Abstract
To adequately prepare against imminent disease outbreaks from diverse and ever-changing viral pathogens, improved experimental models that can accurately recapitulate host-virus responses and disease pathogenesis in human are essential. Organoid platforms have emerged in recent years as amenable in vitro tools that can bridge the limitations of traditional 2D cell lines and animal models for viral disease research. We highlight in this review the key insights that have contributed by organoid models to virus research, the limitations that exist in current platforms, and outline novel approaches that are being applied to address these shortcomings.
Similar content being viewed by others
Introduction
Emerging zoonotic viruses have had a profound impact on the early twenty-first century. Influenza virus, Ebola virus, Zika virus, and the pandemic severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) have collectively exemplified the substantiality of public health threat posed by emerging viruses. At the same time, urbanization1, climate change2 and wildlife trade3 are predicted to increase the propensity for viral disease emergence in the coming decades. As we usher in a new era of infectious disease, novel approaches to establish greater outbreak preparedness need to be employed. Within this framework, development of physiologically relevant laboratory models for studying disease pathophysiology and development of therapeutics to relieve disease burden needs to be prioritized.
In recent years, organoids have gained significant recognition as physiologically relevant in vitro platforms in the realm of virus research, and existing platforms are being refined continuously to better accommodate human viral disease studies. In this review, we will highlight key virological findings that have been uncovered using organoid models. Additionally, we will discuss the limitations of existing organoid platforms, particularly with relevance to virological research, and novel strategies that are being explored to overcome these shortcomings.
Traditional experimental models for viral diseases
Two-dimensional (2D) cultured cell lines have served as tools for virus propagation since the early 1900s4, and remain the most used platform for studying viral diseases. Traditional 2D cell lines are easy to handle, affordable to maintain and amenable to a wide range of experimental techniques. However, most cell lines are genetically immortalized, cancerous, or transformed to enable long-term culturing, thus often possess defects such as dysfunctional innate immune signaling pathways. Inherent differences between 2D cell lines and normal cells in vivo can result in bottlenecks that select for cell line adapted viral mutants with altered entry receptor usage5, transmissibility6 and pathogenicity7,8 compared to their clinically isolated forms. An alternative in vitro platform is primary cells derived directly from patient tissues, which are of greater resemblance to human cells in vivo, but rely on steady sources of fresh tissue to generate as they typically can only be passaged a few times before showing signs of senescence9.
Animal models are used to recapitulate viral disease processes that occur at the whole organism level. Model organisms have served as valuable platforms for determining disease-causative viral agents in fulfillment of Koch’s postulates10,11 and as preclinical platforms for vaccines and antivirals12,13. Certain physiological traits encompassed by individual animal models that are of high resemblance to those in humans have also been harnessed to study viral disease pathogenesis14. For example, ferrets are a favorable model for studying transmission of influenza viruses, owing to the high histological and anatomical similarity of their respiratory tracts to those of human15. However, inherent differences between host species limit clinical translatability of findings from animal studies, since viral disease is the end product of a myriad of virus-host interactions that are fine-tuned by evolution. Moreover, as viruses generally exhibit a narrow host range16, studies of human viruses in animal models often require extensive rounds of virus adaptation in the target species, resulting in adapted variants with mutations that may alter key viral phenotypes that are relevant to human disease17,18,19. Alternatively, specialized transgenic or humanized organisms with engineered susceptibility towards the viral target may be employed, but are generally costly, time consuming and labor intensive to generate20,21,22.
Given the shortcomings of traditional experimental platforms in authentically recapitulating viral disease in human, developing improved models that can better represent cellular phenotype diversity, disease susceptibility and host responses, whilst being amenable to a wide range of experimental techniques, is a priority in virology. In the past decade, human stem cell-derived organoids have proven repeatedly to be an asset to virus research, unraveling critical viral disease mechanisms that were previously undeterminable in traditional experimental platforms, identifying viral and host factors that facilitate infection, and providing tools for propagating viruses that were previously challenging to cultivate in vitro. Applicability of organoid platforms to virus research was further demonstrated when they were rapidly employed for disease modeling early during the COVID-19 pandemic and have since contributed a wealth of knowledge towards the disease. In this review, we discuss how organoids have been employed to fulfill critical knowledge gaps in virology. We focus our discussion on viruses that have been studied extensively using organoid models in recent years.
Organoid models for viral diseases
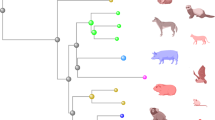
Organoids are defined as multicellular structures or “mini-organs” derived from stem or progenitor cells that are composed of organ-specific cell types with the capacity to self-organize through cell sorting and spatially restricted lineage commitment23,24. Organoid models are generally three-dimensional (3D) structures that are either embedded in an extracellular matrix (ECM) gel or free-floating in medium, and require exogenous supplementation of growth factors that are essential for stem cell proliferation. In recent years, many 2D organoid models composed of cells that are directly grown on a flat surface with or without ECM coating, have also been established25,26,27,28. Organoids can either be derived from embryonic or induced pluripotent stem cells (together referred to as PSCs) or adult stem cells (ASCs) (Fig. 1). PSC-derived organoids from fetal tissues or somatic cells are generated through a stepwise differentiation process that requires supplementation of specific growth factors at each stage29,30. Additionally, PSCs can be induced to differentiate into a wide range of tissue types from all three germ layers. Since PSC organoids are formed through processes that are unique to embryonic development, they make for ideal models to study organogenesis, in vivo development, and viral diseases known to impact these processes. ASC-derived organoids, which were first established for the intestine after the identification of Lgr5 as a marker of adult gut stem cells31,32, can be generated from tissue-resident stem cells in adult tissues or organs through mimicking the adult stem cell niche during tissue renewal or damage repair. Resultant ASC-derived organoids model adult cells in vivo with high accuracy, but generally only contain epithelial cells.
A Embryonic (ESC) or induced pluripotent stem cells (iPSCs), together referred to as PSCs, can be derived from embryonic tissues or reprogrammed from terminally differentiated fibroblasts (and other somatic cells), respectively. These progenitors can be induced to form PSC organoids from all three germ layers through a stepwise differentiation process that requires supplementation of specific growth factors at each stage. PSC organoids contain niche components from the stroma in addition to epithelial cell types, but often present fetal or neonatal phenotypes that closer resemble developing organs rather than adult tissues. B ASC organoids are generated from stem cells isolated from adult tissues that have regenerative potential. Following isolation and expansion, ASCs can be cultured in vitro with specific combinations of growth factors to induce formation of organoids containing diverse epithelial cell types from the tissue origin. Figure created with BioRender.
