Featured
-
-
Article
| Open Access Harnessing noncanonical crRNA for highly efficient genome editing
Harnessing noncanonical crRNA for highly efficient genome editing
The inclusion of base Z has the potential to heighten the binding affinity between complementary nucleic acids. Here, the authors integrated base Z into CRISRP-Cas12a crRNA to augment the interaction between the crRNA and the target DNA, resulting in a significant enhancement of editing efficiency.
- Guanhua Xun
- , Zhixin Zhu
- & Huimin Zhao
-
Article
| Open Access In vivo genome editing via CRISPR/Cas9-mediated homology-independent targeted integration for Bietti crystalline corneoretinal dystrophy treatment
In vivo genome editing via CRISPR/Cas9-mediated homology-independent targeted integration for Bietti crystalline corneoretinal dystrophy treatment
Bietti crystalline corneoretinal dystrophy (BCD) is an autosomal recessive chorioretinal degenerative disease without approved therapeutic drug. Here, the authors show a promising CRISPR/Cas9 mediated homology-independent targeted integration therapy in patient derived cells and humanized mice carrying BCD mutations.
- Xiang Meng
- , Ruixuan Jia
- & Liping Yang
-
Article
| Open Access Unraveling the mechanisms of PAMless DNA interrogation by SpRY-Cas9
Unraveling the mechanisms of PAMless DNA interrogation by SpRY-Cas9
CRISPR-Cas9 is a powerful tool, but the strict requirement for an “NGG” protospacer-adjacent motif (PAM) sequence limits the number of editable genes. Here the authors combine enzyme kinetics, cryo-EM, and single-molecule imaging to determine how SpRY interrogates DNA and recognises target sites for cleavage.
- Grace N. Hibshman
- , Jack P. K. Bravo
- & David W. Taylor
-
Article
| Open Access Efficient gene knockout and genetic interaction screening using the in4mer CRISPR/Cas12a multiplex knockout platform
Efficient gene knockout and genetic interaction screening using the in4mer CRISPR/Cas12a multiplex knockout platform
Paralog synthetic lethals have been assessed with multiple CRISPR-based methods, but systematic comparison among these platforms is unavailable. Here, the authors systematically compare combinatorial perturbation platforms and establish the in4mer CRISPR/Cas12a multiplex knockout platform.
- Nazanin Esmaeili Anvar
- , Chenchu Lin
- & Traver Hart
-
Article
| Open Access Flexible TAM requirement of TnpB enables efficient single-nucleotide editing with expanded targeting scope
Flexible TAM requirement of TnpB enables efficient single-nucleotide editing with expanded targeting scope
Here the authors report that a thermophilic archaeal TnpB enables efficient gene editing in the natural host: they see that the TnpB has different TAM requirements for eliciting cell death and for facilitating gene editing. They show that TnpB can be harnessed for flexible single-nucleotide editing with templated repair.
- Xu Feng
- , Ruyi Xu
- & Qunxin She
-
Article
| Open Access CoCas9 is a compact nuclease from the human microbiome for efficient and precise genome editing
CoCas9 is a compact nuclease from the human microbiome for efficient and precise genome editing
Cas9 nucleases hold clinical significance for genome editing therapies. Here the authors characterize CoCas9, a compact, efficient and precise Cas9 from the human microbiome, and show that delivery via AAV vectors enables efficient editing in the mouse retina, expanding the genome editing toolbox.
- Eleonora Pedrazzoli
- , Michele Demozzi
- & Anna Cereseto
-
Article
| Open Access Splice modulators target PMS1 to reduce somatic expansion of the Huntington’s disease-associated CAG repeat
Splice modulators target PMS1 to reduce somatic expansion of the Huntington’s disease-associated CAG repeat
Somatic expansion of a CAG repeat in HTT drives onset of Huntington’s disease. Using a human cell line model and splice modulators, here the authors show that PMS1 is an enhancer of CAG repeat expansion, making it a target for therapeutic intervention.
