Comment
|
Open Access
Featured
-
-
Article
| Open Access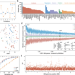 Genetically encoded transcriptional plasticity underlies stress adaptation in Mycobacterium tuberculosis
Genetically encoded transcriptional plasticity underlies stress adaptation in Mycobacterium tuberculosis
Transcriptional plasticity (TP) governs gene expression variability, yet remains unexplored in prokaryotes. This study examines Mycobacterium tuberculosis genes’ TP via RNA-seq meta-analysis, uncovering genetic and functional traits impacting mycobacterial TP.
- Cheng Bei
- , Junhao Zhu
- & Qingyun Liu
-
Article
| Open Access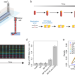 Real-time monitoring of replication errors’ fate reveals the origin and dynamics of spontaneous mutations
Real-time monitoring of replication errors’ fate reveals the origin and dynamics of spontaneous mutations
An interdisciplinary approach following replication errors in Escherichia coli unveils that many spontaneous mutations originate from inefficient repair, and that repair capacity is variable between single cells within a bacterial population.
- Chiara Enrico Bena
- , Jean Ollion
- & Marina Elez
-
Article
| Open Access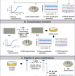 Bacteria can compensate the fitness costs of amplified resistance genes via a bypass mechanism
Bacteria can compensate the fitness costs of amplified resistance genes via a bypass mechanism
Antibiotic heteroresistance, in which a susceptible bacterial population includes a small resistant subpopulation, can arise by tandem amplification of resistance genes, which often carry fitness costs. Here, Pal and Andersson show that these fitness costs can be ameliorated by the acquisition of compensatory mutations and a reduction in copy number of the resistance genes.
- Ankita Pal
- & Dan I. Andersson
-
Article
| Open Access Inter-species gene flow drives ongoing evolution of Streptococcus pyogenes and Streptococcus dysgalactiae subsp. equisimilis
Inter-species gene flow drives ongoing evolution of Streptococcus pyogenes and Streptococcus dysgalactiae subsp. equisimilis
Streptococcus dysgalactiae subsp. equisimilis (SDSE) is an emerging cause of human infection closely related to Streptococcus pyogenes. Here the authors investigate the degree of genomic similarity between the two species and assess implications for development of vaccines.
- Ouli Xie
- , Jacqueline M. Morris
- & Mark R. Davies
-
Article
| Open Access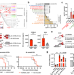 Dependency on host vitamin B12 has shaped Mycobacterium tuberculosis Complex evolution
Dependency on host vitamin B12 has shaped Mycobacterium tuberculosis Complex evolution
Campos-Pardos et al show that the virulence of Mycobacterium tuberculosis is dependent on sufficient uptake of exogenous vitamin B12 from host serum and this phenotype is not conserved in environmental, opportunistic and ancestral lineages.
- Elena Campos-Pardos
- , Santiago Uranga
- & Jesús Gonzalo-Asensio
-
Article
| Open Access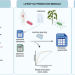 bacLIFE: a user-friendly computational workflow for genome analysis and prediction of lifestyle-associated genes in bacteria
bacLIFE: a user-friendly computational workflow for genome analysis and prediction of lifestyle-associated genes in bacteria
Many bacteria live in close association with eukaryotic hosts, exhibiting detrimental, neutral or beneficial effects on host growth and health. Here, the authors present a streamlined computational workflow for bacterial genome annotation, large-scale comparative genomics, and prediction of genes potentially involved in niche adaptation.
- Guillermo Guerrero-Egido
- , Adrian Pintado
- & Víctor J. Carrión
-
Article
| Open Access ProTInSeq: transposon insertion tracking by ultra-deep DNA sequencing to identify translated large and small ORFs
ProTInSeq: transposon insertion tracking by ultra-deep DNA sequencing to identify translated large and small ORFs
Identifying small proteins is challenging. ProTInSeq uses modified transposons to express markers inserted in-frame to protein-coding genes. This method identifies 153 unannotated small proteins in M. pneumoniae and additional proteomic information.
