Featured
-
-
Article
| Open Access Genome-wide detection of cytosine methylations in plant from Nanopore data using deep learning
Genome-wide detection of cytosine methylations in plant from Nanopore data using deep learning
Existing methods cannot profile genome-wide cytosine DNA methylations (5mCs) in all three contexts with acceptable accuracy. Here, the authors develop a deep learning tool to detect genome-wide 5mCs of all three contexts in plants with high accuracy from Nanopore reads.
- Peng Ni
- , Neng Huang
- & Jianxin Wang
-
Article
| Open Access Epigenetic modifications affect the rate of spontaneous mutations in a pathogenic fungus
Epigenetic modifications affect the rate of spontaneous mutations in a pathogenic fungus
While a correlation between epigenetic modifications and mutation rates has been observed, experimental evidence of causality is limited. Here the authors measure the mutation rate in fungal mutants lacking histone modifications and confirm experimentally a causal effect of epigenetic modifications on mutation rates.
- Michael Habig
- , Cecile Lorrain
- & Eva H. Stukenbrock
-
Article
| Open Access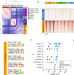 Subtype heterogeneity and epigenetic convergence in neuroendocrine prostate cancer
Subtype heterogeneity and epigenetic convergence in neuroendocrine prostate cancer
Neuroendocrine carcinomas (NECs) arise from different anatomic sites, but have similar histological and clinical features. Here, the authors show that the epigenetic landscape of a range of NECs converges towards a common epigenetic state, while distinct subtypes occur within neuroendocrine prostate cancer contributing to intratumor heterogeneity in clinical samples.
- Paloma Cejas
- , Yingtian Xie
- & Henry W. Long
-
Article
| Open Access SPOP mutation induces DNA methylation via stabilizing GLP/G9a
SPOP mutation induces DNA methylation via stabilizing GLP/G9a
The molecular mechanism underlying the DNA hypermethylation phenotype observed in the SPOP-mutant prostate cancers is unclear. Here, the authors show that mutant SPOP induces global aberrant DNA methylation patterns through GLP/G9a and renders prostate cancer cells susceptible to DNA demethylating agents.
- Jianong Zhang
- , Kun Gao
- & Haojie Huang
-
Article
| Open Access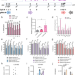 Unraveling the functional role of DNA demethylation at specific promoters by targeted steric blockage of DNA methyltransferase with CRISPR/dCas9
Unraveling the functional role of DNA demethylation at specific promoters by targeted steric blockage of DNA methyltransferase with CRISPR/dCas9
The causal relationship between DNA demethylation and gene expression regulation has not yet been fully resolved. Here the authors develop a nuclease-dead Cas9 (dCas9) and gRNA site-specific targeting approach to physically block DNA methylation at specific promoters to cause DNA demethylation in cells and tackle this question.
- Daniel M. Sapozhnikov
- & Moshe Szyf
-
Article
| Open Access CHARGE syndrome protein CHD7 regulates epigenomic activation of enhancers in granule cell precursors and gyrification of the cerebellum
CHARGE syndrome protein CHD7 regulates epigenomic activation of enhancers in granule cell precursors and gyrification of the cerebellum
CHARGE syndrome that affects cerebellar development can be caused by haploinsufficiency of the chromatin remodeling enzyme CHD7; however the precise role of CHD7 remains unknown. Here the authors show CHD7 promotes chromatin accessibility and enhancer activity in granule cell precursors and regulates morphogenesis of the cerebellar cortex, where loss of CHD7 triggers cerebellar polymicrogyria.
- Naveen C. Reddy
- , Shahriyar P. Majidi
- & Harrison W. Gabel
-
Article
| Open Access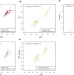 Identical twins carry a persistent epigenetic signature of early genome programming
Identical twins carry a persistent epigenetic signature of early genome programming
The mechanisms underlying how monozygotic (or identical) twins arise are yet to be determined. Here, the authors investigate this in an epigenome-wide association study, showing that monozygotic twinning has a characteristic DNA methylation signature in adult somatic tissues.