For 3D organoids, the cell surface is often embedded in the interior and inaccessible to viruses. As such, viral infection studies often require additional steps such as disrupting organoids into fragments, precision slicing of organoids33, microinjecting viruses into the organoid lumen34, or generating inverted organoids that adopt an “apical out” conformation in 3D35. In contrast, 2D organoids such as those differentiated on a Transwell at air-liquid interface (ALI), do not have this limitation and thus are frequently used as tools in virology. Comparisons of these available approaches have previously been reviewed36 and is illustrated in Fig. 2. In recent years, a wide range of organoid systems have been generated and applied to model human viral diseases.
Airway organoids are depicted as an example in this figure. Differentiated organoids generated for viral disease studies can be cultured as (A). 3D structures embedded in an ECM scaffold or (B). a 2D pseudostratified epithelial layer at air-liquid interface. 3D organoids typically require additional processing prior to infection to enable access to the apical layer that is embedded in the interior. This can be achieved through (C). enzymatically or mechanically disrupting organoids into fragments and re-embedding into ECM scaffolds prior to infection or (D). generating inverted apical-out organoids. Alternatively, viruses can be (E). microinjected into the lumen of intact 3D organoids. Figure created with BioRender.
Norovirus
Human noroviruses (HuNoVs) are the leading cause of acute and food-borne gastroenteritis worldwide37. Unsuccessful attempts in cultivating HuNoVs using primary and continuous cell lines have hindered understanding of disease pathogenesis and development of therapeutics for over three decades. Human intestinal organoids (hIOs) derived from human ASCs provided the first platform that allowed for in vitro HuNoV replication in a reproducible manner, and further aided identification of enterocytes as the predominant intestinal target for infection and replication38. Importantly, the study pinpointed bile as a strain-dependent required factor or enhancer for in vitro viral replication, the mechanism of which requires further investigation38. hIOs also provided a platform for assessing the effectiveness of virus inactivation strategies as disease countermeasures against different HuNoV strains39. Consistent with findings from animal and human studies40,41, hIOs demonstrated strain-dependent HuNoV binding specificity for histo-blood group antigens expressed on the mucosal surface of the gastrointestinal tract, matchedness of which was necessary to enable viral replication in vitro42. Following successes in HuNoVs studies, hIOs have also been adopted to compare infectivity and host responses elicited by other enteric viruses such as human rotavirus43, echovirus (E11)44, coxsackie B virus (CVB)45 and enterovirus 7146. In contrast to observations from infected 2D intestinal cell lines conducted in the same study, infectivity and profile of inflammatory responses mounted by each enteric virus in hIOs were distinct. In particular, CVB and E11 were able to replicate in enteroids with greater efficiency, but robust induction of transcripts for ISGs, cytokines and chemokines associated with antiviral signaling was observed only following infection with E1147, collectively demonstrating the promise of hIOs as a human relevant model to unveil critical host-pathogen interactions at the gastro-intestinal cellular barrier.
Influenza virus
Influenza A viruses (IAV) exhibit a diverse host range and display remarkable capacity to rapidly evolve under selection pressure and adapt to new hosts. IAVs possess a segmented genome and a low fidelity RNA polymerase, which continuously generate reassortants and antigenic variants that have the potential to cross species barriers and cause epidemics, and even pandemics. However, robust in vitro models for predicting zoonotic spillover potential of animal harbored IAVs had previously been lacking. A potential recourse for this is human ASC-derived airway organoids (AOs) that morphologically and functionally resemble human airways at near physiological levels. 2D AOs differentiated under defined conditions demonstrated the ability to distinguish the replicative capacity of human-infective IAVs from poorly human-infective avian and swine subtypes48. Importantly, high levels of serine proteases including TMPRSS2, TMPRSS4, HAT and matriptase, which conduct proteolytic cleavage of the virus surface hemagglutinin preceding viral-host membrane fusion, were expressed intrinsically upon AO differentiation48. Another study comparing replication capacity and host innate immune responses elicited by avian H5 and H7 viruses isolated from human patients, and human pandemic H1N1, identified higher replication capacity of H1N1 and avian H7N9 in AO cultures, whereas secretion of pro-inflammatory cytokines such as IL-6 and IFN-β were most pronounced following infection with high-pathogenic avian (HPAI) H5N149, for which cytokine dysregulation has been postulated to contribute to severe disease phenotypes in human patients. For H7 subtypes, higher replication capacity was displayed by the human-infective H7N9/Ah strain compared to the poorly human-infective H7N2 in AO49. These findings were consistent with observations in ex vivo human bronchus explants. In another study, human AO were used to assess the spillover risk of recent HPAI avian H5N6 and H5N8 isolates. Lower replication competence of H5N6/H5N8 isolates compared to human pandemic H1N1 and H5N1 in AO epithelial cells indicated low likelihood of these strains in contributing to severe disease in humans50. Taken together, these studies have substantiated the promise of using AO platforms to assess viral tropism and infectivity in human for pandemic risk assessment of zoonotic IAVs.
Owing to the rapidly evolving nature of IAVs, seasonal influenza vaccine formulations need to be updated annually. Quadrivalent and trivalent inactivated vaccines (IIVs), which are most commonly administered around the world, contain only antigens representative of human IAV and IBV strains, with limited coverage for zoonotic subtypes. While exploration of novel vaccine modalities that can provide broad spectrum protection are underway, majority of pre-clinical vaccine assessments are conducted in mice or non-human primates, and observed responses are often poorly predictive of those in humans51,52. Primary human tonsil organoids were recently explored in proof-of-concept studies as a platform for evaluating humoral responses to influenza vaccines. Tonsil organoids cultured in vitro were able to functionally retain germinal centre (GC) features, including production of antigen specific antibodies, somatic hypermutation and affinity maturation upon stimulation with a live-attenuated influenza vaccine (LAIV)53. In another study, tonsil organoids isolated and expanded from a single individual established distinct cellular and antibody dynamics following stimulation by different antigen modalities originating from IIVs, LAIVs and IAVs, and has been proposed as a blueprint to profile in vivo immune responses in future clinical trials without the need for experimental animals54.
SARS-CoV-2
The COVID-19 pandemic has heralded an unprecedented volume of research activity over a short period of time. Notably, significant insights into SARS-CoV-2 tropism, replication kinetics and mechanisms of disease pathogenesis were elucidated using organoids. Consistent with findings from human COVID-19 cases55,56, ciliated cells expressing ACE2 and TMPRSS2 were identified as the primary cellular target during initial infection in differentiated AOs derived from resected human airway tissues57. In severe COVID-19 cases, patients develop acute respiratory distress syndrome that is characterized by diffuse alveolar damage and formation of hyaline membranes that hinders gas exchange58,59. A recent study in a primary nasal epithelial organoid model identified that SARS-CoV-2 binds to ACE2 receptors expressed on the surface of motile cilia and hijacks the host cell to trigger formation of elongated apical microvilli, which are used to traverse viral progeny across the mucus layer60. Importantly, depletion of epithelial cilia in this model inhibited SARS-CoV-2 infection, suggesting motile cilia as a novel target for blocking entry and spread of viruses in the airway60. Due to the lack of clinical material from early infection timepoints, characterization of COVID-19 pathophysiology in the distal lung during early infection has been a challenge. Studies using 3D distal lung organoids have aided identification of club cells as a cellular target in the lower airways61. Further, transcriptomic and histological analyses on alveolospheres cultured from primary human lung tissue have shed light on crucial constituents of the inflammatory state upon infection of the alveoli, including upregulation of interferon-mediated responses and apoptosis, as well as a loss of mature AT2 phenotype and AT2 cell death, as indicated by downregulated expression of genes encoding for surfactant proteins62,63,64,65, which corroborates with clinical findings in COVID-19 lung tissues66,67,68.