- Zachariah L. McLean
- , Dadi Gao
- & James F. Gusella
-
Article
| Open Access Developmental progression of DNA double-strand break repair deciphered by a single-allele resolution mutation classifier
Developmental progression of DNA double-strand break repair deciphered by a single-allele resolution mutation classifier
DNA double-strand breaks (DSBs) are repaired by a hierarchically regulated network of pathways. Here, authors develop ICP for deciphering somatic DSB repair patterns in multicellular organisms and discover developmental regulation in flies and mosquitoes, enabling tracking of mutant alleles and interhomolog copying of gene cassettes.
- Zhiqian Li
- , Lang You
- & Ethan Bier
-
Article
| Open Access Identifying regulators of aberrant stem cell and differentiation activity in colorectal cancer using a dual endogenous reporter system
Identifying regulators of aberrant stem cell and differentiation activity in colorectal cancer using a dual endogenous reporter system
Aberrant stem cell-like activity and impaired differentiation are central to the development of colorectal cancer. Here, authors develop a dual endogenous reporter system to identify functional regulators of aberrant stem cell and differentiation programs, showing that SMARCB1 restricts differentiation, and nominating other regulators with therapeutic potential.
- Sandor Spisak
- , David Chen
- & Nilay S. Sethi
-
Article
| Open Access Enhancing prime editor activity by directed protein evolution in yeast
Enhancing prime editor activity by directed protein evolution in yeast
Compared to traditional Cas9 nucleases prime editors (PEs) are less active. Here the authors use OrthoRep, a yeast-based platform for directed protein evolution to enhance the editing efficiency of PEs: they identify mutations that have a positive effect on kinetics and use this knowledge to generate an efficient in vivo PE.
- Yanik Weber
- , Desirée Böck
- & Gerald Schwank
-
Article
| Open Access Cas9-assisted biological containment of a genetically engineered human commensal bacterium and genetic elements
Cas9-assisted biological containment of a genetically engineered human commensal bacterium and genetic elements
Engineered biosensing bacteria can potentially probe the human gut microbiome to prevent, diagnose, or treat disease. Here the authors present a robust biocontainment assisted by Cas9 and an engineered gene expression control combined in a genetically engineered human commensal bacterium that successfully functioned in a mouse intestinal tract as well as cell culture condition.
- Naoki Hayashi
- , Yong Lai
- & Timothy K. Lu
-
Article
| Open Access Topological barrier to Cas12a activation by circular DNA nanostructures facilitates autocatalysis and transforms DNA/RNA sensing
Topological barrier to Cas12a activation by circular DNA nanostructures facilitates autocatalysis and transforms DNA/RNA sensing
The authors find that small circular DNA nanostructures which partially match gRNA sequences only minimally activate Cas12a. They report AutoCAR (Autocatalytic Cas12a Circular DNA Amplification Reaction) which allows a single nucleic acid target to activate multiple ribonucleoproteins, and increases reporter cleavage rates.
- Fei Deng
- , Yi Li
- & Ewa M. Goldys
-
Article
| Open Access Development of pathophysiologically relevant models of sickle cell disease and β-thalassemia for therapeutic studies
Development of pathophysiologically relevant models of sickle cell disease and β-thalassemia for therapeutic studies
Sickle cell disease (SCD) and β-thalassemia (BT) are globally prevalent inherited blood disorders but, despite extensive research, no ex vivo system exists for SCD and BT. Here, the authors generate pathophysiologically relevant erythroid progenitor models of SCD and BT.
- Pragya Gupta
- , Sangam Giri Goswami
- & Sivaprakash Ramalingam
-
Article
| Open Access Engineering self-deliverable ribonucleoproteins for genome editing in the brain
Engineering self-deliverable ribonucleoproteins for genome editing in the brain
The delivery of CRISPR RNPs has potential advantages over other genome editing approaches, including reduced off-target editing and reduced immunogenicity. Here the authors report self-deliverable Cas9 RNPs capable of robustly editing cultured cells in vitro and the mouse brain upon direct injections.