- Jenny van Dongen
- , Scott D. Gordon
- & Dorret I. Boomsma
-
Article
| Open Access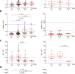 A longitudinal sampling study of transcriptomic and epigenetic profiles in patients with thrombocytopenia syndrome
A longitudinal sampling study of transcriptomic and epigenetic profiles in patients with thrombocytopenia syndrome
Severe fever with thrombocytopenia syndrome (SFTS) is an emerging hemorrhagic fever caused by tick-borne SFTS virus. Here, Wang et al. characterize transcriptomic and epigenetic changes in infected patients and correlate them with clinical parameters to improve the understanding of disease progression.
- Yafen Wang
- , Shaoqing Han
- & Xiang Zhou
-
Article
| Open Access Exosome-mediated stable epigenetic repression of HIV-1
Exosome-mediated stable epigenetic repression of HIV-1
A strategy to control HIV-1 infection is to stably repress HIV-1 and induce “deep latency”. Here the authors show that a recombinant anti-HIV-1-1 protein can be packaged as mRNA into exosomes and delivered systemically to repress HIV-1-1 within the context of virus infected mice and achieve long term silencing of HIV-1-1 expression.
- Surya Shrivastava
- , Roslyn M. Ray
- & Kevin V. Morris
-
Article
| Open Access Mammary-specific expression of Trim24 establishes a mouse model of human metaplastic breast cancer
Mammary-specific expression of Trim24 establishes a mouse model of human metaplastic breast cancer
Human metaplastic breast cancers (MpBC) are a rare, aggressive subclass of triple-negative breast cancers. Here, the authors show over-expression of histone reader TRIM24 is sufficient to generate tumors with a molecular signature of metabolic dysfunction and EMT in a mouse model of human MpBC.
- Vrutant V. Shah
- , Aundrietta D. Duncan
- & Michelle Craig Barton
-
Article
| Open Access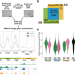 Variant PCGF1-PRC1 links PRC2 recruitment with differentiation-associated transcriptional inactivation at target genes
Variant PCGF1-PRC1 links PRC2 recruitment with differentiation-associated transcriptional inactivation at target genes
Polycomb repressive complexes (PRC1 and PRC2) repress genes that are crucial for development via epigenetic modifications; however, their role in differentiation is not well known. Here the authors reveal that a PCGF1-containing PRC1 variant facilitates exit from pluripotency by downregulating target genes and recruiting PRC2.
- Hiroki Sugishita
- , Takashi Kondo
- & Haruhiko Koseki
-
Article
| Open Access Locus specific epigenetic modalities of random allelic expression imbalance
Locus specific epigenetic modalities of random allelic expression imbalance
Some autosomal genes are expressed in a random monoallelic manner, but its extent and mechanisms have remained unclear. Here the authors show robust monoallelic expression in cell lines and mice, where the silent allele can be reexpressed using epidrugs. Further, they find these genes display various modalities of allelic expression with different degrees of allelic imbalance.
- Lucile Marion-Poll
- , Benjamin Forêt
- & Edith Heard
-
Article
| Open Access The concurrence of DNA methylation and demethylation is associated with transcription regulation
The concurrence of DNA methylation and demethylation is associated with transcription regulation
The global pattern of the mammalian methylome is formed by changes in methylation and demethylation. Here the authors describe a metric methylation concurrence that measures the ratio of unmethylated CpGs inside the partially methylated reads and show that methylation concurrence is associated with epigenetically regulated tumour suppressor genes.
- Jiejun Shi
- , Jianfeng Xu
- & Wei Li
-
Article
| Open Access Genome-wide sequencing-based identification of methylation quantitative trait loci and their role in schizophrenia risk
Genome-wide sequencing-based identification of methylation quantitative trait loci and their role in schizophrenia risk
The authors provide a comprehensive, single base resolution view of association between genetic variation and DNA methylation in human brain. They also show that heritability attributed to schizophrenia GWAS-associated variants reflects the epigenetic plasticity of the brain.
