« Prev Next »
In addition to genetic insults caused by the environment, the very process of DNA replication during cell division is prone to error. The rate at which DNA polymerase adds incorrect nucleotides during DNA replication is a major factor in determining the spontaneous mutation rate in an organism. While a "proofreading" enzyme normally recognizes and corrects many of these errors, some mutations survive this process. Estimates of the frequency at which human DNA undergoes lasting, uncorrected errors range from 1 x 10-4 to 1 x 10-6 mutations per gamete for a given gene. A rate of 1 x 10-6 means that a scientist would expect to find one mutation at a specific locus per one million gametes. Mutation rates in other organisms are often much lower (Table 1).
One way scientists are able to estimate mutation rates is by considering the rate of new dominant mutations found at different loci. For example, by examining the number of individuals in a given population who were diagnosed with neurofibromatosis (NF1, a disease caused by a spontaneous—or noninherited—dominant mutation), scientists determined that the spontaneous mutation rate of the gene responsible for this disease averaged 1 x 10-4 mutations per gamete (Crowe et al., 1956). Other researchers have found that the mutation rates of other genes, like that for Huntington's disease, are significantly lower than the rate for NF1. The fact that investigators have reported different mutation rates for different genes suggests that certain loci are more prone to damage or error than others.
DNA Repair Mechanisms and Human Disease
Defects in DNA repair underlie a number of human genetic diseases that affect a wide variety of body systems but share a constellation of common traits, most notably a predisposition to cancer (Table 2). These disorders include ataxia-telangiectasia (AT), a degenerative motor condition caused by failure to repair oxidative damage in the cerebellum, and xeroderma pigmentosum (XP), a condition characterized by sensitivity to sunlight and linked to a defect in an important ultraviolet (UV) damage repair pathway. In addition, a number of genes that have been implicated in cancer, such as the RAD group, have also been determined to encode proteins critical for DNA damage repair.
UV Damage, Nucleotide Excision Repair, and Photoreactivation
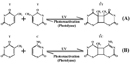
UV radiation causes two classes of DNA lesions: cyclobutane pyrimidine dimers (CPDs, Figure 1) and 6-4 photoproducts (6-4 PPs, Figure 2). Both of these lesions distort DNA's structure, introducing bends or kinks and thereby impeding transcription and replication. Relatively flexible areas of the DNA double helix are most susceptible to damage. In fact, one "hot spot" for UV-induced damage is found within a commonly mutated oncogene, the p53 gene.
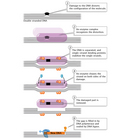
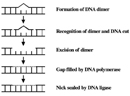
CPDs and 6-4 PPs are both repaired through a process known as nucleotide excision repair (NER). In eukaryotes, this complex process relies on the products of approximately 30 genes. Defects in some of these genes have been shown to cause the human disease XP, as well as other conditions that share a risk of skin cancer that is elevated about a thousandfold over normal. More specifically, eukaryotic NER is carried out by at least 18 protein complexes via four discrete steps (Figure 3): detection of damage; excision of the section of DNA that includes and surrounds the error; filling in of the resulting gap by DNA polymerase; and sealing of the nick between the newly synthesized and older DNA (Figure 4). In bacteria (which are prokaryotes), however, the process of NER is completed by only three proteins, named UvrA, UvrB, and UvrC.
Bacteria and several other organisms also possess another mechanism to repair UV damage called photoreactivation. This method is often referred to as "light repair," because it is dependent on the presence of light energy. (In comparison, NER and most other repair mechanisms are frequently referred to as "dark repair," as they do not require light as an energy source.) During photoreactivation, an enzyme called photolyase binds pyrimidine dimer lesions; in addition, a second molecule known as chromophore converts light energy into the chemical energy required to directly revert the affected area of DNA to its undamaged form. Photolyases are found in numerous organisms, including fungi, plants, invertebrates such as fruit flies, and vertebrates including frogs. They do not appear to exist in humans, however (Sinha & Hader, 2002).
Additional DNA Repair mechanisms
Yet another form of DNA damage is double-strand breaks, which are caused by ionizing radiation, including gamma rays and X-rays. These breaks are highly deleterious. In addition to interfering with transcription or replication, they can lead to chromosomal rearrangements, in which pieces of one chromosome become attached to another chromosome. Genes are disrupted in this process, leading to hybrid proteins or inappropriate activation of genes. A number of cancers are associated with such rearrangements. Double-strand breaks are repaired through one of two mechanisms: nonhomologous end joining (NHEJ) or homologous recombination repair (HRR). In NHEJ, an enzyme called DNA ligase IV uses overhanging pieces of DNA adjacent to the break to join and fill in the ends. Additional errors can be introduced during this process, which is the case if a cell has not completely replicated its DNA in preparation for division. In contrast, during HRR, the homologous chromosome itself is used as a template for repair.
Mutations in an organism's DNA are a part of life. Our genetic code is exposed to a variety of insults that threaten its integrity. But, a rigorous system of checks and balances is in place through the DNA repair machinery. The errors that slip through the cracks may sometimes be associated with disease, but they are also a source of variation that is acted upon by longer-term processes, such as evolution and natural selection.
References and Recommended Reading
Branze, D., & Foiani, M. Regulation of DNA repair throughout the cell cycle.
Nature Reviews Molecular Cell Biology 9, 297–308 (2008) doi:10.1038/nrm2351.pdf (link to article)
Crowe, F. W., et al. A Clinical, Pathological, and Genetic Study of Multiple Neurofibromatosis (Springfield, Illinois, Charles C. Thomas, 1956)
Lodish, H., et al. Molecular Biology of the Cell, 5th ed. (New York, Freeman, 2004)
Sinha, R. P., & Häder, D. P. UV-induced DNA damage and repair: A review. Photochemical and Photobiological Sciences 1, 225–236 (2002)




 Table 1
Table 1