Article
|
Open Access
Featured
-
-
Article |
 Profiling the genetic determinants of chromatin accessibility with scalable single-cell CRISPR screens
Profiling the genetic determinants of chromatin accessibility with scalable single-cell CRISPR screens
The effects of gene knockouts on chromatin accessibility are measured with single-cell CRISPR screens.
- Noa Liscovitch-Brauer
- , Antonino Montalbano
- & Neville E. Sanjana
-
Article |
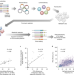 Predicting the efficiency of prime editing guide RNAs in human cells
Predicting the efficiency of prime editing guide RNAs in human cells
Prime editing is optimized by a method to choose the most efficient guide RNA.
- Hui Kwon Kim
- , Goosang Yu
- & Hyongbum Henry Kim
-
Article |
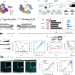 Massively parallel kinetic profiling of natural and engineered CRISPR nucleases
Massively parallel kinetic profiling of natural and engineered CRISPR nucleases
The enzymatic properties of RNA-guided nucleases are revealed through massively parallel analysis.
- Stephen K. Jones Jr
- , John A. Hawkins
- & Ilya J. Finkelstein
-
Article |
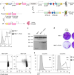 Inducible de novo expression of neoantigens in tumor cells and mice
Inducible de novo expression of neoantigens in tumor cells and mice
A system to express neoantigens in tumor cells and mice circumvents central T cell tolerance.
- Martina Damo
- , Brittany Fitzgerald
- & Nikhil S. Joshi
-
Letter |
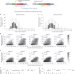 Sequence-specific prediction of the efficiencies of adenine and cytosine base editors
Sequence-specific prediction of the efficiencies of adenine and cytosine base editors
The activity of adenine or cytosine base editors at specific target nucleotides is predicted computationally.
- Myungjae Song
- , Hui Kwon Kim
- & Hyongbum Henry Kim
-
Article |
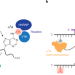 Programmable m6A modification of cellular RNAs with a Cas13-directed methyltransferase
Programmable m6A modification of cellular RNAs with a Cas13-directed methyltransferase
The most abundant RNA modification in humans, m6A, can be installed at specified RNA sequences in cells, enabling functional studies.
- Christopher Wilson
- , Peter J. Chen
- & David R. Liu
-
Review Article |
 Genome editing with CRISPR–Cas nucleases, base editors, transposases and prime editors
Genome editing with CRISPR–Cas nucleases, base editors, transposases and prime editors
A growing arsenal of CRISPR-based tools enables increasingly sophisticated genome editing applications.
- Andrew V. Anzalone
- , Luke W. Koblan
- & David R. Liu
-
Article |
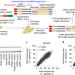 Prediction of the sequence-specific cleavage activity of Cas9 variants
Prediction of the sequence-specific cleavage activity of Cas9 variants
Deep-learning models predict the Cas9 variant with optimal activity and specificity for any target sequence.
- Nahye Kim
- , Hui Kwon Kim
- & Hyongbum Henry Kim
-
Brief Communication |
 Base editors for simultaneous introduction of C-to-T and A-to-G mutations
Base editors for simultaneous introduction of C-to-T and A-to-G mutations
Base editors that modify both adenosine and cytosine broaden the potential applications of base editing.
- Rina C. Sakata
- , Soh Ishiguro
- & Nozomu Yachie
-
Brief Communication |
 Dual base editor catalyzes both cytosine and adenine base conversions in human cells
Dual base editor catalyzes both cytosine and adenine base conversions in human cells
Base editors that modify both adenine and cytosine broaden the potential applications of base editing.
- Xiaohui Zhang
- , Biyun Zhu
- & Dali Li
-
Brief Communication |
 A dual-deaminase CRISPR base editor enables concurrent adenine and cytosine editing
A dual-deaminase CRISPR base editor enables concurrent adenine and cytosine editing
A base editor that concurrently modifies both adenine and cytosine broadens the potential applications of base editing.
- Julian Grünewald
- , Ronghao Zhou
- & J. Keith Joung
-
Review Article |
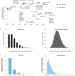 Design and analysis of CRISPR–Cas experiments
Design and analysis of CRISPR–Cas experiments
Hanna and Doench review the computational methods and tools that have become indispensable for planning and analyzing CRISPR experiments.