- Kira A. Perzel Mandell
- , Nicholas J. Eagles
- & Andrew E. Jaffe
-
Article
| Open Access Placental DNA methylation signatures of maternal smoking during pregnancy and potential impacts on fetal growth
Placental DNA methylation signatures of maternal smoking during pregnancy and potential impacts on fetal growth
Maternal smoking during pregnancy contributes to poor birth outcomes. Here the authors perform a meta-analysis of the associations between maternal smoking during pregnancy and placental DNA methylation and identify links between these and poor birth outcomes, which may better inform the mechanisms through which smoking impacts placental function and fetal growth.
- Todd M. Everson
- , Marta Vives-Usano
- & Mariona Bustamante
-
Article
| Open Access Nuclear condensates of p300 formed though the structured catalytic core can act as a storage pool of p300 with reduced HAT activity
Nuclear condensates of p300 formed though the structured catalytic core can act as a storage pool of p300 with reduced HAT activity
The histone acetyltransferase p300 mostly localizes to active chromatin; however, some repressed genes marked with H3K27me3 are also bound by p300. Here the authors show p300 is capable of phase separation, which relies on its catalytic core, and that p300 catalytic activity is decreased in phase-separated droplets that co-localize with H3K27me3-marked chromatin.
- Yi Zhang
- , Kyle Brown
- & Tatiana G. Kutateladze
-
Article
| Open Access PALI1 facilitates DNA and nucleosome binding by PRC2 and triggers an allosteric activation of catalysis
PALI1 facilitates DNA and nucleosome binding by PRC2 and triggers an allosteric activation of catalysis
The polycomb repressive complex 2 (PRC2) is a histone methyltransferase regulating cell differentiation and identity. Here, the authors show that the vertebrate-specific PRC2 accessory subunit PALI1 facilitates substrate binding by the complex and elucidate the allosteric mechanism of PALI1- mediated PRC2 activation.
- Qi Zhang
- , Samuel C. Agius
- & Chen Davidovich
-
Article
| Open Access Functional and epigenetic phenotypes of humans and mice with DNMT3A Overgrowth Syndrome
Functional and epigenetic phenotypes of humans and mice with DNMT3A Overgrowth Syndrome
Germline mutations in the DNMT3A gene can cause an overgrowth syndrome associated with behavioural and hematopoietic phenotypes. Here the authors describe a mouse model of this syndrome that recapitulates many of these features, including conserved alterations in DNA methylation in the blood cells of both species.
- Amanda M. Smith
- , Taylor A. LaValle
- & Timothy J. Ley
-
Article
| Open Access A machine learning approach to brain epigenetic analysis reveals kinases associated with Alzheimer’s disease
A machine learning approach to brain epigenetic analysis reveals kinases associated with Alzheimer’s disease
Array-based epigenome-wide association studies only test about 2% of the CpG sites in the genome. Here, the authors describe EWASplus, a supervised machine learning strategy that extends EWAS coverage to the entire genome, and use it to identify novel brain CpGs associated with Alzheimer’s disease.
- Yanting Huang
- , Xiaobo Sun
- & Zhaohui S. Qin
-
Article
| Open Access Neonatal thyroxine activation modifies epigenetic programming of the liver
Neonatal thyroxine activation modifies epigenetic programming of the liver
Whether thyroid hormones affect gene expression via DNA methylation is not well known. Here the authors show that type 2 deiodinase (D2) converts T4 to produce T3, which prevents DNA methylation of discrete areas in the neonatal liver. In the absence of D2, DNA methylation occurs and is associated with reduced chromatin accessibility in promoters and enhancers and affects gene expression.
- Tatiana L. Fonseca
- , Tzintzuni Garcia
- & Antonio C. Bianco
-
Article
| Open Access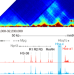 Reactivation of a developmentally silenced embryonic globin gene
Reactivation of a developmentally silenced embryonic globin gene
Globin loci harbor genes that are expressed embryonically and silenced postnatally. Here the authors show that zeta-globin silencing depends upon selective hypoacetylation of its TAD subdomain, which blocks its interaction with the alpha-globin super-enhancer, and zeta-globin can be reactivated by acetylation.
