Abstract
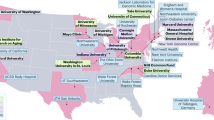
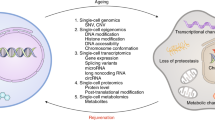
Cellular senescence is a well-established driver of aging and age-related diseases. There are many challenges to mapping senescent cells in tissues such as the absence of specific markers and their relatively low abundance and vast heterogeneity. Single-cell technologies have allowed unprecedented characterization of senescence; however, many methodologies fail to provide spatial insights. The spatial component is essential, as senescent cells communicate with neighboring cells, impacting their function and the composition of extracellular space. The Cellular Senescence Network (SenNet), a National Institutes of Health (NIH) Common Fund initiative, aims to map senescent cells across the lifespan of humans and mice. Here, we provide a comprehensive review of the existing and emerging methodologies for spatial imaging and their application toward mapping senescent cells. Moreover, we discuss the limitations and challenges inherent to each technology. We argue that the development of spatially resolved methods is essential toward the goal of attaining an atlas of senescent cells.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Gorgoulis, V. et al. Cellular senescence: defining a path forward. Cell 179, 813–827 (2019).
Robbins, P. D. et al. Senolytic drugs: reducing senescent cell viability to extend health span. Annu. Rev. Pharmacol. Toxicol. 61, 779–803 (2021).
Demaria, M. et al. An essential role for senescent cells in optimal wound healing through secretion of PDGF-AA. Dev. Cell 31, 722–733 (2014).
Born, E. et al. Eliminating senescent cells can promote pulmonary hypertension development and progression. Circulation 147, 650–666 (2023).
Reyes, N. S. et al. Sentinel p16INK4a+ cells in the basement membrane form a reparative niche in the lung. Science 378, 192–201 (2022).
Severino, J., Allen, R. G., Balin, S., Balin, A. & Cristofalo, V. J. Is β-galactosidase staining a marker of senescence in vitro and in vivo? Exp. Cell Res. 257, 162–171 (2000).
Hall, B. M. et al. Aging of mice is associated with p16Ink4a- and β-galactosidase-positive macrophage accumulation that can be induced in young mice by senescent cells. Aging 8, 1294–1315 (2016).
Ogrodnik, M., Salmonowicz, H., Jurk, D. & Passos, J. F. Expansion and cell-cycle arrest: common denominators of cellular senescence. Trends Biochem. Sci. 44, 996–1008 (2019).
Biran, A. et al. Quantitative identification of senescent cells in aging and disease. Aging Cell 16, 661–671 (2017).
Tabula Muris, C. et al. Single-cell transcriptomics of 20 mouse organs creates a Tabula Muris. Nature 562, 367–372 (2018).
Tabula Sapiens, C. et al. The Tabula Sapiens: a multiple-organ, single-cell transcriptomic atlas of humans. Science 376, eabl4896 (2022).
Zhang, X. et al. Characterization of cellular senescence in aging skeletal muscle. Nat. Aging 2, 601–615 (2022).
Ogrodnik, M. et al. Whole-body senescent cell clearance alleviates age-related brain inflammation and cognitive impairment in mice. Aging Cell 20, e13296 (2021).
Acosta, J. C. et al. A complex secretory program orchestrated by the inflammasome controls paracrine senescence. Nat. Cell Biol. 15, 978–990 (2013).
Xu, M. et al. Senolytics improve physical function and increase lifespan in old age. Nat. Med. 24, 1246–1256 (2018).
Lagnado, A. et al. Neutrophils induce paracrine telomere dysfunction and senescence in ROS-dependent manner. EMBO J. 40, e106048 (2021).
Laberge, R. M., Awad, P., Campisi, J. & Desprez, P. Y. Epithelial–mesenchymal transition induced by senescent fibroblasts. Cancer Microenviron. 5, 39–44 (2012).
