Abstract
Insect wing polyphenism is characterized by its ability to produce two or more distinct wing morphs from a single genotype in response to changing environments. However, the molecular basis of this phenomenon remains poorly understood. Here, we identified a zinc finger homeodomain transcription factor Zfh1 that acts as an upstream regulator for the development of long-winged (LW) or shorted-winged (SW) morphs in planthoppers. Knockdown of Zfh1 directs SW-destined nymphs to develop into LW morphs by down-regulating the transcriptional level of FoxO, a prominent downstream effector of the insulin/IGF signaling (IIS) pathway. The balance between transcriptional regulation via the Zfh1-FoxO cascade and post-translational regulation via the IIS-FoxO cascade provides a flexible regulatory mechanism for the development of alternative wing morphs. These findings help us understand how phenotypic diversity is generated by altering the activity of conserved proteins, and provide an extended framework for the evolution of wing morphological diversity in insects.
Similar content being viewed by others
Introduction
The evolution of wings was one of the most important events in the diversification of insects1,2. Yet despite the evolutionary advantages of flight, nearly all originally winged insect orders possess many partially winged or secondarily wingless lineages3,4,5. Wing polymorphism is an especially common version of this phenomenon, in which both flying and flightless forms are contained within the same population. Typically, the flying morph possesses fully developed wings and flight muscles, suited for long-distance dispersal. By contrast, the flightless morph possesses vestigial or no wings as well as undeveloped flight muscles, and does not disperse. Winged and wingless morphs maintain high fitness by specializing on different life histories: a winged morph able to disperse when local conditions are declining, and a wingless and more sedentary morph able to invest heavily into reproduction when local conditions are favorable6,7. Wing polymorphic insects therefore provide many promising opportunities to investigate the adaptive significance of dispersal polymorphisms and more generally the evolution of alternative life histories. Doing so, however, is hampered by our incomplete understanding of the molecular basis underlying the development of alternative winged and wingless morphs.
Studies in diverse taxa including aphids, crickets, and planthoppers indicate that insect wing polymorphism can be caused by environmental cues encountered during particular developmental stages (polyphenism), by different genotypes, or by a combination of both7,8,9. Juvenile hormone (JH), in particular, has been the main subject of studies into the endocrine regulation of wing polyphenism in diverse insect species, yet persuasive direct evidence documenting a functional role of JH in wing polyphenisms is still lacking10,11,12. In contrast, growing evidence indicates that insect wing polyphenism might be regulated by diverse genes and gene regulatory networks whose identities may depend greatly on the taxa under study: for example, winged or wingless morphs are regulated, at least in part, by ecdysone signaling in female aphids Acyrthosiphon pisum13, whereas the male dimorphism is genetically controlled by a single locus on the X chromosome14,15,16. In another aphid species, Aphis citricidus, small RNAs (e.g., miR-9b) also contribute significantly to wing dimorphism17.
These results contrast to findings in the wing polymorphic brown planthopper (BPH), Nilaparvata lugens (Hemiptera: Delphacidae), a destructive rice pest in Asia. During nymphal development, comprised of five nymphal stages, BPH wing buds grow roughly proportionally with body size. Nymphs eventually mature into distinct, short (SW)- and long-winged (LW) morphs which become externally distinguishable for the first time following the terminal nymphal-to-adult molt. SW adults possess vestigial forewings and rudimentary hindwing buds, in contrast to fully developed forewings and hindwings typical of LW adults (Fig. 1a, Supplementary Fig. 1a). Diverse environmental cues, including crowding, host plant quality, photoperiod, and temperature, have been shown to influence BPH wing-morph switching, although dominant environmental regulators—if they indeed exist—remain yet to be identified12,18. The ability to develop into the LW morph enables BPHs to migrate northward from tropical to subtropical areas in the spring, followed by returning migrations in the autumn, causing severe rice yield loss throughout Asia19,20,21. Previous studies showed that the forkhead transcription factor subgroup O (FoxO) serves as a key regulator that directs the development of LW or SW morphs22,23, and may play an important role in interfacing with environmental cues such as wounding and stress24,25. Additional studies indicated that the insulin/IGF signaling (IIS) pathway affects BPH wing dimorphism via modulating the phosphorylation level of FoxO22,26,27, resulting in phosphorylated FoxO to be excluded from the nucleus and thus unable to inhibit wing development, thereby leading to LW morphs. Interestingly, the IIS pathway has also been implicated in the regulation of wing dimorphism in both the red-shouldered soapberry bug Jadera haematoloma28 and the European firebug Pyrrhocoris apterus29, likely constituting independent yet parallel co-option events. Yet despite the prominence of the IIS-FoxO signaling axis in the regulation of wing dimorphism, we still know little about how it interfaces with the gene regulatory networks underpinning wing formation in insects.
a Morphology of wild-type long-winged (LW) and short-winged (SW) female adults. Arrows, forewings. Arrowheads, hindwings. b Differentially expressed transcription factors between LW-destined (WtLW) and SW-destined (WtSW) fifth-instar nymphs. Thoracic nota were dissected from fifth-instar nymphs for RNA sequencing. Genes differentially expressed between WtLW and WtSW with a false discovery rate (FDR) < 0.01 are considered to be significant. The Zfh1 protein contains a homeodomain (orange rectangle) and nine zinc-finger motifs (blue boxes). Regions targeted by dsRNAs (dsZfh1 and dsZfh1_2) are underlined. The PCR amplified region used to examine RNAi efficiency is indicated by arrows. c RNAi efficiency of dsZfh1 and dsZfh1_2. Third-instar WtSW nymphs were microinjected with dsZfh1 or dsZfh1_2. Individuals (n = 5) were collected for qRT-PCR assay two days after microinjection. The relative expression of Zfh1 was normalized to the expression of the ribosomal protein S11 gene (rps11). The experiments were repeated three times with similar results (circles). Data are presented as mean ± SEM. Two-tailed unpaired t-test was used for the statistical analysis (** P = 0.0028 for dsZfh1 vs dsGfp and ** P = 0.0011 for dsZfh1_2 vs dsGfp). d Morphology of dsGfpSW, dsZfh1LW, and dsZfh1IMW females. dsGfpSW, dsGfp-treated BPHs with short wings (SW). dsZfh1LWand dsZfh1IMW, dsZfh1-treated BPHs with long and intermediate-size wings (IMW), respectively. Scale bars in (a) and (c), 1 mm. e Number of females with different wing morphs following dsRNA treatments. The LW ratio is compared between two groups using Pearson χ2 test (****P = 1.1716E-56, χ2 = 251.586 and df = 1 for dsZfh1 vs dsGfp; ****P = 4.9769E-45, χ2 = 198.2772 and df = 1 for dsZfh1_2 vs dsGfp). f Wing size and hind tibia length in dsZfh1LW, WtLW and WtSW females. Each circle represents an individual female (n = 20). g Vein patterning on forewings from WtLW and dsZfh1LW females. Twenty samples were repeated independently with similar results. Scale bars, 500 μm. Source data are provided as a Source Data file.
In this work, we performed comparative transcriptomic analysis on SW-destined and LW-destined BPH nymphs, and identified a zinc-finger homeodomain transcription factor Zfh1 that acts as a wing-morph switch in planthoppers. We find that knockdown of Zfh1 directs SW-destined nymphs to develop into LW morphs by affecting the transcription of FoxO. In addition, we find that Zfh1 also acts independently of both the IIS and Ultrabithorax pathways in regulating wing-morph development. These findings deepen our understanding of the genetic bases underpinning insect wing polymorphism.