- Andrew J. King
- , Duantida Songdej
- & Christian Babbs
-
Article
| Open Access Early-life social experience affects offspring DNA methylation and later life stress phenotype
Early-life social experience affects offspring DNA methylation and later life stress phenotype
Early social experience can alter epigenetic patterns and stress responses later in life. A study on wild spotted hyenas finds that maternal care and social connections after leaving the den influence DNA methylation and contribute to a developmentally plastic stress response.
- Zachary M. Laubach
- , Julia R. Greenberg
- & Wei Perng
-
Article
| Open Access Complete loss of H3K9 methylation dissolves mouse heterochromatin organization
Complete loss of H3K9 methylation dissolves mouse heterochromatin organization
Histone H3K9 methylation (H3K9me) states define repressed chromatin in eukaryotic cells. Here the authors reveal complete loss of all H3K9me in mammalian cells through successive deletion of H3K9 methyltransferase genes that results in the dissolution of heterochromatin and the derepression of nearly all repeat families.
- Thomas Montavon
- , Nicholas Shukeir
- & Thomas Jenuwein
-
Article
| Open Access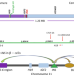 Large parental differences in chromatin organization in pancreatic beta cell line explaining diabetes susceptibility effects
Large parental differences in chromatin organization in pancreatic beta cell line explaining diabetes susceptibility effects
A SNP distant from the human insulin (INS) gene near the KRTAP5-6 gene confers increased susceptibility to type 2 diabetes when present on the paternal allele while decreased susceptibility when on the maternal allele. Here the authors show that long-range contacts between the INS locus and the KRTAP5-6 gene locus distinguish paternal and maternal alleles.
- Xing Jian
- & Gary Felsenfeld
-
Article
| Open Access Arabidopsis MORC proteins function in the efficient establishment of RNA directed DNA methylation
Arabidopsis MORC proteins function in the efficient establishment of RNA directed DNA methylation
MORC ATPases are required for transposable element silencing and heterochromatin condensation in plants and animals. Here the authors show that Arabidopsis MORCs colocalize with sites of RNA-directed DNA methylation and provide evidence that they act as molecular tethers to efficiently establish DNA methylation.
- Yan Xue
- , Zhenhui Zhong
- & Steven E. Jacobsen
-
Article
| Open Access Epigenetic memories and the evolution of infectious diseases
Epigenetic memories and the evolution of infectious diseases
The virulence of some infectious diseases seems to depend on the sex of the host the infection came from, as well as that of the current host. Here, McLeod et al. develop an epidemiological model to investigate the evolution of virulence when pathogens can retain epigenetic memories of their previous host.
- David V. McLeod
- , Geoff Wild
- & Francisco Úbeda
-
Article
| Open Access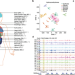 Tissue-specific 5-hydroxymethylcytosine landscape of the human genome
Tissue-specific 5-hydroxymethylcytosine landscape of the human genome
Charting the landscape of 5hmC in human tissues is fundamental to understanding its regulatory functions. Here, we systematically profiled the whole-genome 5hmC landscape at single-base resolution for 19 types of human tissues and found 5hmC shows tissue-specific patterns.
- Bo He
- , Chao Zhang
- & Chengqi Yi
-
Article
| Open Access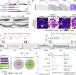 Mating can initiate stable RNA silencing that overcomes epigenetic recovery
Mating can initiate stable RNA silencing that overcomes epigenetic recovery
Stable epigenetic changes are relatively rare. Here the authors report that mating induces stable silencing of a single-copy transgene in C. elegans. Components of small RNA silencing are required for this stable silencing.
- Sindhuja Devanapally
- , Pravrutha Raman
- & Antony M. Jose
-
Article
| Open Access EZH2-induced lysine K362 methylation enhances TMPRSS2-ERG oncogenic activity in prostate cancer
EZH2-induced lysine K362 methylation enhances TMPRSS2-ERG oncogenic activity in prostate cancer
Although the TMPRSS2-ERG gene fusion is the most common alteration in human prostate cancer, its involvement in disease progression remains unclear. Here, the authors demonstrate that ERG is methylated by Enhancer of zest homolog 2 leading to enhanced transcriptional and oncogenic activity.
