Article
|
Open Access
Featured
-
-
Article
| Open Access Colonisation of hospital surfaces from low- and middle-income countries by extended spectrum β-lactamase- and carbapenemase-producing bacteria
Colonisation of hospital surfaces from low- and middle-income countries by extended spectrum β-lactamase- and carbapenemase-producing bacteria
In hospitals, surfaces present as a reservoir for bacteria pathogens, potentially leading to nosocomial infections. In this work, authors aim to profile extended-spectrum β lactamase- and carbapenemase-carrying bacterial species colonising neonatal hospital wards and causing neonatal sepsis.
- Maria Nieto-Rosado
- , Kirsty Sands
- & Timothy R. Walsh
-
Article
| Open Access Microevolution, reinfection and highly complex genomic diversity in patients with sequential isolates of Mycobacterium abscessus
Microevolution, reinfection and highly complex genomic diversity in patients with sequential isolates of Mycobacterium abscessus
Mycobacterium abscessus is considered an emerging pathogen, given its prevalence in patients with pulmonary diseases, such as cystic fibrosis. Here, authors perform a genomic analysis on sequential isolates obtained from patients with persistent infections of M. abscessus.
- Sergio Buenestado-Serrano
- , Miguel Martínez-Lirola
- & Darío García de Viedma
-
Article
| Open Access Inter-species gene flow drives ongoing evolution of Streptococcus pyogenes and Streptococcus dysgalactiae subsp. equisimilis
Inter-species gene flow drives ongoing evolution of Streptococcus pyogenes and Streptococcus dysgalactiae subsp. equisimilis
Streptococcus dysgalactiae subsp. equisimilis (SDSE) is an emerging cause of human infection closely related to Streptococcus pyogenes. Here the authors investigate the degree of genomic similarity between the two species and assess implications for development of vaccines.
- Ouli Xie
- , Jacqueline M. Morris
- & Mark R. Davies
-
Article
| Open Access The defensome of complex bacterial communities
The defensome of complex bacterial communities
Bacteria have evolved numerous innate and adaptive defence mechanisms. Here, Beavogui et al characterise the impact of biogeography, genetic mobility, and clustering in defense islands, on the defence systems of soil, marine, and human gut bacterial populations genomes.
- Angelina Beavogui
- , Auriane Lacroix
- & Pedro H. Oliveira
-
Article
| Open Access The global speciation continuum of the cyanobacterium Microcoleus
The global speciation continuum of the cyanobacterium Microcoleus
The relative importance of the various mechanisms that can drive microbial speciation is poorly understood. Here, Stanojković et al. explore the diversification of the soil cyanobacterium Microcoleus, showing that this genus represents a global speciation continuum of at least 12 lineages, with lineage divergence driven by selection, geographical distance, and the environment.
- Aleksandar Stanojković
- , Svatopluk Skoupý
- & Petr Dvořák
-
Article
| Open Access bacLIFE: a user-friendly computational workflow for genome analysis and prediction of lifestyle-associated genes in bacteria
bacLIFE: a user-friendly computational workflow for genome analysis and prediction of lifestyle-associated genes in bacteria
Many bacteria live in close association with eukaryotic hosts, exhibiting detrimental, neutral or beneficial effects on host growth and health. Here, the authors present a streamlined computational workflow for bacterial genome annotation, large-scale comparative genomics, and prediction of genes potentially involved in niche adaptation.
- Guillermo Guerrero-Egido
- , Adrian Pintado
- & Víctor J. Carrión
-
Article
| Open Access Global emergence of a hypervirulent carbapenem-resistant Escherichia coli ST410 clone
Global emergence of a hypervirulent carbapenem-resistant Escherichia coli ST410 clone
In this work, the authors identified a hypervirulent carbapenem-resistant Escherichia coli ST410 clone which carries a high pathogenicity island and an O-antigen gene cluster. The findings highlight the ongoing evolution of ST410 towards increased resistance and virulence.
