Article
|
Open Access
Featured
-
-
Article
| Open Access Prediction of base editor off-targets by deep learning
Prediction of base editor off-targets by deep learning
Base editors can induce unwanted off-target effects. Here the authors design libraries of gRNA-off-target pairs and perform a screen to obtain editing efficiencies for ABE and CBE: they use the datasets to train DL models (ABEdeepoff and CBEdeepoff) which can predict mutation tolerance at potential off-targets.
- Chengdong Zhang
- , Yuan Yang
- & Yongming Wang
-
Article
| Open Access Whole genomic analysis reveals atypical non-homologous off-target large structural variants induced by CRISPR-Cas9-mediated genome editing
Whole genomic analysis reveals atypical non-homologous off-target large structural variants induced by CRISPR-Cas9-mediated genome editing
The safety of CRISPR-Cas9 editing is a concern. Here the authors use whole genomic analysis by 10x linked-read sequencing and optical genome mapping to interrogate the genome integrity after editing: they see large structural variants at on-target sites and unexpected large chromosomal deletions.
- Hsiu-Hui Tsai
- , Hsiao-Jung Kao
- & John Yu
-
Article
| Open Access One-step generation of tumor models by base editor multiplexing in adult stem cell-derived organoids
One-step generation of tumor models by base editor multiplexing in adult stem cell-derived organoids
CRISPR base editing technologies can be used for disease modelling. Here the authors use various base editing tools to generate tumour models in human adult stem cell-derived hepatocyte, endometrial and intestinal organoids.
- Maarten H. Geurts
- , Shashank Gandhi
- & Hans Clevers
-
Article
| Open Access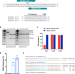 Treatment of monogenic and digenic dominant genetic hearing loss by CRISPR-Cas9 ribonucleoprotein delivery in vivo
Treatment of monogenic and digenic dominant genetic hearing loss by CRISPR-Cas9 ribonucleoprotein delivery in vivo
Liposome-mediated gene editing was used to abolish a mutation in gene Atp2b2 and recover hearing in a mouse model of dominant deafness. Editing was also used to target two mutations to recover hearing. The study detected large deletions due to editing.
- Yong Tao
- , Veronica Lamas
- & Zheng-Yi Chen
-
Article
| Open Access Simultaneous inhibition of DNA-PK and Polϴ improves integration efficiency and precision of genome editing
Simultaneous inhibition of DNA-PK and Polϴ improves integration efficiency and precision of genome editing
Low efficiency of target DNA integration remains a challenge in genome engineering. Here the authors perform large-scale compound library and genetic screens to identify targets that enhance gene editing: they see that combined DNA-PK and Polϴ inhibition with potent compounds increases editing efficiency and precision.
- Sandra Wimberger
- , Nina Akrap
- & Marcello Maresca
-
Article
| Open Access Cas9-mediated knockout of Ndrg2 enhances the regenerative potential of dendritic cells for wound healing
Cas9-mediated knockout of Ndrg2 enhances the regenerative potential of dendritic cells for wound healing
Chronic wounds impose a significant burden to a broad patient population. Here, the authors use CRISPR/Cas9 to enhance the regenerative capacity of dendritic cells by knocking out the gene Ndrg2, and show that seeding these engineered dendritic cells on hydrogels constitutes an effective therapy for chronic wounds in diabetic and non-diabetic conditions.
- Dominic Henn
- , Dehua Zhao
- & Geoffrey C. Gurtner
-
Article
| Open Access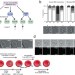 Sequential roles for red blood cell binding proteins enable phased commitment to invasion for malaria parasites
Sequential roles for red blood cell binding proteins enable phased commitment to invasion for malaria parasites
Malaria parasites invade erythrocytes to proliferate, but visualizing this rapid process is challenging. Here the authors use live imaging and genome-editing of P. knowlesi to dissect invasion and establish the roles of two vital parasite proteins.
