Abstract
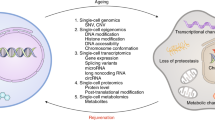
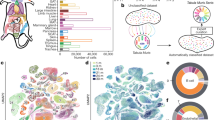
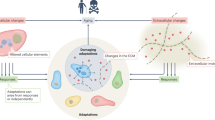
Organismal aging is often characterized as a steady, monotonic decline of organ and tissue function. However, recent studies indicate spatial and temporal variations of aging rates across the lifespan. We consider these variations from the perspective of underlying cellular changes. Cells in certain tissues may age earlier and produce signals that accelerate the aging of other cells, locally or distantly, acting as drivers for organismal aging and leading to a lack of synchronous aging between tissues. As cells adopt new homeostatic states, cellular aging can be viewed, at least in part, as a quantal process we refer to as digital aging. Analog declines of tissue function with age may be the sum of underlying digital events. Cellular aging, digital or otherwise, is not uniform across time or space within organisms or between organisms of the same species. Advanced systems-level and single-cell methodologies will refine our understanding of cell and tissue aging, and how these processes integrate to produce the complexities of individual, organismal aging.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Flatt, T. & Partridge, L. Horizons in the evolution of aging. BMC Biol. 16, 93 (2018).
Medawar, P. B. An Unsolved Problem of Biology (H. K. Lewis, 1952).
Williams, G. C. Pleiotropy, natural selection, and the evolution of senescence. Evolution 11, 398–411 (1957).
Kirkwood, T. B. Evolution of ageing. Nature 270, 301–304 (1977).
Kennedy, B. K. et al. Geroscience: linking aging to chronic disease. Cell 159, 709–713 (2014).
Gladyshev, V. N. The ground zero of organismal life and aging. Trends Mol. Med. 27, 11–19 (2020).
Rose, M. R. Adaptation, aging, and genomic information. Aging 1, 444–450 (2009).
Gladyshev, V. N. Aging: progressive decline in fitness due to the rising deleteriome adjusted by genetic, environmental, and stochastic processes. Aging Cell 15, 594–602 (2016).
Tissenbaum, H. A. & Guarente, L. Model organisms as a guide to mammalian aging. Dev. Cell 2, 9–19 (2002).
Piper, M. D. W. & Partridge, L. Drosophila as a model for ageing. Biochim. Biophys. Acta Mol. Basis Dis. 1864, 2707–2717 (2018).
Singh, P. P., Demmitt, B. A., Nath, R. D. & Brunet, A. The genetics of aging: a vertebrate perspective. Cell 177, 200–220 (2019).
Gonzalez-Freire, M. et al. The road ahead for health and lifespan interventions. Ageing Res. Rev. 59, 101037 (2020).
Fontana, L., Partridge, L. & Longo, V. D. Extending healthy life span—from yeast to humans. Science 328, 321–326 (2010).
Hayflick, L. Entropy explains aging, genetic determinism explains longevity, and undefined terminology explains misunderstanding both. PLoS Genet. 3, e220 (2007).
Goodell, M. A. & Rando, T. A. Stem cells and healthy aging. Science 350, 1199–1204 (2015).
Brett, J. O. & Rando, T. A. Alive and well? Exploring disease by studying lifespan. Curr. Opin. Genet. Dev. 26, 33–40 (2014).
Bansal, A., Zhu, L. J., Yen, K. & Tissenbaum, H. A. Uncoupling lifespan and healthspan in Caenorhabditis elegans longevity mutants. Proc. Natl Acad. Sci. USA 112, E277–E286 (2015).
Pearl, R. The Rate of Living (University of London Press, 1928).
Rando, T. A. Stem cells, ageing and the quest for immortality. Nature 441, 1080–1086 (2006).
Butler, P. G., Wanamaker, A. D., Scourse, J. D., Richardson, C. A. & Reynolds, D. J. Variability of marine climate on the North Icelandic Shelf in a 1357-year proxy archive based on growth increments in the bivalve Artica islandica. Palaeogeogr. Palaeoclimatol. Palaeoecol. 373, 141–151 (2013).
Ailshire, J. A., Beltran-Sanchez, H. & Crimmins, E. M. Becoming centenarians: disease and functioning trajectories of older US adults as they survive to 100. J. Gerontol. A Biol. Sci. Med. Sci. 70, 193–201 (2015).