- Xiaoliang Ba
- , Yingyi Guo
- & Chao Zhuo
-
Article
| Open Access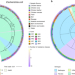 Convergence of resistance and evolutionary responses in Escherichia coli and Salmonella enterica co-inhabiting chicken farms in China
Convergence of resistance and evolutionary responses in Escherichia coli and Salmonella enterica co-inhabiting chicken farms in China
Bacteria in the same environment can share genetic material but the extent to which this influences development of antimicrobial resistance is unclear. Here, the authors investigate the evidence for co-evolution of antimicrobial resistance in bacteria found coexisting in animals and the environment in chicken farms and slaughterhouses in China.
- Michelle Baker
- , Xibin Zhang
- & Tania Dottorini
-
Article
| Open Access Increase in antioxidant capacity associated with the successful subclone of hypervirulent carbapenem-resistant Klebsiella pneumoniae ST11-KL64
Increase in antioxidant capacity associated with the successful subclone of hypervirulent carbapenem-resistant Klebsiella pneumoniae ST11-KL64
Plasmid acquisition imposes an adaptive burden, which can be ameliorated by host-plasmid coevolution. Here, the authors characterise virulence plasmids of carbapenem-resistant Klebsiella pneumoniae, and show the discard of certain sequences to enhance survival, conferring an evolutionary advantage.
- Ruobing Wang
- , Anru Zhang
- & Hui Wang
-
Article
| Open Access The Helicobacter pylori Genome Project: insights into H. pylori population structure from analysis of a worldwide collection of complete genomes
The Helicobacter pylori Genome Project: insights into H. pylori population structure from analysis of a worldwide collection of complete genomes
The bacterium Helicobacter pylori, often found in the human stomach, can be classified into distinct subpopulations associated with the geographic origin of the host. Here, the authors provide insights into H. pylori population structure by collecting over 1,000 clinical strains from 50 countries and generating and analyzing high-quality bacterial genome sequences.
- Kaisa Thorell
- , Zilia Y. Muñoz-Ramírez
- & Charles S. Rabkin
-
Article
| Open Access National genomic surveillance integrating standardized quantitative susceptibility testing clarifies antimicrobial resistance in Enterobacterales
National genomic surveillance integrating standardized quantitative susceptibility testing clarifies antimicrobial resistance in Enterobacterales
Kayama et al. present a blueprint for a national genomic surveillance study that conducts genome sequencing of thousands of strains, integrates standardized quantitative antimicrobial susceptibility testing, and characterizes antimicrobial resistance determinants.
- Shizuo Kayama
- , Koji Yahara
- & Motoyuki Sugai
-
Article
| Open Access Distributed genotyping and clustering of Neisseria strains reveal continual emergence of epidemic meningococcus over a century
Distributed genotyping and clustering of Neisseria strains reveal continual emergence of epidemic meningococcus over a century
Core genome multilocus sequence typing (cgMLST) is used to classify bacterial strains for epidemiological applications. Here, the authors describe a distributed cgMLST scheme that does not require a central database of allelic sequences, and apply it to study evolutionary patterns of epidemic and endemic strains of the genus Neisseria.
- Ling Zhong
- , Menghan Zhang
- & Zhemin Zhou
-
Article
| Open Access Taxonomic and environmental distribution of bacterial amino acid auxotrophies
Taxonomic and environmental distribution of bacterial amino acid auxotrophies
Many microorganisms are auxotrophic, that is, unable to synthesize the compounds they require for growth. Here, Ramoneda et al. predict amino acid biosynthetic capabilities of over 26,000 bacterial genomes using a metabolic pathway model validated with empirical data, and identify ecological contexts in which auxotrophy can be a successful strategy.
- Josep Ramoneda
- , Thomas B. N. Jensen
- & Noah Fierer
-
Article
| Open Access Bacterial genome size and gene functional diversity negatively correlate with taxonomic diversity along a pH gradient
Bacterial genome size and gene functional diversity negatively correlate with taxonomic diversity along a pH gradient
Bacterial functional diversity does not necessarily correlate with taxonomic diversity because average genome size may vary by community. Here, Wang et al. investigate bacterial communities along a natural pH gradient in forest soils, and find that average genome size and functional diversity decrease, whereas taxonomic diversity increases, as soil pH rises from acid to neutral.