Results
Knockdown of Zfh1 results in LW morph development
To screen potential wing-morph regulators, we conducted RNA sequencing (RNA-seq) on thoracic nota (mesonotum and metanotum) dissected from 0–72 h fifth-instar nymphs of wild-type SW-BPH (WtSW, SW ratio > 90%) and LW-BPH (WtLW, LW ratio > 80%) strains, representing SW-destined and LW-destined BPHs, respectively. The fifth-instar stage was chosen because it is the instar during which SW and LW developmental fates are determined30. We identified 123 putative transcription factors (TFs) including FoxO and Zfh1 (Fig. 1b, Nl.chr09.837 in Supplementary Data 1) that were significantly differentially expressed in WtLW versus WtSW nymphs. Next, we performed RNA interference (RNAi)-mediated gene silencing by microinjection of WtSW nymphs with double-stranded RNA (dsRNA) targeting each TF gene except for FoxO which has been examined previously22,26. Microinjection of dsRNA targeting Zfh1 (dsZfh1, Fig. 1b) reduced the transcriptional level of Zfh1 by ~40% compared to dsRNA targeting green fluorescent protein (dsGfp, Fig. 1c). Both female (Fig. 1d, e) and male (Supplementary Fig. 1b, c) nymphs treated with dsZfh1 exhibited a strong and comparable bias towards LW morphs (dsZfh1LW), relative to control individuals injected with dsGfp (dsGfpSW). In contrast, no effect on wing morph determination was detected following dsRNA-mediated knockdown of any of the other 121 TF genes (Supplementary Data 2). Hypomorphic knockdown phenotypes were detectable in a small fraction of dsZfh1-treated adults with intermediate-size wings (dsZfh1IMW, Fig. 1d, e, Supplementary Fig. 1b, c). Notably, dsZfh1 had no detrimental effect on nymph development and survival (Supplementary Fig. 2). Additionally, dsZfh1LW BPHs exhibited hind tibia lengths, a common metric for body size in BPH, comparable to WtLW BPHs (Fig. 1f, Supplementary Fig. 1d), suggesting that dsZfh1 had no obvious effect on body size. However, the forewing size of dsZfh1LW BPHs was slightly but significantly smaller than that of WtLW BPHs although the vein patterning was within the normal range of variation observed in WtLW BPHs (Fig. 1f, g, Supplementary Fig. 1d, e). In addition to wings, dsZfh1LW BPHs resembled WtLW BPHs with regards to indirect flight muscle (IFM) development. Specially, immunohistochemistry (IHC) staining showed that IFM was abundant only in the thorax of dsZfh1LW and WtLW BPHs, but not in WtSW BPHs (Fig. 2a). Further, examination by transmission electron microscopy (TEM) showed that dsZfh1LW and WtLW but not WtSW BPHs contained well-organized sarcomeres with visible Z and M discs (Fig. 2b). This observation indicates that Zfh1 inhibits IFM differentiation in BPHs, in line with the inhibitory role of Zfh1 in muscle differentiation described in Drosophila and vertebrates31,32,33.
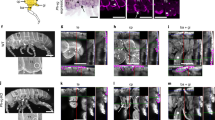
a Immunohistochemistry staining of indirect flight muscles (IFM). Third-instar WtSW nymphs were microinjected with dsZfh1. Thorax and abdomen of emerged female adults were longitudinally cut into sections, followed by staining with rhodamine-labeled phalloidin (red) and DAPI (blue). The IFM is framed by a dotted line, and the trochanteral depressor muscle is indicated by stars. The part of muscles used for transmission electron microscopy is indicated by rectangles. Scale bars, 200 μm. b Examination of IFM by transmission electron microscopy. Thoraxes were dissected from female adults and then cut into sections for IFM examination. Arrows and arrowheads represent Z and M discs of sarcomeres, respectively. Scale bars, 1 μm. The experiments in (a) and (b) were repeated six times independently with similar results. dsZfh1LW, dsZfh1-treated BPHs with long wings. WtLW and WtSW, wild-type long-winged and short-winged BPHs, respectively.
To confirm the dsZfh1 phenotype, we conducted Zfh1 knockdown using a second non-overlapping dsRNA targeting Zfh1 (dsZfh1_2, Fig. 1b). Microinjection with dsZfh1_2 reduced the transcriptional level of Zfh1 by ~50% compared to dsGfp treatment (Fig. 1c). As with dsZfh1, the majority of dsZfh1_2-treated WtSW nymphs developed into LW adults (Fig. 1e, Supplementary Fig. 1c). Additionally, quantitative real-time PCR (qRT-PCR) analysis showed that Zfh1 expression exhibited a distinct spatiotemporal pattern, with low level of Zfh1 detected in the thorax and thoracic nota of fifth-instar WtLW relative to WtSW nymphs (Supplementary Fig. 3), indicating that the expression of Zfh1 is inversely correlated with LW development.
To investigate whether Zfh1 was functionally conserved in the planthopper family Delphacidae, we performed RNAi-mediated knockdown of the Zfh1 homolog in the planthopper Laodelphax striatellus (LsZfh1). Microinjection with dsRNA targeting LsZfh1 (dsLsZfh1) significantly decreased the transcriptional level of LsZfh1 (Supplementary Fig. 4a), and significantly increased the LW ratio (dsLsZfh1LW, Fig. 3a, b) relative to dsGfp treatment, consistent with the phenotype observed in BPHs. Notably, hypomorphic knockdown of LsZfh1 also led to adults with intermediate-size wings (dsLsZfh1IMW). Together, these findings indicate that Zfh1 acts as a molecular switch that regulates alternative wing morphs in planthoppers.
Third-instar L. striatellus nymphs were microinjected with dsLsZfh1 or dsGfp. a Morphology of dsGfpSW, dsLsZfh1LW, and dsLsZfh1IMW females. dsGfpSW, dsGfp-treated L. striatellus planthoppers with short wings. dsLsZfh1LW and dsLsZfh1IMW, dsLsZfh1-treated L. striatellus planthoppers with LW and intermediate-size wings (IMW), respectively. b Number of L. striatellus planthoppers with different wing morphs upon dsRNA treatments. The LW ratio is compared between two groups using Pearson χ2 test (****P = 6.6841E-13, χ2 = 51.635 and df = 1 for females; ****P = 0.000001, and χ2 = 23.413 and df = 1 for males). The experiment was repeated three times independently with similar results. Source data are provided as a Source Data file.
Functional specificity of Zfh1 among BPH homeobox genes
BPH Zfh1 contains a homeodomain in addition to nine Cys2His2 type zinc-finger motifs (Fig. 1b), and as such belongs to the zfh family of homeobox genes across metazoan phyla34. As does Drosophila, BPH possesses two zfh family members, Zfh1 and Zfh2, which differ in the number and position of homeodomains and zinc-finger motifs. Studies in Drosophila indicated that both Zfh1 and Zfh2 are involved in neurogenesis and cell differentiation35,36,37. In addition, Drosophila Zfh1 functions in the positioning of mesoderm derivatives including somatic musculature38, whereas Zfh2 is involved in the proximal-distal patterning of Drosophila wing discs39,40. Phylogenetic analyses show that both Zfh1 and Zfh2 are conserved across animals (Supplementary Fig. 5a): invertebrate Zfh1 homologs cluster with two vertebrates Zfh1 orthologues (Zeb1 and Zeb2)38, whereas Zfh2 and other zfh family members cluster separately (Supplementary Fig. 5a).
To test whether BPH Zfh2 is functionally redundant to Zfh1 with respect to the regulation of wing dimorphism, we conducted Zfh2-specific knockdowns by microinjection of fourth-instar WtSW nymphs with dsRNA targeting Zfh2 (dsZfh2). Notably, Zfh2 knockdown caused ~50% mortality of nymphs before adult eclosion, while surviving adults exhibited curved wings (Supplementary Fig. 5b). However, we found that surviving dsZfh2-treated BPHs produced a high proportion of SW morphs, similar to WtSW BPHs (Supplementary Fig. 5c), indicating that Zfh2 is not involved in BPH wing dimorphism.
To assess whether any additional homeobox genes might be involved in LW development, we conducted RNAi targeting the remaining 87 potential BPH homeobox genes in WtSW nymphs. We found that knockdown of any of these 87 genes did not change wing-morph ratios (Supplementary Data 3). Thus, our results suggest that Zfh1 may be the sole BPH homeobox gene that functions specifically in the regulation of wing dimorphism.
Zfh1 regulates FoxO transcriptional activity
To further detail the role of Zfh1 in the development of wing dimorphism, we collected the mesonotum and metanotum of dsZfh1- and dsGfp-treated fifth-instar nymphs for RNA-seq (Fig. 4a). Interestingly, FoxO was one of the 478 genes downregulated following dsZfh1 injection (Fig. 4b, Supplementary Data 4). This is notable because dsZfh1 phenocopied the effects of FoxO knockdown (dsFoxO) on LW development22. qRT-PCR analysis confirmed that dsZfh1 significantly reduced FoxO mRNA level (Fig. 4c).