- Ruth E. Hanna
- & John G. Doench
-
Matters Arising |
Reply to: Fitness effects of CRISPR/Cas9-targeting of long noncoding RNA genes
- Ying Liu
- , Zhiheng Liu
- & Wensheng Wei
-
Article |
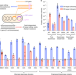 Evaluation and minimization of Cas9-independent off-target DNA editing by cytosine base editors
Evaluation and minimization of Cas9-independent off-target DNA editing by cytosine base editors
Methods to efficiently detect Cas9-independent cytosine base editor off-target activity enable the identification and development of variants with minimal off-target editing and robust on-target editing.
- Jordan L. Doman
- , Aditya Raguram
- & David R. Liu
-
Article |
 Continuous evolution of SpCas9 variants compatible with non-G PAMs
Continuous evolution of SpCas9 variants compatible with non-G PAMs
PAM sequences without G bases can be edited with SpCas9 variants that were continuously evolved in the laboratory.
- Shannon M. Miller
- , Tina Wang
- & David R. Liu
-
-
Letter |
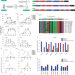 Targeted, random mutagenesis of plant genes with dual cytosine and adenine base editors
Targeted, random mutagenesis of plant genes with dual cytosine and adenine base editors
Saturation mutagenesis using dual base editors improves the herbicide resistance of rice.
- Chao Li
- , Rui Zhang
- & Caixia Gao
-
News & Views |
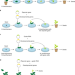 Fast-track for engineered plants
Fast-track for engineered plants
A rapid method to introduce gene edits advances plant science.
- Oliver Xiaoou Dong
- & Pamela C. Ronald
-
Brief Communication |
 Efficient, continuous mutagenesis in human cells using a pseudo-random DNA editor
Efficient, continuous mutagenesis in human cells using a pseudo-random DNA editor
Efficient diversification of DNA sequences is enabled by a T7 RNA polymerase fused to a cytidine deaminase.
- Haiqi Chen
- , Sophia Liu
- & Fei Chen
-
Letter |
 Polymer-stabilized Cas9 nanoparticles and modified repair templates increase genome editing efficiency
Polymer-stabilized Cas9 nanoparticles and modified repair templates increase genome editing efficiency
Precise genome editing is made more efficient by stabilizing Cas9 and enhancing shuttling to the nucleus.
- David N. Nguyen
- , Theodore L. Roth
- & Alexander Marson
-
Letter |
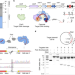 Harnessing type I CRISPR–Cas systems for genome engineering in human cells
Harnessing type I CRISPR–Cas systems for genome engineering in human cells
A type I CRISPR–Cas system, rather than the conventional type II CRISPR systems, is adapted for gene editing by fusing Cascade to the FokI nuclease domain.
- Peter Cameron
- , Mary M. Coons
- & Samuel H. Sternberg
-
Letter |
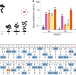 Adenine base editors catalyze cytosine conversions in human cells
Adenine base editors catalyze cytosine conversions in human cells
In addition to causing A-to-G base transitions, adenine base editors also cause C base substitutions
- Heon Seok Kim
- , You Kyeong Jeong
- & Sangsu Bae
-
Article |
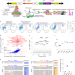 In vivo CRISPR screening in CD8 T cells with AAV–Sleeping Beauty hybrid vectors identifies membrane targets for improving immunotherapy for glioblastoma
In vivo CRISPR screening in CD8 T cells with AAV–Sleeping Beauty hybrid vectors identifies membrane targets for improving immunotherapy for glioblastoma
A hybrid AAV–transposon CRISPR approach enables large-scale in vivo screens in T cells and identifies potential targets for immunotherapy.
- Lupeng Ye
- , Jonathan J. Park
- & Sidi Chen
-
Letter |
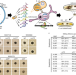 Hidden antibiotics in actinomycetes can be identified by inactivation of gene clusters for common antibiotics
Hidden antibiotics in actinomycetes can be identified by inactivation of gene clusters for common antibiotics
Dereplication by inactivation of antibiotic production clusters in actinomycetes enables antibiotic discovery.
- Elizabeth J. Culp
- , Grace Yim
- & Gerard D. Wright
-
News & Views |
 Jumping at the chance for precise DNA integration
Jumping at the chance for precise DNA integration
CRISPR–Cas-associated transposases enable targeted integration of new sequences into genomes.