- Melissa N. Hart
- , Franziska Mohring
- & Robert W. Moon
-
Article
| Open Access Cell cycle arrest and p53 prevent ON-target megabase-scale rearrangements induced by CRISPR-Cas9
Cell cycle arrest and p53 prevent ON-target megabase-scale rearrangements induced by CRISPR-Cas9
ON-target genotoxicity in gene editing is generally underestimated. Here the authors report Fluorescence-Assisted Megabase-scale Rearrangements Detection (FAMReD) systems to detect and characterize rare large loss of heterozygosity: they show that ON-target genotoxicity can be prevented by p53 and cell cycle arrest.
- G. Cullot
- , J. Boutin
- & A. Bedel
-
Article
| Open Access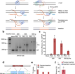 Template-jumping prime editing enables large insertion and exon rewriting in vivo
Template-jumping prime editing enables large insertion and exon rewriting in vivo
Retrotransposons replicate their genetic information through target-primed reverse transcription (TPRT). Here the authors report a template-jumping prime editor (TJ-PE) to act similarly to TPRT and achieve insertions of large DNA fragments at endogenous sites: they rewrite a mutated exon in the mouse liver.
- Chunwei Zheng
- , Bin Liu
- & Wen Xue
-
Article
| Open Access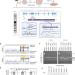 Increased renal elimination of endogenous and synthetic pyrimidine nucleosides in concentrative nucleoside transporter 1 deficient mice
Increased renal elimination of endogenous and synthetic pyrimidine nucleosides in concentrative nucleoside transporter 1 deficient mice
Concentrative nucleoside transporters (CNTs) are cellular nucleoside influx systems, but their in vivo roles are poorly defined. By generating CNT1 knockout (KO) mice, here the authors show a role of CNT1 in the renal reabsorption of endogenous and synthetic nucleosides.
- Avinash K. Persaud
- , Matthew C. Bernier
- & Rajgopal Govindarajan
-
Article
| Open Access HMGN1 enhances CRISPR-directed dual-function A-to-G and C-to-G base editing
HMGN1 enhances CRISPR-directed dual-function A-to-G and C-to-G base editing
Limited work has been done on concurrent C-to-G and A-to-G base editing. Here the authors test how a number of chromatin-associated factors affect base editing and show that HMGN1 enhanced the efficiency; by fusing HMGN1 to GBE and ABE they develop a CRISPR-based dual-function A-to-G and C-to-G base editor (GGBE).
- Chao Yang
- , Zhenzhen Ma
- & Xueli Zhang
-
Article
| Open Access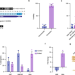 Direct correction of haemoglobin E β-thalassaemia using base editors
Direct correction of haemoglobin E β-thalassaemia using base editors
The authors demonstrate efficient and direct correction of the DNA mutation causing Haemoglobin E β-thalassaemia with CRISPR Cas9 base editors. The work includes profiling of off-target effects using deep neural networks.
- Mohsin Badat
- , Ayesha Ejaz
- & James O. J. Davies
-
Article
| Open Access ARRDC5 expression is conserved in mammalian testes and required for normal sperm morphogenesis
ARRDC5 expression is conserved in mammalian testes and required for normal sperm morphogenesis
Male organisms produce huge numbers of sperm each day, with defects in the process resulting infertility. Here they show that arrestin domain-containing 5 (ARRDC5) is a testis-specific molecule required for mammalian spermatogenesis.
- Mariana I. Giassetti
- , Deqiang Miao
- & Jon M. Oatley
-
Article
| Open Access Prime editing with genuine Cas9 nickases minimizes unwanted indels
Prime editing with genuine Cas9 nickases minimizes unwanted indels
Cas9 nickases (nCas9s) produce nicks or single-strand breaks in the DNA. Here the authors analyse the on- and off-target nicks generated by these nickases, and show that nCas9 (H840A) but not nCas9 (D10A) can cleave both strands and produce unwanted DNA double-strand breaks.