a Scheme depicting mesonotum and metanotum (yellow) used for RNA-seq (illustrated by Miss Yi Wan). b Differentially expressed genes in nota of dsZfh1- and dsGfp-treated fifth-instar nymphs. FoxO is indicated by an arrow. c qRT-PCR analysis of FoxO expression in nota of dsZfh1- and dsGfp-treated fifth-instar nymphs. The experiment was repeated five times with similar results (dots). d Scheme of construction of FoxO::HA mutants using CRISPR/Cas9-mediated homology-directed repair. The target site for Cas9 and protospacer-adjacent motif (PAM) are indicated. An ssDNA donor contains a left homology arm, a HA-tag (HA), and a right homology arm. Ex 1–5, exons 1–5 of FoxO. FoxO-KI-F and FoxO-KI-R, primers used for FoxO::HA verification. e Confirmation of FoxO::HA mutants by western blot using an HA antibody. f Confirmation of FoxO::HA mutants by Sanger sequencing with primers of FoxO-KI-F and FoxO-KI-R. g dsZfh1 decreases FoxO::HA protein level. FoxO::HA mutants were microinjected with dsZfh1 or dsGfp, followed by western blot using an HA antibody. β-Actin is used to show equal protein loading in (e) and (g). The experiments in (e) and (g) were repeated three times independently with similar results. h Scheme of immunoprecipitation assay. Red vertical lines and letters, putative Zfh1 consensus binding sites in the Foxo promoter (ProFoxO). TSS, the transcription start site of FoxO. The HEK293T cells are transfected with the ProFoxO fragment mixed with either pCMV-Zfh1-His or pCMV plasmid. i Relative binding of the Foxo promoter to Zfh1 by immunoprecipitation assays. The ProFoxO fragment was immunoprecipitated with antibodies against His tag. The experiments were repeated three times with similar results (dots). Two-tailed unpaired t-test was used for the statistical analysis (c, ****P = 3.48132E−06) and (i, **P = 0.0075). Data are presented as mean ± SEM in (c) and (i). j Venn diagram and heat map of thoracic genes commonly regulated by Zfh1 and FoxO. Thoraxes treated with dsZfh1, dsFoxO, and dsGfp are denoted as Zfh1-T, FoxO-T, and Gfp-T, respectively. Expression levels are indicated by FPKM (fragments per kilobase of transcript per million mapped reads). Source data are provided as a Source Data file.
To further assess whether FoxO is indeed a target gene transcriptionally regulated by Zfh1, we examined FoxO translation following Zfh1 knockdown, using a fusion protein consisting of FoxO and Human influenza hemagglutinin (HA) as a reporter. To do so, we microinjected eggs with a mix of Cas9, single-guide RNA (sgRNA), and single-strand DNA (ssDNA) donors to construct a knock-in BPH strain (FoxO::HA) using clustered regularly interspaced palindromic repeats/CRISPR-associated (CRISPR/Cas9)-mediated homology-directed repair. The resulting FoxO::HA mutant then allowed us to detect FoxO protein levels using commercially available anti-HA antibodies. We found that 13.2% (n = 1023) injected egg successfully molted into nymphs (G0), among which 11 out of 44 examined nymphs (25%, Supplementary Table 1) contained an HA-tag at the C terminus of FoxO. To examine germline transmission, G0 adults were mated with WtSW adults to produce G1 offspring. We found that 26.7% G1 individuals (32 out of 120, Supplementary Table 1) contained FoxO with an HA insertion (Fig. 4d). Crossing G1 siblings then allowed us to obtain a FoxO::HA homozygous mutant. To verify the FoxO::HA mutant, whole bodies of FoxO::HA mutants and WtSW adults were homogenized for western blot assay using an HA antibody. We found that the FoxO-HA fusion protein was specifically detected in FoxO::HA mutants rather than in WtSW BPHs (Fig. 4e). To further confirm the FoxO::HA mutant, genomic DNA of FoxO::HA mutants was extracted and then used for Sanger sequencing with primers (FoxO-KI-F/FoxO-KI-R, Fig. 4d) located outside of homology arms. Sanger sequencing showed that the HA-tag was correctly inserted before the stop codon (TAG) of FoxO (Fig. 4f). Next, we performed Zfh1 knockdown in FoxO::HA mutants, followed by western blot assay using an HA antibody. In line with the qRT-PCR data, dsZfh1 significantly reduced the FoxO::HA protein expression compared to dsGfp (Fig. 4g).
Given that FoxO is transcriptionally regulated by Zfh1, we asked whether Zfh1 protein could bind to the FoxO promoter, similar to the vertebrate Zfh1 homolog which represses gene expression by binding to the consensus sequence (CACCT and CACCTG)41. We identified three high-affinity Zfh1 binding sites in the first 2.5 kb fragment of the FoxO promoter (ProFoxO, Fig. 4h). Using an immunoprecipitation (IP) protein-expressing Zfh1-his in human embryonic kidney 293 cells (HEK293T), we found that the FoxO promoter sequence was significantly enriched by anti-His antibodies when ProFoxO was co-transfected with pCMV-Zfh1-His (Fig. 4i), confirming Zfh1 binding to the FoxO promoter.
To further ascertain whether FoxO is positioned downstream of Zfh1, thoraxes dissected from dsFoxO-, dsZfh1-, and dsGfp-treated fifth-instar nymphs were subjected to RNA-seq. The comparative transcriptomic analysis revealed that 55% of genes (546 out of 997) regulated by dsFoxO were also regulated by dsZfh1 (Fig. 4j, Supplementary Data 5). Among these, 96% (522 out of 546) showed comparable changes in expression (Fig. 4j). Gene ontology (GO) analysis showed important similarities as well as differences among the genes whose expression was significantly altered following dsFoxO and/or dsZfh1: genes up-regulated by both dsFoxO and dsZfh1 were significantly enriched for 9 GO terms associated with cellular processes (Supplementary Fig. 6a), whereas genes downregulated by both dsFoxO and dsZfh1 were significantly enriched for 10 GO terms associated with chitin metabolic processes (Supplementary Fig. 6b). In contrast, the top GO terms specific to genes whose expression was altered following dsFoxO were ‘mitochondrial ribosome’, ‘organellar ribosome’ and ‘mitochondrial translation’, whereas the top-ranked GO terms specific to dsZfh1 regulation were ‘structural constituent of cuticle’, ‘extracellular region’ and ‘muscle system process’ (Supplementary Fig. 6c, d). Taken together, these findings suggest that Zfh1 operates as an upstream regulator for wing dimorphism by affecting FoxO transcriptional activity.
Zfh1 regulates wing morphs in parallel to the IIS pathway
Inactivation of genes encoding positively acting components (InR1 or Akt) of the IIS pathway can lead to SW morphs by dephosphorylation of FoxO; however, inactivation of InR2, a negative regulator of InR1, enhances the phosphorylation of FoxO, leading to LW morphs22. Because FoxO is an effector downstream of the IIS pathway and because FoxO is transcriptionally regulated by Zfh1, we asked whether Zfh1 regulates the expression of FoxO by relaying IIS activity. If this is the case, the inactivation of Zfh1 would abolish any effects of the IIS pathway on wing dimorphism in the same manner as done by FoxO22. Thus, we performed a double-gene knockdown using dsFoxO and dsZfh1 in combination with a dsRNA targeting InR1 (dsInR1) or Akt (dsAkt). Although dsZfh1 combined with dsGfp led to LW morphs, WtSW nymphs developed into SW adults when dsZfh1 was combined with either dsInR1 or dsAkt (Fig. 5a, Supplementary Fig. 7). As a control, WtSW nymphs developed into LW adults when dsFoxO was combined with dsInR1 or dsAkt (Fig. 5a, Supplementary Fig. 7). These data indicate that Zfh1 is not a TF downstream of the IIS pathway.
a Number of females with different wing morphs upon double-gene knockdown. The LW ratio is compared between two groups using Pearson χ2 test (dsInR1 + dsFoxO vs dsInR1 + dsGfp: **** P = 5.0866E-47, χ2 = 207.131, df = 1; dsAkt + dsFoxO vs dsAkt + dsGfp: ****P = 4.763E-45, χ2 = 198.359, df = 1; n.s. non-significant). b The FoxO::HA protein level in FoxO::HA mutants following dsRNA treatments. Treatment with dsZfh1 only lowers the level of FoxO proteins. c Construction of Zfh1::HA mutants using CRISPR/Cas9-mediated homology-directed repair. The target site for Cas9, protospacer-adjacent motif (PAM), and exons 1–5 are indicated. An ssDNA donor contains a left homology arm, a HA-tag (HA), and a right homology arm. The primers of Zfh1-KI-F and Zfh-KI-R are designed for Sanger sequencing. d Confirmation of Zfh1::HA mutants by western blot. Zfh1::HA fusion protein was detected by an HA antibody. e Confirmation of Zfh1::HA mutants by Sanger sequencing. PCR products amplified from genomic DNA with Zfh1-KI-F and Zfh-KI-R were exposed to Sanger sequencing. The HA sequence is in blue. f The Zfh1::HA protein level in the context of perturbation of the IIS pathway in Zfh1::HA mutants. β-actin is used to show equal protein loading in (b), (d), and (f), and the experiments were repeated three times independently with similar results. g Relative expression of components of the IIS pathway in dsZfh1-treated BPHs. The experiments were repeated five times with similar results (dots or circles). Data are presented as mean ± SEM. Two corresponding columns are compared using two-tailed unpaired t-test (n.s. non-significant). h Morphology of an InR2-null mutant (InR2E4)43. i dsZfh1 increases the forewing size of InR2E4 mutants. InR2E4 nymphs were microinjected with dsGfp (InR2E4::dsGfp) or dsZfh1 (InR2E4::dsGfp), and then emerged adults were collected for wing size measurement. The whiskers of box plot represent the quantile percentile, from bottom to top are minima, 25%, median, 75%, and maxima, respectively. Two corresponding boxes are compared using two-tailed unpaired t-test (*P = 0.0222 for male and ****P = 5.70008E−05 for female). Source data are provided as a Source Data file.
