Abstract
The adult mouse heart responds to injury by scarring with consequent loss of contractile function, whereas the neonatal heart possesses the ability to regenerate. Activation of the immune system is among the first events upon tissue injury. It has been shown that immune response kinetics differ between regeneration and pathological remodeling, yet the underlying mechanisms of the distinct immune reactions during tissue healing remain unclear. Here we show that the immunomodulatory PD-1–PD-L1 pathway is highly active in regenerative neonatal hearts but rapidly silenced later in life. Deletion of the PD-1 receptor or inactivation of its ligand PD-L1 prevented regeneration of neonatal hearts after injury. Disruption of the pathway during neonatal cardiac injury led to increased inflammation and aberrant T cell activation, which ultimately impaired cardiac regeneration. Our findings reveal an immunomodulatory and cardioprotective role for the PD-1–PD-L1 pathway in heart regeneration and offer potential avenues for the control of adult tissue regeneration.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Data availability
Code availability
All code used in this study is freely available on GitHub (https://github.com/CodeUTSW/Code_for_Paper/tree/main).
References
Porrello, E. R. et al. Transient regenerative potential of the neonatal mouse heart. Science 331, 1078–1080 (2011).
Cui, M. et al. Dynamic transcriptional responses to injury of regenerative and non-regenerative cardiomyocytes revealed by single-nucleus RNA sequencing. Dev. Cell 53, 102–116 (2020).
Sadek, H. & Olson, E. N. Toward the goal of human heart regeneration. Cell Stem Cell 26, 7–16 (2020).
Liu, X. et al. Lymphoangiocrine signals promote cardiac growth and repair. Nature 588, 705–711 (2020).
Cui, M. et al. Nrf1 promotes heart regeneration and repair by regulating proteostasis and redox balance. Nat. Commun. 12, 5270 (2021).
Aurora, A. B. et al. Macrophages are required for neonatal heart regeneration. J. Clin. Invest. 124, 1382–1392 (2014).
Lavine, K. J. et al. Distinct macrophage lineages contribute to disparate patterns of cardiac recovery and remodeling in the neonatal and adult heart. Proc. Natl Acad. Sci. USA 113, E1414 (2016).
Wang, Z. et al. Resource cell-type-specific gene regulatory networks underlying murine neonatal heart regeneration at single-cell resolution. Cell. Rep. 33, 108472 (2020).
Bajpai, G. et al. Tissue resident CCR2− and CCR2+ cardiac macrophages differentially orchestrate monocyte recruitment and fate specification following myocardial injury. Circ. Res. 124, 263–278 (2019).
Dick, S. A. et al. Self-renewing resident cardiac macrophages limit adverse remodeling following myocardial infarction. Nat. Immunol. 20, 29–39 (2019).
Tan, Y., Duan, X., Wang, B., Liu, X. & Zhan, Z. Murine neonatal cardiac B cells promote cardiomyocyte proliferation and heart regeneration. NPJ Regen. Med. 8, 7 (2023).
Wang, G. et al. ‘Default’ generation of neonatal regulatory T cells. J. Immunol. 185, 71–78 (2010).
Zacchigna, S. et al. Paracrine effect of regulatory T cells promotes cardiomyocyte proliferation during pregnancy and after myocardial infarction. Nat. Commun. 9, 2432 (2018).
Li, J. et al. Regulatory T-cells regulate neonatal heart regeneration by potentiating cardiomyocyte proliferation in a paracrine manner. Theranostics 9, 4324–4341 (2019).
Blanton, R. M., Carrillo-Salinas, F. J. & Alcaide, P. T-cell recruitment to the heart: friendly guests or unwelcome visitors? Am. J. Physiol. Heart Circ. Physiol. 317, H124–H140 (2019).
Dolejsi, T. et al. Adult T-cells impair neonatal cardiac regeneration. Eur. Heart J. https://doi.org/10.1093/eurheartj/ehac153 (2022).
Li, J. et al. Specific ablation of CD4+ T-cells promotes heart regeneration in juvenile mice. Theranostics 10, 8018–8035 (2020).
Wang, Z. et al. Mechanistic basis of neonatal heart regeneration revealed by transcriptome and histone modification profiling. Proc. Natl Acad. Sci. USA 116, 18455–18465 (2019).