- Samuel Miravet-Verde
- , Rocco Mazzolini
- & Luis Serrano
-
Article
| Open Access Transmission and dynamics of mother-infant gut viruses during pregnancy and early life
Transmission and dynamics of mother-infant gut viruses during pregnancy and early life
Gut ecosystem colonization impacts lifelong health. Here, authors track mother-infant gut viruses over time, reveal feeding’s influence on early viral colonization, and demonstrate the co-transmission of bacteriophages and bacteria from mothers to infants.
- Sanzhima Garmaeva
- , Trishla Sinha
- & Alexandra Zhernakova
-
Article
| Open Access The Deinococcus protease PprI senses DNA damage by directly interacting with single-stranded DNA
The Deinococcus protease PprI senses DNA damage by directly interacting with single-stranded DNA
Lu et al. show that single-stranded DNA produced as a result of DNA damage may directly activate PprI in Deinococcus species, triggering the DNA damage response.
- Huizhi Lu
- , Zijing Chen
- & Yuejin Hua
-
Article
| Open Access TcrXY is an acid-sensing two-component transcriptional regulator of Mycobacterium tuberculosis required for persistent infection
TcrXY is an acid-sensing two-component transcriptional regulator of Mycobacterium tuberculosis required for persistent infection
Stupar et al. describe a new role for the Mycobacterium tuberculosis two-component system, TcrXY, in the modulation of up to 70 genes, including two effectors, TarA and TarB which mitigate intracellular redox stress.
- Miljan Stupar
- , Lendl Tan
- & Nicholas P. West
-
Article
| Open Access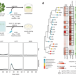 Commensal lifestyle regulated by a negative feedback loop between Arabidopsis ROS and the bacterial T2SS
Commensal lifestyle regulated by a negative feedback loop between Arabidopsis ROS and the bacterial T2SS
The plant immune output reactive oxygen species tames a detrimental bacterial commensal from native microbiota by suppressing a bacterial secretion system, allowing the co-existence and turning it into a beneficial bacterium to the host.
- Frederickson Entila
- , Xiaowei Han
- & Kenichi Tsuda
-
Article
| Open Access CRISPR-Cas-based identification of a sialylated human milk oligosaccharides utilization cluster in the infant gut commensal Bacteroides dorei
CRISPR-Cas-based identification of a sialylated human milk oligosaccharides utilization cluster in the infant gut commensal Bacteroides dorei
Human milk oligosaccharides (HMOs) utilization by Bacteroides species remains poorly understood. Here, the authors describe a single specific gene cluster responsible for sialylated HMOs utilization in a B. dorei natural isolate and prove its functionality in vivo using CRISPR-Cas12a.
- Sivan Kijner
- , Dena Ennis
- & Moran Yassour
-
Article
| Open Access The Helicobacter pylori Genome Project: insights into H. pylori population structure from analysis of a worldwide collection of complete genomes
The Helicobacter pylori Genome Project: insights into H. pylori population structure from analysis of a worldwide collection of complete genomes
The bacterium Helicobacter pylori, often found in the human stomach, can be classified into distinct subpopulations associated with the geographic origin of the host. Here, the authors provide insights into H. pylori population structure by collecting over 1,000 clinical strains from 50 countries and generating and analyzing high-quality bacterial genome sequences.
- Kaisa Thorell
- , Zilia Y. Muñoz-Ramírez
- & Charles S. Rabkin
-
Article
| Open Access TRS: a method for determining transcript termini from RNAtag-seq sequencing data
TRS: a method for determining transcript termini from RNAtag-seq sequencing data
TRS is a new method for determining 3’ transcript termini in bacteria, using data generated by the RNAtag-seq protocol. This methodology opens the door to study the evolution of transcription termini and their condition-dependent dynamics.