- Jennifer B. Kwon
- & Charles A. Gersbach
-
Article |
 Continuous evolution of base editors with expanded target compatibility and improved activity
Continuous evolution of base editors with expanded target compatibility and improved activity
Improved base editors are generated by continuous evolution.
- B W. Thuronyi
- , Luke W. Koblan
- & David R. Liu
-
Article |
 Programmable RNA editing by recruiting endogenous ADAR using engineered RNAs
Programmable RNA editing by recruiting endogenous ADAR using engineered RNAs
Cellular RNAs are edited with high specificity using engineered ADAR-recruiting RNAs.
- Liang Qu
- , Zongyi Yi
- & Wensheng Wei
-
Review Article |
 Synthetic evolution
Synthetic evolution
From unbiased mutagenesis to precision modification, in genes or whole genomes, researchers have a panoply of tools to direct evolution.
- Anna J. Simon
- , Simon d’Oelsnitz
- & Andrew D. Ellington
-
News & Views |
 CRISPR–Cas9 strikes out in cassava
CRISPR–Cas9 strikes out in cassava
Transgenic cassava expressing Cas9 is not protected from geminivirus infection.
- Edward P. Rybicki
-
Review Article |
 Gene editing for immune cell therapies
Gene editing for immune cell therapies
Bailey and Maus discuss cutting-edge developments in engineering immune cells towards expanding the reach and efficacy of adoptive cell therapies in cancer and beyond.
- Stefanie R. Bailey
- & Marcela V. Maus
-
Letter |
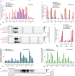 Circularly permuted and PAM-modified Cas9 variants broaden the targeting scope of base editors
Circularly permuted and PAM-modified Cas9 variants broaden the targeting scope of base editors
Wider editing windows and different PAM requirements enable a broad set of genomic positions to be targeted with A and C base editors.
- Tony P. Huang
- , Kevin T. Zhao
- & David R. Liu
-
Letter |
 Engineered CRISPR–Cas12a variants with increased activities and improved targeting ranges for gene, epigenetic and base editing
Engineered CRISPR–Cas12a variants with increased activities and improved targeting ranges for gene, epigenetic and base editing
Structure-guided protein engineering of Cas12a yields variants that have increased activity and that can edit sites with previously inaccessible PAMs.
- Benjamin P. Kleinstiver
- , Alexander A. Sousa
- & J. Keith Joung
-
Article |
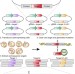 Predicting the mutations generated by repair of Cas9-induced double-strand breaks
Predicting the mutations generated by repair of Cas9-induced double-strand breaks
Analysis of more than a billion repair outcomes yields an algorithm that predicts Cas9-induced mutations.
- Felicity Allen
- , Luca Crepaldi
- & Leopold Parts
-
Correspondence |
A call for science-based review of the European court's decision on gene-edited crops
- Fyodor D Urnov
- , Pamela C Ronald
- & Dana Carroll
-
Brief Communication |
 Improving cytidine and adenine base editors by expression optimization and ancestral reconstruction
Improving cytidine and adenine base editors by expression optimization and ancestral reconstruction
Optimization of cytidine and adenine base editors increases editing efficiency.
- Luke W Koblan
- , Jordan L Doman
- & David R Liu
-
Letter |
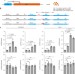 Genome editing of upstream open reading frames enables translational control in plants
Genome editing of upstream open reading frames enables translational control in plants
CRISPR/Cas9 editing of upstream open reading frames (uORFs) in Arabidopsis thaliana and lettuce enables upregulation of translation of plant genes.
- Huawei Zhang
- , Xiaomin Si
- & Caixia Gao
-
Letter |
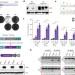 Optimized base editors enable efficient editing in cells, organoids and mice
Optimized base editors enable efficient editing in cells, organoids and mice
The efficiency of base editing is substantially increased by optimizing expression and nuclear localization of the editing enzymes.
- Maria Paz Zafra
- , Emma M Schatoff
- & Lukas E Dow
-
Article
| Open Access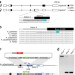 A CRISPR–Cas9 gene drive targeting doublesex causes complete population suppression in caged Anopheles gambiae mosquitoes
A CRISPR–Cas9 gene drive targeting doublesex causes complete population suppression in caged Anopheles gambiae mosquitoes
Complete population collapse of malaria vector Anopheles gambiae in cages is achieved using a gene drive that targets doublesex.