- Cong Wang
- , Qing-Yi Yu
- & Cheng Gao
-
Article
| Open Access Functional annotation of enzyme-encoding genes using deep learning with transformer layers
Functional annotation of enzyme-encoding genes using deep learning with transformer layers
Functional annotation of open reading frames in microbial genomes remains substantially incomplete. Here, Kim et al. present a deep learning model that utilizes transformer layers as a neural network architecture to predict specific catalytic functions for enzyme-encoding genes of unknown function.
- Gi Bae Kim
- , Ji Yeon Kim
- & Sang Yup Lee
-
Article
| Open Access Bacteria can maintain rRNA operons solely on plasmids for hundreds of millions of years
Bacteria can maintain rRNA operons solely on plasmids for hundreds of millions of years
Bacteria usually have at least one rRNA operon on the chromosome, suggesting that the exclusive presence of rRNA operons on a plasmid is rare and unlikely to be stably maintained. Here, Anda et al. find that at least four bacterial clades in different phyla lost their chromosomal rRNA operons independently, and one of the clades has maintained this peculiar genome organization for hundreds of millions of years.
- Mizue Anda
- , Shun Yamanouchi
- & Wataru Iwasaki
-
Article
| Open Access Essential gene complement of Planctopirus limnophila from the bacterial phylum Planctomycetes
Essential gene complement of Planctopirus limnophila from the bacterial phylum Planctomycetes
Bacteria of the phylum Planctomycetes display unique cell biology features but are relatively understudied. Here, the authors report a genome-wide analysis of essential gene content in a planctomycete, providing insights into the divergent molecular and cell biology of these organisms.
- Elena Rivas-Marin
- , David Moyano-Palazuelo
- & Damien P. Devos
-
Article
| Open Access Cryptic susceptibility to penicillin/β-lactamase inhibitor combinations in emerging multidrug-resistant, hospital-adapted Staphylococcus epidermidis lineages
Cryptic susceptibility to penicillin/β-lactamase inhibitor combinations in emerging multidrug-resistant, hospital-adapted Staphylococcus epidermidis lineages
Staphylococcus epidermidis can cause invasive infections that are difficult to treat due to multi-resistance to most clinically relevant drugs, including methicillin and other β-lactam antibiotics, vancomycin, and rifampicin. In this work, the authors use in vitro assays and a mouse infection model to explore cryptic susceptibility and development of resistance to penicillin/β-lactamase combinations.
- Xiaoliang Ba
- , Claire L. Raisen
- & Jesper Larsen
-
Article
| Open Access Herbarium specimen sequencing allows precise dating of Xanthomonas citri pv. citri diversification history
Herbarium specimen sequencing allows precise dating of Xanthomonas citri pv. citri diversification history
Herbarium collections are an important source of historical DNA, whose analysis can shed light on the evolutionary history of plant pathogens. Here, Campos et al. reconstruct historical genomes of the bacterial crop pathogen Xanthomonas citri pv. citri from citrus herbarium specimens, estimating that the pathogen originated in Southern Asia ~11,500 years ago and diversified during the beginning of the 13th century.
- Paola E. Campos
- , Olivier Pruvost
- & Lionel Gagnevin
-
Article
| Open Access Decoding a cryptic mechanism of metronidazole resistance among globally disseminated fluoroquinolone-resistant Clostridioides difficile
Decoding a cryptic mechanism of metronidazole resistance among globally disseminated fluoroquinolone-resistant Clostridioides difficile
Detection of resistance to the antibiotic metronidazole in C. difficile often requires the presence of heme in the media, for unclear reasons. Here, the authors show that most metronidazole-resistant strains carry a mutation that promotes expression of a heme-dependent enzyme that degrades nitroimidazoles, and the mutation often co-occurs with an amino-acid substitution in DNA gyrase that confers resistance to another class of antibiotics, fluoroquinolones.