Dong, H., Zhu, G., Tamada, K. & Chen, L. B7-H1, a third member of the B7 family, co-stimulates T-cell proliferation and interleukin-10 secretion. Nat. Med. 5, 1365–1369 (1999).
Freeman, G. J. et al. Engagement of the PD-1 immunoinhibitory receptor by a novel B7 family member leads to negative regulation of lymphocyte activation. J. Exp. Med. 192, 1027–1034 (2000).
Latchman, Y. et al. PD-L2 is a second ligand for PD-1 and inhibits T cell activation. Nat. Immunol. 2, 261–268 (2001).
Laroumanie, F. et al. CD4+ T cells promote the transition from hypertrophy to heart failure during chronic pressure overload. Circulation 129, 2111–2124 (2014).
Nevers, T. et al. Left ventricular T-cell recruitment contributes to the pathogenesis of heart failure. Circ. Heart Fail. 8, 776–787 (2015).
Kallikourdis, M. et al. T cell costimulation blockade blunts pressure overload-induced heart failure. Nat. Commun. 8, 14680 (2017).
Aghajanian, H. et al. Targeting cardiac fibrosis with engineered T cells. Nature 573, 430–433 (2019).
Rurik, J. G. et al. CAR T cells produced in vivo to treat cardiac injury. Science 375, 91–96 (2022).
Hayashi, T. et al. The programmed death-1 signaling axis modulates inflammation and LV structure/function in a stress-induced cardiomyopathy model. JACC Basic Transl. Sci. 7, 1120–1139 (2022).
Grabie, N. et al. Endothelial programmed death-1 ligand 1 (PD-L1) regulates CD8+ T-cell-mediated injury in the heart. Circulation 116, 2062–2071 (2007).
Spitzer, M. H. & Nolan, G. P. Mass cytometry: single cells, many features. Cell 165, 780–791 (2016).
Quaife-Ryan, G. A. et al. Multicellular transcriptional analysis of mammalian heart regeneration. Circulation 136, 1123–1139 (2017).
Keir, M. E., Freeman, G. J. & Sharpe, A. H. PD-1 regulates self-reactive CD8+ T cell responses to antigen in lymph nodes and tissues. J. Immunol. 179, 5064–5070 (2007).
Gao, S., Ho, D., Vatner, D. E. & Vatner, S. F. Echocardiography in mice. Curr. Protoc. Mouse Biol. 1, 71–83 (2011).
Suthahar, N., Meijers, W. C., Silljé, H. H. W. & de Boer, R. A. From inflammation to fibrosis—molecular and cellular mechanisms of myocardial tissue remodelling and perspectives on differential treatment opportunities. Curr. Heart Fail. Rep. 14, 235–250 (2017).
Kanisicak, O. et al. Genetic lineage tracing defines myofibroblast origin and function in the injured heart. Nat. Commun. 7, 12260 (2016).
Morath, A. & Schamel, W. W. αβ and γδ T cell receptors: similar but different. J. Leukoc. Biol. 107, 1045–1055 (2020).
Geltink, R. I. K., Kyle, R. L. & Pearce, E. L. Unraveling the complex interplay between T cell metabolism and function. Annu. Rev. Immunol. 36, 461–488 (2018).
Won, T. et al. Cardiac myosin-specific autoimmune T cells contribute to immune-checkpoint-inhibitor-associated myocarditis. Cell Rep. 41, 111611 (2022).
Axelrod, M. L. et al. T cells specific for α-myosin drive immunotherapy-related myocarditis. Nature 611, 818–826 (2022).
Zhang, W. et al. Necrotic myocardial cells release damage-associated molecular patterns that provoke fibroblast activation in vitro and trigger myocardial inflammation and fibrosis in vivo. J. Am. Heart Assoc. 4, e001993 (2015).
Sattler, S. & Rosenthal, N. The neonate versus adult mammalian immune system in cardiac repair and regeneration. Biochim. Biophys. Acta Mol. Cell Res. 1863, 1813–1821 (2016).