- Kyros Kyrou
- , Andrew M Hammond
- & Andrea Crisanti
-
Brief Communication |
 Efficient base editing in methylated regions with a human APOBEC3A-Cas9 fusion
Efficient base editing in methylated regions with a human APOBEC3A-Cas9 fusion
Increased efficiency of base editing in methylated DNA using human APOBEC3A as a deaminase.
- Xiao Wang
- , Jianan Li
- & Li Yang
-
News & Views |
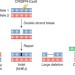 Unexpected CRISPR on-target effects
Unexpected CRISPR on-target effects
Cas9 can induce extensive on-target damage, including large deletions, inversions, and insertions.
- Hyunji Lee
- & Jin-Soo Kim
-
Article |
 Meganuclease targeting of PCSK9 in macaque liver leads to stable reduction in serum cholesterol
Meganuclease targeting of PCSK9 in macaque liver leads to stable reduction in serum cholesterol
Genome editing in non-human primates with a PCSK9-targeted meganuclease lowers cholesterol levels.
- Lili Wang
- , Jeff Smith
- & James M Wilson
-
Brief Communication |
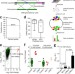 Genome-scale engineering of Saccharomyces cerevisiae with single-nucleotide precision
Genome-scale engineering of Saccharomyces cerevisiae with single-nucleotide precision
Efficient genome-scale precision editing in yeast is enabled using CRISPR–Cas9 and homology-directed-repair.
- Zehua Bao
- , Mohammad HamediRad
- & Huimin Zhao
-
Letter |
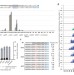 Adenine base editing in mouse embryos and an adult mouse model of Duchenne muscular dystrophy
Adenine base editing in mouse embryos and an adult mouse model of Duchenne muscular dystrophy
Adenine base editing is used to treat a mouse model of Duchenne muscular dystrophy and to create defined mutations in mouse embryos.
- Seuk-Min Ryu
- , Taeyoung Koo
- & Jin-Soo Kim
-
Letter |
 Simultaneous lineage tracing and cell-type identification using CRISPR–Cas9-induced genetic scars
Simultaneous lineage tracing and cell-type identification using CRISPR–Cas9-induced genetic scars
LINNAEUS reconstructs developmental lineages using RNA sequencing data and lineage markers from the same single cells.
- Bastiaan Spanjaard
- , Bo Hu
- & Jan Philipp Junker
-
Article |
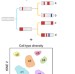 Simultaneous single-cell profiling of lineages and cell types in the vertebrate brain
Simultaneous single-cell profiling of lineages and cell types in the vertebrate brain
scGESTALT enables large-scale characterization of cell types and lineage relationships during vertebrate brain development.
- Bushra Raj
- , Daniel E Wagner
- & Alexander F Schier
-
Brief Communication |
 Base editing with a Cpf1–cytidine deaminase fusion
Base editing with a Cpf1–cytidine deaminase fusion
A new fusion protein enables precise editing of single bases in A/T-rich regions of the human genome.
- Xiaosa Li
- , Ying Wang
- & Jia Chen
-
Letter |
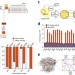 A highly specific SpCas9 variant is identified by in vivo screening in yeast
A highly specific SpCas9 variant is identified by in vivo screening in yeast
Evolved SpCas9 variant evoCas9 has improved specificity and retains near wild-type on-target activity.
- Antonio Casini
- , Michele Olivieri
- & Anna Cereseto
-
Brief Communication |
 Deep learning improves prediction of CRISPR–Cpf1 guide RNA activity
Deep learning improves prediction of CRISPR–Cpf1 guide RNA activity
Using deep learning to combine target sequence and chromatin accessibility data boosts the accuracy of CRISPR–Cpf1 guide RNA activity
- Hui Kwon Kim
- , Seonwoo Min
- & Hyongbum (Henry) Kim
-
News & Views |
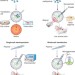 Viruses leave their stamp on single cells
Viruses leave their stamp on single cells
A method for 'stamping' viruses achieves efficient transduction of single target cells in cultured tissues and in the mouse brain.
- Ede A Rancz
- & Andreas T Schaefer

 In vivo adenine base editing of PCSK9 in macaques reduces LDL cholesterol levels
In vivo adenine base editing of PCSK9 in macaques reduces LDL cholesterol levels Genome editing in animals: why FDA regulation matters
Genome editing in animals: why FDA regulation matters