- Kai Chen
- , Elizabeth C. Stahl
- & Jennifer A. Doudna
-
Article
| Open Access Phage-assisted evolution of highly active cytosine base editors with enhanced selectivity and minimal sequence context preference
Phage-assisted evolution of highly active cytosine base editors with enhanced selectivity and minimal sequence context preference
Existing TadA-derived CBEs exhibit residual A•T-to-G•C editing activity and suffer from lower activity at several sequence contexts and with non-SpCas9 targeting domains. Here, the authors use phage-assisted evolution to evolve CBE6 variants that address these limitations.
- Emily Zhang
- , Monica E. Neugebauer
- & David R. Liu
-
Article
| Open Access Orthogonal inducible control of Cas13 circuits enables programmable RNA regulation in mammalian cells
Orthogonal inducible control of Cas13 circuits enables programmable RNA regulation in mammalian cells
The lack of control over Cas13 activity has limited its utility. Here the authors report Control of RNA with Inducible SpliT CAs13 Orthologs and Exogenous Ligands (CRISTAL), controlled by orthogonal split inducible Cas13 effectors that can be turned ON or OFF, providing precise temporal control.
- Yage Ding
- , Cristina Tous
- & Wilson W. Wong
-
Article
| Open Access Domain-inlaid Nme2Cas9 adenine base editors with improved activity and targeting scope
Domain-inlaid Nme2Cas9 adenine base editors with improved activity and targeting scope
Nme2Cas9 has been well established as a genome editing platform. Here the authors engineer Nme2Cas9 to further increase the activity and targeting scope of compact Nme2Cas9 base editors and validate domain-inlaid Nme2-ABEs for single-AAV delivery in vivo.
- Nathan Bamidele
- , Han Zhang
- & Erik J. Sontheimer
-
Article
| Open Access Compact zinc finger architecture utilizing toxin-derived cytidine deaminases for highly efficient base editing in human cells
Compact zinc finger architecture utilizing toxin-derived cytidine deaminases for highly efficient base editing in human cells
The most recent class of base editors utilize DddAtox, a deaminase domain that can act upon double-stranded DNA. Here the authors target DddAtox fragments and a FokI-based nickase to the human CIITA gene by fusing these domains to arrays of engineered zinc fingers; they also identify a variety of DddAtox orthologues.
- Friedrich Fauser
- , Bhakti N. Kadam
- & Jeffrey C. Miller
-
Article
| Open Access Logical design of synthetic cis-regulatory DNA for genetic tracing of cell identities and state changes
Logical design of synthetic cis-regulatory DNA for genetic tracing of cell identities and state changes
Descriptive data in biomedical research are expanding rapidly, but functional validation methods lag behind. Here, authors present Logical Synthetic cis-regulatory DNA, a framework to design reporters that mark cellular states and pathways, showcasing its applicability to complex phenotypic states.
- Carlos Company
- , Matthias Jürgen Schmitt
- & Gaetano Gargiulo
-
Article
| Open Access Anti-CRISPR Anopheles mosquitoes inhibit gene drive spread under challenging behavioural conditions in large cages
Anti-CRISPR Anopheles mosquitoes inhibit gene drive spread under challenging behavioural conditions in large cages
CRISPR-based gene drives have the potential to spread within populations and are considered as promising vector control tools. Here the authors show an anti-drive mosquito strain that prevents the spread and collapse of a population suppression gene drive in laboratory Anopheles mosquito large cage trials in complex ecological and behavioral conditions.
- Rocco D’Amato
- , Chrysanthi Taxiarchi
- & Ruth Müller
-
Article
| Open Access BacPE: a versatile prime-editing platform in bacteria by inhibiting DNA exonucleases
BacPE: a versatile prime-editing platform in bacteria by inhibiting DNA exonucleases
Prime editing in bacteria is currently inefficient. Here the authors report BacPE, a versatile prime editing platform in Escherichia coli that works by inhibiting 3′→5′ DNA exonucleases, highlighting the intrinsic genetic factors that are adverse to efficient prime editing.