Kirkwood, T. B. et al. What accounts for the wide variation in life span of genetically identical organisms reared in a constant environment? Mech. Ageing Dev. 126, 439–443 (2005).
Khan, S. S., Singer, B. D. & Vaughan, D. E. Molecular and physiological manifestations and measurement of aging in humans. Aging Cell 16, 624–633 (2017).
Bocklandt, S. et al. Epigenetic predictor of age. PLoS ONE 6, e14821 (2011).
Stubbs, T. M. et al. Multi-tissue DNA methylation age predictor in mouse. Genome Biol. 18, 68 (2017).
Heidinger, B. J. et al. Telomere length in early life predicts lifespan. Proc. Natl Acad. Sci. USA 109, 1743–1748 (2012).
Steenstrup, T. et al. Telomeres and the natural lifespan limit in humans. Aging 9, 1130–1142 (2017).
Lehallier, B. et al. Undulating changes in human plasma proteome profiles across the lifespan. Nat. Med. 25, 1843–1850 (2019).
Chen, W. et al. Three-dimensional human facial morphologies as robust aging markers. Cell Res. 25, 574–587 (2015).
Herndon, L. A. et al. Stochastic and genetic factors influence tissue-specific decline in ageing C. elegans. Nature 419, 808–814 (2002).
Tricoire, H. & Rera, M. A new, discontinuous 2 phases of aging model: lessons from Drosophila melanogaster. PLoS ONE 10, e0141920 (2015).
Schaum, N. et al. Ageing hallmarks exhibit organ-specific temporal signatures. Nature 583, 596–602 (2020).
Isildak, U., Somel, M., Thornton, J. M. & Donertas, H. M. Temporal changes in the gene expression heterogeneity during brain development and aging. Sci. Rep. 10, 4080 (2020).
Horvath, S. DNA methylation age of human tissues and cell types. Genome Biol. 14, R115 (2013).
Li, X. et al. Longitudinal trajectories, correlations and mortality associations of nine biological ages across 20-years follow-up. eLife 9, e51507 (2020).
Marquez, E. J. et al. Sexual-dimorphism in human immune system aging. Nat. Commun. 11, 751 (2020).
Holzscheck, N. et al. Multi-omics network analysis reveals distinct stages in the human aging progression in epidermal tissue. Aging 12, 12393–12409 (2020).
Edde, M., Leroux, G., Altena, E. & Chanraud, S. Functional brain connectivity changes across the human life span: from fetal development to old age. J. Neurosci. Res. 99, 236–262 (2020).
Biteau, B., Hochmuth, C. E. & Jasper, H. JNK activity in somatic stem cells causes loss of tissue homeostasis in the aging Drosophila gut. Cell Stem Cell 3, 442–455 (2008).
Clark, R. I. et al. Distinct shifts in microbiota composition during Drosophila aging impair intestinal function and drive mortality. Cell Rep. 12, 1656–1667 (2015).
Demontis, F. & Perrimon, N. FOXO/4E-BP signaling in Drosophila muscles regulates organism-wide proteostasis during aging. Cell 143, 813–825 (2010).
Owusu-Ansah, E., Song, W. & Perrimon, N. Muscle mitohormesis promotes longevity via systemic repression of insulin signaling. Cell 155, 699–712 (2013).
Ori, A. et al. Integrated transcriptome and proteome analyses reveal organ-specific proteome deterioration in old rats. Cell Syst. 1, 224–237 (2015).
Ahadi, S. et al. Personal aging markers and ageotypes revealed by deep longitudinal profiling. Nat. Med. 26, 83–90 (2020).
Bahar, R. et al. Increased cell-to-cell variation in gene expression in ageing mouse heart. Nature 441, 1011–1014 (2006).
Arrojo e Drigo, R. et al. Age mosaicism across multiple scales in adult tissues. Cell Metab. 30, 343–351 (2019).
Martinez-Jimenez, C. P. et al. Aging increases cell-to-cell transcriptional variability upon immune stimulation. Science 355, 1433–1436 (2017).
Enge, M. et al. Single-cell analysis of human pancreas reveals transcriptional signatures of aging and somatic mutation patterns. Cell 171, 321–330 (2017).
Tabula Muris Consortium. A single-cell transcriptomic atlas characterizes ageing tissues in the mouse. Nature 583, 590–595 (2020).
Ma, S. et al. Caloric restriction reprograms the single-cell transcriptional landscape of Rattus norvegicus aging. Cell 180, 984–1001 (2020).