Krizhanovsky, V. et al. Senescence of activated stellate cells limits liver fibrosis. Cell 134, 657–667 (2008).
Lee, P. J. et al. NIH SenNet Consortium to map senescent cells throughout the human lifespan to understand physiological health. Nat. Aging 2, 1090–1100 (2022).
Dimri, G. P. et al. A biomarker that identifies senescent human cells in culture and in aging skin in vivo. Proc. Natl Acad. Sci. USA 92, 9363–9367 (1995).
Georgakopoulou, E. A. et al. Specific lipofuscin staining as a novel biomarker to detect replicative and stress-induced senescence. A method applicable in cryo-preserved and archival tissues. Aging 5, 37–50 (2013).
Wang, C. et al. DNA damage response and cellular senescence in tissues of aging mice. Aging Cell 8, 311–323 (2009).
Freund, A., Laberge, R. M., Demaria, M. & Campisi, J. Lamin B1 loss is a senescence-associated biomarker. Mol. Biol. Cell 23, 2066–2075 (2012).
Davalos, A. R. et al. p53-dependent release of alarmin HMGB1 is a central mediator of senescent phenotypes. J. Cell Biol. 201, 613–629 (2013).
Sun, J. et al. A Glb1-2A-mCherry reporter monitors systemic aging and predicts lifespan in middle-aged mice. Nat. Commun. 13, 7028 (2022).
Hewitt, G. et al. Telomeres are favoured targets of a persistent DNA damage response in ageing and stress-induced senescence. Nat. Commun. 3, 708 (2012).
Herbig, U., Ferreira, M., Condel, L., Carey, D. & Sedivy, J. M. Cellular senescence in aging primates. Science 311, 1257 (2006).
Ogrodnik, M. et al. Cellular senescence drives age-dependent hepatic steatosis. Nat. Commun. 8, 15691 (2017).
Farr, J. N. et al. Targeting cellular senescence prevents age-related bone loss in mice. Nat. Med. 23, 1072–1079 (2017).
Swanson, E. C., Manning, B., Zhang, H. & Lawrence, J. B. Higher-order unfolding of satellite heterochromatin is a consistent and early event in cell senescence. J. Cell Biol. 203, 929–942 (2013).
Ogrodnik, M. et al. Obesity-induced cellular senescence drives anxiety and impairs neurogenesis. Cell Metab. 29, 1061–1077 (2019).
Valli, J. et al. Seeing beyond the limit: a guide to choosing the right super-resolution microscopy technique. J. Biol. Chem. 297, 100791 (2021).
Boumendil, C., Hari, P., Olsen, K. C. F., Acosta, J. C. & Bickmore, W. A. Nuclear pore density controls heterochromatin reorganization during senescence. Genes Dev. 33, 144–149 (2019).
Rodier, F. et al. DNA-SCARS: distinct nuclear structures that sustain damage-induced senescence growth arrest and inflammatory cytokine secretion. J. Cell Sci. 124, 68–81 (2011).
Anderson, R. et al. Length-independent telomere damage drives post-mitotic cardiomyocyte senescence. EMBO J. 38, e100492 (2019).
Rocha, A., Dalgarno, A. & Neretti, N. The functional impact of nuclear reorganization in cellular senescence. Brief. Funct. Genomics 21, 24–34 (2022).
Ricci, M. A., Manzo, C., García-Parajo, M. F., Lakadamyali, M. & Cosma, M. P. Chromatin fibers are formed by heterogeneous groups of nucleosomes in vivo. Cell 160, 1145–1158 (2015).
Shah, P. P. et al. Lamin B1 depletion in senescent cells triggers large-scale changes in gene expression and the chromatin landscape. Genes Dev. 27, 1787–1799 (2013).
De Cecco, M. et al. Genomes of replicatively senescent cells undergo global epigenetic changes leading to gene silencing and activation of transposable elements. Aging Cell 12, 247–256 (2013).
Chandra, T. et al. Global reorganization of the nuclear landscape in senescent cells. Cell Rep. 10, 471–483 (2015).