- Hongyuan Zhang
- , Jiacheng Ma
- & Quanjiang Ji
-
Article
| Open Access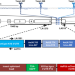 A multiplexed, confinable CRISPR/Cas9 gene drive can propagate in caged Aedes aegypti populations
A multiplexed, confinable CRISPR/Cas9 gene drive can propagate in caged Aedes aegypti populations
Aedes aegypti is the main vector of several major pathogens including dengue, Zika and chikungunya viruses. Here the authors find that a CRISPR/Cas9 based split gene drive in Aedes aegypti could successfully bias inheritance up to 89% over successive generations in a multi-cage trial with further deep sequencing suggesting that the multiplexing design could mitigate resistance allele formation.
- Michelle A. E. Anderson
- , Estela Gonzalez
- & Luke Alphey
-
Article
| Open Access Programmable RNA base editing with photoactivatable CRISPR-Cas13
Programmable RNA base editing with photoactivatable CRISPR-Cas13
Cas13 systems suffer from a lack of spatiotemporal control. Here the authors report paCas13, a light-inducible Cas13 system created by fusing Magnet with fragment pairs; they also report padCas13, a light-inducible base-editing system by fusing ADAR2 to catalytically inactive paCas13 fragments.
- Jeonghye Yu
- , Jongpil Shin
- & Won Do Heo
-
Article
| Open Access The low-density lipoprotein receptor promotes infection of multiple encephalitic alphaviruses
The low-density lipoprotein receptor promotes infection of multiple encephalitic alphaviruses
Ma et al. identify LDLR as an entry receptor for Eastern equine encephalitis virus (EEEV) and other alphaviruses. Soluble decoy proteins with multiple LA domain 3 repeats of LDLR inhibit EEEV infection in cell culture and mice.
- Hongming Ma
- , Lucas J. Adams
- & Michael S. Diamond
-
Article
| Open Access CRISPR-Cas-based identification of a sialylated human milk oligosaccharides utilization cluster in the infant gut commensal Bacteroides dorei
CRISPR-Cas-based identification of a sialylated human milk oligosaccharides utilization cluster in the infant gut commensal Bacteroides dorei
Human milk oligosaccharides (HMOs) utilization by Bacteroides species remains poorly understood. Here, the authors describe a single specific gene cluster responsible for sialylated HMOs utilization in a B. dorei natural isolate and prove its functionality in vivo using CRISPR-Cas12a.
- Sivan Kijner
- , Dena Ennis
- & Moran Yassour
-
Article
| Open Access Transient inhibition of 53BP1 increases the frequency of targeted integration in human hematopoietic stem and progenitor cells
Transient inhibition of 53BP1 increases the frequency of targeted integration in human hematopoietic stem and progenitor cells
Here the authors demonstrate that the frequency of HDR in human hematopoietic stem and progenitor cells is increased by the delivery of an inhibitor of 53BP1 as a recombinant peptide. This approach is applicable for a variety of therapeutically relevant loci in HSPCs as well in other primary human cell types.
- Ron Baik
- , M. Kyle Cromer
- & Matthew H. Porteus
-
Article
| Open Access Shuttle peptide delivers base editor RNPs to rhesus monkey airway epithelial cells in vivo
Shuttle peptide delivers base editor RNPs to rhesus monkey airway epithelial cells in vivo
Gene editing strategies for cystic fibrosis are challenging. Here the authors improve on their previously reported shuttle peptide noncovalently combined with Cas ribonucleoprotein (RNP), and derive the S315 peptide for delivery: they show base editing in the respiratory tract of the rhesus macaques.
- Katarina Kulhankova
- , Soumba Traore
- & Paul B. McCray Jr.
-
Article
| Open Access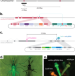 CRISPR-based gene drives generate super-Mendelian inheritance in the disease vector Culex quinquefasciatus
CRISPR-based gene drives generate super-Mendelian inheritance in the disease vector Culex quinquefasciatus
Culex mosquitoes are carriers of major diseases like West Nile virus and are a public health concern. Here the authors present a CRISPR-Cas9 gene drive as a control technology in the Culex quinquefasciatus mosquito species.
