factor that has known similarity to GTPases. Here, the authors report the Tsr1 crystal structure and show that it is similar to GTPases but that active site residues are not conserved; modelling of the structure into the pre-40S maps allows inferences on ribosomal maturation to be drawn.
- Urszula M. McCaughan
- , Uma Jayachandran
- & Atlanta G. Cook
-
Article
| Open Access Genome-wide assessment of differential translations with ribosome profiling data
Genome-wide assessment of differential translations with ribosome profiling data
The global measurement of ribosome occupancy on mRNAs is commonly used as a proxy in estimating rates of protein synthesis. Here the authors describe Xtail, a computational approach that facilitates the extraction of accurate quantitative insight from ribosome profiling data (Ribo-Seq).
- Zhengtao Xiao
- , Qin Zou
- & Xuerui Yang
-
Article
| Open Access Steric interactions lead to collective tilting motion in the ribosome during mRNA–tRNA translocation
Steric interactions lead to collective tilting motion in the ribosome during mRNA–tRNA translocation
During protein elongation, the translocation of mRNA and tRNA molecules across the 30S ribosomal subunit is associated with large-scale motions of the 30S head domain. Here the authors carry out MD simulations to probe the associated steric interactions and identify novel tilting motions during the late stages of translocation.
- Kien Nguyen
- & Paul C. Whitford
-
Article
| Open Access Structures of the E. coli translating ribosome with SRP and its receptor and with the translocon
Structures of the E. coli translating ribosome with SRP and its receptor and with the translocon
The co-translational insertion of proteins into membranes requires interaction between a ribosome-bound signal recognition particle (SRP) and a membrane-bound translocon. Here the authors use cryo-EM and single particle reconstructions to obtain a comprehensive view of the co-translational protein targeting process.
- Ahmad Jomaa
- , Daniel Boehringer
- & Nenad Ban
-
Article
| Open Access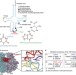 Novel base-pairing interactions at the tRNA wobble position crucial for accurate reading of the genetic code
Novel base-pairing interactions at the tRNA wobble position crucial for accurate reading of the genetic code
The anticodon loops of almost all tRNAs contain modifications known to be important for their function. Here the authors use crystallography to provide new mechanistic insights into how the modification at the wobble position of the E. coli tRNALysUUUassists in discrimination between cognate and near-cognate codons.
- Alexey Rozov
- , Natalia Demeshkina
- & Gulnara Yusupova
-
Article
| Open Access Mammalian SRP receptor switches the Sec61 translocase from Sec62 to SRP-dependent translocation
Mammalian SRP receptor switches the Sec61 translocase from Sec62 to SRP-dependent translocation
Sec62 is a membrane-bound protein that is involved in the translocation of proteins via the signal recognition particle-independent pathway. Here, the authors show that the receptor SRα displaces Sec62 from the translocon and isolate the domain on SRα that is responsible for this.
- Bhalchandra Jadhav
- , Michael McKenna
- & Martin R. Pool
-
Article
| Open Access Cryo-EM structure of Hepatitis C virus IRES bound to the human ribosome at 3.9-Å resolution
Cryo-EM structure of Hepatitis C virus IRES bound to the human ribosome at 3.9-Å resolution
The Hepatitis C virus (HCV) relies on an internal ribosome entry site (IRES) for translation of all the proteins encoded by its single-stranded RNA genome. Here the authors present a near-atomic cryo-EM structure of the HCV IRES bound to the human ribosome, shedding light on the initiation mechanism of HCV's and related IRESs.
- Nick Quade
- , Daniel Boehringer
- & Nenad Ban
-
Article
| Open Access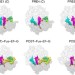 Fluctuations between multiple EF-G-induced chimeric tRNA states during translocation on the ribosome
Fluctuations between multiple EF-G-induced chimeric tRNA states during translocation on the ribosome
EF-G enhances the rate of tRNA–mRNA translocation on the ribosome. Here the authors use single-molecule FRET to follow tRNA translocation in real time, identifying new chimeric intermediates and suggesting how EF-G binding and GTP hydrolysis change the energetic landscape of translocation to accelerate forward tRNA movement.
- Sarah Adio
- , Tamara Senyushkina
- & Marina V. Rodnina
-
Article
| Open Access Structural insights into the translational infidelity mechanism
Structural insights into the translational infidelity mechanism
Translation of mRNA into proteins is the least accurate process during genetic information transfer. Here the authors suggest—based on 11 high-resolution ribosome crystal structures—that the origin of protein missense errors involves molecular mimicry via tautomerism or ionization.
- Alexey Rozov
- , Natalia Demeshkina
- & Gulnara Yusupova
-
Article |
 Maintenance of protein synthesis reading frame by EF-P and m1G37-tRNA
Maintenance of protein synthesis reading frame by EF-P and m1G37-tRNA
Slippery mRNA sequences CC[C/U]-[C/U] are prone to +1 frameshift (+1FS) errors during mRNA translation. Here, the authors show that +1FS errors occur predominantly when CC[C/U]-[C/U] are placed at the second sense codon, and that error suppression requires m1G37-tRNA and the translation factor EF-P.
