Featured
-
-
Article |
 SlyB encapsulates outer membrane proteins in stress-induced lipid nanodomains
SlyB encapsulates outer membrane proteins in stress-induced lipid nanodomains
SlyB, a lipoprotein in the PhoPQ stress regulon in Gram-negative bacteria, forms stable stress-induced complexes with the outer membrane proteome.
- Arne Janssens
- , Van Son Nguyen
- & Han Remaut
-
Article |
 Nucleic-acid-triggered NADase activation of a short prokaryotic Argonaute
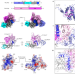
Nucleic-acid-triggered NADase activation of a short prokaryotic Argonaute
Cryo-electron microscopy structures of a short prokaryotic Argonaute (pAgo) and the associated TIR-APAZ proteins bound by guide RNA and target DNA shed light on the mechanisms of assembly and activation of pAgo systems.
- Xiaopan Gao
- , Kun Shang
- & Sheng Cui
-
Article |
 Structure of an endogenous mycobacterial MCE lipid transporter
Structure of an endogenous mycobacterial MCE lipid transporter
Proteins of the Mycobacterium smegmatis Mce1 system assemble to form an elongated ABC transporter complex that is long enough to span the impermeable mycobacterial cell envelope.
- James Chen
- , Alice Fruhauf
- & Damian C. Ekiert
-
Article |
 An E1–E2 fusion protein primes antiviral immune signalling in bacteria
An E1–E2 fusion protein primes antiviral immune signalling in bacteria
This study demonstrates that bacteria use a ubiquitin transferase-like enzyme to prime cGAS/DncV-like nucleotidyltransferase activation and use a deubiquitinase-like enzyme to decrease activity.
- Hannah E. Ledvina
- , Qiaozhen Ye
- & Aaron T. Whiteley
-
Article |
 Antiviral signalling by a cyclic nucleotide activated CRISPR protease
Antiviral signalling by a cyclic nucleotide activated CRISPR protease
A CalpL–CalpT–CalpS cascade mediated by cyclic oligoadenylates is identified as a mechanism to detect viral RNA and activate subsequent antivirus responses in microorganisms.
- Christophe Rouillon
- , Niels Schneberger
- & Gregor Hagelueken
-
Article
| Open Access Visualizing translation dynamics at atomic detail inside a bacterial cell
Visualizing translation dynamics at atomic detail inside a bacterial cell
Cryo-electron tomography is used to reveal the structural dynamics and functional diversity of translating ribosomes in Mycoplasma pneumoniae, providing insight into the translation elongation cycle inside cells and how it is reshaped by antibiotics.
- Liang Xue
- , Swantje Lenz
- & Julia Mahamid
-
Article |
 Cyclic nucleotide-induced helical structure activates a TIR immune effector
Cyclic nucleotide-induced helical structure activates a TIR immune effector
A bacterial antiviral defence system generates a cyclic tri-adenylate that binds to a TIR–SAVED effector, inducing formation of a superhelical structure with adjacent TIR domains organizing into an active site, allowing NAD+ degradation.
- Gaëlle Hogrel
- , Abbie Guild
- & Malcolm F. White
-
Article |
 Structural basis of lipopolysaccharide maturation by the O-antigen ligase
Structural basis of lipopolysaccharide maturation by the O-antigen ligase
Cryo-electron microscopy structures of the bacterial O-antigen ligase WaaL, combined with genetics, biochemistry and molecular dynamics simulations, provide insight into the mechanism by which WaaL catalyses the biosynthesis of lipopolysaccharide.
- Khuram U. Ashraf
- , Rie Nygaard
- & Filippo Mancia
-
Article
| Open Access Structure and dynamics of a mycobacterial type VII secretion system
Structure and dynamics of a mycobacterial type VII secretion system
A cryo-electron microscopy structure of the inner membrane complex of the ESX-5 type VII secretion system of Mycobacterium tuberculosis reveals an important role of interactions with MycP5 protease for complex integrity.
- Catalin M. Bunduc
- , Dirk Fahrenkamp
- & Thomas C. Marlovits
-
Article |
 Molecular basis for control of antibiotic production by a bacterial hormone
Molecular basis for control of antibiotic production by a bacterial hormone
X-ray crystallography and cryo-electron microscopy structures of the transcriptional repressor of the methylomycin gene cluster, MmfR, reveal the molecular basis for regulation of antibiotic biosynthesis by AHFCA hormones in Actinobacteria.