- Abiola O. Olaitan
- , Chetna Dureja
- & Julian G. Hurdle
-
Article
| Open Access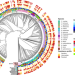 Roving methyltransferases generate a mosaic epigenetic landscape and influence evolution in Bacteroides fragilis group
Roving methyltransferases generate a mosaic epigenetic landscape and influence evolution in Bacteroides fragilis group
Here, Tisza, Dekker, and colleagues perform large scale analysis of genome methylation in the gut commensal and pathogen, Bacteroides fragilis group, revealing immense methyl motif diversity and evidence of widespread methyltransferase exchange among phages.
- Michael J. Tisza
- , Derek D. N. Smith
- & John P. Dekker
-
Article
| Open Access Genomic epidemiology of Vibrio cholerae during a mass vaccination campaign of displaced communities in Bangladesh
Genomic epidemiology of Vibrio cholerae during a mass vaccination campaign of displaced communities in Bangladesh
The Cox’s Bazar area of Bangladesh has received a large number of Forcibly Displaced Myanmar Nationals. Cholera outbreaks have been detected in the area, and here, the authors perform genomic surveillance of cholera in the refugee and non-refugee population to infer the risk of epidemic spread.
- Alyce Taylor-Brown
- , Mokibul Hassan Afrad
- & Firdausi Qadri
-
Article
| Open Access Evolutionary and functional history of the Escherichia coli K1 capsule
Evolutionary and functional history of the Escherichia coli K1 capsule
Little is known about the distribution, evolution and functions of the K1 capsule at a population level, despite the important role in the pathogenesis of E. coli; authors explore this through the utilisation of over 5,000 clinical isolates in population genomics studies and statistical modelling.
- Sergio Arredondo-Alonso
- , George Blundell-Hunter
- & Alex J. McCarthy
-
Article
| Open Access De novo cholesterol biosynthesis in bacteria
De novo cholesterol biosynthesis in bacteria
Production of highly modified sterols, such as cholesterol, is essential to eukaryotic physiology but has not been yet reported for bacteria. Here, Lee et al. show that a marine myxobacterium produces cholesterol, and provide evidence for further downstream modifications in this and other bacterial species.
- Alysha K. Lee
- , Jeremy H. Wei
- & Paula V. Welander
-
Article
| Open Access Roles of adenine methylation in the physiology of Lacticaseibacillus paracasei
Roles of adenine methylation in the physiology of Lacticaseibacillus paracasei
The bacterium Lacticaseibacillus paracasei is used in the food industry and as a probiotic. Here, the authors use multi-omics and high-throughput chromosome conformation capture analyses to investigate the roles of a type of DNA methylation (N6-methyladenine modification) in this organism.
- Jie Zhao
- , Meng Zhang
- & Wenyi Zhang
-
Article
| Open Access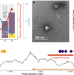 Mutation-induced infections of phage-plasmids
Mutation-induced infections of phage-plasmids
Phage-plasmids are bacterial extrachromosomal elements that act both as plasmids and as viruses. Here, Shan et al. show that segregational drift and loss-of-function mutations play key roles in the infection dynamics of a cosmopolitan phage-plasmid, allowing it to create continuous productive infections in marine bacteria.
- Xiaoyu Shan
- , Rachel E. Szabo
- & Otto X. Cordero
-
Article
| Open Access The relative transmission fitness of multidrug-resistant Mycobacterium tuberculosis in a drug resistance hotspot
The relative transmission fitness of multidrug-resistant Mycobacterium tuberculosis in a drug resistance hotspot
Geographical hotspots with high frequency of multi-drug resistant tuberculosis (MDR-TB) have been observed in several locations, such as the country of Georgia. Here, the authors analyse genomic sequences from tuberculosis bacteria isolated from Georgia to show that the transmission fitness of MDR-TB strains is heterogeneous, and highly drug-resistant and transmissible isolates contribute to the emergence and maintenance of MDR-TB hotspots.
- Chloé Loiseau
- , Etthel M. Windels
- & Sebastien Gagneux
-
Article
| Open Access The genomic epidemiology of Escherichia albertii infecting humans and birds in Great Britain
The genomic epidemiology of Escherichia albertii infecting humans and birds in Great Britain
Escherichia albertii is an emerging gastrointestinal pathogen that causes disease in humans and animals, notably birds. In this genomic epidemiology study, the authors investigate characteristics of isolates sampled from humans and birds in Great Britain and find that they tend to cluster separately.