- Jaesuk Lee
- , Kayeong Lim
- & Jin-Soo Kim
-
Article
| Open Access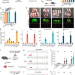 Cytosine base editors induce off-target mutations and adverse phenotypic effects in transgenic mice
Cytosine base editors induce off-target mutations and adverse phenotypic effects in transgenic mice
The potential off-target effects of long-term expression of base editors in vivo are unclear. Here the authors report SAFETI, Systematic evaluation Approach For gene Editing tools by Transgenic mIce, to examine off-target effects of base editors over time in mice, and see abnormal side effects.
- Nana Yan
- , Hu Feng
- & Erwei Zuo
-
Comment
| Open AccessA gene drive is a gene drive: the debate over lumping or splitting definitions
We address a controversy over use of the term “gene drive” to include both natural and synthetic genetic elements that promote their own transmission within a population, arguing that this broad definition is both practical and has advantages for risk analysis.
- Stephanie L. James
- , David A. O’Brochta
- & Omar S. Akbari
-
Article
| Open Access Limitations of gene editing assessments in human preimplantation embryos
Limitations of gene editing assessments in human preimplantation embryos
DNA repair in response to DSBs in the preimplantation embryo is hard to analyze. Here the authors show that over 25% of pre-existing heterozygous loci in control single blastomere samples appeared as homozygous after whole genome amplification, therefore, they validated gene editing seen in human embryos in ESCs.
- Dan Liang
- , Aleksei Mikhalchenko
- & Shoukhrat Mitalipov
-
Article
| Open Access Improving adenine and dual base editors through introduction of TadA-8e and Rad51DBD
Improving adenine and dual base editors through introduction of TadA-8e and Rad51DBD
There is a low efficiency of A-to-G base conversion in at specific positions using base editors. Here the authors fuse ABE8e with the Rad51 DNA-binding domain to generate a hyperactive ABE allowing improved A-to-G editing efficiency at the region proximal to the PAM and improved simultaneous A/C conversion efficiency.
- Niannian Xue
- , Xu Liu
- & Xiaohui Zhang
-
Article
| Open Access A versatile, high-efficiency platform for CRISPR-based gene activation
A versatile, high-efficiency platform for CRISPR-based gene activation
The generation of CRISPR-mediated transcriptional activation (CRISPRa)-competent cell lines pose significant technical challenges. Here the authors report a platform for production of CRISPRa-ready cell populations which they combine with optimised expressed and synthetic gRNA scaffolds to enhance functionality.
- Amy J. Heidersbach
- , Kristel M. Dorighi
- & Benjamin Haley
-
Article
| Open Access DddA homolog search and engineering expand sequence compatibility of mitochondrial base editing
DddA homolog search and engineering expand sequence compatibility of mitochondrial base editing
There is a need to improve and expand mitochondrial base editing tools. Here the authors identify a DddA homolog from Simiaoa sunii (Ddd_Ss) which can efficiently deaminate cytosine in dsDNA; they develop cytosine base editors and introduce mutations at multiple mitochondrial DNA loci.
- Li Mi
- , Ming Shi
- & Yangming Wang
-
Article
| Open Access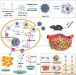 A cooperative nano-CRISPR scaffold potentiates immunotherapy via activation of tumour-intrinsic pyroptosis
A cooperative nano-CRISPR scaffold potentiates immunotherapy via activation of tumour-intrinsic pyroptosis
Delivery of immune therapy drugs to tumours might be hampered by their limited bioavailability and the difficulty of targeting complex exogenous compounds. Here authors trigger immunologic cell death, via activating tumour-cell-intrinsic pathways via CRISPR-based nanotechnology to enable efficient anti-tumour immune response in mouse models of melanoma.