- Marita Zoma
- , Laura Curti
- & Giuseppina M. Carbone
-
Article
| Open Access Chromatin states shaped by an epigenetic code confer regenerative potential to the mouse liver
Chromatin states shaped by an epigenetic code confer regenerative potential to the mouse liver
Few studies have provided functional analysis of the epigenetic landscape in the regenerating liver. Here the authors define chromatin states in the quiescent vs. regenerating mouse liver through integration of genome wide profiles of DNA methylation, histone modifications, and chromatin accessibility, identifying H3K27me3 as an epigenetic mark conferring regenerative potential.
- Chi Zhang
- , Filippo Macchi
- & Kirsten C. Sadler
-
Article
| Open Access Redirected nuclear glutamate dehydrogenase supplies Tet3 with α-ketoglutarate in neurons
Redirected nuclear glutamate dehydrogenase supplies Tet3 with α-ketoglutarate in neurons
α-ketoglutarate (αKG) is an intermediate in the tricarboxylic acid cycle that is required in the nucleus for genomic DNA demethylation by Tet3. Here, the authors show that the enzyme glutamate dehydrogenase, which converts glutamate to αKG, is redirected from the mitochondria to the nucleus.
- Franziska R. Traube
- , Dilara Özdemir
- & Thomas Carell
-
Article
| Open Access High-resolution characterization of gene function using single-cell CRISPR tiling screen
High-resolution characterization of gene function using single-cell CRISPR tiling screen
Identifying functional domains and genetic regulatory mechanisms is essential for developing new therapies. Here the authors present sc-Tiling, single-cell high-density CRISPR tiling screening for functional domain characterization.
- Lu Yang
- , Anthony K. N. Chan
- & Chun-Wei Chen
-
Article
| Open Access A multi-ethnic epigenome-wide association study of leukocyte DNA methylation and blood lipids
A multi-ethnic epigenome-wide association study of leukocyte DNA methylation and blood lipids
Abnormal blood lipid levels are important risk factors for cardiovascular and other various diseases. Here the authors conduct a large-scale multi-ethnic epigenome-wide association study combined with epigenetic (cis-QTL and eQTM) data, and identify CpG-lipid traits associations that are specific to or common across racial/ethnic groups.
- Min-A Jhun
- , Michael Mendelson
- & Themistocles L. Assimes
-
Article
| Open Access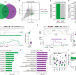 Neuronal and glial 3D chromatin architecture informs the cellular etiology of brain disorders
Neuronal and glial 3D chromatin architecture informs the cellular etiology of brain disorders
The cellular heterogeneity in brain obscures the identification of robust cellular regulatory networks. Here the authors integrate genome-wide chromosome conformation data from sorted neurons and glia, with transcriptomic and enhancer profiles, to characterize cell-type-specific gene regulatory landscapes in the human brain, and provide insights into cell-type-specific gene regulatory networks in brain disorders.
- Benxia Hu
- , Hyejung Won
- & Daniel H. Geschwind
-
Article
| Open Access Environmental enrichment preserves a young DNA methylation landscape in the aged mouse hippocampus
Environmental enrichment preserves a young DNA methylation landscape in the aged mouse hippocampus
Decline of brain function during aging is associated with epigenetic changes, including DNA methylation. Here the authors provide evidence that environmental enrichment delays age-related DNA methylation alterations in the mouse hippocampus.
- Sara Zocher
- , Rupert W. Overall
- & Gerd Kempermann
-
Article
| Open Access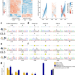 Genomic imprinting in mouse blastocysts is predominantly associated with H3K27me3
Genomic imprinting in mouse blastocysts is predominantly associated with H3K27me3
In most mammals, imprinted genes contain epigenetic marks that differ in each parental genome and control their parent-of-origin-specific expression. Here, the authors map imprinted genes in mouse preimplantation embryos and find that imprinted gene expression in blastocysts is mainly dependent on Polycomb-mediated H3K27me3-associated gene silencing.