Next, we sought to determine whether Zfh1 is positioned upstream of the IIS pathway. If correct, inactivation of Akt would abolish the regulatory role of Zfh1 on FoxO protein expression. To test these predictions, we microinjected FoxO::HA nymphs with dsZfh1, dsAkt, a dsRNA targeting InR2 (dsInR2), a dsRNA mixture targeting Zfh1 and Akt (dsZfh1;dsAkt), or dsGfp, followed by the dissection of the nota of fifth-instar nymphs for western blot assay. Compared to dsGfp, double-gene knockdown using dsZfh1;dsAkt significantly reduced FoxO::HA fusion protein levels, as did dsZfh1 alone (Fig. 5b), indicating that Zfh1 is also not a factor upstream of the IIS pathway. Notably, both dsInR2 and dsAkt resulted in comparable levels of FoxO::HA protein relative to dsGfp (Fig. 5b). This supports previous reports that the IIS pathway may regulate FoxO protein function via post-translational modification42.
To exclude any possible crosstalk between the IIS pathway and Zfh1, we generated a Zfh1::HA knock-in mutant using CRISPR/Cas9-mediated homology-directed repair, which would allow us to detect Zfh1 protein with anti-HA antibodies since Zfh1 was fused an HA-tag with at the C terminus (Fig. 5c). The CRISP/Cas9-medicated mutagenesis rate and germline transmission for Zfh1 reached 11.6% (n = 64) and 18.9% (n = 90), respectively. The Zfh1::HA homozygous mutant was verified by western blot using anti-HA antibodies (Fig. 5d) and by Sanger sequencing (Fig. 5e) with primers (Zfh1-KI-F/Zfh1-KI-R, Fig. 5c) located outside of the homology arms. The Zfh1::HA mutant allowed us to detect Zfh1 protein in the context of perturbation of the IIS pathway using anti-HA antibodies. We found that knockdown of components of the IIS pathway (InR1, InR2, and Akt) did not affect Zfh1 protein level (Fig. 5f). Meanwhile, knockdown of Zfh1 did not affect the transcriptional level of InR1, InR2, and Akt (Fig. 5g). Thus, these findings indicate that Zfh1 regulates wing dimorphism independent of the IIS pathway in BPHs.
Zfh1 functions synergistically with the IIS pathway
Given that Zfh1 regulates LW development in parallel to the IIS pathway, we asked whether Zfh1 and the IIS pathway may interact synergistically in the regulation of LW development. For this purpose, we performed Zfh1 knockdown in the context of an InR2-null mutant (InR2E4)43, which by itself results in 100% LW morphs (Fig. 5h) due to accumulated phosphorylated FoxO in the cytoplasm via activating the IIS pathway. We found that dsZfh1-treated InR2E4 mutants (InR2E4::dsZfh1) developed forewings that were 7% larger than the InR2E4 mutants treated with dsGfp (InR2E4::dsGfp, Fig. 5i), indicating a moderate synergistic effect of Zfh1 and the IIS pathway on LW development.
Zfh1 is functionally distinct from Ultrabithorax
The presence of a homeodomain in BPH Zfh1 combined with the Zfh1 knockdown phenotype are reminiscent of the ‘four-winged Drosophila’ phenotype derived from Ultrabithorax (Ubx) mutants44. This phenotype results from the homeotic transformation of the third thoracic segment (T3) to the identity of the second thoracic segment (T2), and the concomitant transformation of halters into hindwings in Drosophila44, or membranous hindwings into elytra in the flour beetle Tribolium castaneum45.
To exclude the possibility that dsZfh1 produced LW morphs via homeotic transformation, we compared and contrasted the morphological characteristics of wings and thoracic segments between dsZfh1 and Ubx-RNAi (dsUbx) adults, focusing on three aspects: (i) dsUbx adults that were derived from RNAi in SW-destined nymphs developed only small wing-like appendages on T3 and SW forewings at T2 (Fig. 6a), in strikingly contrast to the fully developed forewings and hindwings of dsZfh1 BPHs (Fig. 1d); (ii) dsZfh1 adults that were derived from RNAi in LW-destined nymphs developed membranous and fully transparent hindwings similar to the control BPHs (dsGfp, Fig. 6b), whereas dsUbx adults that were derived from RNAi in LW-destined nymphs developed heavily-pigmented hindwings (Fig. 6b), resembling forewings; and (iii) dsZfh1 adults developed T2 and T3 nota identical to those of dsGfp-treated BPHs (Fig. 6c), whereas Ubx knockdown led to an expansion of the T3 notum (Fig. 6c), thereby causing it to resemble the characteristics of the T2 notum. In addition, Zfh1 and Ubx differed strikingly in terms of tissue-specific expression. qRT-PCR assay showed that hindwing buds had a slightly but significantly higher level of Zfh1 relative to forewing buds (Fig. 6d). In contrast, Ubx expression was strongly biased toward hindwing buds relative to forewing duds (Fig. 6d). Together, these observations indicate that the SW-to-LW transition derived from Zfh1 knockdown is most likely not due to homeotic transformation.
a Morphology of short-winged (SW) BPHs microinjected with dsUbx. Fourth-instar SW-destined (WtSW) nymphs were used for microinjection with dsUbx. Small wing-like appendages (hindwings, arrow) protrude from the third thoracic (T3) segment of dsUbx-treated BPHs. Scale bars, 750 μm. b Morphology of long-winged (LW) BPHs microinjected with dsUbx, dsZfh1, and dsGfp. LW-destined (WtLW) nymphs were used for microinjection with dsUbx, dsZfh1, and dsGfp. dsUbx-treated BPHs contain heavily-pigmented hindwings compared to transparently membranous hindwings of dsZfh1- and dsGfp-treated BPHs. Arrowheads, hindwings. Scale bars, 750 μm. c Scanning electron microscope observation of the second thoracic (T2) and T3 nota. Fourth-instar WtLW nymphs were microinjected with dsUbx, dsZfh1 or dsGfp, and emerged adults were directly used for scanning electron microscope. Scale bars, 200 μm. Twenty samples in a–c were repeated independently with similar results. d Examination of the expression of Ubx and Zfh1 in forewing buds (FWB) and hindwing buds (HWB) of fifth-instar nymphs by qRT-PCR. The experiments were repeated three times with similar results (dots). Data are presented as mean ± SEM. Two-tailed unpaired t-test was used for the statistical analysis (**P = 0.004267 for Zfh1 and ****P = 8.11903E-07 for Ubx). Source data are provided as a Source Data file.
A molecular model of wing dimorphism in planthoppers
Our data revise and significantly extend our understanding of how distinct wing morphs may result from modifications of the same gene regulatory networks (Fig. 7). Under normal conditions, BPHs maintain a basic level of Zfh1, which in turn maintains FoxO transcription at low levels. As a result, low activity of the Zfh1-FoxO signaling cascade prevents FoxO from inhibiting wing development, thus leading to LW morphs. However, in response to certain environmental inputs, BPHs raise Zfh1 transcription, thereby raising FoxO levels, which in turn suppresses LW development and therefore leads to SW morphs. In addition to this Zfh1-FoxO signaling cascade, wing dimorphism is regulated by a second, and independent, signaling axis at the post-translational level: activation of the IIS pathway phosphorylates FoxO and thus exports FoxO from the nucleus to the cytoplasm, leading to LW morphs. In contrast, inactivation of the IIS pathway dephosphorylates FoxO and thus accumulates FoxO in the nucleus, leading to SW morphs.
The balance of alternative wing morphs is regulated (i) transcriptionally by the Zfh1-FoxO cascade or (ii) post-translationally by the IIS-FoxO cascade. Components with increased expression levels are shown in bold and red. Components with decreased expression levels are shown in gray. FoxO-P, phosphorylated FoxO. The BPH images were illustrated by Miss Xin-Qiu Wang.