Sati, S. et al. 4D genome rewiring during oncogene-induced and replicative senescence. Mol. Cell 78, 522–538 (2020).
Hickey, J. W. et al. Spatial mapping of protein composition and tissue organization: a primer for multiplexed antibody-based imaging. Nat. Methods 19, 284–295 (2022).
Taylor, M. J., Liyu, A., Vertes, A. & Anderton, C. R. Ambient single-cell analysis and native tissue imaging using laser-ablation electrospray ionization mass spectrometry with increased spatial resolution. J. Am. Soc. Mass Spectrom. 32, 2490–2494 (2021).
Buczak, K. et al. Spatially resolved analysis of FFPE tissue proteomes by quantitative mass spectrometry. Nat. Protoc. 15, 2956–2979 (2020).
Lin, J. R., Fallahi-Sichani, M., Chen, J. Y. & Sorger, P. K. Cyclic immunofluorescence (CycIF), a highly multiplexed method for single-cell imaging. Curr. Protoc. Chem. Biol. 8, 251–264 (2016).
Gut, G., Herrmann, M. D. & Pelkmans, L. Multiplexed protein maps link subcellular organization to cellular states. Science 361, eaar7042 (2018).
Stallaert, W. et al. The structure of the human cell cycle. Cell Syst. 13, 230–240 (2022).
Eng, C.-H. L. et al. Transcriptome-scale super-resolved imaging in tissues by RNA seqFISH+. Nature 568, 235–239 (2019).
Xia, C., Fan, J., Emanuel, G., Hao, J. & Zhuang, X. Spatial transcriptome profiling by MERFISH reveals subcellular RNA compartmentalization and cell cycle-dependent gene expression. Proc. Natl Acad. Sci. USA 116, 19490–19499 (2019).
He, S. et al. High-plex imaging of RNA and proteins at subcellular resolution in fixed tissue by spatial molecular imaging. Nat. Biotechnol. 40, 1794–1806 (2022).
Stringer, C., Wang, T., Michaelos, M. & Pachitariu, M. Cellpose: a generalist algorithm for cellular segmentation. Nat. Methods 18, 100–106 (2021).
Kiss, T. et al. Spatial transcriptomic analysis reveals inflammatory foci defined by senescent cells in the white matter, hippocampi and cortical grey matter in the aged mouse brain. GeroScience 44, 661–681 (2022).
Xu, P. et al. The landscape of human tissue and cell type specific expression and co-regulation of senescence genes. Mol. Neurodegener. 17, 5 (2022).
Tower, R. J. et al. Spatial transcriptomics reveals metabolic changes underly age-dependent declines in digit regeneration. eLife 11, e71542 (2022).
Gracia Villacampa, E. et al. Genome-wide spatial expression profiling in formalin-fixed tissues. Cell Genom. 1, 100065 (2021).
Merritt, C. R. et al. Multiplex digital spatial profiling of proteins and RNA in fixed tissue. Nat. Biotechnol. 38, 586–599 (2020).
Liu, Y. et al. High-spatial-resolution multi-omics sequencing via deterministic barcoding in tissue. Cell 183, 1665–1681 (2020).
Liu, Y. et al. High-plex protein and whole transcriptome co-mapping at cellular resolution with spatial CITE-seq. Nat. Biotechnol., https://doi.org/10.1038/s41587-023-01676-0 (2023).
Deng, Y. et al. Spatial-CUT&Tag: spatially resolved chromatin modification profiling at the cellular level. Science 375, 681–686 (2022).
Liu, Y., Enninful, A., Deng, Y. & Fan, R. Spatial transcriptome sequencing of FFPE tissues at cellular level. Preprint at bioRxiv https://doi.org/10.1101/2020.10.13.338475 (2020).
Cho, C.-S. et al. Microscopic examination of spatial transcriptome using Seq-Scope. Cell 184, 3559–3572 (2021).