- Rebecca J. Bengtsson
- , Kate S. Baker
- & Becki Lawson
-
Article
| Open Access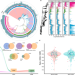 The genomic landscape of reference genomes of cultivated human gut bacteria
The genomic landscape of reference genomes of cultivated human gut bacteria
Here, the authors present an expanded version of the Cultivated Genome Reference (CGR), termed CGR2, a catalog that includes 3324 high-quality draft genomes based on gut bacterial isolates from Chinese individuals, and classifies 527 species from 8 phyla, including 179 previously unidentified species, and provides information of secondary metabolite biosynthetic gene clusters and gut phage-bacteria interactions.
- Xiaoqian Lin
- , Tongyuan Hu
- & Yuanqiang Zou
-
Article
| Open Access Genomic diversity of non-diarrheagenic fecal Escherichia coli from children in sub-Saharan Africa and south Asia and their relatedness to diarrheagenic E. coli
Genomic diversity of non-diarrheagenic fecal Escherichia coli from children in sub-Saharan Africa and south Asia and their relatedness to diarrheagenic E. coli
Escherichia coli is a human pathogen and a member of the healthy microbiota. Here, using samples from children of low and middle-income living in sub-Saharan Africa and south Asia countries, the authors characterize the genomic landscape of non-diarrheagenic fecal E. coli, finding similarities to diarrheagenic pathotype E. coli.
- Tracy H. Hazen
- , Jane M. Michalski
- & David. A. Rasko
-
Article
| Open Access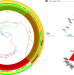 Genomic attributes of Vibrio cholerae O1 responsible for 2022 massive cholera outbreak in Bangladesh
Genomic attributes of Vibrio cholerae O1 responsible for 2022 massive cholera outbreak in Bangladesh
Vibrio cholerae has undergone continuous evolution, and differing strains have caused numerous outbreaks. Here, the authors present a genomic study of Vibrio cholerae O1 responsible for a 2022 outbreak in Dhaka.
- Md Mamun Monir
- , Mohammad Tarequl Islam
- & Munirul Alam
-
Article
| Open Access Rapid emergence of extensively drug-resistant Shigella sonnei in France
Rapid emergence of extensively drug-resistant Shigella sonnei in France
There have been increasing reports of extensively drug-resistant (XDR) Shigella sonnei infections in recent years. In this laboratory surveillance study from France, the authors document the rise of XDR isolates from 2005 to 2021 and perform whole genome sequencing to investigate their genomic diversity and evolutionary history.
- Sophie Lefèvre
- , Elisabeth Njamkepo
- & François-Xavier Weill
-
Article
| Open Access Resolving colistin resistance and heteroresistance in Enterobacter species
Resolving colistin resistance and heteroresistance in Enterobacter species
Taxonomical complexity has muddled the classification of clinically relevant Enterobacter species. Authors carry out a genome-based study on clinical isolates to investigate colistin resistance and heteroresistance in Enterobacter.
- Swapnil Prakash Doijad
- , Nicolas Gisch
- & Trinad Chakraborty
-
Article
| Open Access Paratype: a genotyping tool for Salmonella Paratyphi A reveals its global genomic diversity
Paratype: a genotyping tool for Salmonella Paratyphi A reveals its global genomic diversity
The bacterium Salmonella Paratyphi A causes paratyphoid fever. Here, the authors sequence over 800 isolates from South Asia, build a global database representing 37 countries, and develop a genotyping tool that identifies genomic variation and antimicrobial resistance markers for surveillance studies.
- Arif M. Tanmoy
- , Yogesh Hooda
- & Senjuti Saha
-
Article
| Open Access Dynamics of extended-spectrum cephalosporin resistance genes in Escherichia coli from Europe and North America
Dynamics of extended-spectrum cephalosporin resistance genes in Escherichia coli from Europe and North America
Extended-spectrum cephalosporin resistance genes in Escherichia coli have spread worldwide. Here, the authors dissect the emergence and distribution of these genes over time, and across geographic location and host species, to better understand their dynamics and mechanisms of transmission.