- Amir Bar
- , Liron Argaman
- & Hanah Margalit
-
Article
| Open Access The genomic epidemiology of shigellosis in South Africa
The genomic epidemiology of shigellosis in South Africa
As a leading cause of diarrhoeal mortality and morbidity, authors examine the epidemiology and genome dynamics of shigellosis in South Africa, utilising whole genome sequence analysis.
- George E. Stenhouse
- , Karen H. Keddy
- & Kate S. Baker
-
Article
| Open Access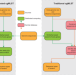 Distributed genotyping and clustering of Neisseria strains reveal continual emergence of epidemic meningococcus over a century
Distributed genotyping and clustering of Neisseria strains reveal continual emergence of epidemic meningococcus over a century
Core genome multilocus sequence typing (cgMLST) is used to classify bacterial strains for epidemiological applications. Here, the authors describe a distributed cgMLST scheme that does not require a central database of allelic sequences, and apply it to study evolutionary patterns of epidemic and endemic strains of the genus Neisseria.
- Ling Zhong
- , Menghan Zhang
- & Zhemin Zhou
-
Article
| Open Access Global pathogenomic analysis identifies known and candidate genetic antimicrobial resistance determinants in twelve species
Global pathogenomic analysis identifies known and candidate genetic antimicrobial resistance determinants in twelve species
A global analysis of antimicrobial resistance (AMR) across 27,155 genomes and 69 drugs reveals patterns in AMR gene transfer between species and identifies 142 AMR gene candidates, two of which were tested and confirmed as contributing to AMR.
- Jason C. Hyun
- , Jonathan M. Monk
- & Bernhard O. Palsson
-
Article
| Open Access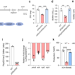 Regulation of the physiology and virulence of Ralstonia solanacearum by the second messenger 2′,3′-cyclic guanosine monophosphate
Regulation of the physiology and virulence of Ralstonia solanacearum by the second messenger 2′,3′-cyclic guanosine monophosphate
Nucleotide second messengers are employed by many bacterial species to regulate various cellular processes. Here, the authors demonstrate that 2',3'-cyclic guanosine monophosphate (2',3'-cGMP) controls the important biological functions and virulence in Ralstonia solanacearum by abolishing the interaction between a transcriptional regulator and the promoters of target genes.
- Xia Li
- , Wenfang Yin
- & Yinyue Deng
-
Article
| Open Access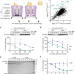 The heat shock protein LarA activates the Lon protease in response to proteotoxic stress
The heat shock protein LarA activates the Lon protease in response to proteotoxic stress
The Lon protease is an important protein degradation machine and is conserved across the three domains of life. Here, the authors describe a small proteotoxic stress-induced protein that functions as an allosteric activator of Lon.
- Deike J. Omnus
- , Matthias J. Fink
- & Kristina Jonas
-
Article
| Open Access High-resolution temporal profiling of E. coli transcriptional response
High-resolution temporal profiling of E. coli transcriptional response
Understanding how cells dynamically adapt to their environment is important, but temporal information about cellular behaviour is often limited. Here, Miano et al. apply unsupervised machine learning to a dataset describing the activity of over 1,800 promoters in E. coli, measured every 10 minutes, defining three primary stages of promoter activation in response to heavy metal stress.
- Arianna Miano
- , Kevin Rychel
- & Jeff Hasty
-
Article
| Open Access A smooth tubercle bacillus from Ethiopia phylogenetically close to the Mycobacterium tuberculosis complex
A smooth tubercle bacillus from Ethiopia phylogenetically close to the Mycobacterium tuberculosis complex
The Mycobacterium tuberculosis complex (MTBC) includes several pathogens thought to have originated in East Africa from an ancestor closely related to Mycobacterium canettii. Here, the authors describe a clinical tuberculosis strain isolated in Ethiopia that has typical M. canettii features but is phylogenetically much closer to the MTBC clade, supporting that the emergence of MTBC pathogens is a recent evolutionary event.