Chen, A. et al. Spatiotemporal transcriptomic atlas of mouse organogenesis using DNA nanoball-patterned arrays. Cell 185, 1777–1792 (2022).
Fu, X. et al. Polony gels enable amplifiable DNA stamping and spatial transcriptomics of chronic pain. Cell 185, 4621–4633 (2022).
Zhang, W. et al. Identification of cell types in multiplexed in situ images by combining protein expression and spatial information using CELESTA. Nat. Methods 19, 759–769 (2022).
Heckenbach, I. et al. Nuclear morphology is a deep learning biomarker of cellular senescence. Nat. Aging 2, 742–755 (2022).
Sage, D. & Unser, M. A. Teaching image-processing programming in Java. IEEE Signal Process. Mag. 20, 43–52 (2003).
Bannon, D. et al. DeepCell Kiosk: scaling deep learning-enabled cellular image analysis with Kubernetes. Nat. Methods 18, 43–45 (2021).
Schmidt, U., Weigert, M., Broaddus, C. & Myers, G. Cell detection with star-convex polygons. In Medical Image Computing and Computer Assisted Intervention – MICCAI 2018 (MICCAI 2018. Lecture Notes in Computer Science), vol 11071) (eds Frangi, A. et al.) 265–273 (Springer, 2018).
Ronneberger, O., Fischer, P. & Brox, T. U-Net: convolutional networks for biomedical image segmentation. In Medical Image Computing and Computer-Assisted Intervention—MICCAI 2015 (MICCAI 2015. Lecture Notes in Computer Science, vol 9351) (eds. Navab, N. et al.) 234–241 (Springer, 2015).
Hollandi, R. et al. nucleAIzer: a parameter-free deep learning framework for nucleus segmentation using image style transfer. Cell Syst. 10, 453–458 (2020).
Dayao, M. T., Brusko, M., Wasserfall, C. & Bar-Joseph, Z. Membrane marker selection for segmenting single cell spatial proteomics data. Nat. Commun. 13, 1999 (2022).
Dimopoulos, S., Mayer, C. E., Rudolf, F. & Stelling, J. Accurate cell segmentation in microscopy images using membrane patterns. Bioinformatics 30, 2644–2651 (2014).
Czech, E., Aksoy, B. A., Aksoy, P. & Hammerbacher, J. Cytokit: a single-cell analysis toolkit for high dimensional fluorescent microscopy imaging. BMC Bioinformatics 20, 448 (2019).
Petukhov, V. et al. Cell segmentation in imaging-based spatial transcriptomics. Nat. Biotechnol. 40, 345–354 (2022).
Qian, X. et al. Probabilistic cell typing enables fine mapping of closely related cell types in situ. Nat. Methods 17, 101–106 (2020).
Littman, R. et al. Joint cell segmentation and cell type annotation for spatial transcriptomics. Mol. Syst. Biol. 17, e10108 (2021).
Park, J. et al. Cell segmentation-free inference of cell types from in situ transcriptomics data. Nat. Commun. 12, 3545 (2021).
Li, D. et al. TraSig: inferring cell–cell interactions from pseudotime ordering of scRNA-seq data. Genome Biol. 23, 73 (2022).
Ding, J., Sharon, N. & Bar-Joseph, Z. Temporal modelling using single-cell transcriptomics. Nat. Rev. Genet. 23, 355–368 (2022).
Song, Q., Wang, J. & Bar-Joseph, Z. scSTEM: clustering pseudotime ordered single-cell data. Genome Biol. 23, 150 (2022).
Li, D., Ding, J. & Bar-Joseph, Z. Identifying signaling genes in spatial single-cell expression data. Bioinformatics 37, 968–975 (2021).
Yuan, Y. & Bar-Joseph, Z. GCNG: graph convolutional networks for inferring gene interaction from spatial transcriptomics data. Genome Biol. 21, 300 (2020).