- Shanshan Zhou
- , Hussain Bhukya
- & Christophe Corre
-
Article |
 The Card1 nuclease provides defence during type III CRISPR immunity
The Card1 nuclease provides defence during type III CRISPR immunity
Structural analyses of the type III CRISPR accessory protein Card1, which induces dormancy in infected hosts to provide immunity against phage infection, reveal the mechanisms by which it cleaves single-stranded RNA and DNA.
- Jakob T. Rostøl
- , Wei Xie
- & Luciano A. Marraffini
-
Article |
 Action of a minimal contractile bactericidal nanomachine
Action of a minimal contractile bactericidal nanomachine
The authors report near-atomic resolution structures of the R-type bacteriocin from Pseudomonas aeruginosa in the pre-contraction and post-contraction states, and these structures provide insight into the mechanism of action of molecular syringes.
- Peng Ge
- , Dean Scholl
- & Z. Hong Zhou
-
Article |
 A mycobacterial ABC transporter mediates the uptake of hydrophilic compounds
A mycobacterial ABC transporter mediates the uptake of hydrophilic compounds
Analysis of cryo-electron microscopy structures of the Mycobacterium tuberculosis ABC transporter Rv1819c suggests that it is a multi-solute transporter for hydrophilic molecules.
- S. Rempel
- , C. Gati
- & D. J. Slotboom
-
Article |
 An interbacterial toxin inhibits target cell growth by synthesizing (p)ppApp
An interbacterial toxin inhibits target cell growth by synthesizing (p)ppApp
The bacterium Pseudomonas aeruginosa attacks competing bacteria using the toxin Tas1, which pyrophosphorylates adenosine nucleotides to generate (p)ppApp, thereby depleting ATP and disrupting multiple cellular functions.
- Shehryar Ahmad
- , Boyuan Wang
- & John C. Whitney
-
Article |
 Structural basis of lipopolysaccharide extraction by the LptB2FGC complex
Structural basis of lipopolysaccharide extraction by the LptB2FGC complex
Cryo-electron microscopy structures of LptB2FGC, in nucleotide-free and vanadate-trapped states, reveal the mechanism of lipopolysaccharide extraction from the inner membrane of Gram-negative bacteria and a role for LptC in efficient lipopolysaccharide transport.
- Yanyan Li
- , Benjamin J. Orlando
- & Maofu Liao
-
Letter |
 Mechanism of phosphoribosyl-ubiquitination mediated by a single Legionella effector
Mechanism of phosphoribosyl-ubiquitination mediated by a single Legionella effector
Crystal structures of the Legionella effectors SdeA and SdeD uncover the mechanism of a unique phosphoribosyl-ubiquitination reaction.
- Anil Akturk
- , David J. Wasilko
- & Yuxin Mao
-
Article |
 Structural basis of ubiquitin modification by the Legionella effector SdeA
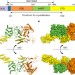
Structural basis of ubiquitin modification by the Legionella effector SdeA
Crystal structures of the Legionella effector SdeA in a ligand-free state and in complex with ubiquitin and NADH provide insight into SdeA-mediated phosphoribosyl-linked ubiquitination.
- Yanan Dong
- , Yajuan Mu
- & Yue Feng
-
Article |
 Structural basis of MsbA-mediated lipopolysaccharide transport
Structural basis of MsbA-mediated lipopolysaccharide transport
Cryo-electron microscopy snapshots of the E. coli flippase MsbA at discrete functional states reveal a ‘trap and flip’ mechanism for lipopolysaccharide flipping and the conformational transitions of MsbA during its substrate transport cycle.
- Wei Mi
- , Yanyan Li
- & Maofu Liao
-
Letter |
 Structure of the Cpf1 endonuclease R-loop complex after target DNA cleavage
Structure of the Cpf1 endonuclease R-loop complex after target DNA cleavage
The structure of Cpf1, a CRISPR–Cas/RNA-guided nuclease, is presented with a three-stranded RNA–DNA loop after cleavage, providing insight into its working mechanism.
- Stefano Stella
- , Pablo Alcón
- & Guillermo Montoya
-
Letter |
 Structural basis for nutrient acquisition by dominant members of the human gut microbiota
Structural basis for nutrient acquisition by dominant members of the human gut microbiota
The authors present structures of nutrient transport complexes of the commensal bacterium Bacteroides thetaiotaomicron and the mechanism by which it imports glycans.
- Amy J. Glenwright
- , Karunakar R. Pothula
- & Bert van den Berg
-
Letter |
 Mechanistic insights into the alternative translation termination by ArfA and RF2
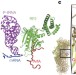
Mechanistic insights into the alternative translation termination by ArfA and RF2
The structure of the bacterial 70S ribosome in complex with ArfA, the release factor RF2, a short non-stop mRNA and a cognate P-site tRNA is presented, revealing how ArfA and RF2 facilitate alternative translation termination of the non-stop ribosomal complex using a stop-codon surrogate mechanism.