- Tim Harvey-Samuel
- , Xuechun Feng
- & Valentino M. Gantz
-
Article
| Open Access Asymmetric CRISPR enabling cascade signal amplification for nucleic acid detection by competitive crRNA
Asymmetric CRISPR enabling cascade signal amplification for nucleic acid detection by competitive crRNA
New strategies are being developed to simplify CRISPR-based nucleic acid detection. By investigating the competitive reaction between a full-sized crRNA and split crRNA for CRISPR-Cas12a, the authors develop an asymmetric CRISPR assay for amplification-free, cascade signal amplification detection of nucleic acids.
- Jeong Moon
- & Changchun Liu
-
Article
| Open Access Lung SORT LNPs enable precise homology-directed repair mediated CRISPR/Cas genome correction in cystic fibrosis models
Lung SORT LNPs enable precise homology-directed repair mediated CRISPR/Cas genome correction in cystic fibrosis models
Roughly 10% of Cystic Fibrosis (CF) patients still have no effective medicine to take. Lung Selective Organ Targeting (SORT) Lipid Nanoparticles can efficiently deliver Cas9 mRNA, sgRNA, and donor ssDNA templates for precise homology-directed repair-mediated gene correction in ex vivo and in vivo CF models.
- Tuo Wei
- , Yehui Sun
- & Daniel J. Siegwart
-
Article
| Open Access Specific heterozygous variants in MGP lead to endoplasmic reticulum stress and cause spondyloepiphyseal dysplasia
Specific heterozygous variants in MGP lead to endoplasmic reticulum stress and cause spondyloepiphyseal dysplasia
Biallelic loss-of-function variants in the gene encoding Matrix Gla Protein (MGP) are known to cause a recessive disorder called Keutel syndrome. Here, the authors report that heterozygous missense variants affecting one particular cysteine residue of MGP can cause a clinically distinct, dominant disorder, likely via impaired signal peptide processing leading to cellular stress and apoptosis.
- Ophélie Gourgas
- , Gabrielle Lemire
- & Monzur Murshed
-
Article
| Open Access Hidden prevalence of deletion-inversion bi-alleles in CRISPR-mediated deletions of tandemly arrayed genes in plants
Hidden prevalence of deletion-inversion bi-alleles in CRISPR-mediated deletions of tandemly arrayed genes in plants
The multiplex CRISPR system is the tool of choice for creating targeted tandemly arrayed genes (TAGs) deletions in plants. Here, the authors show that up to 80% of CRISPR-mediated TAG knockout alleles in Arabidopsis and rice are deletion-inversion bi-alleles, an unwanted products of targeted TAG deletions.
- Jiuer Liu
- , Feng-Zhu Wang
- & Jian-Feng Li
-
Article
| Open Access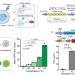 Sonogenetic control of multiplexed genome regulation and base editing
Sonogenetic control of multiplexed genome regulation and base editing
Exogenous control of genes in vivo is important. Here the authors report a system that can be inducibly activated through thermal energy produced by ultrasound absorption and use this to control induction of gene activation and base editing: they apply this in cell lines and in a mouse model.
- Pei Liu
- , Josquin Foiret
- & Lei S. Qi
-
Article
| Open Access Mitigating a TDP-43 proteinopathy by targeting ataxin-2 using RNA-targeting CRISPR effector proteins
Mitigating a TDP-43 proteinopathy by targeting ataxin-2 using RNA-targeting CRISPR effector proteins
TDP43 proteinopathies are a devastating group of neurodegenerative disorders. Here the authors show that RNA-targeting CRISPR effector proteins can be used to mitigate TDP-43 pathology when targeting ataxin-2, a modifier of TDP-43-associated toxicity, and apply this to a mouse model.
- M. Alejandra Zeballos C.
- , Hayden J. Moore
- & Thomas Gaj
-
Article
| Open Access Lipid nanoparticles with PEG-variant surface modifications mediate genome editing in the mouse retina
Lipid nanoparticles with PEG-variant surface modifications mediate genome editing in the mouse retina
There is a need for development of efficient delivery vehicles for the treatment of inherited retinal degeneration with gene therapy. Here, Gautam et al., show that surface modifications of lipid nanoparticles with PEG variants alters their cellular tropism allowing gene editing in diverse retinal cell types in mice.