Discussion
We identified the zinc-finger homeodomain transcription factor Zfh1 as a wing-morph molecular switch that determines long or short wings by regulating the transcriptional level of a second transcription factor FoxO in BPHs. Previous studies indicated that Zfh1 is an evolutionarily conserved gene critical for counteracting the myogenic differentiation program31,34,46. In Drosophila, embryos with a Zfh1 loss-of-function mutation show alterations in somatic muscles38. In the same way, a vertebrate homolog of Zfh1 (ZEB) acts as a negative regulator of muscle differentiation31. Here, we found that knockdown of BPH Zfh1 promoted IFM development, consistent with myogenesis suppression by Zfh1 in Drosophila and vertebrates. However, in addition to these common phenotypes reported previously, this study demonstrates for the first time that knockdown of Zfh1 directs SW-destined nymphs to develop into LW in wing dimorphic BPHs, and likely planthoppers broadly. Given that the development of LW in BPHs mainly relies on cell proliferation23,43, this phenotype indicates that Zfh1 might act as an inhibitor of wing cell proliferation. However, whether suppression of wing cell via Zfh1 is species-dependent or conserved among insects remains unknown. Additionally, a recent study showed that the maintenance of Drosophila muscle progenitors involves a switch between Zfh1-long and Zfh1-short RNA isoforms33. The Zfh1-long isoform is subjected to miR-8 microRNA-mediated downregulation that enables muscle progenitors to differentiate in early pupal stages, whereas the Zfh1-short isoform cannot be targeted by miR-8, thus allowing these cells to persist as muscle progenitors in the adult stage33. In this study, only one Zfh1 isoform was identified after examining all BPH transcriptomes prepared by our lab. Thus, whether multiple BPH Zfh1 isoforms, if they do indeed exist, possess distinct roles in the regulation of wing development and muscle homeostasis needs to be clarified in further studies.
Our previous studies showed that FoxO plays a pivotal role in controlling the development of alternative wing morphs in BPHs via relaying the activity of the IIS pathway22. Activation of the IIS pathway phosphorylates and deactivates FoxO, leading to LW morphs, while IIS inactivation and FoxO dephosphorylation promote SW morphs. Here, we found that knockdown of Zfh1 stimulates LW development by decreasing FoxO transcriptional activity. These findings highlight that the Zfh1 and IIS cascades may converge on FoxO to regulate wing dimorphism in BPHs. This assumption is supported by a synergistic effect of both pathways on LW development. However, exactly how BPHs balance the relative activities of the Zfh1-FoxO and IIS-FoxO cascades to regulate wing developmental plasticity remains to be determined. More generally, our results suggest that this regulatory system may provide a multi-tiered regulatory machinery able to facilitate evolutionary changes in threshold responses separating wing morphs and increase the ways in which population and species may diverge genetically in the precise regulation of wing polyphenism.
Methods
Insects
Laboratory populations of N. lugens were established from colonies collected in Hangzhou, China, in 2008. The WtSW colony was purified by inbreeding for more than 13 generations47. The WtLW strain was provided by Dr. Hong-Xia Hua (Huazhong Agricultural University, China). The planthopper L. striatellus was collected in Hangzhou, China, 2018. All insects were reared at 26 ± 0.5 °C under a photoperiod of 16:8 h (L:D) at a relative humidity of 50 ± 5% on rice seedlings (rice variety: Xiushui 11).
Sample preparation for RNA-seq
To screen for potential wing-morph regulators, mesonotum and metanotum were dissected from 0–72 h fifth-instar WtSW (n = 20) and WtLW nymphs (n = 20), denoted as 5th-SW and 5th-LW respectively (BioProject ID: PRJNA805393). To identify downstream effectors targeted by Zfh1, third-instar WtSW nymphs were microinjected with 50 ng dsZfh1 or dsGfp. Subsequently, mesonotum and metanotum were dissected from 36–48 h fifth-instar nymphs with three replicates (n = 30 for each replicate), denoted as Zfh1-nota and Gfp-nota (BioProject ID: PRJNA805395). To investigate common genes regulated by Zfh1 and FoxO, third-instar WtSW nymphs were microinjected with 50 ng dsZfh1, dsFoxO, or dsGfp. Thoraxes were dissected from 48–54 h fifth-instar nymphs with three replicates (n = 30 for each replicate), denoted as Zfh1-T, FoxO-T and Gfp-T (BioProject ID: PRJNA805400). Samples were homogenized for total RNA isolation using RNAiso Plus (Cat#9109, Takara) according to the manufacturer’s protocol.
cDNA library preparation and Illumina sequencing
A total of 1 µg RNA per sample was used to construct a sequencing library using a NEBNext Ultra RNA library prep kit for Illumina (Cat#E7770, NEB) according to the manufacturer’s recommendations, and index codes were added to each sequence. Library fragments of 250–300 bp in length were preferentially purified using an AMPure XP system (Beckman Coulter). Clustering of the index-coded samples was performed on a cBot Cluster generation system using a TruSeq PE cluster kit v3-cBot-HS (Cat#PE-401-3001, Illumina) according to the manufacturer’s instructions. The cDNA libraries were sequenced on an Illumina Novaseq 6000 platform and 150 bp paired-end reads were generated.
Read mapping and differentially expressed genes (DEGs)
After Illumina sequencing, clean reads were generated by removing adapters, poly-N, and low-quality reads from the raw data using fastp algorithm (v0.12.4)48. Clean reads were mapped against the N. lugenes genome v4.047 using hisat2 (v2.1.0)49, and transcript abundance was quantified using StringTie (v1.3.5)50. Fragments Per Kilobase of transcript per Million mapped reads (FPKM) was used to quantify the expression level of each transcript. The read count information of each transcript was extracted directly from the files generated by StringTie using a python script of prepDE.py (http://ccb.jhu.edu/software/stringtie/dl/prepDE.py). Differential expression analysis of 5th-SW and 5th-LW samples was conducted using the edgeR package (v3.38.4)51, and genes with a false discovery rate (FDR) < 0.01 were considered to be significant. Differential expression analysis of Zfh1-nota versus Gfp-nota, Zfh1-T versus Gfp-T, and FoxO-T versus Gfp-T was conducted using the DESeq2 package (v1.36.0)52 and gene expression change was considered to be significant under the FDR < 0.05. To generate heat map, FPKM of each replicate of Zfh1-T, FoxO-T and Gfp-T were z-score transformed and clustered using the online OmicShare tool (https://www.omicshare.com/tools/Home/Soft/heatmap). GO enrichment analysis of DEGs was performed using the online OmicShare tool (https://www.omicshare.com/tools/home/report/goenrich.html), and the enriched GO terms with FDR ≤ 0.05 were considered as significant.
RNAi and RNAi efficiency
DsRNA synthesis and injection were performed as previously described22. Briefly, primers were synthesized with T7 RNA polymerase promoter at both ends (Supplementary Data 6), and then dsRNAs were synthesized using a T7 high yield RNA transcription kit (Cat#TR101-02, Vazyme) according to the manufacturer’s instructions. Microinjection was performed using a FemtoJet microinjection system (Eppendorf). Each third-instar or fourth-instar nymph was microinjected with approximately 50 ng or 100 ng dsRNAs, respectively. Two days after microinjection, insects (n = 5 for each of three replicates) were collected for RNAi efficiency examination by qRT-PCR. The ribosomal protein S11 gene (rps11)53 and ribosomal protein L5 gene (rpl5)54 were used as the internal reference gene in BPH and L. striatellus planthopper, respectively.
qRT-PCR assay
Total RNAs were isolated from BPHs using RNAiso Plus (Cat#9109, Takara) according to the manufacturer’s protocol. The first-strand cDNA was synthesized from total RNAs (900 ng) using HiScript QRT super mix (Cat#R123-01, Vazyme). The qRT-PCR was conducted on a CFX96 real-time PCR detection system (Bio-Rad) with the following conditions: denaturation for 3 min at 95 °C, followed by 40 cycles at 95 °C for 10 s, and then 60 °C for 30 s. The rps11 and rpl5 were used as the internal reference gene in BPH and L. striatellus planthopper, respectively. The 2−∆∆Ct method (Ct represents the cycle threshold) was used to measure relative expression levels. Three biological replicates were used for statistical comparison between samples.
Nymphal duration and survival rate
Third-instar WtSW nymphs were microinjected with 50 ng dsZfh1 or dsGfp. The developmental times of fourth- and fifth-instar stages and survival rate were monitored every 12 h. To determining the nymphal duration, 13 females and 19 males were used for dsZfh1, and 17 females and 11 males were used for dsGfp. To conduct the survival assay, 50 nymphs were used for dsZfh1 and dsGfp.
Spatiotemporal expression of Zfh1
To investigate the temporal expression of Zfh1 in WtLW and WtSW BPHs, total RNAs were isolated from the thorax of first-instar (n = 100), second-instar (n = 50), third-instar (n = 50), fourth-instar (n = 30), fifth-instar nymphs (n = 15), and female adults (n = 15). To investigate the spatial expression of Zfh1, fifth-instar WtLW and WtSW nypmphs (n = 50) were collected for tissue dissection, and then total RNA was isolated from head, fat body, the abdominal cuticle, six legs, the whole digestive tract (gut), and nota (mesonotum and metanotum), respectively. To investigate the expression pattern of Zfh1 across the fifth-instar stage, mesonotum and metanotum were dissected from 24 h-, 48 h-, and 72 h-fifth-instar WtLW and WtSW nymphs (n = 50), and then used for total RNA isolation. Three independent biological replicates were used for RNA isolation, and the first-strand cDNA was synthesized for Zfh1 quantification. The relative expression level of Zfh1 was normalized to that of the rps11 gene.