- Bazezew Yenew
- , Arash Ghodousi
- & Daniela Maria Cirillo
-
Article
| Open Access The metabolic, virulence and antimicrobial resistance profiles of colonising Streptococcus pneumoniae shift after PCV13 introduction in urban Malawi
The metabolic, virulence and antimicrobial resistance profiles of colonising Streptococcus pneumoniae shift after PCV13 introduction in urban Malawi
Pneumococcal vaccination has been shown to promote emergence of non-vaccine S. pneumoniae serotypes. Here, the authors use data from Malawi to investigate whether vaccine introduction also results in changes in metabolic, virulence, and antimicrobial resistance profiles of circulating strains.
- Uri Obolski
- , Todd D. Swarthout
- & Robert S. Heyderman
-
Article
| Open Access Mutational spectra are associated with bacterial niche
Mutational spectra are associated with bacterial niche
Mutagens and DNA repair defects generate context-specific mutational signatures in cancer cells. Here, Ruis et al. provide evidence of similar processes in bacteria, showing that mutational spectra may be associated with sites of bacterial replication when mutagen exposures differ, and can be used in these cases to infer transmission routes.
- Christopher Ruis
- , Aaron Weimann
- & Julian Parkhill
-
Article
| Open Access Inferring bacterial transmission dynamics using deep sequencing genomic surveillance data
Inferring bacterial transmission dynamics using deep sequencing genomic surveillance data
Studying rare genetic changes that arose as an infectious bacterium spread between lab mice, here the authors show that using the relative abundance of any changes rather than just whether they occurred can more precisely identify who likely infected who.
- Madikay Senghore
- , Hannah Read
- & Siouxsie Wiles
-
Article
| Open Access A genomic appraisal of invasive Salmonella Typhimurium and associated antibiotic resistance in sub-Saharan Africa
A genomic appraisal of invasive Salmonella Typhimurium and associated antibiotic resistance in sub-Saharan Africa
Invasive Salmonella Typhimurium bloodstream infection causes a significant public health burden in sub-Saharan Africa. Here, the authors analyse whole genome sequences of 1,302 S. Typhimurium isolates from Africa and describe its evolution, geographic spread, and antimicrobial resistance characteristics.
- Sandra Van Puyvelde
- , Tessa de Block
- & Octavie Lunguya
-
Article
| Open Access The ClpX protease is essential for inactivating the CI master repressor and completing prophage induction in Staphylococcus aureus
The ClpX protease is essential for inactivating the CI master repressor and completing prophage induction in Staphylococcus aureus
Prophage induction is a fundamental process in the phage life cycle. Here the authors show that the final stage of prophage induction in Staphylococcus aureus is controlled by the ClpX protease, unveiling and unexpected role for ClpX in phage biology.
- Mohammed A. Thabet
- , José R. Penadés
- & Andreas F. Haag
-
Article
| Open Access Mechanism of outer membrane destabilization by global reduction of protein content
Mechanism of outer membrane destabilization by global reduction of protein content
The outer membrane (OM) of Gram-negative bacteria is an asymmetric bilayer, with phospholipids in the inner leaflet. Here the authors show that a reduction in OM proteins and the subsequent mislocalization of phospholipids weaken the OM and alter growth rate and cell shape, emphasizing the role of OM proteins in OM stiffness and cell shape.
- Irina V. Mikheyeva
- , Jiawei Sun
- & Thomas J. Silhavy
-
Article
| Open Access A linear and circular dual-conformation noncoding RNA involved in oxidative stress tolerance in Bacillus altitudinis
A linear and circular dual-conformation noncoding RNA involved in oxidative stress tolerance in Bacillus altitudinis
The presence and/or biological functionality of circular RNAs in bacteria are unclear. Here, the authors identify a dual-conformation (linear and circular) noncoding RNA that promotes tolerance to oxidative stress in Bacillus altitutidinis, and provide evidence for the existence of other circular RNAs in diverse bacterial species.