- Milan Gautam
- , Antony Jozic
- & Gaurav Sahay
-
Article
| Open Access Next-generation CRISPR gene-drive systems using Cas12a nuclease
Next-generation CRISPR gene-drive systems using Cas12a nuclease
One method for reducing the impact of vector-borne diseases is through the use of CRISPR-based gene drives, which manipulate insect populations due to their ability to rapidly propagate desired genetic traits into a target population. Here the authors describe a Cas12a gene drive system whose activity can be finetuned in a temperature-dependent manner.
- Sara Sanz Juste
- , Emily M. Okamoto
- & Víctor López Del Amo
-
Article
| Open Access PAM-flexible genome editing with an engineered chimeric Cas9
PAM-flexible genome editing with an engineered chimeric Cas9
CRISPR enzymes require a defined protospacer adjacent motif (PAM) which can be limiting for editing applications. Here the authors recombine the PAM-interacting domain of SpRY with the N-terminus of Sc + + to generate a chimeric enzyme with highly flexible PAM preference: SpRYc.
- Lin Zhao
- , Sabrina R. T. Koseki
- & Pranam Chatterjee
-
Article
| Open Access A thermostable type I-B CRISPR-Cas system for orthogonal and multiplexed genetic engineering
A thermostable type I-B CRISPR-Cas system for orthogonal and multiplexed genetic engineering
Thermophilic genetic manipulation tools are limited. Here the authors report a thermophilic type I-B CRISPR-Cas system and show it displays efficient transcriptional repression or DNA cleavage activity: they develop a tool for genome editing and transcriptional repression in both thermophile and mesophile hosts.
- Zhiheng Yang
- , Zilong Li
- & Weishan Wang
-
Article
| Open Access Fast, multiplexable and efficient somatic gene deletions in adult mouse skeletal muscle fibers using AAV-CRISPR/Cas9
Fast, multiplexable and efficient somatic gene deletions in adult mouse skeletal muscle fibers using AAV-CRISPR/Cas9
Methods for somatic gene perturbation would offer advantages for screening multiple muscle gene candidates. Here the authors couple Cre-mediated skeletal muscle fiber-specific Cas9 expression with myotropic adeno-associated virus-mediated sgRNA delivery and report a system for effective somatic gene deletions in mice.
- Marco Thürkauf
- , Shuo Lin
- & Markus A. Rüegg
-
Article
| Open Access Efficient plant genome engineering using a probiotic sourced CRISPR-Cas9 system
Efficient plant genome engineering using a probiotic sourced CRISPR-Cas9 system
In the field of plant genome engineering, new nucleases with improved editing efficiency and alterative PAM requirements are needed. Here, the authors report a probiotic sourced CRISPR-LrCas9 system with similar PAM requirement to Cas12a and show its high efficiencies in various genome editing applications.
- Zhaohui Zhong
- , Guanqing Liu
- & Yong Zhang
-
Article
| Open Access A cleavage rule for selection of increased-fidelity SpCas9 variants with high efficiency and no detectable off-targets
A cleavage rule for selection of increased-fidelity SpCas9 variants with high efficiency and no detectable off-targets
SpCas9 off-targets are a safety concern. Here the authors report a cleavage rule that governs the on-target and off-target cleavage of increased(/high)-fidelity SpCas9 variants: the variants have differences in fidelity small enough to comprise an optimal variant for each target, irrespective of its cleavability ranking.
- Péter István Kulcsár
- , András Tálas
- & Ervin Welker
-
Article
| Open Access Inducing multiple nicks promotes interhomolog homologous recombination to correct heterozygous mutations in somatic cells
Inducing multiple nicks promotes interhomolog homologous recombination to correct heterozygous mutations in somatic cells
Gene editing is still hampered by unintended genomic alterations. Here the authors propose a method for correcting heterozygous mutations that employs multiple nicks induced by Cas9 nickase and the homologous chromosome as an endogenous repair template (NICER): this rarely induces unintended genomic alterations.