- Chengying Ma
- , Daisuke Kurita
- & Ning Gao
-
Letter |
 Structural basis for ArfA–RF2-mediated translation termination on mRNAs lacking stop codons
Structural basis for ArfA–RF2-mediated translation termination on mRNAs lacking stop codons
The structure of the bacterial ribosome stalled on a truncated mRNA in complex with ArfA and the release factor RF2 is presented, revealing how ArfA recruits RF2 to the ribosome and induces conformational changes within RF2 to enable translation termination in the absence of a stop codon.
- Paul Huter
- , Claudia Müller
- & Daniel N. Wilson
-
Article |
 Structural insight into the role of the Ton complex in energy transduction
Structural insight into the role of the Ton complex in energy transduction
Structural studies shed light on the function and stoichiometry of the Ton complex, which harnesses the proton motive force across the bacterial inner membrane to transduce energy to the outer membrane.
- Hervé Celia
- , Nicholas Noinaj
- & Susan K. Buchanan
-
Letter |
 Ribosome-dependent activation of stringent control
Ribosome-dependent activation of stringent control
The structure of a bacterial ribosome–RelA complex reveals that RelA, a protein recruited to the ribosome in the case of scarce amino acids, binds in a different location to translation factors, and that this binding event suppresses auto-inhibition to activate synthesis of the (p)ppGpp secondary messenger, thus initiating stringent control.
- Alan Brown
- , Israel S. Fernández
- & V. Ramakrishnan
-
Letter |
 The crystal structure of Cpf1 in complex with CRISPR RNA
The crystal structure of Cpf1 in complex with CRISPR RNA
The crystal structure of monomeric Lachnospiraceae bacterium Cpf1 protein bound to CRISPR RNA is presented, establishing a framework for engineering LbCpf1 to improve its efficiency and specificity for genome editing.
- De Dong
- , Kuan Ren
- & Zhiwei Huang
-
Article |
 Priming and polymerization of a bacterial contractile tail structure
Priming and polymerization of a bacterial contractile tail structure
A combination of X-ray crystallography, electron microscopy, functional assays and time-lapse fluorescence microscopy shows that a protein of previously unknown function, TssA, forms a dodecameric complex that interacts with components of the tube and sheath of the type VI secretion system of bacteria, and that it primes and coordinates biogenesis of both the tail tube and the sheath.
- Abdelrahim Zoued
- , Eric Durand
- & Eric Cascales
-
Article |
 Structural basis of outer membrane protein insertion by the BAM complex
Structural basis of outer membrane protein insertion by the BAM complex
Two crystal structures of the Escherichia coli β-barrel assembly machinery (BAM complex) are presented, one of which includes all five subunits (BamA–BamE), in two distinct conformational states; together with functional assays and molecular dynamics stimulations, these structures help to generate a model for outer membrane protein insertion.
- Yinghong Gu
- , Huanyu Li
- & Changjiang Dong
-
Letter |
 Structural basis for promiscuous PAM recognition in type I–E Cascade from E. coli
Structural basis for promiscuous PAM recognition in type I–E Cascade from E. coli
The structure of E. coli Cascade bound to foreign target DNA is presented, revealing the basis of the relaxed Cascade PAM recognition specificity, which results from its interaction with the minor groove, and demonstrating how a wedge in Cascade forces the directional pairing of the target strand with CRISPR RNA while stabilizing the non-target displaced strand.
- Robert P. Hayes
- , Yibei Xiao
- & Ailong Ke
-
Article |
 Selective small-molecule inhibition of an RNA structural element
Selective small-molecule inhibition of an RNA structural element
A novel drug, ribocil, is shown to mimic the binding of a natural ligand to a bacterial riboflavin riboswitch (a non-coding stretch of messenger RNA whose structure is affected by a ligand—usually one related to the function of the protein encoded by the messenger RNA) to cause inhibition of bacterial growth; the ability to target an RNA structural element with a synthetic small molecule may expand our view of the target space susceptible to therapeutic intervention.
- John A. Howe
- , Hao Wang
- & Terry Roemer
-
Letter |
 Structure of the E. coli ribosome–EF-Tu complex at <3 Å resolution by Cs-corrected cryo-EM
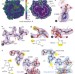
Structure of the E. coli ribosome–EF-Tu complex at <3 Å resolution by Cs-corrected cryo-EM
A single particle cryo-EM structure of the 70S ribosome in complex with the elongation factor Tu breaks the 3 Å resolution barrier of the technique and locally exceeds the resolution of previous crystallographic studies, revealing all modifications in rRNA and explaining their roles in ribosome function and antibiotic binding.