Trajano, L. F. & Smart, N. Immunomodulation for optimal cardiac regeneration: insights from comparative analyses. NPJ Regen. Med. https://doi.org/10.1038/s41536-021-00118-2 (2021).
Forte, E., Furtado, M. B. & Rosenthal, N. The interstitium in cardiac repair: role of the immune–stromal cell interplay. Nat. Rev. Cardiol. 15, 601–616 (2018).
Chen, Z., Huang, J., Kwak-Kim, J. & Wang, W. Immune checkpoint inhibitors and reproductive failures. J. Reprod. Immunol. 156, 103799 (2023).
Montero, A. M. & Huang, A. H. The regenerative capacity of neonatal tissues. Development 149, dev199819 (2022).
Nishimura, H. et al. Developmentally regulated expression of the PD-1 protein on the surface of double-negative (CD4−CD8−) thymocytes. Int. Immunol. 8, 773–780 (1996).
Rodríguez-Rodríguez, N. et al. Pro-inflammatory self-reactive T cells are found within murine TCR-αβ+CD4−CD8−PD-1+ cells. Eur. J. Immunol. 46, 1383–1391 (2016).
Sadasivam, M. et al. Activation and proliferation of PD-1+ kidney double-negative T Cells is dependent on nonclassical MHC proteins and IL-2. J. Am. Soc. Nephrol. 30, 277–292 (2019).
Velikkakam, T., Gollob, K. J. & Dutra, W. O. Double-negative T cells: setting the stage for disease control or progression. Immunology 165, 371–385 (2022).
Qin, W. et al. The diverse function of PD-1/PD-L pathway beyond cancer. Front. Immunol. 10, 1–16 (2019).
Lv, H. J. et al. Impaired thymic tolerance to α-myosin directs autoimmunity to the heart in mice and humans. J. Clin. Invest. 121, 1561–1573 (2011).
Gabrielsen, I. S. M. et al. Transcriptomes of antigen presenting cells in human thymus. PLoS ONE 14, e0218858 (2019).
Nishimura, H. et al. Autoimmune dilated cardiomyopathy in PD-1 receptor-deficient mice. Science 291, 319–322 (2001).
Okazaki, T. et al. Autoantibodies against cardiac troponin I are responsible for dilated cardiomyopathy in PD-1-deficient mice. Nat. Med. 9, 1477–1483 (2003).
Mahmoud, A. I., Porrello, E. R., Kimura, W., Olson, E. N. & Sadek, H. A. Surgical models for cardiac regeneration in neonatal mice. Nat. Protoc. 9, 305–311 (2014).
Porrello, E. R. et al. Regulation of neonatal and adult mammalian heart regeneration by the miR-15 family. Proc. Natl Acad. Sci. USA 110, 187–192 (2013).
Aronoff, L., Epelman, S. & Clemente-Casares, X. Isolation and identification of extravascular immune cells of the heart. J. Vis. Exp. 1, e58114 (2018).
Becht, E. et al. Dimensionality reduction for visualizing single-cell data using UMAP. Nat. Biotechnol. 37, 38–44 (2019).
Butler, A., Hoffman, P., Smibert, P., Papalexi, E. & Satija, R. Integrating single-cell transcriptomic data across different conditions, technologies, and species. Nat. Biotechnol. 36, 411–420 (2018).
Büttner, M., Ostner, J., Müller, C. L., Theis, F. J. & Schubert, B. scCODA is a Bayesian model for compositional single-cell data analysis. Nat. Commun. 12, 6876 (2021).
Kleshchevnikov, V. et al. Cell2location maps fine-grained cell types in spatial transcriptomics. Nat. Biotechnol. 40, 661–671 (2022).
Lim, J. F., Berger, H. & Su, I.-H. Isolation and activation of murine lymphocytes. J. Vis. Exp. 2016, e54596 (2016).