Rodriques, S. G. et al. Slide-seq: a scalable technology for measuring genome-wide expression at high spatial resolution. Science 363, 1463–1467 (2019).
Elosua-Bayes, M., Nieto, P., Mereu, E., Gut, I. & Heyn, H. SPOTlight: seeded NMF regression to deconvolute spatial transcriptomics spots with single-cell transcriptomes. Nucleic Acids Res. 49, e50 (2021).
Cable, D. M. et al. Robust decomposition of cell type mixtures in spatial transcriptomics. Nat. Biotechnol. 40, 517–526 (2022).
Biancalani, T. et al. Deep learning and alignment of spatially resolved single-cell transcriptomes with Tangram. Nat. Methods 18, 1352–1362 (2021).
Ma, Y. & Zhou, X. Spatially informed cell-type deconvolution for spatial transcriptomics. Nat. Biotechnol. 40, 1349–1359 (2022).
Chu, T., Wang, Z., Pe’er, D. & Danko, C. G. Cell type and gene expression deconvolution with BayesPrism enables Bayesian integrative analysis across bulk and single-cell RNA sequencing in oncology. Nat. Cancer 3, 505–517 (2022).
Han, R., Luo, Y., Wang, M. & Zhang, A. R. Exact clustering in tensor block model: statistical optimality and computational limit. J. R. Stat. Soc. B Stat. Methodol. 84, 1666–1698 (2022).
Wu, M., Huang, J. & Ma, S. Identifying gene–gene interactions using penalized tensor regression. Stat. Med. 37, 598–610 (2018).
Burlingame, E. A. et al. SHIFT: speedy histological-to-immunofluorescent translation of a tumor signature enabled by deep learning. Sci. Rep. 10, 17507 (2020).
Zeira, R., Land, M., Strzalkowski, A. & Raphael, B. J. Alignment and integration of spatial transcriptomics data. Nat. Methods 19, 567–575 (2022).
Palla, G. et al. Squidpy: a scalable framework for spatial omics analysis. Nat. Methods 19, 171–178 (2022).
Tian, T., Yang, Z. & Li, X. Tissue clearing technique: recent progress and biomedical applications. J. Anat. 238, 489–507 (2021).
Kiemen, A. L. et al. CODA: quantitative 3D reconstruction of large tissues at cellular resolution. Nat. Methods 19, 1490–1499 (2022).
Sandkühler, R., Jud, C., Andermatt, S. & Cattin, P. C. AirLab: Autograd Image Registration Laboratory. Preprint at arXiv https://doi.org/10.48550/arXiv.1806.09907 (2018).
Klein, S., Staring, M., Murphy, K., Viergever, M. A. & Pluim, J. P. elastix: a toolbox for intensity-based medical image registration. IEEE Trans. Med. Imaging 29, 196–205 (2010).
Chiaruttini, N. et al. An open-source whole slide image registration workflow at cellular precision using Fiji, QuPath and elastix. Front. Comput. Sci. 3, https://doi.org/10.3389/fcomp.2021.780026 (2022).
Kusumoto, D. et al. Anti-senescent drug screening by deep learning-based morphology senescence scoring. Nat. Commun. 12, 257 (2021).
Wang, X. et al. Three-dimensional intact-tissue sequencing of single-cell transcriptional states. Science 361, eaat5691 (2018).
Black, S. et al. CODEX multiplexed tissue imaging with DNA-conjugated antibodies. Nat. Protoc. 16, 3802–3835 (2021).
Keren, L. et al. MIBI-TOF: a multiplexed imaging platform relates cellular phenotypes and tissue structure. Sci. Adv. 5, eaax5851 (2019).
Giesen, C. et al. Highly multiplexed imaging of tumor tissues with subcellular resolution by mass cytometry. Nat. Methods 11, 417–422 (2014).
Kinkhabwala, A. et al. MACSima imaging cyclic staining (MICS) technology reveals combinatorial target pairs for CAR T cell treatment of solid tumors. Sci. Rep. 12, 1911 (2022).