IHC staining
IHC staining was performed as previously described55. Briefly, third-instar nymphs were microinjected with 50 ng dsZfh1. Samples were fixed using 4% paraformaldehyde in PBS overnight at 4 °C, following by blocking with Tissue-Tek O.C.T. Compound (Cat#4583, Sakura Finetek) at −80 °C. Samples were longitudinally cut into ~30 μm sections using a Lecia CM1900 cryotome (Leica Microsystems) at −20 °C, and then transferred to Superfrost+ slides (Cat#12-550-15, Thermo Fisher Scientific). The cytoskeleton and nucleus were stained using 100 nM rhodamine-labeled phalloidin (Cat# 40734ES75, Yeasen) and 100 nM DAPI (Cat#D9542, Sigma Aldrich), respectively. Fluorescence images were acquired using a Zeiss LSM 800 confocal microscopy (Carl Zeiss MicroImaging).
TEM assay
Third-instar nymphs were microinjected with dsZfh1. Thoraxes were dissected from 24 h dsZfh1LW females for transmission electron microscopy as previously described55. Briefly, samples were fixed in 2.5% glutaraldehyde overnight at 4 °C, followed by post-fixation in 1% osmium tetroxide for 1.5 h. The samples were then dehydrated in a standard ethanol/acetone series, infiltrated and embedded in Spurr medium, and then superthin sections (~70 nm) were cut. The sections were stained with 5% uranyl acetate followed by Reynolds’ lead citrate solution and observed under a JEM-1230 transmission electron microscope (JEOL) at the Analysis Center of Agrobiology and Environmental Sciences of Zhejiang University.
Phylogenetic analysis of Zfh1 and Zfh2
BlastP (v2.2.31) was used to search Zfh1 and Zfh2 homologs using the N. lugens Zfh1 and Zfh2 as query sequences. Multiple amino acid sequence alignment was performed using the MUSCLE algorithm with MEGA-X (v10.1.8) software. The phylogenetic tree was constructed by MEGA-X using maximum-likelihood methods under the JTT + G model with 1000 bootstrap replicates. Phylogenetic tree was annotated and edited using the iTOL tools (https://itol.embl.de/) and Adobe Photoshop CC (v19.1.9) program.
In vitro synthesis of sgRNA and Cas9 mRNA
sgRNA was designed as previously reported56. Briefly, sgRNA was designed by manually searching genomic sequence around the region of the FoxO and Zfh1 stop codon for the sequences corresponding to 5′-N17-20NGG-3′, where NGG is the protospacer-adjacent motif (PAM) of SpCas9 and N is any nucleotide. For in vitro transcription of sgRNA, a DNA fragment was amplified by PCR from pMD19-T sgRNA scaffold vector with a forward primer containing a T7 promoter and a reverse primer containing a partial sgRNA sequence. The PCR products were used as a template for in vitro transcription using a T7 high yield RNA transcription kit (Cat#TR101-02, Vazyme) at 37 °C overnight according to the manufacturer’s instruction.
In vitro synthesis of Cas9 mRNA was prepared as previously reported57. Cas9 mRNA was transcribed from plasmid pSP6-2sNLS-SpCas9 vector using the mMESSAGE mMACHINE SP6 transcription kit (Cat#AM1340, Thermo Scientific) and Poly(A) tailing kit (Cat#AM1350, Thermo Scientific).
Construction of ssDNA donors
The ssDNA was produced via lambda exonuclease digestion from PCR amplicons according to a previous description58 with minor modifications. Briefly, a dsDNA fragment containing the whole ssDNA donor cassette for HA-tag knock-in was de novo synthesized and then subcloned into a cloning vector. For FoxO::HA knock-in mutants, the ssDNA donor cassette consisted of a 125 nt left homology arm (LHA) upstream of the FoxO stop codon, a 136 nt right homology arm (RHA) downstream of the FoxO stop codon, and a HA-tag (5′-TACCCATACGATGTTCCAGATTACGCT-3′) in the middle region. For Zfh1::HA knock-in mutant, the ssDNA donor cassette consisted of a 142 nt LHA upstream of the Zfh1 stop codon, a 133 nt RHA downstream of the Zfh1 stop codon, and a HA-tag in the middle region. Then, the dsDNA fragments were used as a template for PCR amplification with a forward primer and a 5′ phosphorylated reverse primer (Supplementary Data 6). PCR products were treated with lambda exonuclease (Cat#EN6501, Thermo Scientific) to remove the 5′-phosphorylated DNA. The ssDNAs were purified by phenol-chloroform extraction, followed by isopropanol precipitation, and then dissolved nuclease-free water.
Embryo microinjection and CRISPR/Cas9-mediated homology-directed repair
Microinjection of BPH embryos was performed as previously described57 with minor modifications. Briefly, pre-blastoderm eggs were collected within 1 h after oviposition, and microinjected with a solution mixture (~0.5 nl) containing 500 ng/μl Cas9 mRNA, 300 ng/μl sgRNA and 300 ng/μl ssDNA using a FemtoJet microinjection system (Eppendorf). To minimize mechanical damage, the posterior end of the eggs was pierced with glass needles. After microinjection, eggs were transferred to a sterile petri dish covered with filter papers that were moistened with an antibiotic solution containing tebuconazole (20 ng/ml) and kanamycin (50 ng/ml). The eggs were incubated at 30 °C and allowed to hatch into nymphs (G0).
Crossing and genotyping
DNA typing of FoxO:: HA and Zfh1::HA heterozygous or homozygous mutants was determined as described previously57. Newly enclosed G0 adults were mated with WtSW adults to produce G1 progeny. After mating, one individual G0 adult was homogenized in a 1.5-ml sterile Eppendorf tube containing 100 μl of DNA extraction buffer (10 mM Tris-HCl pH 8.2, 1 mM EDTA, 25 mM NaCl, and 0.2 mg/ml proteinase K) as reported previously59. The body lysate was incubated at 37 °C for 30 min, followed by incubation at 95 °C for 2 min to inactivate proteinase K. Then, the precipitation was removed by centrifuging at 15,000 × g for 2 min at room temperature, and the supernatant was kept for PCR amplification.
To determine the genotyping of G1 BPHs, wings were dissected from a single adult and then incubated in 100 μl wing gDNA extraction buffer (10 mM Tris-HCl pH 8.0, 10 mM EDTA, 100 mM NaCl, 39 mM DTT, 2% SDS, and 0.02 mg/ml proteinase K) overnight at 37°C. gDNA was precipitated in 100 μl isopropanol, followed by washing with 500 μl ethanol (80%). The gDNA precipitation was dissolved in ddH2O, and then used for PCR amplification.
To confirm the correct insertion of the HA sequence in FoxO::HA mutants, PCR was conducted with primer pairs of FoxO-KI-F/HA-R or FoxO-KI-F/FoxO-KI-R (Supplementary Data 6). To confirm the correct insertion of the HA sequence in Zfh1::HA mutants, PCR was conducted with primer pairs of Zfh1-KI-F/HA-R or Zfh1-KI-F/Zfh1-KI-R (Supplementary Data 6). The FoxO-KI-F/FoxO-KI-R and Zfh1-KI-F/Zfh1-KI-R primer pairs were located outside of homology arms and the HA-R primer was located inside the HA encoding sequence. The PCR products were either directly used for Sanger sequencing or subcloned into cloning vectors, and then single clones were picked for Sanger sequencing. Those G1 individuals with the correct HA-tag insertion were collected and sibling-crossed to obtain homozygous FoxO::HA and Zfh1::HA strains.
Western blot analysis
To detect FoxO::HA or Zfh1:HA fusion protein, whole body of fourth-instar nymphs (n = 40) or thorax (mesonotum and metanotum) dissected from 48 h fifth-instar nymphs (n = 35) was pooled for homogenization. Samples were homogenized in RIPA lysis and extraction buffer (Cat#89900, Thermo Scientific), and then denatured in SDS-PAGE loading buffer. Equal amounts of protein were loaded for each lane on SDS-PAGE gel and then transferred to a polyvinylidene difluoride membrane (Cat#IPVH00010, Millipore). The membranes were blocked in TBST (TBS with 0.1% Tween-20) containing 5% non-fat dried milk powder overnight at 4 °C. After washing, the membranes were incubated with anti-HA mAb (Cat# M180-3, MBL, 1: 2000) for 1.5 h at 35 °C, followed by incubation with horseradish peroxidase (HRP) conjugated goat anti-mouse antibody (Cat#SA00001-1, Proteintech, 1: 5000) for 1 h at 35 °C. The antibody against β-actin (Cat#M1210-2, Huabio, 1:5000) was used as the loading control. Secondary antibody binding was detected using the chemiluminescence HRP substrate detection method (Cat#34577, Thermo Scientific) and imaged with the Molecular Imager ChemiDoc XRS system (Bio-Rad).