- Ting-Ting He
- , Yun-Fan Xu
- & Hai-Yan Wang
-
Article
| Open Access Ancient Clostridium DNA and variants of tetanus neurotoxins associated with human archaeological remains
Ancient Clostridium DNA and variants of tetanus neurotoxins associated with human archaeological remains
The analysis of microbial genomes from human archaeological samples offers a snapshot of ancient pathogens. Here, Hodgins et al. analyze metagenomic datasets from 38 human archaeological samples and identify bacterial genomic sequences related to modern-day Clostridium tetani, encoding tetanus neurotoxins.
- Harold P. Hodgins
- , Pengsheng Chen
- & Andrew C. Doxey
-
Article
| Open Access Genomic dissection of endemic carbapenem resistance reveals metallo-beta-lactamase dissemination through clonal, plasmid and integron transfer
Genomic dissection of endemic carbapenem resistance reveals metallo-beta-lactamase dissemination through clonal, plasmid and integron transfer
Resistance to carbapenems, a class of last-line antibiotics, is a global health threat. This study analysed a two-decade history of carbapenem resistance and identified complex, multi-level (bacterial strain, plasmid, gene) transmission dynamics.
- Nenad Macesic
- , Jane Hawkey
- & Anton Y. Peleg
-
Article
| Open Access An RNA modification enzyme directly senses reactive oxygen species for translational regulation in Enterococcus faecalis
An RNA modification enzyme directly senses reactive oxygen species for translational regulation in Enterococcus faecalis
Here the authors found an RNA modifying enzyme that serves as a molecular switch, directly relaying the sensing of reactive oxygen species to chemical modifications in rRNA and tRNA for increasing stress response proteins in the proteome.
- Wei Lin Lee
- , Ameya Sinha
- & Peter Dedon
-
Article
| Open Access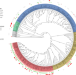 Epistatic interactions between the high pathogenicity island and other iron uptake systems shape Escherichia coli extra-intestinal virulence
Epistatic interactions between the high pathogenicity island and other iron uptake systems shape Escherichia coli extra-intestinal virulence
The virulence of extra-intestinal pathogenic Escherichia coli is associated with multiple different genes in different lineages. Here, Royer et al. show that the emergence of virulence is associated with acquisition of the siderophore-encoding high-pathogenicity island (HPI), and full virulence is associated with the additional presence of the aer or sit operons.
- Guilhem Royer
- , Olivier Clermont
- & Erick Denamur
-
Article
| Open Access Culturing of a complex gut microbial community in mucin-hydrogel carriers reveals strain- and gene-associated spatial organization
Culturing of a complex gut microbial community in mucin-hydrogel carriers reveals strain- and gene-associated spatial organization
The organization of gut microbes in lumen and mucosa and the microbial genes regulating this organization remain poorly understood. Here, using in vitro cultures incorporating a complex gut microbial community in mucin-hydrogel carriers, the authors show greater richness and strain-specific spatial organization, enabling discovery of associated genes.
- Xiaofan Jin
- , Feiqiao B. Yu
- & Katherine S. Pollard
-
Article
| Open Access A global genomic analysis of Salmonella Concord reveals lineages with high antimicrobial resistance in Ethiopia
A global genomic analysis of Salmonella Concord reveals lineages with high antimicrobial resistance in Ethiopia
Authors carry out a longitudinal genomic analysis of Salmonella enterica serovar Concord isolates from various geographical locations, to reconstruct population diversity, evolution and antimicrobial resistance distribution.
- Wim L. Cuypers
- , Pieter Meysman
- & Sandra Van Puyvelde
-
Article
| Open Access The divisome but not the elongasome organizes capsule synthesis in Streptococcus pneumoniae
The divisome but not the elongasome organizes capsule synthesis in Streptococcus pneumoniae
The bacterial cell envelope consists of multiple layers, the synthesis of which is coordinated through unclear mechanisms. Here, Nakamoto et al. reveal a mechanism linking the synthesis of capsular polysaccharides and cell wall peptidoglycan in pneumococci.