Though COVID-19 primarily presents clinically as a respiratory disease, systemic symptoms that manifest during early infection or consequently as disease sequelae have also been reported widely69. Moreover, as ACE2 is expressed across various mammalian tissues, SARS-CoV-2 can potentially spread to other organs in the body following initial infection of the respiratory tract. Enterocytes in the intestinal epithelium, which express ACE2 abundantly, were shown to support robust SARS-CoV-2 replication in a small intestine organoid (hSIO) model. Genetic signatures induced by infection in hSIO included cytokines and IFN-stimulated genes associated with IFN type I and III responses70,71, which may contribute to gastrointestinal inflammation and in part give rise to gastrointestinal symptoms observed in COVID-19 patients72. The type II transmembrane serine protease (TTSP)-dependent entry of SARS-CoV-2 had been identified early during the pandemic in a human AO model, which further pinpointed the multibasic cleavage motif (MBCS) on the virus spike protein to be an adaptation feature that facilitates efficient TTSP usage73. Importantly, loss of the MBCS motif is observed upon viral propagation in cells that lack TTSPs, but not in AOs differentiated at ALI74. A follow-up study conducted in CRISPR/Cas9 modified hIOs further demonstrated that SARS-CoV-2, as well as MERS-CoV and SARS-CoV, specifically utilize TMPRSS2 for entry, making it an attractive target for pan-coronavirus therapeutics. During the emergence of the Omicron variant, several studies reported a switch in viral entry pathway towards TMPRSS2-independent endosomal entry75,76. This was later demonstrated in AOs and CRISPR/Cas9 modified hSIOs to be a cell line-specific phenomenon, as TTSP-mediated entry was still efficiently utilized by Omicron in organoids77.
Besides elucidating host-virus interaction and pathophysiological mechanisms, organoids have also shown promise as a translational platform for drug efficacy determination and identification of novel therapeutics for SARS-CoV-2. In particular, cardiac, lung and hIOs have been adopted for drug screening, from which a broad spectrum of SARS-CoV-2 entry inhibitors78, replication suppressors63,79, as well as cardioprotective drugs80 that could be utilized as COVID-19 therapeutics had been identified. Additionally, innovative efforts to combine organoid models with gene editing techniques such as CRISPR/Cas9 to identify host genetic factors implied in SARS-CoV-2 infection have recently been established in the form of a CRISPR-knockout biobank81. Combination of CRISPR/Cas9 technology with human PSC-derived hepatocyte and blood vessel organoids have also been implemented in studies for viruses beyond SARS-CoV-2, such as filoviruses, which have identified the guanine nucleoside exchange factor CCZ1 as an essential host factor that controls early replication of Ebola and Marburg viruses82. Such technique could be applied to future studies of other novel and emerging viruses to enable rapid characterization of host factors involved in pathogen interaction, and identify targets for antiviral development. Additionally, organoids can also be readily cultured from diverse host species, thus hold promise to model differences in virus susceptibility between hosts. New animal derived organoid models ranging from snake venom glands83 to bat intestines70 are being established rapidly, some of which had already been applied to study virus tropism and host responses, such as for SARS-CoV-270.
Zika virus
Since its identification in Africa, the mosquito-borne Zika virus (ZIKV) has spread extensively across the globe. Global footprint of ZIKV has increased drastically in the last two decades, and has caused large outbreaks in the Yap Islands in 200784, in French Polynesia in 201385 and in South America in 201586. While majority of ZIKV cases are asymptomatic or mildly symptomatic, concern of ZIKV-associated severe pathogenic effects became prominent during the 2015 outbreak in Brazil, where a substantial number of infections during pregnancy was associated with neurological abnormalities and in particular, microencephaly in newborns87. Due to the lack of accessibility to live infected human fetal tissues, clinical understanding of ZIKV pathogenesis in the fetal central nervous system were limited. Using human PSC-derived cerebral organoids and pregnant mouse models, multiple groups have demonstrated that ZIKV infection results in disruption of the cerebral cortical layers and decline in proliferative zones, as well as reduced levels of functional neurones88,89. In accordance with these findings, another study that used brain region-specific organoids grown in spinning bioreactors showed that both Asian and African ZIKV strains preferentially and productively infected neural progenitors and triggered premature differentiation90, which closely resembled infection patterns observed in human fetuses91. Subsequent studies in human PSC-derived brain organoids identified alterations in the DNA methylome of neural progenitors and differentiated neurons following ZIKV infection at genes implicated in neuropsychiatric disorders92, suggesting neuropsychiatric complications as a potential downstream effect of fetal ZIKV infection. Additionally, human cortical organoids have been adopted in drug repurposing screens for therapeutics that can limit ZIKV induced progenitor cell death from FDA-approved drug libraries93 as well as in characterizing the mechanism of action of novel anti-ZIKV compounds94.
Limitations of organoid platforms in virology
As organoid platforms gain increasing popularity as in vitro tools for modeling viral diseases, it is important to recognize the advantages and address the limitations of each organoid system to achieve improved models with optimal physiological relevance, and increase translational applicability of findings. For example, ASC-derived organoids can well represent the architecture and functional aspects of the adult tissue epithelium, and stably retain epigenetic signatures from the original tissue throughout passages95. As such, they are able to model responses to viral infections within different genetic backgrounds and disease states. However, ASC-organoids generally lack representation beyond epithelial phenotypes, which include stromal, immune and vascular niches, and thus are of limited representativeness of epithelial-microenvironment interactions in the context of viral diseases. On the other hand, PSCs are valuable for culturing organoids from tissues with stem cells that are difficult to access or are of embryonic derivative, such as the brain. PSC-derived organoids typically contain both epithelial and mesenchymal layers, making them of greater complexity than ASC-derived organoids and can support investigations of epithelial-mesenchymal interactions during viral infections. However, even in a fully differentiated state, PSC models often retain cellular phenotypes that are fetal or neonatal, which limits their representativeness of viral-host interactions in adult functional tissues96,97.