Hennig, C., Adams, N. & Hansen, G. A versatile platform for comprehensive chip-based explorative cytometry. Cytometry A 75, 362–370 (2009).
Lebrigand, K. et al. The spatial landscape of gene expression isoforms in tissue sections. Nucleic Acids Res. 51, e47 (2023).
Stickels, R. R. et al. Highly sensitive spatial transcriptomics at near-cellular resolution with Slide-seqV2. Nat. Biotechnol. 39, 313–319 (2021).
Vickovic, S. et al. High-definition spatial transcriptomics for in situ tissue profiling. Nat. Methods 16, 987–990 (2019).
Acknowledgements
This research was supported by the NIH Common Fund, through the Office of Strategic Coordination and the Office of the NIH Director under awards UG3CA268103 (J.F.P., A.B.L., S.V., D.J.), U54AG075931 (A.U.G., A.L.M., D.B., H.C., J.K.A., M.B., M. Königshoff., O.E., Q.H., T.K.), UG3CA268202 (N.N.), U54AG075941 (J.H.C.), UG3CA268091 (J.H.L.), UG3CA268096 (R.D., X.F.), U54AG075931 (Z.B.-J.), U24CA268108 (Z.B.-J.), U54AG076041 (L.J.N., A.C.N., E.L.T., S.P., F.R., G.B.), U54AG075936 (A.R.Z., Z.J.), U54AG079758 (P.D.A., Q.Z., R.A.P.), U54AG076040 (H.P., J.P.), U54 AG075932 (D.F., A.A.G., C.B., F.W., I.H.), UG3CA268112 (V.S., M. Kumar.). We thank A. Impagliazzo for her work on the illustrations.
Author information
Authors and Affiliations
Contributions
All authors contributed to writing the manuscript and reviewed and approved of its submission for publication. A.U.G., M.B., V.S., O.E., J.K.A. and J.F.P. contributed to the Main; J.F.P., D.J., Q.H., D.B., S.V., N.N., M.S., J.H., J.P. and H.P. contributed to Low-plex imaging methods; G.B., F.R., A.C.N., A.B.L., M. Kumar., L.J.N., S.P. and E.L.T. contributed to High-plex imaging methods; A.E., J.H.L., T.K., A.L.M., X.F., R.D., A.C.N., P.D.A., Q.Z. and R.A.P. contributed to Spatial transcriptomics; J.H.C., H.C., T.P., Z.J., Z.B.-J and A.A.G. contributed to Image data analysis of senescent cells; A.A.G., C.B., D.F., F.W., M.S.K. and I.H. contributed to Use of deep learning methods to identify senescent cells. J.F.P. designed and coordinated the writing of the Review.
Corresponding author
Ethics declarations
Competing interests
J.H.L. is an inventor on pending patent applications related to Seq-Scope. All other authors declare no competing interests.
Peer review
Peer review information
Nature Aging thanks Jesus Gil, Kristina Kirschner, and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Gurkar, A.U., Gerencser, A.A., Mora, A.L. et al. Spatial mapping of cellular senescence: emerging challenges and opportunities. Nat Aging 3, 776–790 (2023). https://doi.org/10.1038/s43587-023-00446-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s43587-023-00446-6
This article is cited by
-
Global Healthspan Summit 2023: closing the gap between healthspan and lifespan
Nature Aging (2024)
-
Telomeres, cellular senescence, and aging: past and future
Biogerontology (2024)
-
Molecular hallmarks of ageing in amyotrophic lateral sclerosis
Cellular and Molecular Life Sciences (2024)
-
Oncogene-induced senescence in meningiomas—an immunohistochemical study
Journal of Neuro-Oncology (2024)
-
Senolytic treatment does not mitigate oxidative stress-induced muscle atrophy but improves muscle force generation in CuZn superoxide dismutase knockout mice
GeroScience (2024)