- Rei Nakamoto
- , Sarp Bamyaci
- & Lok-To Sham
-
Article
| Open Access Associate toxin-antitoxin with CRISPR-Cas to kill multidrug-resistant pathogens
Associate toxin-antitoxin with CRISPR-Cas to kill multidrug-resistant pathogens
CRISPR-regulated toxin-antitoxin (CreTA), safeguards CRISPR-Cas immune systems. Here the authors characterize a bacterial CreTA and use this to generate a proof-of-concept antimicrobial strategy, ATTACK, which associates TA and CRISPR-Cas to kill multidrug resistant pathogens.
- Rui Wang
- , Xian Shu
- & Ming Li
-
Article
| Open Access The genomic landscape of reference genomes of cultivated human gut bacteria
The genomic landscape of reference genomes of cultivated human gut bacteria
Here, the authors present an expanded version of the Cultivated Genome Reference (CGR), termed CGR2, a catalog that includes 3324 high-quality draft genomes based on gut bacterial isolates from Chinese individuals, and classifies 527 species from 8 phyla, including 179 previously unidentified species, and provides information of secondary metabolite biosynthetic gene clusters and gut phage-bacteria interactions.
- Xiaoqian Lin
- , Tongyuan Hu
- & Yuanqiang Zou
-
Article
| Open Access The evolution of antibiotic resistance is associated with collateral drug phenotypes in Mycobacterium tuberculosis
The evolution of antibiotic resistance is associated with collateral drug phenotypes in Mycobacterium tuberculosis
Here using drug susceptibility profiling, genomics and evolutionary studies the authors provide strategies to exploit collateral drug responses in Mycobacterium tuberculosis to prevent the emergence of drug resistance.
- Natalie J. E. Waller
- , Chen-Yi Cheung
- & Matthew B. McNeil
-
Article
| Open Access Genomic attributes of Vibrio cholerae O1 responsible for 2022 massive cholera outbreak in Bangladesh
Genomic attributes of Vibrio cholerae O1 responsible for 2022 massive cholera outbreak in Bangladesh
Vibrio cholerae has undergone continuous evolution, and differing strains have caused numerous outbreaks. Here, the authors present a genomic study of Vibrio cholerae O1 responsible for a 2022 outbreak in Dhaka.
- Md Mamun Monir
- , Mohammad Tarequl Islam
- & Munirul Alam
-
Article
| Open Access Host-microbe co-metabolism via MCAD generates circulating metabolites including hippuric acid
Host-microbe co-metabolism via MCAD generates circulating metabolites including hippuric acid
Here, using a mouse model, the authors report a previously undescribed role for medium-chain acyl-CoA dehydrogenase in host metabolism of gut microbiota metabolites, and show that circulating compounds, including the abundant organic acid hippurate, depend on host-microbe co-metabolism of phenylalanine by Clostridium sporogenes.
- Kali M. Pruss
- , Haoqing Chen
- & Dylan Dodd
-
Article
| Open Access Real-time visualisation of the intracellular dynamics of conjugative plasmid transfer
Real-time visualisation of the intracellular dynamics of conjugative plasmid transfer
Conjugation is a contact-dependent mechanism for the transfer of plasmid DNA between bacterial cells. Here, Couturier et al. use live-cell microscopy to visualise the intracellular dynamics of conjugation in real time, revealing a molecular strategy that allows the sequential production of factors involved in establishing, maintaining and disseminating the plasmid.
- Agathe Couturier
- , Chloé Virolle
- & Christian Lesterlin
-
Article
| Open Access Harnessing gut microbes for glycan detection and quantification
Harnessing gut microbes for glycan detection and quantification
Detecting distinct glycans within heterogeneous mixtures is hindered by glycan structural complexity and diversity. Here the authors exploit the ability of gut microbes to sense different glycan structures in order to develop quantitative glycan biosensors by coupling bacterial detection machinery to an optimised luciferase reporter.