A shortcoming that is common in both ASC and PSC models is the general lack of immune cell populations, which is pivotal in the context of viral disease modeling, since excessive inflammation or dysfunctional immune responses often play an equally or more important role in inducing host damage than direct infection58,98,99. This infection-induced hyperinflammatory state has yet to be thoroughly modeled in any organoid platforms. Another aspect with room for improvement in both models is better definition of growth conditions to achieve greater reproducibility of findings. This may entail adopting culture methods that can limit artefacts from ECM derivatives, such as inflammatory proteins, which can lead to heterogeneity in organoid formation from study to study100. In terms of integrating organoid platforms into translational research, a major challenge is to generate high-throughput platforms that are complementary with organoid technology. In one study, human PSC-derived lung organoid cultured in a 384-well format was used for antiviral screening, and identified FDA approved drugs such as imatinib and mycophenolic acid to be able to inhibit SARS-CoV-2 entry101. Other models have additionally incorporated custom-engineered platforms, such as a 96-well microchannel based ALI system, that is designed to be compatible with high resolution in situ imaging and real-time sensing. The platform supported replication of IAV and the human seasonal coronavirus NL63, for which detected viral copies could be reduced using oseltamivir and camostat mesylate respectively102. As of present, organoid platforms for certain high-risk pathogens such as causative agents for viral hemorrhagic fevers remain scarce, in part due to the stringency of BSL-4 containment facilities. However, research on these viruses —particularly those featured in the WHO priority list103—is critical for epidemic and pandemic preparedness. Finally, since viral disease pathophysiology rarely localizes in a single organ, employing novel strategies to capture inter-organ interactions would be essential to understand systemic effects of viral infections. Approaches to address these existing limitations are currently being explored progressively in organoid research.
“Next-generation” organoid platforms
Immune co-culture
For viral disease studies, devising methods to include immune cell populations in organoid platforms is a critical consideration going forward, since exacerbated or dysfunctional immune responses are thought to underlie severe disease for many viruses, including SARS-CoV-2 58,99,104, HPAI H5N1105 and Ebola106. Acute cardiac injury was shown to be associated with a significantly greater mortality rate in COVID-19 patients, but causes of myocardial histopathology is poorly understood. Previously, studies from post-mortem heart samples from COVID-19 patients have identified increased inflammatory infiltration of CD11b+ and CD68+ macrophages107,108,109,110. To model the effects of macrophage-mediated hyperinflammation in cardiomyocytes (CM), an immuno-cardiac co-culture platform was generated using human PSC-derived CMs and monocytes/macrophages. The study identified CCL2 as the primary chemokine secreted by CMs to recruit monocytes upon SARS-CoV-2 infection, and presence of macrophages to significantly reduce the incidence of infected CMs111. In a separate co-culture study, IL-6 and TNF-ɑ generated by recruited macrophages during SARS-CoV-2 infection resulted in increased levels reactive oxygen species and apoptosis of CMs. Inhibition of IL-6 and TNF-ɑ through drug blockage of the JAK/STAT pathway was able to protect CMs from macrophage-induced hyperinflammation112.
A recent complex lung organoid model generated from intact human lung fragments was shown to retain differentiated lung epithelial lineages, mesenchymal components as well as tissue-resident immune cell subsets. Innate and adaptive immune responses following SARS-CoV-2 infection of the complex organoid mirrored clinical observations in vivo113. Importantly, virus-specific memory T cell responses was observed following infection in lung samples from SARS-CoV-2 seropositive donors, demonstrating the capacity for mounting of lung resident T cell memory in absence of peripheral lymphoid components in this co-culture system. While maintaining immune cell longevity and phenotype preservation is a major challenge for organoid immune co-cultures, the study observed that T cell-tropic cytokines helped to sustain organoid immune content over time114.
Studies of other respiratory viruses beyond SARS-CoV-2 have also adopted immune co-culture models. Differentiated nasal epithelium co-cultured with human PBMCs were used to characterize epithelium-leukocyte crosstalk during exposure to the influenza H3N2 strain, identifying monocytes, NK cells and innate T cells as the first to be activated at early stages of infection115. By partitioning epithelial cells and PBMCs in culture with a Transwell insert, direct PBMC infection was prevented, allowing for identification of immune responses triggered by soluble factors that are released by the infected epithelium115. Co-cultures of human pulmonary organoids and neutrophils have also been established previously to investigate neutrophil migration patterns in the context of inflammation, and been adapted to study neutrophil-epithelium interactions during RSV infection, identifying cytokines such as IP-10 and RANTES to have a crucial role in recruiting leukocytes during early infection34.
Recently, a co-culture study established using PSC-derived liver organoids (LOs) comprised of hepatocytes and Kupffer-like cells, and macrophages from the same donor, was used to elucidate the role of hepatitis C virus (HCV) infection in the pathogenesis of non-alcoholic fatty liver disease (NAFLD)116. In particular, infection with HCV upregulated host lipogenesis resulting in fatty acid accumulation in LOs, which was accentuated in the presence of macrophages. Moreover, therapeutics that are used to target non-alcoholic steatohepatitis and are currently in late-stage clinical trials elicited similar outcomes in the LO platform and in human cohorts, demonstrating the potential for the co-culture system to be used for future evaluation of NAFLD pathogenesis, and potentially other chronic liver diseases, as well as for evaluation of therapeutic efficacy116. Currently, novel organoids platforms to model immune responses to viral infections in the lymphoid microenvironment are also being established. Lymphoid organoids proliferated from murine splenocytes has established a basis for in vitro induction of GC-like B cells, which could be applied to elucidate mechanisms of mounting humoral B cell immunity during viral infections in humans, as well as development of vaccines and antibody-based immunotherapy against viral diseases117. As immune co-culture organoid systems advance rapidly and become increasingly incorporated into viral disease studies, additional considerations need to be accounted for, such as high variability of immune responses between individuals to viral infections, depending on their immunological history118. For this, individualized co-culture platforms generated from patient tissues and paired-blood samples can be utilized (Fig. 3). Additionally, future studies on immunological responses to viral infection will have to optimize co-culture systems to faithfully recapitulate immune cell states during infection, and support long-term preservation of immune cells in culture.
A Organ-on-a-chip. Microfluidic-based single or multiple organ-on-a-chip platforms engineered to contain vascular flow and spatiotemporal mechanical and chemical gradients can be used to model complex interactions between the epithelium, stroma, and circulating immune components during virus infection. Multi-organ “body” chips can additionally capture inter-organ crosstalk during infection to model systemic disease. B Immune co-culture. ASC or PSC organoids can be cultured with immune cells isolated from a paired-blood sample or generated from PSCs to capture immune-epithelial crosstalk during viral infections. C CRISPR/Cas9-gene edited organoids. Knock-out organoid biobanks can be used to screen for and identify host factors utilized by novel viruses for entry. Biobanks can be generated from CRISPR/Cas9-edited ASC organoids, or edited iPSC progenitors that are subsequently expanded clonally into PSC organoids. Figure created with BioRender.