- Laura Santini
- , Florian Halbritter
- & Martin Leeb
-
Article
| Open Access A gain-of-function single nucleotide variant creates a new promoter which acts as an orientation-dependent enhancer-blocker
A gain-of-function single nucleotide variant creates a new promoter which acts as an orientation-dependent enhancer-blocker
The role of promoters as potential insulator elements has been largely unexplored in mammals. Here the authors show that a single nucleotide variant in the α-globin locus forms a new promoter and acts as an orientation-dependent enhancer-blocking insulator element.
- Yavor K. Bozhilov
- , Damien J. Downes
- & Douglas R. Higgs
-
Article
| Open Access Hakai is required for stabilization of core components of the m6A mRNA methylation machinery
Hakai is required for stabilization of core components of the m6A mRNA methylation machinery
The E3 ligase Hakai can interact with the m6A methylation machinery but its function is still unclear. Here, the authors show that Hakai is a conserved component of the m6A methyltransferase complex and provide functional and molecular insights into its role in regulating m6A levels in Drosophila.
- Praveen Bawankar
- , Tina Lence
- & Jean-Yves Roignant
-
Article
| Open Access Defective folate metabolism causes germline epigenetic instability and distinguishes Hira as a phenotype inheritance biomarker
Defective folate metabolism causes germline epigenetic instability and distinguishes Hira as a phenotype inheritance biomarker
Abnormal folate metabolism in mice results in transgenerational epigenetic inheritance of congenital malformations. Here, the authors provide evidence that defective folate metabolism causes germline epigenetic instability and observe multigenerational misexpression of Hira in embryos, implicating Hira transcript levels as a biomarker of maternal phenotypic inheritance.
- Georgina E. T. Blake
- , Xiaohui Zhao
- & Erica D. Watson
-
Article
| Open Access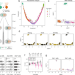 Integrated analysis of Xist upregulation and X-chromosome inactivation with single-cell and single-allele resolution
Integrated analysis of Xist upregulation and X-chromosome inactivation with single-cell and single-allele resolution
X-chromosome inactivation (XCI) ensures dosage compensation between the sexes. Here the authors perform allele-specific single-cell RNA sequencing in differentiating mouse embryonic stem cells to provide a detailed profile of the onset of XCI.
- Guido Pacini
- , Ilona Dunkel
- & Edda G. Schulz
-
Article
| Open Access Mapping chromatin accessibility and active regulatory elements reveals pathological mechanisms in human gliomas
Mapping chromatin accessibility and active regulatory elements reveals pathological mechanisms in human gliomas
Gliomas are tumors often associated with epigenetics-related gene deregulation. Here the authors reveal an atlas of active enhancers and promoters in benign and malignant gliomas by performing whole-genome mapping of chromatin accessibility, histone modifications, DNA methylation patterns and transcriptome analysis simultaneously in multiple tumor samples.
- Karolina Stępniak
- , Magdalena A. Machnicka
- & Bartek Wilczyński
-
Article
| Open Access E2F6 initiates stable epigenetic silencing of germline genes during embryonic development
E2F6 initiates stable epigenetic silencing of germline genes during embryonic development
DNA methylation targets CpG island promoters of germline genes to repress their expression in mouse somatic cells. Here the authors show that a transcription factor E2F6 is required to target CpG island DNA methylation and epigenetic silencing to germline genes during early mouse development.
- Thomas Dahlet
- , Matthias Truss
- & Michael Weber
-
Article
| Open Access A meta-analysis of epigenome-wide association studies in Alzheimer’s disease highlights novel differentially methylated loci across cortex
A meta-analysis of epigenome-wide association studies in Alzheimer’s disease highlights novel differentially methylated loci across cortex
Although epigenome-wide association studies of Alzheimer’s disease have highlighted neuropathology-associated DNA methylation differences, previous studies have been limited in sample size and brain region used. Here, the authors combine data from six DNA methylomic studies of Alzheimer’s disease (N = 1453 unique individuals) to identify differentially methylated loci across cortex.