Determination of the full-length cDNA of Zfh1
Total RNA was isolated from the whole body of fifth-instar WtSW BPHs (n = 10) using RNAiso plus (Cat#9109, Takara). First-strand cDNA was synthesized using a HiScript-TS 5′/3′ RACE kit (Cat#RA101-01, Vazyme) according to the manufacturer’s instruction. To obtain the 5′ end sequence of Zfh1, PCR was conducted with primers of Universal-primer and Zfh1-5RACE-GSP1, followed by a nested PCR with primers of nested-primer and Zfh1-5RACE-GSP2 (Supplementary Data 6). To obtain the 3′ end sequence of Zfh1, PCR was conducted with primer of Universal-primer and Zfh1-3RACE-GSP1, followed by a nested PCR with primers of Nested-primer and Zfh1-3RACE-GSP2 (Supplementary Data 6). All amplicons were subcloned into cloning vectors and confirmed by Sanger sequencing (Tsingke Biological Technology). The cDNA sequence of Zfh1 was deposited in GenBank with accession number OM283826.
Double-gene RNAi
Third-instar WtSW nymphs were microinjected with 50 ng dsZfh1 or dsFoxO, followed by microinjection with 50 ng dsInR1, dsAkt, or dsGfp at the fourth-instar stage. In parallel, fourth-instar WtSW nymphs were microinjected with dsRNA mixtures of dsAkt and dsGfp (50 ng each) or dsInR1 and dsGfp (50 ng each). All dsRNA-treated nymphs were raised to adults, and BPHs with different wing morphs (LW, SW, and IMW) were collected for statistical analysis.
IP assay
To express Zfh1 in HEK293T cell, the recombinant fragment contained Zfh1 ORF and 6*His tag were cloned downstream of a CMV promoter, designed pCMV-Zfh1-His. A genomic DNA fragment (ProFoxO) located 4,606-bp upstream of the start codon (ATG) of Zfh1 was amplified by PCR with primers of ProFoxO-F and ProFoxO-R (Supplementary Data 6), which contains a 607-bp 5ʹUTR of Zfh1 and putative FoxO promotor region. IP assays were performed using IP assay kit (Cat#P2078, Beyotime) as per the manufacturer’s protocol. Briefly, HEK293T cells were transfected with ProFoxO in combination with pCMV-Zfh1-his or pCMV vector alone. After 36 h, the transfected cells were cross linked using 1% formaldehyde before cell lysis. Sonicated lysates were incubated overnight at 4°C with 6*His tag monoclonal antibody (Cat#66005-1-Ig, 1: 650 (4 μg), Proteintech). The ProFoxO fragments were quantified by qRT-PCR with ChamQ SYBR color qPCR master mix (Cat# Q411-02, Vazyme) using primers of qChIP-F and qChIP-R (Supplementary Data 6).
Relative expression of InR1, InR2, Akt, and FoxO in the context of Zfh1 knockdown
Third-instar nymphs were microinjected with 50 ng dsZfh1 or dsGfp. Mesonotum and metanotum were dissected from 36–48 h fifth-instar nymphs (n = 25) and then used for total RNA isolation. The first-strand cDNA was synthesized with random primers and then used as templates for qRT-PCR. The relative expression level of InR1, InR2, Akt, and FoxO was normalized to that of the rps11 gene. Five independent biological replicates with three technical replicates were conducted for each experiment.
Relative expression of Zfh1 and Ubx in wing buds
Wing buds on the second and third thoracic segments were dissected from 48 h fifth-instar WtSW nymphs (n = 150), and then used for RNA isolation. The first-strand cDNA was synthesized with random primers and then used as templates for qRT-PCR with primers specific to Zfh1 and Ubx (Supplementary Data 6). The relative expression level of Zfh1 was normalized to that of the rps11 gene. Three independent biological replicates with three technical replicates were conducted for each experiment.
Scanning electron microscope (SEM)
Fourth-instar WtLW nymphs were microinjected with dsUbx, dsZfh1, or dsGfp, and allowed to molting into adults. After removing wings, female adults were placed on a stub and dried in an ion sputter (Hitachi) under vacuum. After gold sputtering, the samples were observed under SEM (TM-1000, Hitachi).
Image acquisition
Images of insects were taken using a DVM6 digital microscope (Leica Microsystems) with LAS X software. Images of wings and tibia were captured with a DFC320 digital camera attached to a Leica S8AP0 stereomicroscope using the LAS (v. 3.8) digital imaging system. Digital images of forewings (n = 20) and hind tibias (n = 20) were collected for measurement of forewing size and hind tibia length using ImageJ (v. 1.47). Plotting and statistical analysis were performed with GraphPad Prism (v8.0.1).
Reporting summary
Further information on research design is available in the Nature Research Reporting Summary linked to this article.
Data availability
All data are available in the manuscript or the supplementary materials. The DNA sequencing data generated in this study have been deposited in the GenBank database under the following accession codes: OM283826 (the Zfh1 gene), XP_039284941.1 (the Zfh2 gene), OM676634 (the LsZfh1 gene). The RNA sequencing data generated in this study have been deposited under the following accession coded: PRJNA805393, PRJNA805395, and PRJNA805400. Source data are provided with this paper.
References
Grimaldi, D. & Engel, M. S. Evolution of the Insects, Ch.6 (Cambridge Univ. Press, New York, 2005).
Wootton, R. J. & Kukalová-peck, J. Flight adaptation in Palaeozoic Palaeoptera (Insecta). Biol. Rev. Camb. Philos. Soc. 75, 129–167 (2000).
Roff, D. A. The evolution of flightlessness: is history important? Evol. Ecol. 8, 639–657 (1994).
Wagner, D. L. & Liebherr, J. K. Flightlessness in insects. Trends Ecol. Evol. 7, 216–220 (1992).
Whiting, M. F., Bradler, S. & Maxwell, T. Loss and recovery of wings in stick insects. Nature 421, 264–267 (2003).
Harrison, R. G. Dispersal polymorphisms in insects. Annu. Rev. Ecol. Syst. 11, 95–118 (1980).
Zera, A. J. & Denno, R. F. Physiology and ecology of dispersal polymorphism in insects. Annu. Rev. Entomol. 42, 207–230 (1997).
Abouheif, E. & Wray, A. Evolution of the gene network underlying wing polyphenism in ants. Science 297, 249–252 (2002).
Roff, D. A. & Fairbairn, D. J. Wing dimorphisms and the evolution of migratory polymorphisms among the insecta. Am. Zool. 31, 243–251 (1991).
Simpson, S. J., Sword, G. A. & Lo, N. Polyphenism in insects. Curr. Biol. 21, R738–R749 (2011).
Nijhout, H. F. & Wheeler, D. E. Juvenile hormone and the physiological basis of insect polymorphisms. Q. Rev. Biol. 57, 109–133 (1982).
Zhang, C. X., Brisson, J. A. & Xu, H. J. Molecular mechanisms of wing polymorphism in insects. Annu. Rev. Entomol. 64, 297–314 (2019).
Vellichirammal, N. N., Gupta, P., Hall, T. A. & Brisson, J. A. Ecdysone signaling underlies the pea aphid transgenerational wing polyphenism. Proc. Natl Acad. Sci. USA 114, 1419–1423 (2017).
Braendle, C., Caillaud, M. C. & Stern, D. L. Genetic mapping of aphicarus – a sex-linked locus controlling a wing polymorphism in the pea aphid (Acyrthosiphon pisum). Heredity 94, 435–442 (2005).
Caillaud, M. C., Boutin, M., Braendle, C. & Simon, J. C. A sex-lined locus controls wing polymorphism in males of the pea aphid, Acyrthosiphon pisum (Harris). Heredity 89, 346–352 (2002).
Li, B. et al. A large genomic insertion containing a duplicated follistatin gene is linked to the pea aphid male wing dimorphism. eLife 9, e50608 (2020).
Shang, F. et al. The miR-9b microRNA mediates dimorphism and development of wing in aphids. Proc. Nat. Acad. Sci. USA 117, 8404–8409 (2020).
Xu, H. J. & Zhang, C. X. Insulin receptors and wing dimorphism in rice planthoppers. Philos. Trans. R. Soc. B 372, 20150489 (2017).
Riley, J. R. et al. The long distance migration of Nilaparvata lugens (Stål) (Delphacidae) in China: radar observations of mass return flight in the autumn. Ecol. Entomol. 16, 471–489 (1991).
Riley, J. R., Reynolds, D. R., Mukhopadhyay, S., Ghosh, M. R. & Sarkar, T. K. Long-distance migration of aphids and other small insects in northeast India. Eur. J. Entomol. 92, 639–653 (1995).