- Ning Wang
- , Chao Liu
- & Changyang Gong
-
Article
| Open Access Modeling CRISPR-Cas13d on-target and off-target effects using machine learning approaches
Modeling CRISPR-Cas13d on-target and off-target effects using machine learning approaches
Application of CRISPR-Cas13d is limited by the inability to predict on- and off-targets. Here the authors perform CRISPR-Cas13d proliferation screens followed by modeling of Cas13d on- and off-targets; they design a deep learning model, DeepCas13, to predict the on-target activity of a gRNA.
- Xiaolong Cheng
- , Zexu Li
- & Wei Li
-
Article
| Open Access Systematically attenuating DNA targeting enables CRISPR-driven editing in bacteria
Systematically attenuating DNA targeting enables CRISPR-driven editing in bacteria
Genome editing in bacteria normally requires efficient recombination and high transformation efficiencies, which often isn’t. Here the authors report that systematically attenuating DNA targeting activity enables RecA-mediated repair in different bacteria, allowing chromosomal cleavage to drive editing.
- Daphne Collias
- , Elena Vialetto
- & Chase L. Beisel
-
Article
| Open Access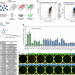 B1 SINE-binding ZFP266 impedes mouse iPSC generation through suppression of chromatin opening mediated by reprogramming factors
B1 SINE-binding ZFP266 impedes mouse iPSC generation through suppression of chromatin opening mediated by reprogramming factors
Induced pluripotent stem cell (iPSC) reprogramming is inherently inefficient. Here the authors identify 24 reprogramming roadblock genes through a CRISPR/Cas9-mediated genome-wide knockout screen including a KRAB-ZFP Zfp266, knockout of which consistently enhances murine iPSC generation.
- Daniel F. Kaemena
- , Masahito Yoshihara
- & Keisuke Kaji
-
Article
| Open Access TadA orthologs enable both cytosine and adenine editing of base editors
TadA orthologs enable both cytosine and adenine editing of base editors
Properties of cytidine and adenosine deaminases lead to off-target effects for cytosine base editors (CBEs) and adenine base editors (ABEs). Here the authors report that 25 TadA orthologs could be engineered to generate functional ABEs, CBEs or ACBEs via single/double mutations with minimised off-targets.
- Shuqian Zhang
- , Bo Yuan
- & Tian-Lin Cheng
-
Article
| Open Access TadA reprogramming to generate potent miniature base editors with high precision
TadA reprogramming to generate potent miniature base editors with high precision
Hypercompact CRISPR-Cas12f systems have been engineered to generate miniABEs but these have limitations. Here the authors generate Cas12f-derived miniCBEs and develop miniABEs with improved editing and targeting scopes; they use these to correct pathogenic mutations in cell lines and introduce mutations in vivo.
- Shuqian Zhang
- , Liting Song
- & Tian-Lin Cheng
-
Article
| Open Access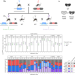 Closing the gap to effective gene drive in Aedes aegypti by exploiting germline regulatory elements
Closing the gap to effective gene drive in Aedes aegypti by exploiting germline regulatory elements
CRISPR/Cas9-based homing gene drives have emerged as a potential new approach to mosquito control. Here the authors use transgenic lines with germline-specific regulatory elements to express Cas9 and achieve up to 94% inheritance bias, closing the gap between A. aegyptidrives and the highly efficient drives observed in Anopheles species.
- Michelle A. E. Anderson
- , Estela Gonzalez
- & Luke Alphey
-
Article
| Open Access Development of a versatile nuclease prime editor with upgraded precision
Development of a versatile nuclease prime editor with upgraded precision
Strategies to improve the specificity of nuclease-based prime editor (PEn) are needed. Here the authors report a 53BP1-inhibitory ubiquitin variant-assisted PEn platform (uPEn) to inhibit NHEJ and enable precise prime editing for generation of insertions, deletions and replacements.