- Akiko Tomita
- , Hiroyuki Sasanuma
- & Shinichiro Nakada
-
Article
| Open Access A split and inducible adenine base editor for precise in vivo base editing
A split and inducible adenine base editor for precise in vivo base editing
TadA deaminases widely used in many base editors lack post-translational control in cells. Here the authors report a split adenine base editor (sABE) using chemically induced dimerisation (CID) to control the catalytic activity of TadA8e and show this can be used for PCSK9 gene editing in the mouse liver.
- Hongzhi Zeng
- , Qichen Yuan
- & Xue Gao
-
Article
| Open Access A strategy for Cas13 miniaturization based on the structure and AlphaFold
A strategy for Cas13 miniaturization based on the structure and AlphaFold
Small Cas enzymes are required for therapeutic use. Here the authors report an Interaction, Dynamics and Conservation (IDC) strategy for protein miniaturisation and use this to generate five compact variants of Cas13 based on a combination of IDC strategy and AlphaFold2.
- Feiyu Zhao
- , Tao Zhang
- & Zhanjun Li
-
Article
| Open Access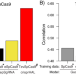 A generalizable Cas9/sgRNA prediction model using machine transfer learning with small high-quality datasets
A generalizable Cas9/sgRNA prediction model using machine transfer learning with small high-quality datasets
Current bacterial sgRNA activity models struggle with accurate predictions and generalizations. Here the authors report crisprHAL, a machine learning architecture that can be trained on existing datasets, and shows good sgRNA activity prediction accuracy can generalize predictions to different bacteria.
- Dalton T. Ham
- , Tyler S. Browne
- & David R. Edgell
-
Article
| Open Access A Type II-B Cas9 nuclease with minimized off-targets and reduced chromosomal translocations in vivo
A Type II-B Cas9 nuclease with minimized off-targets and reduced chromosomal translocations in vivo
SpCas9 unintended editing is a major concern. Here the authors report an off-target method using Duplex Sequencing with increased sensitivity for Cas9 mutation detection; they also identify a Cas9 variant of the II-B subfamily with intrinsic high fidelity (PsCas9) and see improved specificity.
- Burcu Bestas
- , Sandra Wimberger
- & Marcello Maresca
-
Article
| Open Access Programmable RNA detection with CRISPR-Cas12a
Programmable RNA detection with CRISPR-Cas12a
Cas12a is widely used in diagnostic platforms. Here the authors show that Cas12a can be programmed to directly detect RNA substrates, this is due to the 3’-end of the crRNA tolerating both RNA and DNA substrates: they use this to report a method, SAHARA, to detect RNA sequences.
- Santosh R. Rananaware
- , Emma K. Vesco
- & Piyush K. Jain
-
Article
| Open Access Prediction of base editor off-targets by deep learning
Prediction of base editor off-targets by deep learning
Base editors can induce unwanted off-target effects. Here the authors design libraries of gRNA-off-target pairs and perform a screen to obtain editing efficiencies for ABE and CBE: they use the datasets to train DL models (ABEdeepoff and CBEdeepoff) which can predict mutation tolerance at potential off-targets.
- Chengdong Zhang
- , Yuan Yang
- & Yongming Wang
-
Article
| Open Access Whole genomic analysis reveals atypical non-homologous off-target large structural variants induced by CRISPR-Cas9-mediated genome editing
Whole genomic analysis reveals atypical non-homologous off-target large structural variants induced by CRISPR-Cas9-mediated genome editing
The safety of CRISPR-Cas9 editing is a concern. Here the authors use whole genomic analysis by 10x linked-read sequencing and optical genome mapping to interrogate the genome integrity after editing: they see large structural variants at on-target sites and unexpected large chromosomal deletions.
- Hsiu-Hui Tsai
- , Hsiao-Jung Kao
- & John Yu

 Enhancing genome editing in hPSCs through dual inhibition of DNA damage response and repair pathways
Enhancing genome editing in hPSCs through dual inhibition of DNA damage response and repair pathways