Acknowledgements
We thank S. Bezprozvannaya at the University of Texas Southwestern Medical Center (UTSW) for technical help; J. Cabrera (UTSW) for assistance on graphic design; A. Mobley and the Flow Cytometry Core at UTSW for assistance with CyTOF; the Moody Foundation Flow Cytometry Facility for assistance with cell sorting; J. Xu and Y. Jung Kim from the Children’s Research Institute for performing the Illumina sequencing; L. Straub from the Touchstone Diabetes Center at UTSW for experimental advice and scientific discussions; K. Chen for support on the analysis of CyTOF data; L. Pathira Kankanamge and S. Miriyala from Boston Children’s Hospital for assistance with scCODA single-cell and spatial transcriptome analysis, respectively; B. Li from the Lyda Hill Department of Bioinformatics; Z. Zhong from the Department of Immunology for scientific discussions; and K. O’Donnell from Department of Molecular Biology at UTSW for scientific discussions and reagents. S.V.A. is supported by a Postdoctoral Fellowship from the American Heart Association (no. 916883) and M.C. is supported by grants from the National Institutes of Health (K99HL153683 and R00HL1536833). This work was supported by grants from the National Institutes of Health (HL130253 and HL157281, to E.N.O. and R.B.-D.), the Robert A. Welch Foundation (1-0025, to E.N.O.) and the Leducq Foundation Transatlantic Network of Excellence (20CVD04, to E.N.O.).
Author information
Authors and Affiliations
Contributions
S.V.A., M.C. and E.N.O. were responsible for conceptualization. S.V.A., M.C., W.T. and E.S.-O. were responsible for the methodology. S.V.A. and M.C. were responsible for investigation. S.V.A. and M.C. were responsible for visualization. E.N.O., R.B.-D., S.V.A. and M.C. were responsible for funding acquisition. E.N.O., N.L. and R.B.-D. were responsible for supervision. S.V.A., M.C., N.L., E.N.O. and R.B.-D. were responsible for writing.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Cardiovascular Research thanks Sikander Hayat, Douglas Mann, Javid Moslehi, Nadia Rosenthal and Han Zhu for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 PD-1 and PD-L1 are expressed in the neonatal heart.
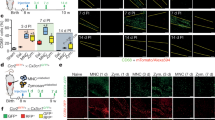
a, Expression of PD-L1 and PD-L2 in P1 mouse hearts at steady state, as measured by qPCR. Fold change expression normalized to PD-L1 is shown. n = 3 mice per group. p = 0.0026 (PD-L1 vs. PD-L2). b, t-SNE visualization of cardiac cell types identified by CyTOF represented as a composite of all samples combined. c, Median intensity of marker genes for each cardiac cell type (scaled across individual columns) in combined samples, as measured by CyTOF. d, t-SNE plots depicting PD-L1, CD45, and CD31 expression in cardiac cell types from hearts at d3 after MI or sham performed at P2 (P2-MI or P2-sham) or at P8 (P8-MI or P8-sham), as measured by CyTOF. e, Immunofluorescence of PD-L1 (red), MOMA-2 (green) and DAPI (blue) on heart sections at d1 after MI performed at P1 (P1-MI). Sections were collected at 200μm below the ligation. Scale bar, 50μm. f, Immunofluorescence of PD-L1 (red), PECAM-1 (green) and DAPI (blue) on heart sections at d1 after P8-sham. Scale bar, 50μm. g, Violin plots of PD-L1 expression in individual cells in the conditions depicted, measured by CyTOF. These values represent the raw data used to create the heat map in Fig. 1d. The boxplots contain the 25th (Q1) to 75th (Q3) percentile of the dataset (IQR, interquartile range), and the center line denotes the median value. Whiskers mark the minimum (Q1-1.5*IQR) and maximum (Q3 + 1.5*IQR) values. Statistics were calculated using a two-sided Wilcoxon test and are shown only for selected comparisons. h, PD-L1 (left) and PD-1 (right) expression in cardiomyocytes, fibroblasts, immune cells, and endothelial cells from hearts collected at d3 after MI performed at P1 (P1-MI) or at P56 (P56-MI), measured by RNA Sequencing. Data have been previously published as GSE95755. Expression is shown as count per million reads (CPM). n = 4 biologically independent samples. **p < 0.01 by two-tailed Student’s t-test, compared to the expression in corresponding cell type from P1 hearts. p = 0.0027 (PD-L1 Immune cell: P1 vs. P56), p = 0.0085 (PD-L1 Endothelial cell: P1 vs. P56), p = 0.0038 (PD-1 Immune cell: P1 vs. P56). i, t-SNE plots depicting CD3 (upper panel) and PD-1 (lower panel) expression, as measured by CyTOF. t-SNE plots are composites of all samples combined. Red circle denotes CD3+ cells. j, Absolute counts of PD-1+ T cells and PD-1+ macrophages/monocytes/neutrophils in hearts at d3 after P2-MI (n = 7 mice per group) or P8-MI (n = 9 mice per group), as measured by flow cytometry. p = 9.4 ×10−8 (P2-MI PD-1+ T cells vs. P2-MI PD-1+ macrophages/monocytes/neutrophils), p = 1.74 ×10−7 (P2-MI PD-1+ T cells vs. P8-MI PD-1+ T cells). Results are shown as mean ± s.d (h) or mean ± s.e.m (a, j); nsp > 0.05, **p < 0.01, ****p < 0.0001 by two-tailed Student’s t-test (a) and by two-way ANOVA with Fisher’s LSD test on multiple comparisons (j).