Organ-on-a-chip
While existing organoid platforms show promise in representing viral disease pathogenesis in individual tissues or organs, they lack the ability to recapitulate systemic disease, which involves signaling cues and crosstalk between epithelium components and their surrounding microenvironment. A possible avenue to bridge these limitations is microfluidics-based organ chips. Minimalistic organ chips are generally composed of cells from a tissue of interest, an ECM interface and a neighboring vascular and/or connective tissues which are seeded onto interconnected flow-through chambers of a microfluidic device119,120,121. Since the first development of a lung alveolus organ chip, numerous designs have become available at pre-commercial and commercial stages, allowing for prototyping of cellular, tissue, and organ level responses for different viral diseases122,123,124,125.
Perfused human gut chip platforms have been applied to study enteric virus infection. In addition to an intestinal epithelial layer, a model additionally incorporated a vascular channel that is separated from the epithelium by a porous matrix-coated membrane to generate continuous fluid flow through the intestinal lumen126. Under consistent perfusion and cyclic mechanical strain, spontaneous formation of a differentiated 3D epithelial layer with villus-like structures containing proliferative crypts, mucus-secreting, Paneth and enteroendocrine cells that resemble the in vivo intestinal epithelium was enabled. Reconstituted human airways consisting of differentiated airway epithelium cultured in parallel channels with pulmonary microvascular components was used to recapitulate epithelial and endothelium dysfunction as in human airways in vivo when inoculated with IAV. The airway chip has also been utilized to study human rhinovirus-induced exacerbation of asthma, characterizing immune cell transendothelial migration and hallmark inflammatory markers of the disease state127. Perfusable, endothelialized microvessel chips have shown promise for studying vascular dysfunction resulting from infection with haemorrhagic viruses. In addition to identifying vascular integrity loss and the resultant increase in vascular permeability following infusion of Ebola virus-like particles, the platform also identified activation of the Rho/ROCK pathway to be a mediator of cytoskeleton remodeling which resulted in albumin leakage from the engineered vessels128. These chip platforms can potentially offer a new avenue for uncovering pathophysiological mechanisms and provide a basis for identifying candidate therapeutics for haemorrhagic viruses, which otherwise would typically require non-human primate model studies in specialized animal BSL-4 facilities.
Organ-on-a-chip technology can imbue drawbacks of existing organoid platforms, such as presence tissue barriers, vascular flow, spatiotemporal mechanical and chemical gradients as well as recruitment of circulating immune cells (Fig. 3). Recent efforts have further built upon existing chip technology to integrate multiple organs on the same platform to form “body chips”, allowing simulation of inter-organ communication and systemic function129,130. However, existing microfluidic chip platforms still have drawbacks, particularly since they often contain immortalized cell lines. Increasing effort towards advancing chip platforms to contain organoid-derived cells would help attain greater physiological likedness. Achieving in vitro platforms that can model viral disease to high physiological relevance with reproducibility is of great priority, particularly as organoids and organ-on-a-chip models are gaining increased traction in translational research as a substitute for animal testing131,132. Integrating human-derived stem cells into chip compositions could also allow for generation of individualized organ chips composed of homogenous genotypes, which can serve as platforms for personalized therapeutic development.
Conclusion
Over the past decade, organoid platforms have contributed important findings to virus research by addressing critical knowledge gaps in viral-host interaction and disease pathophysiology. These models have shown to be advantageous in their tractability and high human physiological relevance compared to other traditional experimental platforms. As organoids become increasingly prominent as in vitro models for virology studies, crucial shortcomings that exist in platforms need to be addressed. Novel approaches to improve existing organoid systems are currently being explored widely. Next-generation organoid platforms such as immune co-culture models that can emulate epithelial-immune crosstalk during viral infections, microfluidics-based organ chips that can capture systemic effects of viral disease, and CRISPR-edited organoids that allow for precise identification of host targets that modulate viral infections, show great promise to help refine organoid technology to better accommodate viral research studies.
References
Rulli, M. C. et al. Land-use change and the livestock revolution increase the risk of zoonotic coronavirus transmission from rhinolophid bats. Nat. Food 2, 409–416 (2021).
Carlson, C. J. et al. Climate change increases cross-species viral transmission risk. Nature 607, 555–562 (2022).
Jones, K. et al. Global trends in emerging infectious diseases. Nature 451, 990–993 (2008).
Scherer, W. F., Syverton, J. T. & Gey, G. O. Studies on the propagation in vitro of poliomyelitis viruses. J. Exp. Med. 97, 695–710 (1953).
Sasaki, M. et al. SARS-CoV-2 variants with mutations at the S1/S2 cleavage site are generated in vitro during propagation in TMPRSS2-deficient cells. PLOS Pathog. 17, e1009233 (2021).
Le Sage, V. et al. Cell-culture adaptation of H3N2 influenza virus impacts acid stability and reduces airborne transmission in ferret model. Viruses 13, 719 (2021).
Johnson, B. A. et al. Loss of furin cleavage site attenuates SARS-CoV-2 pathogenesis. Nature 591, 293–299 (2021).
Bankamp, B., Fontana, J. M., Bellini, W. J. & Rota, P. A. Adaptation to cell culture induces functional differences in measles virus proteins. Virol. J. 5, 129 (2008).
Hayflick, L. & Moorhead, P. S. The serial cultivation of human diploid cell strains. Exp. Cell Res. 25, 585–621 (1961).
van den Hoogen, B. G. et al. A newly discovered human pneumovirus isolated from young children with respiratory tract disease. Nat. Med. 7, 719–724 (2001).
Fouchier, R. A. M. et al. Koch’s postulates fulfilled for SARS virus. Nature 423, 240–240 (2003).
Mukherjee, P. et al. Role of animal models in biomedical research: a review. Lab. Anim Res. 38, 18 (2022).
Chaudhari, U. et al. An overview of preclinical animal models for SARS-CoV-2 pathogenicity. Indian J. Med. Res. 153, 17 (2021).
Carrion, R. et al. Lassa virus infection in experimentally infected marmosets: liver pathology and immunophenotypic alterations in target tissues. J. Virol. 81, 6482–6490 (2007).
Margine, I. & Krammer, F. Animal models for influenza viruses: implications for universal vaccine development. Pathogens 3, 845–874 (2014).
Rothenburg, S. & Brennan, G. Species-specific host–virus interactions: implications for viral host range and virulence. Trends Microbiol. 28, 46–56 (2020).
Gu, H. et al. Adaptation of SARS-CoV-2 in BALB/c mice for testing vaccine efficacy. Science 369, 1603–1607 (2020).