Heong, K. L. & Hardy, B. Planthoppers: New Threats to the Sustainability of Intensive Rice Production System in Asia (International Rice Research Institute, Los Baños, 2009).
Xu, H. J. et al. Two insulin receptors determine alternative wing morphs in planthoppers. Nature 519, 464–467 (2015).
Lin, X. et al. Cell cycle progress determines wing morph in the polyphonic insect Nilaparvata lugens. iScience 23, 101040 (2020).
Lin, X. et al. JNK signaling mediates wing form polymorphism in brown planthoppers (Nilaparvata lugens). Insect Biochem. Mol. Biol. 73, 55–61 (2016).
Lin, X., Yao, Y., Wang, B., Lavine, M. D. & Lavine, L. C. FOXO links wing form polyphenism and wound healing the brown planthopper, Nilaparvata lugens. Insect Biochem. Mol. Biol. 70, 24–31 (2016).
Hayes, A. M., Lavine, M. D., Gotoh, H., Lin, X. & Lavine, L. C. Mechanisms regulating phenotypic plasticity in wing polyphenic insect. Adv. Insect Phys. 56, 43–72 (2019).
Lin, X., Yao, Y., Wang, B., Emlen, D. J. & Lavine, L. C. Ecological trade-offs between migration and reproduction are mediated by the nutrition-sensitive inulin-signaling pathway. Int. J. Biol. Sci. 12, 607–616 (2016).
Fawcett, M. M. et al. Manipulation of insulin signaling phenocopies evolution of a host-associated polyphenism. Nat. Commun. 9, 1699 (2018).
Smýkal, V. et al. Complex evolution of insect insulin receptors and homologous decoy receptors, and functional significance of their multiplicity. Mol. Biol. Evol. 37, 1775–1789 (2020).
Zhang, J. L. et al. Vestigial mediates the effect of insulin signaling pathway on wing-morph switching in planthoppers. PLoS Genet 17, e1009312 (2021).
Postigo, A. A. & Dean, D. C. ZEB, a vertebrate homolog of Drosophila Zfh-1, is a negative regulator of muscle differentiation. EMBO J. 16, 3935–3943 (1997).
Postigo, A. A., Ward, E., Skeath, J. B. & Dean, D. C. zfh-1, the Drosophila homologue of ZEB, is a transcriptional repressor that regulates somatic myogenesis. Mol. Cell. Biol. 19, 7255–7263 (1999).
Bray, S. & Boukhatmi, H. A population of adult satellite-like cells in Drosophila is maintained through a switch in RNA-isoforms. eLife 7, e35954 (2018).
Pearson, J. C., Lemons, D. & McGinnis, W. Modulating Hox gene functions during animal body patterning. Nat. Rev. Genet. 6, 893–904 (2005).
Lai, Z., Fortini, M. E. & Rubin, G. M. The embryonic expression patterns of Zfh-1 and Zfh-2, two Drosophila genes encoding novel zinc-finger homeodomain proteins. Mech. Dev. 34, 123–134 (1991).
Lundell, M. J. & Hirsh, J. The zfh-2 gene product is a potential regulator of neuron-specific DOPA decarboxylase gene expression in Drosophila. Dev. Biol. 154, 84–94 (1992).
Villa, S. E. R., Meng, F. W. & Biteau, B. zfh2 controls progenitor cell activation and differentiation in the adult Drosophila intestinal absorptive lineage. PLoS Genet. 15, e1008553 (2019).
Lai, Z., Rushton, E., Bate, M. & Rubin, G. M. Loss of function of the Drosophila zfh-1 gene results in abnormal development of mesodermally derived tissues. Proc. Natl Acad. Sci. USA 90, 4122–4126 (1993).
Perea, D., Molohon, K., Edwards, K. & Diaz-Benjumea, F. J. Multiple roles of the gene zinc finger homeodomain-2 in the development of the Drosophila wing. Mech. Dev. 130, 467–481 (2013).
Postigo, A. A. & Dean, D. C. Differential expression and function of members of the zfh-1 family of zinc finger/homeodomain repressors. Proc. Natl Acad. Sci. USA 97, 6391–6396 (2000).
Remacle, J. E. et al. New mode of DNA binding of multi-zinc finger transcription factors: deltaEF1 family members bind with two hands to two target sites. EMBO J. 18, 5073–5084 (1999).
Calnan, D. R. & Brunet, A. The FoxO code. Oncogene 27, 2276–2288 (2008).
Xue, W. H. et al. Neofunctionalization of a second insulin receptor gene in the wing-dimorphic planthoppers, Nilaparvata lugens. PLoS Genet. 17, e1009653 (2021).
Lewis, E. B. A gene complex controlling segmentation in Drosophila. Nature 276, 565–570 (1978).
Tomoyasu, Y., Wheeler, S. R. & Denell, R. E. Ultrabithorax is required for membranous wing identity in the beetle Tribolium castaneum. Nature 433, 643–647 (2005).
Gunage, R. D., Dhanyasi, N., Reichert, H. & VijayRaghavan, K. Drosophila adult muscle development and regeneration. Semin. Cell Dev. Biol. 72, 56–66 (2017).
Ye, Y. X. et al. Chromosome-level assembly of the brown planthopper genome with a characterized Y chromosome. Mol. Ecol. Resour. 21, 1287–1298 (2021).
Chen, S., Zhou, Y., Chen, Y. & Gu, J. fastp: an ultra-fast all-in-one FASTQ preprocessor. Bioinformatics 345, i884–i890 (2018).
Kim, D., Langmead, B. & Salzberg, S. L. HISAT: a fast spliced aligner with low memory requirements. Nat. Methods 12, 357 (2015).
Pertea, M. et al. StringTie enables improved reconstruction of a transcriptome from RNA-seq reads. Nat. Biotechnol. 33, 290–295 (2015).
Robinson, M. D., McCarthy, D. J. & Smyth, G. K. edgeR: a Bioconductor package for differential expression analysis of digital gene expression data. Bioinformatics 26, 139–140 (2010).
Love, M. I., Huber, W. & Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 15, 550 (2014).
Yuan, M. et al. Selection and evaluation of potential references genes for gene expression analysis in the brown planthopper, Nilaparvata lugens (Hemiptera: Delphacidae) using reverse-transcription quantitative PCR. PLoS ONE 9, e86503 (2014).
Wu, W. et al. Determination of suitable RT-qPCR reference genes for studies of gene funcitons in Laodelphax striatellus (Fallén). Genes 10, 887 (2019).
Xue, J. et al. Molecular characterization of the flightin gene in the wing-dimorphic planthopper, Nilaparvata lugens, and its evolution in Pancrustacea. Insect Biochem. Mol. Biol. 43, 433–443 (2013).
Miura, H., Quadros, R. M., Gurumurthy, C. B. & Ohtsuka, M. Easi-CRISPR for creating knock-in and conditional knockout mouse models using long ssDNA donors. Nat. Protoc. 13, 195–215 (2018).
Xue, W. H. et al. CRISPR/Cas9-mediated knockout of two eye pigmentation genes in the brown planthopper, Nilaparvata lugens (Hemiptera: Delphacidae). Insect Biochem. Mol. Biol. 93, 19–26 (2018).
Kanca, O. et al. An efficient CRISPR-based strategy to insert small and large fragments of DNA using short homology arms. eLife 8, e51539 (2019).
Reding, K. & Pick, L. High-efficiency CRISPR/Cas9 mutagenesis of the white gene in the milkweed bug Oncopeltus fasciatus. Genetics 215, 1027–1037 (2020).
Acknowledgements
H.J.X. was supported by the National Natural Science Foundation of China (31972261, 32272519, and 31772158), Key Projects of Natural Science Foundation of Zhejiang Province (LZ21C140002), and Development and Demonstration of Comprehensive Technology for Prevention and Control of Major Diseases and Pests (2021YFD1401100). J.L.Z. was supported by the National Natural Science Foundation of China (32102186). We thank Prof. Hongxia Hua in Huazhong Agricultural University for providing the WtLW strain.
Author information
Authors and Affiliations
Contributions
J.L.Z. and H.J.X. conceived the study and developed the initial study design. J.L.Z., S.J.C., and X.Y.L. performed the experiments. J.L.Z. and H.J.X. analyzed the experiments. A.P.M. and H.J.X. wrote the manuscript. All authors contributed intellectually to the experiments as well as editing and approval of the final version of the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Takahiro Ohde and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Peer reviewer reports are available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Zhang, JL., Chen, SJ., Liu, XY. et al. The transcription factor Zfh1 acts as a wing-morph switch in planthoppers. Nat Commun 13, 5670 (2022). https://doi.org/10.1038/s41467-022-33422-6
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41467-022-33422-6
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.