Extended Data Fig. 2 Gating strategies used for flow cytometry analyses throughout the study.
a, Gating strategy used to detect depicted innate immune cells in the heart. Tissue-resident macrophages, TRMs. b, Gating strategy used to detect depicted immune cells in the spleen. c, Gating strategy used to detect depicted T cell populations in the heart.
Extended Data Fig. 3 Ablation of the PD-1/PD-L1 pathway impairs neonatal heart regeneration.
a-c, Ejection fraction (a), left ventricular end-diastolic volume (b) and left ventricular end-systolic volume (c) of IgG- or anti-PD-L1-treated mice at 1, 2, and 3 weeks after P3-MI. IgG sham: n = 6, 5, and 5 mice at 1, 2, 3 weeks, respectively. anti-PD-L1 sham: n = 6 mice. IgG MI: n = 5, 6, and 6 mice at 1, 2, 3 weeks, respectively. anti-PD-L1 MI: n = 7 mice (a, b c). p = 0.00052 (1 week), p = 0.000008 (2 weeks), p = 0.000009 (3 weeks) (a). p = 0.004638 (2 weeks), p = 0.000823 (3 weeks) (b). p = 0.002964 (2 weeks), p = 0.000145 (3 weeks) (c). d-f, Ejection fraction (d), left ventricular end-diastolic volume (e) and left ventricular end-systolic volume (f) of Pd-1+/− and Pd-1−/− mice at 1, 2, and 3 weeks after P2-MI. Pd-1+/− sham: n = 5, 5, and 4 mice at 1, 2, 3 weeks, respectively. Pd-1−/− sham: n = 5 mice. Pd-1+/− MI: n = 4 mice. Pd-1−/− MI: n = 6, 5, and 5 mice at 1, 2, 3 weeks, respectively (d, e, f). p = 0.000703 (1 week), p = 0.001559 (2 weeks), p = 0.021383 (3 weeks) (d). p = 0.01633 (2 weeks), p = 0.019505 (3 weeks) (e). p = 0.004214 (2 weeks), p = 0.042845 (3 weeks) (f). The same cohort of mice was measured every week. Results are shown as mean ± s.e.m; *p < 0.05, **p < 0.01, ***p < 0.001, ****p < 0.0001 by two-tailed Student’s t-test.
Extended Data Fig. 4 Ablation of PD-1 promotes cardiomyocyte apoptosis post-MI.
a,b, Immunofluorescence of TUNEL (magenta), cardiac troponin T (cTnT, green) and DAPI (blue) on heart sections from Pd-1+/− (a) and Pd-1−/− (b) mice at d7 after MI performed at P2. Sections were collected at 200μm below the ligation. Scale bar, 100μm. Four Pd-1+/− biological replicates and three Pd-1-/- biological replicates are shown. Replicate, rep.
Extended Data Fig. 5 Ablation of PD-1 impairs cardiomyocyte proliferation post-MI.
a,b, Immunofluorescence of phosphorylated histone H3 (pH3, red), cardiac troponin T (cTnT, green) and DAPI (blue) on heart sections from Pd-1+/- (a) and Pd-1-/- (b) mice at d7 after MI performed at P2. Sections were collected at 200μm below the ligation. Scale bar, 100μm. Four Pd-1+/- biological replicates and three PD-1-/- biological replicates are shown. Replicate, rep.