- Xiangyang Li
- , Guiquan Zhang
- & Xingxu Huang
-
Article
| Open Access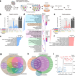 CRISPR screens reveal genetic determinants of PARP inhibitor sensitivity and resistance in prostate cancer
CRISPR screens reveal genetic determinants of PARP inhibitor sensitivity and resistance in prostate cancer
Identifying prostate cancer patients who may respond well to PARP inhibitors is important for their success in the clinic. Here, using a genome-wide CRISPR-Cas9 knockout screen, the authors identify MMS22L as a biomarker for sensitivity to PARP inhibition in BRCA1/2-proficient prostate cancer.
- Takuya Tsujino
- , Tomoaki Takai
- & Li Jia
-
Article
| Open Access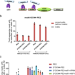 Enhancement of a prime editing system via optimal recruitment of the pioneer transcription factor P65
Enhancement of a prime editing system via optimal recruitment of the pioneer transcription factor P65
Prime editing represents a great advance to the genome editing field but is currently limited by the editing efficiency. Here the authors look to improve the efficiency by recruiting target proteins and show that the transcription factor P65 could enhance the desired editing outcomes at different gene loci.
- Ronghao Chen
- , Yu Cao
- & Xueli Zhang
-
Review Article
| Open Access Assessing and advancing the safety of CRISPR-Cas tools: from DNA to RNA editing
Assessing and advancing the safety of CRISPR-Cas tools: from DNA to RNA editing
CRISPR-Cas tools have shown exceptional promise in genome engineering over the past decade. Here the authors review the development of CRISPR-Cas9/Cas12/Cas13 nucleases, DNA base editors, prime editors, and RNA base editors, as well as their editing precision, off-target effects, and clinical considerations.
- Jianli Tao
- , Daniel E. Bauer
- & Roberto Chiarle
-
Article
| Open Access Therapeutic adenine base editing of human hematopoietic stem cells
Therapeutic adenine base editing of human hematopoietic stem cells
Here, Liao and colleagues apply adenine base editor ABE8e and its PAM-less variant ABE8e-SpRY to β-thalassemia patient hematopoietic stem cells in the form of ribonucleoprotein complexes, resulting in efficient long-term editing and β-thalassemia alleviation.
- Jiaoyang Liao
- , Shuanghong Chen
- & Yuxuan Wu
-
Article
| Open Access Genetic conversion of a split-drive into a full-drive element
Genetic conversion of a split-drive into a full-drive element
CRISPR-based gene-drives can carry the Cas9 and guide RNA (gRNA) components in a single-linked cassette or in separate elements inserted into different genomic loci. Here the authors genetically transform and compare full versus split drives, with the former performing less efficiently than predicted.
- Gerard Terradas
- , Jared B. Bennett
- & Ethan Bier
-
Article
| Open Access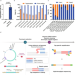 TAPE-seq is a cell-based method for predicting genome-wide off-target effects of prime editor
TAPE-seq is a cell-based method for predicting genome-wide off-target effects of prime editor
Methods to predict genome-wide off-target activities of prime editors (PEs) are currently lacking. Here the authors report a cell-based assay, TAgmentation of Prime Editor sequencing (TAPE-seq), that provides genome-wide off-target candidates for PEs.
- Jeonghun Kwon
- , Minyoung Kim
- & Jungjoon K. Lee
-
Article
| Open Access PEAC-seq adopts Prime Editor to detect CRISPR off-target and DNA translocation
PEAC-seq adopts Prime Editor to detect CRISPR off-target and DNA translocation
It is still a challenge to accurately identify off-target endonuclease edits. Here the authors report PEAC-seq using a Prime Editor to insert a tag to the editing sites and enrich the tagged regions with site-specific primers for sequencing: they show that PEAC-seq could identify DNA translocations.
- Zhenxing Yu
- , Zhike Lu
- & Lijia Ma
-
Article
| Open Access Automated high-throughput genome editing platform with an AI learning in situ prediction model
Automated high-throughput genome editing platform with an AI learning in situ prediction model
A large number of cell disease models with pathogenic SNVs are needed. Here the authors report an automated high-throughput platform to perform the genome editing process from gRNA design to the analysis of the editing results; they characterise in situ base editing outcomes.