- Jennifer L. Modesto
- , Victoria H. Pearce
- & Guy E. Townsend II
-
Article
| Open Access Quantifying the local adaptive landscape of a nascent bacterial community
Quantifying the local adaptive landscape of a nascent bacterial community
Fitness landscapes largely shape the dynamics of evolution, but it is unclear how they shift upon ecological diversification. By engineering genome-wide knockout libraries of a nascent bacterial community, Ascensao et al. show how ecological and epistatic patterns combine to shape adaptive landscapes.
- Joao A. Ascensao
- , Kelly M. Wetmore
- & Oskar Hallatschek
-
Article
| Open Access Resolving colistin resistance and heteroresistance in Enterobacter species
Resolving colistin resistance and heteroresistance in Enterobacter species
Taxonomical complexity has muddled the classification of clinically relevant Enterobacter species. Authors carry out a genome-based study on clinical isolates to investigate colistin resistance and heteroresistance in Enterobacter.
- Swapnil Prakash Doijad
- , Nicolas Gisch
- & Trinad Chakraborty
-
Article
| Open Access An ISO-certified genomics workflow for identification and surveillance of antimicrobial resistance
An ISO-certified genomics workflow for identification and surveillance of antimicrobial resistance
The implementation of genomics for identification and surveillance of antimicrobial resistance (AMR) in clinical laboratories remains challenging. Here, Sherry et al. present a bioinformatics platform for detection of AMR determinants from whole-genome sequencing data, suitable for clinical and public-health microbiology reporting.
- Norelle L. Sherry
- , Kristy A. Horan
- & Torsten Seemann
-
Article
| Open Access Multistep diversification in spatiotemporal bacterial-phage coevolution
Multistep diversification in spatiotemporal bacterial-phage coevolution
Bacteria and their viruses coexist and coevolve in nature, but maintaining them together in the lab is challenging. Here, a spatially structured environment allowed prolonged coevolution, with bacteria and phage diversifying into multiple ecotypes, uncovering gene mechanisms affecting phage-bacteria interactions.
- Einat Shaer Tamar
- & Roy Kishony
-
Article
| Open Access Structure-function studies reveal ComEA contains an oligomerization domain essential for transformation in gram-positive bacteria
Structure-function studies reveal ComEA contains an oligomerization domain essential for transformation in gram-positive bacteria
ComEA is a DNA-binding protein required for DNA uptake during bacterial transformation. Here, Ahmed et al. determine X-ray crystal structures of ComEA from Gram-positive bacteria, identifying a domain that is absent in Gram-negative bacteria and drives ComEA oligomerization, which is required for transformation.
- Ishtiyaq Ahmed
- , Jeanette Hahn
- & Matthew B. Neiditch
-
Article
| Open Access Dynamics of extended-spectrum cephalosporin resistance genes in Escherichia coli from Europe and North America
Dynamics of extended-spectrum cephalosporin resistance genes in Escherichia coli from Europe and North America
Extended-spectrum cephalosporin resistance genes in Escherichia coli have spread worldwide. Here, the authors dissect the emergence and distribution of these genes over time, and across geographic location and host species, to better understand their dynamics and mechanisms of transmission.
- Roxana Zamudio
- , Patrick Boerlin
- & Alison E. Mather
-
Article
| Open Access Genetic manipulation of the human gut bacterium Eggerthella lenta reveals a widespread family of transcriptional regulators
Genetic manipulation of the human gut bacterium Eggerthella lenta reveals a widespread family of transcriptional regulators
Eggerthella lenta is a prominent human gut bacterium implicated in several physiological processes, but its study has remained limited. Here, by developing a genetic toolbox for E. lenta, the authors provide insights into how the bacterium regulates drug and dietary compound metabolism.
- Xueyang Dong
- , Ben G. H. Guthrie
- & Emily P. Balskus

 All-inclusive nitrifiers in Antarctic soils
All-inclusive nitrifiers in Antarctic soils