- Roxana Zamudio
- , Patrick Boerlin
- & Alison E. Mather
-
Article
| Open Access A comprehensive update to the Mycobacterium tuberculosis H37Rv reference genome
A comprehensive update to the Mycobacterium tuberculosis H37Rv reference genome
H37Rv is the most widely used Mycobacterium tuberculosis strain, and its genome is the reference sequence for this pathogen. Here, Chitale et al. present a bioinformatic pipeline for accurate assembly of bacterial genome sequences, and use it to provide important updates to the M. tuberculosis reference genome.
- Poonam Chitale
- , Alexander D. Lemenze
- & David Alland
-
Article
| Open Access Repeated out-of-Africa expansions of Helicobacter pylori driven by replacement of deleterious mutations
Repeated out-of-Africa expansions of Helicobacter pylori driven by replacement of deleterious mutations
Helicobacter pylori is a major human pathogen whose population structure is similar to that of its host. Here, the authors show that H. pylori has repeatedly spread out of Africa recently, replacing deleterious variants that accumulated during the original out of Africa migrations more than 50,000 years ago.
- Harry A. Thorpe
- , Elise Tourrette
- & Daniel Falush
-
Article
| Open Access Genomic insights into the physiology of Quinella, an iconic uncultured rumen bacterium
Genomic insights into the physiology of Quinella, an iconic uncultured rumen bacterium
Uncultured bacteria of the genus Quinella are found in the rumen of ruminant animals, especially in sheep that emit low amounts of methane. Here, Kumar et al. reconstruct genomic sequences from Quinella cells to provide insights into their metabolic capabilities, including lactate and propionate formation as major fermentation pathways and an apparent lack of production of H2, a major precursor of methane.
- Sandeep Kumar
- , Eric Altermann
- & Peter H. Janssen
-
Article
| Open Access Anaerobic oxidation of propane coupled to nitrate reduction by a lineage within the class Symbiobacteriia
Anaerobic oxidation of propane coupled to nitrate reduction by a lineage within the class Symbiobacteriia
Anaerobic microorganisms can oxidize short-chain gaseous alkanes such as ethane, propane and butane using sulfate as electron acceptor. Here, the authors show that a bioreactor enrichment of a wastewater microbial community can perform anaerobic propane oxidation coupled to nitrate reduction.
- Mengxiong Wu
- , Jie Li
- & Jianhua Guo
-
Article
| Open Access The genus Serratia revisited by genomics
The genus Serratia revisited by genomics
The genus Serratia includes clinically-important and diverse environmental bacteria. Here, Williams et al. assemble and analyse a representative set of 664 genomes from across the genus, including historic isolates, to provide a genome-based phylogenetic framework for a better understanding of the emergence of clinical and environmental lineages of Serratia.
- David J. Williams
- , Patrick A. D. Grimont
- & Sarah J. Coulthurst
-
Article
| Open Access Delineating Mycobacterium abscessus population structure and transmission employing high-resolution core genome multilocus sequence typing
Delineating Mycobacterium abscessus population structure and transmission employing high-resolution core genome multilocus sequence typing
Mycobacterium abscessus is an emerging infection of increasing public health concern due to outbreaks and intrinsic multidrug-resistance. Here, the authors develop and evaluate a core-genome multilocus sequence typing scheme for this pathogen to facilitate standardised molecular surveillance.
- Margo Diricks
- , Matthias Merker
- & Florian P. Maurer
-
Article
| Open Access Combined comparative genomics and clinical modeling reveals plasmid-encoded genes are independently associated with Klebsiella infection
Combined comparative genomics and clinical modeling reveals plasmid-encoded genes are independently associated with Klebsiella infection
Patient variables, such as comorbidities, partially explain which patients will progress to Klebsiella infection, with colonization of the gut acting as a reservoir. Little is known, however, regarding Klebsiella genes that may increase risk of disease in colonized individuals. Here, authors conduct a comparative genomics study to identify genes associated with progression from colonisation to infection.