- Siwei Li
- , Jingjing An
- & Meng Wang
-
Article
| Open Access Adenine base editing efficiently restores the function of Fanconi anemia hematopoietic stem and progenitor cells
Adenine base editing efficiently restores the function of Fanconi anemia hematopoietic stem and progenitor cells
Fanconi Anemia (FA) is caused by deficiencies in DNA repair, making it hard to correct. Here the authors report a therapeutic base editing strategy to address two of the most prevalent FANCA mutations in patient hematopoietic stem and progenitor cells.
- Sebastian M. Siegner
- , Laura Ugalde
- & Jacob E. Corn
-
Article
| Open Access SuperFi-Cas9 exhibits remarkable fidelity but severely reduced activity yet works effectively with ABE8e
SuperFi-Cas9 exhibits remarkable fidelity but severely reduced activity yet works effectively with ABE8e
Increased-fidelity SpCas9 variants have been developed, but often show proportionally reduced activity. Here the authors characterise the on-target activity and off-target propensity of SuperFi-Cas9 in mammalian cells and also see strongly reduced activity but with high fidelity features.
- Péter István Kulcsár
- , András Tálas
- & Ervin Welker
-
Article
| Open Access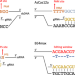 A comprehensive Bioconductor ecosystem for the design of CRISPR guide RNAs across nucleases and technologies
A comprehensive Bioconductor ecosystem for the design of CRISPR guide RNAs across nucleases and technologies
The success of CRISPR experiments relies on the choice of gRNA. Here the authors report crisprVerse, which enables efficient gRNA design and annotation for methods including CRISPRko, CRISPRa, CRISPRi, CRISPRbe and CRISPRkd, enabled for RNA- and DNA-targeting nucleases, including Cas9, Cas12 and Cas13.
- Luke Hoberecht
- , Pirunthan Perampalam
- & Jean-Philippe Fortin
-
Article
| Open Access Automated identification of sequence-tailored Cas9 proteins using massive metagenomic data
Automated identification of sequence-tailored Cas9 proteins using massive metagenomic data
Cas9 proteins require a target-adjacent sequence, the PAM, in order to cleave DNA. Here the authors develop a pipeline to accurately predict PAM sequences in order to facilitate the identification of Cas9s targeting specific sequences, including mutations.
- Matteo Ciciani
- , Michele Demozzi
- & Nicola Segata
-
Article
| Open Access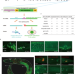 Intestine-specific removal of DAF-2 nearly doubles lifespan in Caenorhabditis elegans with little fitness cost
Intestine-specific removal of DAF-2 nearly doubles lifespan in Caenorhabditis elegans with little fitness cost
The role of lifespan-regulating protein expression in specific tissues in C. elegans is murky. Here, the authors provide clarity by inducible degradation of longevity-related proteins, demonstrating that intestinal DAF-2 is a major determinant of lifespan in the worm.
- Yan-Ping Zhang
- , Wen-Hong Zhang
- & Meng-Qiu Dong
-
Article
| Open Access CRISPR/Cas9-mediated excision of ALS/FTD-causing hexanucleotide repeat expansion in C9ORF72 rescues major disease mechanisms in vivo and in vitro
CRISPR/Cas9-mediated excision of ALS/FTD-causing hexanucleotide repeat expansion in C9ORF72 rescues major disease mechanisms in vivo and in vitro
A hexanucleotide repeat expansion in C9ORF72 is the most common genetic cause of ALS and FTD. Here, the authors demonstrate CRISPR/Cas9 excision of the expansion results in a rescue of disease mechanisms in vivo and in vitro.