Extended Data Fig. 6 PD-1 is required for neonatal heart regeneration.
a, Illustration showing time points of antibody injection, MI or sham and sample collection. b, c, Fractional shortening (b) and ejection fraction (c) of IgG-treated or anti-PD-1 antibody (anti-PD-1)-treated mice at 1, 2 and 3 weeks post-MI or sham. The same cohort of mice was measured every week. IgG/ anti-PD-1 sham: n = 5 mice. IgG MI: n = 8, 7, and 7 mice at 1, 2, 3 weeks, respectively. anti-PD-1 MI: n = 8 mice (b c). p = 0.0022 (1 week), p = 0.0175 (2 weeks), p = 0.0175 (3 weeks) (b). p = 0.0014 (1 week), p = 0.0227 (2 weeks), p = 0.0227 (3 weeks) (c). d, Immunofluorescence of TUNEL (magenta), cardiac troponin T (cTnT, green) and DAPI (blue) on heart sections from IgG-treated or anti-PD-1-treated mice at d7 post-MI with infarct and remote zones depicted. Sections were collected at 200μm below the ligation. Scale bar, 100μm. e, Quantification of TUNEL+ cells in (d). p = 0.002577 (Infarct: IgG vs. anti-PD-1). f, Immunofluorescence of pH3 (red), cardiac troponin T (cTnT, green) and DAPI (blue) in hearts from IgG-treated or anti-PD-1-treated mice at d7 post-MI. Sections were collected at 200μm below the ligation, and pictures were taken at the border zone of the injury. Scale bar, 100μm. g, Quantification of pH3+ cardiomyocytes from (f). p = 0.0395 (IgG vs. anti-PD-1). n = 6 mice per group (3 images counted per mouse) (e, g). Results are shown as mean ± s.e.m; *p < 0.05, **p < 0.01 by two-tailed Student’s t-test (e, g) or two-tailed Student’s t-test with correction for multiple comparisons using the Holm-Šidák method (b, c).
Extended Data Fig. 7 Identification of immune cell types by scRNA-seq.
a, Violin plots showing distribution of numbers of genes (nFeature), numbers of RNA molecules (nCount) and percentage of mitochondrial DNA (percent.mt) that were detected in Pd-1+/- or Pd-1-/- hearts at d7 after MI or sham performed at P2. Sham, Sh. b, UMAP plot of scRNA-seq data showing identified immune cell types in combined samples. c, Heat map showing the expression of top 10 marker genes of each cell type from (b) in combined samples, as measured by scRNA-seq. d, UMAP plot showing macrophage subclusters separated by sample. e, Heat map showing expression of the pro-inflammatory cytokines Il6, Il18, Il23a and Nos2, separated by cell type, genotype, and condition. Sham, Sh.
Extended Data Fig. 8 Ablation of PD-1 causes local increased immune infiltration in the heart post-MI.
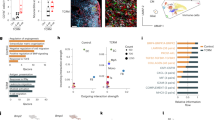
a,b, a, Quantification of spatial transcriptomic data shown in Fig. 3h. Baseline cell abundance was computed by averaging cell abundance across all spatial positions. EndoEC, endocardial cell; Mono, monocyte; Macro, macrophage; DC-like, dendric cell-like cells; FB, fibroblast. b, c, Immunofluorescence of MOMA-2 (green), cardiac troponin T (cTnT, red) and DAPI (blue) on heart sections from Pd-1+/- (b) and Pd-1-/- (c) mice at d7 after MI performed at P2. Absence of cTnT staining indicates damaged myocardium. Sections were collected at 200μm below the ligation. Scale bar, 100μm. Three Pd-1+/- and three Pd-1-/- biological replicates are shown. Replicate, rep. d, Quantification of MOMA-2+ cells from (b) and (c). n = 3 mice per group. p = 0.033 (Pd-1+/- vs. Pd-1-/-). Results are shown as mean ± s.e.m; *p < 0.05, by two-tailed Student’s t-test.
