« Prev Next »
This structure has novel features which are of considerable biological interest . . . It has not escaped our notice that the specific pairing we have postulated immediately suggests a possible copying mechanism for the genetic material.
—Watson & Crick (1953)
Perhaps the most significant aspect of Watson and Crick's discovery of DNA structure was not that it provided scientists with a three-dimensional model of this molecule, but rather that this structure seemed to reveal the way in which DNA was replicated. As noted in their 1953 paper, Watson and Crick strongly suspected that the specific base pairings within the DNA double helix existed in order to ensure a controlled system of DNA replication. However, it took several years of subsequent study, including a classic 1958 experiment by American geneticists Matthew Meselson and Franklin Stahl, before the exact relationship between DNA structure and replication was understood.
Three Proposed Models for DNA Replication
Replication is the process by which a cell copies its DNA prior to division. In humans, for example, each parent cell must copy its entire six billion base pairs of DNA before undergoing mitosis. The molecular details of DNA replication are described elsewhere, and they were not known until some time after Watson and Crick's discovery. In fact, before such details could be determined, scientists were faced with a more fundamental research concern. Specifically, they wanted to know the overall nature of the process by which DNA replication occurs.
Defining the Models
Semiconservative replication was not the only model of DNA replication proposed during the mid-1950s, however. In fact, two other prominent hypotheses were put also forth: conservative replication and dispersive replication. According to the conservative replication model, the entire original DNA double helix serves as a template for a new double helix, such that each round of cell division produces one daughter cell with a completely new DNA double helix and another daughter cell with a completely intact old (or original) DNA double helix. On the other hand, in the dispersive replication model, the original DNA double helix breaks apart into fragments, and each fragment then serves as a template for a new DNA fragment. As a result, every cell division produces two cells with varying amounts of old and new DNA (Figure 1).
Making Predictions Based on the Models
When these three models were first proposed, scientists had few clues about what might be occurring at the molecular level during DNA replication. Fortunately, the models yielded different predictions about the distribution of old versus new DNA in newly divided cells, no matter what the underlying molecular mechanisms. These predictions were as follows:
- According to the semiconservative model, after one round of replication, every new DNA double helix would be a hybrid that consisted of one strand of old DNA bound to one strand of newly synthesized DNA. Then, during the second round of replication, the hybrids would separate, and each strand would pair with a newly synthesized strand. Afterward, only half of the new DNA double helices would be hybrids; the other half would be completely new. Every subsequent round of replication therefore would result in fewer hybrids and more completely new double helices.
- According to the conservative model, after one round of replication, half of the new DNA double helices would be composed of completely old, or original, DNA, and the other half would be completely new. Then, during the second round of replication, each double helix would be copied in its entirety. Afterward, one-quarter of the double helices would be completely old, and three-quarters would be completely new. Thus, each subsequent round of replication would result in a greater proportion of completely new DNA double helices, while the number of completely original DNA double helices would remain constant.
- According to the dispersive model, every round of replication would result in hybrids, or DNA double helices that are part original DNA and part new DNA. Each subsequent round of replication would then produce double helices with greater amounts of new DNA.
Meselson and Stahl’s Elegant Experiment
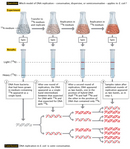
The duo thus began their experiment by choosing two isotopes of nitrogen—the common and lighter 14N, and the rare and heavier 15N (so-called "heavy" nitrogen)—as their labels and a technique known as cesium chloride (CsCl) equilibrium density gradient centrifugation as their sedimentation method. Meselson and Stahl opted for nitrogen because it is an essential chemical component of DNA; therefore, every time a cell divides and its DNA replicates, it incorporates new N atoms into the DNA of either one or both of its two daughter cells, depending on which model was correct. "If several different density species of DNA are present," they predicted, "each will form a band at the position where the density of the CsCl solution is equal to the buoyant density of that species. In this way, DNA labeled with heavy nitrogen (15N) may be resolved from unlabeled DNA" (Meselson & Stahl, 1958).
The scientists then continued their experiment by growing a culture of E. coli bacteria in a medium that had the heavier 15N (in the form of 15N-labeled ammonium chloride) as its only source of nitrogen. In fact, they did this for 14 bacterial generations, which was long enough to create a population of bacterial cells that contained only the heavier isotope (all the original 14N-containing cells had died by then). Next, they changed the medium to one containing only 14N-labeled ammonium salts as the sole nitrogen source. So, from that point onward, every new strand of DNA would be built with 14N rather than 15N.