- Niels Fischer
- , Piotr Neumann
- & Holger Stark
-
Article |
 Structural basis of the non-coding RNA RsmZ acting as a protein sponge
Structural basis of the non-coding RNA RsmZ acting as a protein sponge
A novel combined NMR and EPR spectroscopy approach reveals the structure and assembly mechanism of a 70-kDa bacterial ribonucleoprotein complex acting as a protein sponge in translational regulation.
- Olivier Duss
- , Erich Michel
- & Frédéric H.-T. Allain
-
Letter |
 Structure and mechanism of a bacterial sodium-dependent dicarboxylate transporter
Structure and mechanism of a bacterial sodium-dependent dicarboxylate transporter
The cytosolic concentration of citrate partially depends on its direct import across the plasma membrane by the Na+-dependent citrate transporter (NaCT); here the X-ray crystal structure of a bacterial homologue of NaCT is reported, which, along with transport-activity studies, suggests how specific conformational changes facilitate substrate translocation across the cellular membrane.
- Romina Mancusso
- , G. Glenn Gregorio
- & Da-Neng Wang
-
Letter |
 SbsB structure and lattice reconstruction unveil Ca2+ triggered S-layer assembly
SbsB structure and lattice reconstruction unveil Ca2+ triggered S-layer assembly
Nanobody-aided X-ray crystallography and cryo-electron microscopy are used to describe the Ca2+-dependent polymerization dynamics of the S-layer of the Geobacillus stearothermophilus cell wall.
- Ekaterina Baranova
- , Rémi Fronzes
- & Han Remaut
-
Letter |
 Atomic model of the type III secretion system needle
Atomic model of the type III secretion system needle
The structure of the needle of the type III secretion system of Salmonella typhimurium, used to inject virulence proteins into host cells during infection, has been resolved by a combination of in vitro needle production, solid-state nuclear magnetic resonance, electron microscopy and Rosetta modelling at atomic resolution.
- Antoine Loquet
- , Nikolaos G. Sgourakis
- & Adam Lange
-
Letter |
 Identification and characterization of a bacterial hydrosulphide ion channel
Identification and characterization of a bacterial hydrosulphide ion channel
A channel for the transport of hydrosulphide ions in Clostridium difficile is identified and shown to be polyspecific, being a member of the formate/nitrite transporter family.
- Bryan K. Czyzewski
- & Da-Neng Wang
-
News |
 Study challenges existence of arsenic-based life
Study challenges existence of arsenic-based life
Open-science advocates fail to reproduce controversial findings.
- Erika Check Hayden
-
Letter |
 The NLRC4 inflammasome receptors for bacterial flagellin and type III secretion apparatus
The NLRC4 inflammasome receptors for bacterial flagellin and type III secretion apparatus
- Yue Zhao
- , Jieling Yang
- & Feng Shao
-
Article |
 Crystal structure of a phosphorylation-coupled saccharide transporter
Crystal structure of a phosphorylation-coupled saccharide transporter
- Yu Cao
- , Xiangshu Jin
- & Ming Zhou
-
Letter |
 Crystal structure of the CusBA heavy-metal efflux complex of Escherichia coli
Crystal structure of the CusBA heavy-metal efflux complex of Escherichia coli
Gram-negative bacteria expel toxic chemicals via tripartite efflux pumps spanning both the inner and outer membranes. A crystallographic model of this tripartite efflux complex has been unavailable because co-crystallization of different components of the system has proven to be extremely difficult. The X-ray crystal structure of CusA of the CusCBA tripartite efflux system from Escherichia coli has been reported previously, and here the X-ray crystal structure of the CusBA co-complex is reported. The structure reveals that the trimeric CusA efflux pump interacts with six CusB protein molecules at the upper half of the periplasmic domain, and the predicted structure of the trimeric CusC channel was used to develop a model of for the tripartite efflux complex.
- Chih-Chia Su
- , Feng Long
- & Edward W. Yu
-
News & Views |
 A peep through anion channels
A peep through anion channels
The crystal structure of a protein channel provides clues about the mechanisms that control the closure of pores found in the epidermis of plant leaves. Excitingly, the protein channel folds in a way never seen before. See Article p.1074
- Sébastien Thomine
- & Hélène Barbier-Brygoo

 Structures and activation mechanism of the Gabija anti-phage system
Structures and activation mechanism of the Gabija anti-phage system