Extended Data Fig. 9 TCRγδ+ PD-1+ DN T cells accumulate in hearts injured at P2.
a, Relative composition of T cell populations depicted in Fig. 4a. Double-positive, DP. Double-negative, DN. Plotted values are shown for CD8+, CD4+ and DN T cells. b, t-SNE plots depicting CD4, CD8 and PD-1 expression in T cells from hearts at d3 after MI or sham performed at P2 (P2-MI or P2-sham) or P8 (P8-MI or P8-sham), as measured by CyTOF. c, Expression of TCRγδ, TCRβ and CD90 in DN, CD4+, CD8+ T cells from hearts at d3 after P2-MI, as measured by flow cytometry. d, Percentage of DN T cells among all T cells in spleens at steady state at 2 and 8 days of age, as measured by flow cytometry. n = 5 mice per group. p = 0.0010 (P2 vs. P8). Results are shown as mean ± s.e.m; **p < 0.01 by two-tailed Student’s t-test.
Extended Data Fig. 10 Neonatal DN T cells become aberrantly activated upon disruption of the PD-1/PD-L1 axis during MI.
a,b, Absolute counts of T cell populations (a) and neutrophils (b) in hearts of IgG- or αPD-L1-treated mice, at steady state and at d5 after MI performed at P3 (P3-MI), as measured by flow cytometry. n = 6 mice per group. p = 0.0495 (DN: IgG vs. αPD-L1) (a), p = 0.0004 (IgG vs. αPD-L1) (b). c, Heat map showing expression of marker genes in double-negative (DN), double-positive (DP), CD8+ and CD4+ T cell populations, as measured by scRNA-seq. d, Expression of DN T cell marker genes [log2(fold change) >0.6, p-adj <0.01] in DN, DP, CD8+ and CD4+ T cell populations (left) and their enriched gene ontology (GO) terms (right), as measured by scRNA-seq. A complete gene list is provided in Supplementary Data 3. Red shaded area denotes p-value (-Log10 (p-value)) by two-sided Wilcoxon Rank Sum test. e, Volcano plot depicting differentially expressed genes between Pd-1-/- DN T cells and Pd-1+/- DN T cells upon MI. p-value < 0.01, two-sided Wilcoxon Rank Sum test. A complete gene list is provided in Supplementary Data 4. f, Gene ontology (GO) terms enriched in genes upregulated in Pd-1-/- MI DN T cells vs Pd-1+/- MI DN T cells. Red shaded area denotes p-value (-Log10 (p-value)) by two-sided Wilcoxon Rank Sum test. g, NK1.1 expression in DN T cells in the heart of Pd-1+/- and Pd-1-/- mice at d5 after MI performed at P2 (P2-MI), as measured by flow cytometry. Results are shown as mean ± s.e.m; *p < 0.05, ***p < 0.001 by two-way ANOVA with Šidák correction on multiple comparisons (a) or two-tailed Student’s t-test (b).
Supplementary information
Supplementary Information
Supplementary Tables 1–4 and captions for Supplementary Data files.
Supplementary Data
Supplementary Data 1–4.
Source data
Source Data Fig. 1
Statistical source data.
Source Data Fig. 2
Statistical source data.
Source Data Fig. 3
Statistical source data.
Source Data Fig. 4
Statistical source data.
Source Data Fig. 5
Statistical source data.
Source Data Extended Data Fig. 1
Statistical source data.
Source Data Extended Data Fig. 3
Statistical source data.
Source Data Extended Data Fig. 6
Statistical source data.
Source Data Extended Data Fig. 8
Statistical source data.
Source Data Extended Data Fig. 9
Statistical source data.
Source Data Extended Data Fig. 10
Statistical source data.
Rights and permissions
About this article
Cite this article
Vargas Aguilar, S., Cui, M., Tan, W. et al. The PD-1–PD-L1 pathway maintains an immunosuppressive environment essential for neonatal heart regeneration. Nat Cardiovasc Res 3, 389–402 (2024). https://doi.org/10.1038/s44161-024-00447-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s44161-024-00447-7
This article is cited by
-
PD-1–PD-L1 immunomodulatory pathway regulates cardiac regeneration
Nature Cardiovascular Research (2024)