Just prior to the addition of 14N and periodically thereafter, as the bacterial cells grew and replicated, Meselson and Stahl sampled DNA for use in equilibrium density gradient centrifugation to determine how much 15N (from the original or old DNA) versus 14N (from the new DNA) was present. For the centrifugation procedure, they mixed the DNA samples with a solution of cesium chloride and then centrifuged the samples for enough time to allow the heavier 15N and lighter 14N DNA to migrate to different positions in the centrifuge tube.
By way of centrifugation, the scientists found that DNA composed entirely of 15N-labeled DNA (i.e., DNA collected just prior to changing the culture from one containing only 15N to one containing only 14N) formed a single distinct band, because both of its strands were made entirely in the "heavy" nitrogen medium. Following a single round of replication, the DNA again formed a single distinct band, but the band was located in a different position along the centrifugation gradient. Specifically, it was found midway between where all the 15N and all the 14N DNA would have migrated—in other words, halfway between "heavy" and "light" (Figure 2). Based on these findings, the scientists were immediately able to exclude the conservative model of replication as a possibility. After all, if DNA replicated conservatively, there should have been two distinct bands after a single round of replication; half of the new DNA would have migrated to the same position as it did before the culture was transferred to the 14N-containing medium (i.e., to the "heavy" position), and only the other half would have migrated to the new position (i.e., to the "light" position). That left the scientists with only two options: either DNA replicated semiconservatively, as Watson and Crick had predicted, or it replicated dispersively.
To differentiate between the two, Meselson and Stahl had to let the cells divide again and then sample the DNA after a second round of replication. After that second round of replication, the scientists found that the DNA separated into two distinct bands: one in a position where DNA containing only 14N would be expected to migrate, and the other in a position where hybrid DNA (containing half 14N and half 15N) would be expected to migrate. The scientists continued to observe the same two bands after several subsequent rounds of replication. These results were consistent with the semiconservative model of replication and the reality that, when DNA replicated, each new double helix was built with one old strand and one new strand. If the dispersive model were the correct model, the scientists would have continued to observe only a single band after every round of replication.
Straight or Circular?
Following publication of Meselson and Stahl's results, many scientists confirmed that semiconservative replication was the rule, not just in E. coli, but in every other species studied as well. To date, no one has found any evidence for either conservative or dispersive DNA replication. Scientists have found, however, that semiconservative replication can occur in different ways—for example, it may proceed in either a circular or a linear fashion, depending on chromosome shape.
In fact, in the early 1960s, English molecular biologist John Cairns performed another remarkably elegant experiment to demonstrate that E. coli and other bacteria with circular chromosomes undergo what he termed "theta replication," because the structure generated resembles the Greek letter theta (Θ). Specifically, Cairns grew E. coli bacteria in the presence of radioactive nucleotides such that, after replication, each new DNA molecule had one radioactive (hot) strand and one nonradioactive strand. He then isolated the newly replicated DNA and used it to produce an electron micrograph image of the Θ-shaped replication process (Figure 3; Cairns, 1961).
But how does theta replication work? It turns out that this process results from the original double-stranded DNA unwinding at a single spot on the chromosome known as the replication origin. As the double helix unwinds, it creates a loop known as the replication bubble, with each newly separated single strand serving as a template for DNA synthesis. Replication occurs as the double helix unwinds.
Eukaryotes undergo linear, not circular, replication. As with theta replication, as the double helix unwinds, each newly separated single strand serves as a template for DNA synthesis. However, unlike bacterial replication, because eukaryotic cells carry vastly more DNA than bacteria do (for example, the common house [and laboratory] mouse Mus musculus has about three billion base pairs of DNA, compared to a bacterial cell's one to four million base pairs), eukaryotic chromosomes have multiple replication origins, with multiple replication bubbles forming. For example, M. musculus has as many as 25,000 replication origins, whereas the smaller-genomed fruit fly (Drosophila melanogaster), with its approximately 120 million base pairs of DNA, has only about 3,500 replication origins.
Thus, the discovery of the structure of DNA in 1953 was only the beginning. When Watson and Crick postulated that form predicts function, they provided the scientific community with a challenge to determine exactly how DNA functioned in the cell, including how this molecule was replicated. The work of Meselson and Stahl demonstrates how elegant experiments can distinguish between different hypotheses. Understanding that replication occurs semiconservatively was just the beginning to understanding the key enzymatic events responsible for the physical copying of the genome.
References and Recommended Reading
Cairns, J. The bacterial chromosome and its manner of replication as seen by autoradiography. Journal of Molecular Biology 6, 208–213 (1961)
Meselson, M., & Stahl, F. The replication of DNA in Escherichia coli. Proceedings of the National Academy of Sciences 44, 671–682 (1958)
Watson, J. D., & Crick, F. H. C. A structure for deoxyribose nucleic acid. Nature 171, 737–738 (1953) (link to article).



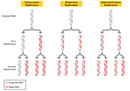
 Figure 1
Figure 1