Roberts, A. et al. A mouse-adapted SARS-coronavirus causes disease and mortality in BALB/c mice. PLoS Pathog. 3, e5 (2007).
Chan, M. et al. Generation and characterization of a mouse-adapted makona variant of Ebola virus. Viruses 11, 987 (2019).
Menachery, V. D. et al. SARS-like WIV1-CoV poised for human emergence. Proc. Natil. Acad. Sci. 113, 3048–3053 (2016).
Sun, S.-H. et al. A mouse model of SARS-CoV-2 infection and pathogenesis. Cell Host Microbe 28, 124–133.e4 (2020).
Chu, H., Chan, J. F. & Yuen, K. Y. Animal models in SARS-CoV-2 research. Nat. Methods 19, 392–394 (2022).
Lancaster, M. A. & Knoblich, J. A. Organogenesis in a dish: modeling development and disease using organoid technologies. Science 345, 1247125 (2014).
Sato, T. & Clevers, H. Growing self-organizing mini-guts from a single intestinal stem cell: mechanism and applications. Science 340, 1190–1194 (2013).
Chiu, M. C. et al. A bipotential organoid model of respiratory epithelium recapitulates high infectivity of SARS-CoV-2 Omicron variant. Cell Discov. 8, 57 (2022).
Braverman, J. & Yilmaz, Ö. H. From 3D organoids back to 2D enteroids. Dev. Cell 44, 533–534 (2018).
Thorne, C. A. et al. Enteroid monolayers reveal an autonomous WNT and BMP circuit controlling intestinal epithelial growth and organization. Dev. Cell 44, 624–633.e4 (2018).
Wang, Y. et al. Self-renewing monolayer of primary colonic or rectal epithelial cells. Cell. Mol. Gastroenterol. Hepatol. 4, 165–182.e7 (2017).
Clevers, H. Modeling development and disease with organoids. Cell 165, 1586–1597 (2016).
Takahashi, K. & Yamanaka, S. Induction of pluripotent stem cells from mouse embryonic and adult fibroblast cultures by defined factors. Cell 126, 663–676 (2006).
Barker, N. et al. Identification of stem cells in small intestine and colon by marker gene Lgr5. Nature 449, 1003–1007 (2007).
Sato, T. et al. Single Lgr5 stem cells build crypt-villus structures in vitro without a mesenchymal niche. Nature 459, 262–265 (2009).
Watanabe, M. et al. Self-organized cerebral organoids with human-specific features predict effective drugs to combat zika virus infection. Cell Rep. 21, 517–532 (2017).
Sachs, N. et al. Long‐term expanding human airway organoids for disease modeling. EMBO J. 38, e100300 (2019).
Stroulios, G. et al. Apical-out airway organoids as a platform for studying viral infections and screening for antiviral drugs. Sci. Rep. 12, 7673 (2022).
Aguilar, C. et al. Organoids as host models for infection biology – a review of methods. Exp. Mol. Med. 53, 1471–1482 (2021).
Lopman, B. A., Steele, D., Kirkwood, C. D. & Parashar, U. D. The vast and varied global burden of norovirus: prospects for prevention and control. PLOS Med. 13, e1001999 (2016).
Ettayebi, K. et al. Replication of human noroviruses in stem cell-derived human enteroids. Science 353, 1387–1393 (2016).
Costantini, V. et al. Human norovirus replication in human intestinal enteroids as model to evaluate virus inactivation. Emerg. Infect. Dis. 24, 1453–1464 (2018).
Rockx, B. H. G., Bogers, W. M. J. M., Heeney, J. L., van Amerongen, G. & Koopmans, M. P. G. Experimental norovirus infections in non‐human primates. J. Med. Virol. 75, 313–320 (2004).
Cheetham, S. et al. Pathogenesis of a genogroup II human norovirus in Gnotobiotic Pigs. J. Virol. 80, 10372–10381 (2006).
Zhang, D. et al. Human intestinal organoids express histo-blood group antigens, bind norovirus VLPs, and support limited norovirus replication. Sci. Rep. 7, 12621 (2017).
Finkbeiner, S. R. et al. Stem cell-derived human intestinal organoids as an infection model for rotaviruses. mBio 3, e00159–12 (2012).
Saxena, K. et al. Human intestinal enteroids: a new model to study human rotavirus infection, host restriction, and pathophysiology. J. Virol. 90, 43–56 (2016).
Villenave, R. et al. Human gut-on-a-chip supports polarized infection of Coxsackie B1 virus in vitro. PLOS ONE 12, e0169412 (2017).
Tsang, J. O.-L. et al. Development of three-dimensional human intestinal organoids as a physiologically relevant model for characterizing the viral replication kinetics and antiviral susceptibility of enteroviruses. Biomedicines 9, 88 (2021).
Drummond, C. G. et al. Enteroviruses infect human enteroids and induce antiviral signaling in a cell lineage-specific manner. Proc. Natl. Acad. Sci. 114, 1672–1677 (2017).
Zhou, J. et al. Differentiated human airway organoids to assess infectivity of emerging influenza virus. Proc. Natl. Acad. Sci. 115, 6822–6827 (2018).
Hui, K. P. Y. et al. Tropism, replication competence, and innate immune responses of influenza virus: an analysis of human airway organoids and ex-vivo bronchus cultures.Lancet Respir. Med. 6, 846–854 (2018).
Bui, C. H. T. et al. Risk assessment for highly pathogenic Avian Influenza A(H5N6/H5N8) Clade 2.3.4.4 viruses. Emerg. Infect. Dis. 27, 2619–2627 (2021).
Rivera, J. & Tessarollo, L. Genetic background and the dilemma of translating mouse studies to humans. Immunity 28, 1–4 (2008).
Watkins, D. I., Burton, D. R., Kallas, E. G., Moore, J. P. & Koff, W. C. Nonhuman primate models and the failure of the Merck HIV-1 vaccine in humans. Nat. Med. 14, 617–621 (2008).
Wagar, L. E. et al. Modeling human adaptive immune responses with tonsil organoids. Nat. Med. 27, 125–135 (2021).
Kastenschmidt, J. M. et al. Influenza vaccine format mediates distinct cellular and antibody responses in human immune organoids. Immunity 56, 1910–1926.e7 (2023).
Hou, Y. J. et al. SARS-CoV-2 reverse genetics reveals a variable infection gradient in the respiratory tract. Cell 182, 429–446.e414 (2020).
Ahn, J. H. et al. Nasal ciliated cells are primary targets for SARS-CoV-2 replication in the early stage of COVID-19. J. Clin. Investig. 131, e148517 (2021).
Milewska, A. et al. Replication of severe acute respiratory syndrome Coronavirus 2 in human respiratory epithelium. J. Virol. 94, e00957–20 (2020).