- Rebecca G. Smith
- , Ehsan Pishva
- & Katie Lunnon
-
Article
| Open Access The epigenetic regulator LSH maintains fork protection and genomic stability via MacroH2A deposition and RAD51 filament formation
The epigenetic regulator LSH maintains fork protection and genomic stability via MacroH2A deposition and RAD51 filament formation
LSH/HELLS is a chromatin remodeler promoting incorporation of histone variant macroH2A. Here the authors reveal a role for LSH in genome stability, in protecting nascent DNA at stalled forks mediated by macroH2A deposition and RAD51 filament formation.
- Xiaoping Xu
- , Kai Ni
- & Kathrin Muegge
-
Article
| Open Access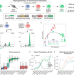 Chromosome compartments on the inactive X guide TAD formation independently of transcription during X-reactivation
Chromosome compartments on the inactive X guide TAD formation independently of transcription during X-reactivation
Both A/B compartments and TADs are thought to be absent from the inactive X chromosome, but to be re-established with transcriptional reactivation and chromatin opening during X-reactivation. Here, the authors characterise gene reactivation, chromatin opening and chromosome topology during X-reactivation, observe A/B-like compartments on the inactive X that guide TAD formation independently of transcription during X-reactivation.
- Moritz Bauer
- , Enrique Vidal
- & Bernhard Payer
-
Article
| Open Access H3K27me3 demethylases alter HSP22 and HSP17.6C expression in response to recurring heat in Arabidopsis
H3K27me3 demethylases alter HSP22 and HSP17.6C expression in response to recurring heat in Arabidopsis
Acclimation to high temperature increases tolerance of heat shock in plants. Here the authors show that JUMONJI H3K27me3 demethylases are needed for heat acclimation in Arabidopsis and act at loci encoding HEAT SHOCK PROTEINS to facilitate induction upon heat stress.
- Nobutoshi Yamaguchi
- , Satoshi Matsubara
- & Toshiro Ito
-
Article
| Open Access Heteromeric HSFA2/HSFA3 complexes drive transcriptional memory after heat stress in Arabidopsis
Heteromeric HSFA2/HSFA3 complexes drive transcriptional memory after heat stress in Arabidopsis
Moderate heat stress primes plants to acquire tolerance to subsequent, more severe heat stress. Here the authors show that the HSFA3 transcription factor forms a heteromeric complex with HSFA2 to sustain activated transcription of genes required for acquired thermotolerance by promoting H3K4 hyper-methylation.
- Thomas Friedrich
- , Vicky Oberkofler
- & Isabel Bäurle
-
Article
| Open Access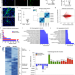 HIRA stabilizes skeletal muscle lineage identity
HIRA stabilizes skeletal muscle lineage identity
The epigenetic mechanisms coordinating the maintenance of adult cellular lineages remain poorly understood. Here the authors demonstrate that HIRA, a H3.3 histone chaperone, establishes the chromatin landscape required for skeletal muscle cell identity.
- Joana Esteves de Lima
- , Reem Bou Akar
- & Frédéric Relaix
-
Article
| Open Access High throughput screening identifies SOX2 as a super pioneer factor that inhibits DNA methylation maintenance at its binding sites
High throughput screening identifies SOX2 as a super pioneer factor that inhibits DNA methylation maintenance at its binding sites
The functional relevance of epigenetic modifications on transcription regulation has been an important question since their discovery. Here, the authors investigate the effect of DNA methylation on Pioneer Transcription Factor (PF) binding and distinguish between PFs that protect their binding sites from methylation and those that bind to methylated DNA and induce DNA demethylation.
- Ludovica Vanzan
- , Hadrien Soldati
- & Rabih Murr

 Histone H4 lysine 16 acetylation controls central carbon metabolism and diet-induced obesity in mice
Histone H4 lysine 16 acetylation controls central carbon metabolism and diet-induced obesity in mice