- Howard B. Gamper
- , Isao Masuda
- & Ya-Ming Hou
-
Article
| Open Access Translational regulation shapes the molecular landscape of complex disease phenotypes
Translational regulation shapes the molecular landscape of complex disease phenotypes
To what extent translational control can contribute to global gene expression patterns in the disease state is poorly defined. Here the authors conduct genome-wide RNA-seq and ribosome profiling in a rat model of hypertension and uncover altered translation patterns in disease associated genes.
- Sebastian Schafer
- , Eleonora Adami
- & Norbert Hubner
-
Article |
 Cytosolic targeting factor AKR2A captures chloroplast outer membrane-localized client proteins at the ribosome during translation
Cytosolic targeting factor AKR2A captures chloroplast outer membrane-localized client proteins at the ribosome during translation
Post-translational import of nuclear-encoded proteins shapes the proteome of organelles. Here, Kim et al.show that AKR2A, a critical targeting factor for chloroplast outer membrane proteins, binds to client proteins co-translationally as they exit the ribosome.
- Dae Heon Kim
- , Jae-Eun Lee
- & Inhwan Hwang
-
Article
| Open Access Methylation of ribosomal RNA by NSUN5 is a conserved mechanism modulating organismal lifespan
Methylation of ribosomal RNA by NSUN5 is a conserved mechanism modulating organismal lifespan
Cellular pathways modulating longevity and stress resistance are known to affect protein translation. Here the authors show that the RNA methyltransferase, Nsun5, or its yeast homologue Rcm1, regulates lifespan of three different model organisms by modifying ribosomal RNA at a specific cytosine residue.
- Markus Schosserer
- , Nadege Minois
- & Johannes Grillari
-
Article |
 Interplay between trigger factor and other protein biogenesis factors on the ribosome
Interplay between trigger factor and other protein biogenesis factors on the ribosome
Ribosome-associated protein biogenesis factors act during protein synthesis to facilitate modification, targeting and folding of the nascent polypeptide. Here, Bornemann et al.establish the dynamic interplay between these factors, thus providing new insight into the early steps of protein biogenesis.
- Thomas Bornemann
- , Wolf Holtkamp
- & Wolfgang Wintermeyer
-
Article |
 60S ribosome biogenesis requires rotation of the 5S ribonucleoprotein particle
60S ribosome biogenesis requires rotation of the 5S ribonucleoprotein particle
Our understanding of ribosome biogenesis is limited by a lack of structural knowledge of assembly intermediates. Here, Leidig et al.report a high-resolution cryo-EM structure of a pre-60S particle that suggests that substantial rearrangements of the 5S RNP are required during ribosome maturation.
- Christoph Leidig
- , Matthias Thoms
- & Roland Beckmann
-
Article
| Open Access Ribosome profiling reveals features of normal and disease-associated mitochondrial translation
Ribosome profiling reveals features of normal and disease-associated mitochondrial translation
Mitochondrial ribosomes are uniquely affected by mutations in the mitochondrial genome. By mapping the position of ribosomes on transcripts, the authors here reveal functional differences between mitochondrial and cytosolic ribosomes, and show that mutations in mitochondrial tRNAs induce ribosome stalling.
- Koos Rooijers
- , Fabricio Loayza-Puch
- & Reuven Agami
-
Article |
 Evolution of the protein stoichiometry in the L12 stalk of bacterial and organellar ribosomes
Evolution of the protein stoichiometry in the L12 stalk of bacterial and organellar ribosomes
The ribosomal stalk L12 is the only multi-copy protein in the ribosome and is essential for translation. Here Davydov et al.use a bioinformatics and mass spectrometry approach to study the evolution of L12 in bacterial ribosomes and predict its stoichiometry in a wide range of species.
- Iakov I. Davydov
- , Ingo Wohlgemuth
- & Marina V. Rodnina
-
Article |
 A structural basis for streptomycin-induced misreading of the genetic code
A structural basis for streptomycin-induced misreading of the genetic code
The antibiotic streptomycin increases errors in protein translation, but it is unclear how streptomycin exerts its effect on the ribosome. Demirci et al. present X-ray crystal structures that reveal conformational changes induced by streptomycin, which may inspire future efforts in antibiotics design.
- Hasan Demirci
- , Frank Murphy IV
- & Gerwald Jogl
-
Article |
 Potential for interdependent development of tRNA determinants for aminoacylation and ribosome decoding
Potential for interdependent development of tRNA determinants for aminoacylation and ribosome decoding
Aminoacyl-transfer RNA synthetases are conserved between bacteria and eukaryotes; however, bacterial enzymes cannot acylate eukaryote tRNAs. Now, fusion of a human and bacterial enzyme is shown to overcome the species barrier and confer tRNA specificity during both codon selection and proofreading on the ribosome.
- Cuiping Liu
- , Howard Gamper
- & Ya-Ming Hou

 UtpA and UtpB chaperone nascent pre-ribosomal RNA and U3 snoRNA to initiate eukaryotic ribosome assembly
UtpA and UtpB chaperone nascent pre-ribosomal RNA and U3 snoRNA to initiate eukaryotic ribosome assembly