Lamers, M. M. & Haagmans, B. L. SARS-CoV-2 pathogenesis. Nat. Rev. Microbiol. 20, 270–284 (2022).
Konopka, K. E. et al. Diffuse alveolar damage (DAD) resulting from coronavirus disease 2019 infection is morphologically indistinguishable from other causes of DAD. Histopathology 77, 570–578 (2020).
Wu, C.-T. et al. SARS-CoV-2 replication in airway epithelia requires motile cilia and microvillar reprogramming. Cell 186, 112–130.e20 (2023).
Salahudeen, A. A. et al. Progenitor identification and SARS-CoV-2 infection in human distal lung organoids. Nature 588, 670–675 (2020).
Katsura, H. et al. Human lung stem cell-based Alveolospheres provide insights into SARS-CoV-2-mediated interferon responses and pneumocyte dysfunction. Cell Stem Cell 27, 890–904.e8 (2020).
Mulay, A. et al. SARS-CoV-2 infection of primary human lung epithelium for COVID-19 modeling and drug discovery. Cell Rep. 35, 109055 (2021).
Youk, J. et al. Three-dimensional human alveolar stem cell culture models reveal infection response to SARS-CoV-2. Cell Stem Cell 27, 905–919.e10 (2020).
Huang, J. et al. SARS-CoV-2 infection of pluripotent stem cell-derived human lung alveolar Type 2 cells elicits a rapid epithelial-intrinsic inflammatory response. Cell Stem Cell 27, 962–973.e7 (2020).
Chen, J., Wu, H., Yu, Y. & Tang, N. Pulmonary alveolar regeneration in adult COVID-19 patients. Cell Res. 30, 708–710 (2020).
Delorey, T. M. et al. COVID-19 tissue atlases reveal SARS-CoV-2 pathology and cellular targets. Nature 595, 107–113 (2021).
Melms, J. C. et al. A molecular single-cell lung atlas of lethal COVID-19. Nature 595, 114–119 (2021).
Ballering, A. V., van Zon, S. K. R., Olde Hartman, T. C. & Rosmalen, J. G. M. Persistence of somatic symptoms after COVID-19 in the Netherlands: an observational cohort study. Lancet 400, 452–461 (2022).
Zhou, J. et al. Infection of bat and human intestinal organoids by SARS-CoV-2. Nat. Med. 26, 1077–1083 (2020).
Lamers, M. M. et al. SARS-CoV-2 productively infects human gut enterocytes. Science 369, 50–54 (2020).
Gu, J., Han, B. & Wang, J. COVID-19: gastrointestinal manifestations and potential Fecal–Oral Transmission. Gastroenterology 158, 1518–1519 (2020).
Mykytyn, A. Z. et al. SARS-CoV-2 entry into human airway organoids is serine protease-mediated and facilitated by the multibasic cleavage site. eLife 10, e64508 (2021).
Lamers, M. M. et al. Human airway cells prevent SARS-CoV-2 multibasic cleavage site cell culture adaptation. eLife 10, e66815 (2021).
Meng, B. et al. Altered TMPRSS2 usage by SARS-CoV-2 omicron impacts infectivity and fusogenicity. Nature 603, 706–714 (2022).
Willett, B. J. et al. SARS-CoV-2 Omicron is an immune escape variant with an altered cell entry pathway.Nat Microbiol. 7, 1161–1179 (2022).
Mykytyn, A. Z. et al. SARS-CoV-2 Omicron entry is type II transmembrane serine protease-mediated in human airway and intestinal organoid models. J. Virol. 97, e0085123 (2023).
Han, Y. et al. Identification of SARS-CoV-2 inhibitors using lung and colonic organoids. Nature 589, 270–275 (2020).
Lee, J.-E. et al. Development of a screening platform to discover natural products active against SARS-CoV-2 infection using lung organoid models. Biomater. Res. 27, 18 (2023).
Mills, R. J. et al. BET inhibition blocks inflammation-induced cardiac dysfunction and SARS-CoV-2 infection. Cell 184, 2167–2182.e22 (2021).
Beumer, J. et al. A CRISPR/Cas9 genetically engineered organoid biobank reveals essential host factors for coronaviruses. Nat. Commun. 12, 5498 (2021).
Monteil, V. et al. Identification of CCZ1 as an essential lysosomal trafficking regulator in Marburg and Ebola virus infections. Nat. Commun. 14, 6785 (2023).
Post, Y. et al. Snake venom gland organoids. Cell 180, 233–247.e21 (2020).
Lanciotti, R. S. et al. Genetic and serologic properties of Zika virus associated with an epidemic, Yap State, Micronesia, 2007. Emerg. Infect. Dis. 14, 1232–1239 (2008).
Oehler, E. et al. Zika virus infection complicated by Guillain-Barré syndrome – case report, French Polynesia, December 2013. Eurosurveillance 19, 20720 (2014).
Dick, G. W. A., Kitchen, S. F. & Haddow, A. J. Zika Virus (I). Isolations and serological specificity. Trans. R. Soc. Trop. Med. Hyg. 46, 509–520 (1952).
Kleber de Oliveira, W. et al. Increase in reported prevalence of microcephaly in infants born to women living in areas with confirmed Zika virus transmission during the first trimester of pregnancy — Brazil, 2015. Morb. Mortal. Weekly Rep. 65, 242–247 (2016).
Garcez, P. P. et al. Zika virus impairs growth in human neurospheres and brain organoids. Science 352, 816–818 (2016).
Cugola, F. R. et al. The Brazilian Zika virus strain causes birth defects in experimental models. Nature 534, 267–271 (2016).
Qian, X. et al. Brain-region-specific organoids using mini-bioreactors for modeling ZIKV exposure. Cell 165, 1238–1254 (2016).
Gabriel, E. et al. Recent Zika virus isolates induce premature differentiation of neural progenitors in human brain organoids. Cell Stem Cell 20, 397–406.e5 (2017).
Janssens, S. et al. Zika Virus alters DNA methylation of neural genes in an organoid model of the developing human brain. mSystems 3, e00219–17 (2018).
Zhou, T. et al. High-content screening in hPSC-neural progenitors identifies drug candidates that inhibit Zika Virus infection in fetal-like organoids and adult brain. Cell Stem Cell 21, 274–283.e5 (2017).
Pettke, A. et al. Broadly active antiviral compounds disturb Zika virus progeny release rescuing virus-induced toxicity in brain organoids. Viruses 13, 37 (2020).
Huch, M. et al. Long-term culture of genome-stable bipotent stem cells from adult human liver. Cell 160, 299–312 (2015).
Camp, J. G. et al. Human cerebral organoids recapitulate gene expression programs of fetal neocortex development. Proc. Natl. Acad. Sci. 112, 15672–15677 (2015).
Hrvatin, S. et al. Differentiated human stem cells resemble fetal, not adult, β cells. Proc. Natl. Acad. Sci. 111, 3038–3043 (2014).