- Katharina E. Meijboom
- , Abbas Abdallah
- & Christian Mueller
-
Article
| Open Access Marker-free co-selection for successive rounds of prime editing in human cells
Marker-free co-selection for successive rounds of prime editing in human cells
Prime editing enables the introduction of precise point mutations, small insertions, or short deletions without requiring donor DNA templates. Here the authors develop a co-selection strategy to facilitate prime editing in human cells and provide design principles to prevent the formation of undesired editing byproducts at the target site.
- Sébastien Levesque
- , Diana Mayorga
- & Yannick Doyon
-
Article
| Open Access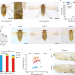 The transcription factor Zfh1 acts as a wing-morph switch in planthoppers
The transcription factor Zfh1 acts as a wing-morph switch in planthoppers
The molecular mechanisms underlying wing polyphenism remain poorly understood. Here the authors use plant hoppers to show that the development of long and short wing morphs is balanced by the relative activities of the Zfh1-FoxO and insulin signaling cascades.
- Jin-Li Zhang
- , Sun-Jie Chen
- & Hai-Jun Xu
-
Article
| Open Access Comprehensive assessment of miniature CRISPR-Cas12f nucleases for gene disruption
Comprehensive assessment of miniature CRISPR-Cas12f nucleases for gene disruption
CRISPR-Cas12f nucleases can be effectively packaged into AAVs for gene therapy, but a systematic evaluation of editing outcomes is lacking. Here the authors perform a comprehensive assessment of 4 Cas12f proteins and compare to Cas9 and two Cas12a proteins at a number of sites.
- Changchang Xin
- , Jianhang Yin
- & Jiazhi Hu
-
Article
| Open Access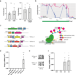 Activation of stably silenced genes by recruitment of a synthetic de-methylating module
Activation of stably silenced genes by recruitment of a synthetic de-methylating module
Stably silenced genes with methylated CpG at the promoter are refractory to current CRISPR activation systems. Here the authors create a more robust activation system, TETact that recruits DNA-demethylating TET1 with transcriptional activators.
- Wing Fuk Chan
- , Hannah D. Coughlan
- & Rhys S. Allan
-
Article
| Open Access Multi-pathway DNA-repair reporters reveal competition between end-joining, single-strand annealing and homologous recombination at Cas9-induced DNA double-strand breaks
Multi-pathway DNA-repair reporters reveal competition between end-joining, single-strand annealing and homologous recombination at Cas9-induced DNA double-strand breaks
Correct repair of broken DNA molecules is required to prevent potentially oncogenic mutations. To study repair fidelity and mechanism, van de Kooij et al. developed single cell reporters that detect if DNA breaks are fixed by error-free or mutagenic repair.
- Bert van de Kooij
- , Alex Kruswick
- & Michael B. Yaffe
-
Article
| Open Access Accounting for small variations in the tracrRNA sequence improves sgRNA activity predictions for CRISPR screening
Accounting for small variations in the tracrRNA sequence improves sgRNA activity predictions for CRISPR screening
Existing methods for generating sgRNA predictions do not account for the tracrRNA sequence. Here the authors report an on-target model, Rule Set 3, to generate optimal predictions for multiple tracrRNA variants, and validate this on a new dataset of sgRNAs showing improvement over prior prediction models.
- Peter C. DeWeirdt
- , Abby V. McGee
- & John G. Doench
-
Article
| Open Access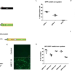 In vivo adenine base editing reverts C282Y and improves iron metabolism in hemochromatosis mice
In vivo adenine base editing reverts C282Y and improves iron metabolism in hemochromatosis mice
Hemochromatosis is a metabolic disorder caused by mutations in the HFE gene. Here, the authors show that a single administration of AAV8 vectors expressing an Adenine Base Editor facilitates efficient in vivo gene correction in hepatocytes and leads to improvement of iron-specific parameters in the liver and the blood in mouse models of the disease.
- Alice Rovai
- , BoMee Chung
- & Michael Ott

 Programmable RNA detection with CRISPR-Cas12a
Programmable RNA detection with CRISPR-Cas12a