- Jay Vornhagen
- , Emily K. Roberts
- & Michael A. Bachman
-
Article
| Open Access Population genomics of Group B Streptococcus reveals the genetics of neonatal disease onset and meningeal invasion
Population genomics of Group B Streptococcus reveals the genetics of neonatal disease onset and meningeal invasion
Group B Streptococcus (GBS) causes neonatal disease and mortality worldwide. Here, the authors use genome-wide association analyses to identify bacterial genetic signatures associated with disease onset time and meningeal tissue infection in acute invasive neonatal GBS disease.
- Chrispin Chaguza
- , Dorota Jamrozy
- & Stephen D. Bentley
-
Article
| Open Access Vibrio cholerae O139 genomes provide a clue to why it may have failed to usher in the eighth cholera pandemic
Vibrio cholerae O139 genomes provide a clue to why it may have failed to usher in the eighth cholera pandemic
The O139 Vibrio cholerae serogroup emerged in the 1990s and spread rapidly but did not become globally dominant. Here, the authors describe the genomic epidemiology of this strain and identify changes in virulence and antimicrobial resistance characteristics that they hypothesise may have contributed to its decline.
- Thandavarayan Ramamurthy
- , Agila Kumari Pragasam
- & Ankur Mutreja
-
Article
| Open Access A comprehensive resource for Bordetella genomic epidemiology and biodiversity studies
A comprehensive resource for Bordetella genomic epidemiology and biodiversity studies
The genus Bordetella includes environmental bacteria as well as human pathogens. Here, the authors present a large database of environmental and clinical Bordetella isolates and genome sequences, and develop genotyping systems to facilitate evolutionary and epidemiological studies.
- Sébastien Bridel
- , Valérie Bouchez
- & Sylvain Brisse
-
Article
| Open Access Genomic dissection of Klebsiella pneumoniae infections in hospital patients reveals insights into an opportunistic pathogen
Genomic dissection of Klebsiella pneumoniae infections in hospital patients reveals insights into an opportunistic pathogen
Klebsiella pneumoniae is an opportunistic pathogen of increasing public health concern due to the prevalence of antimicrobial resistance. Here, the authors provide insight into the resistance profiles, bacterial genome features and virulence genes, in a year-long prospective study of K. pneumoniae clinical isolates.
- Claire L. Gorrie
- , Mirjana Mirčeta
- & Kathryn E. Holt
-
Matters Arising
| Open Access Reply to: Testing the adaptive hypothesis of lagging-strand encoding in bacterial genomes
Reply to: Testing the adaptive hypothesis of lagging-strand encoding in bacterial genomes
- Houra Merrikh
- & Christopher Merrikh
-
Article
| Open Access Genomic diversity across the Rickettsia and ‘Candidatus Megaira’ genera and proposal of genus status for the Torix group
Genomic diversity across the Rickettsia and ‘Candidatus Megaira’ genera and proposal of genus status for the Torix group
The bacterial genus Rickettsia includes vector-borne pathogens and arthropod symbionts that are close relatives of symbionts of microeukaryotes classified under the genus ‘Candidatus Megaira’. Here, Davison et al. clarify the evolutionary relationships between these organisms by assembling 28 genomes of understudied species, and propose that a distinct clade known as Torix Rickettsia should be considered a separate genus.
- Helen R. Davison
- , Jack Pilgrim
- & Stefanos Siozios
-
Article
| Open Access The core and accessory Hfq interactomes across Pseudomonas aeruginosa lineages
The core and accessory Hfq interactomes across Pseudomonas aeruginosa lineages
Protein Hfq regulates bacterial mRNA translation and stability by interacting with mRNAs and small noncoding RNAs. Here, the authors identify Hfq-interacting RNAs in three strains representing major phylogenetic lineages of Pseudomonas aeruginosa, highlighting intra-species diversity in post-transcriptional regulatory networks.
- Julian Trouillon
- , Kook Han
- & Stephen Lory

 Freshwater genome-reduced bacteria exhibit pervasive episodes of adaptive stasis
Freshwater genome-reduced bacteria exhibit pervasive episodes of adaptive stasis