Nishiga, M., Wang, D. W., Han, Y., Lewis, D. B. & Wu, J. C. COVID-19 and cardiovascular disease: from basic mechanisms to clinical perspectives. Nat. Rev. Cardiol. 17, 543–558 (2020).
Liu, J. et al. Longitudinal characteristics of lymphocyte responses and cytokine profiles in the peripheral blood of SARS-CoV-2 infected patients. EBioMedicine 55, 102763 (2020).
Urbischek, M. et al. Organoid culture media formulated with growth factors of defined cellular activity. Sci. Rep. 9, 6193 (2019).
Du, X., Dong, Y., Li, W. & Chen, Y. hPSC-derived lung organoids: potential opportunities and challenges. Heliyon 9, e13498 (2023).
Gard, A. L. et al. High-throughput human primary cell-based airway model for evaluating influenza, coronavirus, or other respiratory viruses in vitro. Sci. Rep. 11, 14961 (2021).
Prioritizing diseases for research and development in emergency contexts. World Health Organization. https://www.who.int/activities/prioritizing-diseases-for-research-and-development-in-emergency-contexts. (2023).
Hu, B. et al. Characteristics of SARS-CoV-2 and COVID-19. Nat. Rev. Microbiol. 19, 141–154 (2021).
Korteweg, C. & Gu, J. Pathology, molecular biology, and pathogenesis of Avian Influenza A (H5N1) infection in humans. Am. J. Pathol. 172, 1155–1170 (2008).
Martines, R. B., Ng, D. L., Greer, P. W., Rollin, P. E. & Zaki, S. R. Tissue and cellular tropism, pathology and pathogenesis of Ebola and Marburg viruses. J. Pathol. 235, 153–174 (2015).
Merad, M. & Martin, J. C. Pathological inflammation in patients with COVID-19: a key role for monocytes and macrophages. Nat. Rev. Immunol. 20, 355–362 (2020).
Lindner, D. et al. Association of cardiac infection with SARS-CoV-2 in confirmed COVID-19 autopsy cases. JAMA Cardiol. 5, 1281 (2020).
Escher, F. et al. Detection of viral SARS‐CoV‐2 genomes and histopathological changes in endomyocardial biopsies. ESC Heart Fail. 7, 2440–2447 (2020).
Tavazzi, G. et al. Myocardial localization of coronavirus in COVID‐19 cardiogenic shock. Eur. J. Heart Fail. 22, 911–915 (2020).
Yang, L. et al. Cardiomyocytes recruit monocytes upon SARS-CoV-2 infection by secreting CCL2. Stem Cell Rep. 16, 2274–2288 (2021).
Yang, L. et al. An immuno-cardiac model for macrophage-mediated inflammation in COVID-19 hearts. Circ. Res. 129, 33–46 (2021).
Bhaskar, S. et al. Cytokine Storm in COVID-19—immunopathological mechanisms, clinical considerations, and therapeutic approaches: the REPROGRAM consortium position paper. Front. Immunol. 11, 1648 (2020).
Choi, SS. et al. Organoid modeling of lung-resident immune responses to SARS-CoV-2 infection. Preprint at: https://www.researchsquare.com/article/rs-2870695/v1 (2023).
Luukkainen, A. et al. A co-culture Model of PBMC and stem cell derived human nasal epithelium reveals rapid activation of NK and Innate T cells upon influenza a virus infection of the nasal epithelium. Front. Immunol. 9, 2514 (2018).
Lee, J. et al. A multicellular liver organoid model for investigating hepatitis C virus infection and non-alcoholic fatty liver disease progression. Hepatology https://journals.lww.com/hep/abstract/9900/a_multicellular_liver_organoid_model_for.655.aspx (2023).
Purwada, A. & Singh, A. Immuno-engineered organoids for regulating the kinetics of B-cell development and antibody production. Nat. Protoc. 12, 168–182 (2016).
Rouse, B. T. & Sehrawat, S. Immunity and immunopathology to viruses: what decides the outcome? Nat. Rev. Immunol. 10, 514–526 (2010).
Sontheimer-Phelps, A., Hassell, B. A. & Ingber, D. E. Modelling cancer in microfluidic human organs-on-chips. Nat. Rev. Cancer 19, 65–81 (2019).
van den Berg, A., Mummery, C. L., Passier, R. & van der Meer, A. D. Personalised organs-on-chips: functional testing for precision medicine. Lab Chip 19, 198–205 (2019).
Bhatia, S. N. & Ingber, D. E. Microfluidic organs-on-chips. Nat. Biotechnol. 32, 760–772 (2014).
Balijepalli, A. & Sivaramakrishan, V. Organs-on-chips: research and commercial perspectives. Drug Discov. Today 22, 397–403 (2017).
Ortega-Prieto, A. M. et al. 3D microfluidic liver cultures as a physiological preclinical tool for hepatitis B virus infection. Nat. Commun. 9, 682 (2018).
Si, L. et al. A human-airway-on-a-chip for the rapid identification of candidate antiviral therapeutics and prophylactics. Nat. Biomed. Eng. 5, 815–829 (2021).
Thacker, V. V. et al. Rapid endotheliitis and vascular damage characterize SARS-CoV-2 infection in a human lung-on-chip model. EMBO Rep. 22, e52744 (2021).
Villenave, R. et al. Human Gut-On-A-Chip Supports Polarized Infection of Coxsackie B1 Virus In Vitro. PLoS One 12, e0169412 (2017)..
Nawroth, J. C. et al. A microengineered airway lung chip models key features of viral-induced exacerbation of asthma. Am. J. Respir. Cell Mol. Biol. 63, 591–600 (2020).
Junaid, A. et al. Ebola Hemorrhagic Shock Syndrome-on-a-Chip. iScience 23, 100765 (2020).
Picollet-D’hahan, N., Zuchowska, A., Lemeunier, I. & Le Gac, S. Multiorgan-on-a-Chip: a systemic approach to model and decipher inter-organ communication. Trends Biotechnol. 39, 788–810 (2021).
Jeon, J., Choi, N., Lee, S. H. & Sung, J. H. Three-tissue microphysiological system for studying inflammatory responses in gut-liver Axis. Biomed. Microdevices 22, 65 (2020).
Adashi, E. Y., O’Mahony, D. P. & Cohen, I. G. The FDA Modernization Act 2.0: drug testing in animals is rendered optional.Am. J. Med. 136, 853–854 (2023).
CONGRESS.GOV. H.R. 2617 - Consolidated Appropriations Act, 2023. Became Public Law No: 117-328. December 29, 2022.
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Chu, J.T.S., Lamers, M.M. Organoids in virology. npj Viruses 2, 5 (2024). https://doi.org/10.1038/s44298-024-00017-5
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s44298-024-00017-5