Kimmel, J. C. et al. Murine single-cell RNA-seq reveals cell-identity- and tissue-specific trajectories of aging. Genome Res. 29, 2088–2103 (2019).
Goronzy, J. J. & Weyand, C. M. T cell development and receptor diversity during aging. Curr. Opin. Immunol. 17, 468–475 (2005).
Tower, J. Programmed cell death in aging. Ageing Res. Rev. 23, 90–100 (2015).
Gorgoulis, V. et al. Cellular senescence: defining a path forward. Cell 179, 813–827 (2019).
Hernandez-Segura, A. et al. Unmasking transcriptional heterogeneity in senescent cells. Curr. Biol. 27, 2652–2660 (2017).
Robbins P. D. et al. Senolytic drugs: reducing senescent cell viability to extend health span. Annu. Rev. Pharmacol. Toxicol. https://doi.org/10.1146/annurev-pharmtox-050120-105018 (2020).
Tyshkovskiy, A. et al. Identification and application of gene expression signatures associated with lifespan extension. Cell Metab. 30, 573–593 (2019).
Acosta, J. C. et al. A complex secretory program orchestrated by the inflammasome controls paracrine senescence. Nat. Cell Biol. 15, 978–990 (2013).
Davis, A. A., Leyns, C. E. G. & Holtzman, D. M. Intercellular spread of protein aggregates in neurodegenerative disease. Annu. Rev. Cell Dev. Biol. 34, 545–568 (2018).
Prusiner, S. B. Biology and genetics of prions causing neurodegeneration. Annu. Rev. Genet. 47, 601–623 (2013).
Guo, J. L. & Lee, V. M. Cell-to-cell transmission of pathogenic proteins in neurodegenerative diseases. Nat. Med. 20, 130–138 (2014).
Morimoto, R. I.Cell-nonautonomous regulation of proteostasis in aging and disease.Cold Spring Harb. Perspect. Biol. 12, a034074 (2020).
Conboy, I. M. et al. Rejuvenation of aged progenitor cells by exposure to a young systemic environment. Nature 433, 760–764 (2005).
Brack, A. S. et al. Increased Wnt signaling during aging alters muscle stem cell fate and increases fibrosis. Science 317, 807–810 (2007).
Villeda, S. A. et al. The ageing systemic milieu negatively regulates neurogenesis and cognitive function. Nature 477, 90–94 (2011).
Ruckh, J. M. et al. Rejuvenation of regeneration in the aging central nervous system. Cell Stem Cell 10, 96–103 (2012).
Salpeter, S. J. et al. Systemic regulation of the age-related decline of pancreatic β-cell replication. Diabetes 62, 2843–2848 (2013).
Katsimpardi, L. et al. Vascular and neurogenic rejuvenation of the aging mouse brain by young systemic factors. Science 344, 630–634 (2014).
Castellano, J. M. et al. Human umbilical cord plasma proteins revitalize hippocampal function in aged mice. Nature 544, 488–492 (2017).
Chen, M. B. et al. Brain endothelial cells are exquisite sensors of age-related circulatory cues. Cell Rep. 30, 4418–4432 (2020).
Niccoli, T. & Partridge, L. Ageing as a risk factor for disease. Curr. Biol. 22, R741–R752 (2012).
Keating, N. L., Norredam, M., Landrum, M. B., Huskamp, H. A. & Meara, E. Physical and mental health status of older long-term cancer survivors. J. Am. Geriatr. Soc. 53, 2145–2152 (2005).
Kirkman, M. S. et al. Diabetes in older adults: a consensus report. J. Am. Geriatr. Soc. 60, 2342–2356 (2012).
High, K. P. et al. HIV and aging: state of knowledge and areas of critical need for research. A report to the NIH Office of AIDS Research by the HIV and Aging Working Group. J. Acquir. Immune Defic. Syndr. 60, S1–S18 (2012).
Longo, V. D. et al. Interventions to slow aging in humans: are we ready? Aging Cell 14, 497–510 (2015).
Schleit, J. et al. Molecular mechanisms underlying genotype-dependent responses to dietary restriction. Aging Cell 12, 1050–1061 (2013).
Rikke, B. A., Liao, C. Y., McQueen, M. B., Nelson, J. F. & Johnson, T. E. Genetic dissection of dietary restriction in mice supports the metabolic efficiency model of life extension. Exp. Gerontol. 45, 691–701 (2010).
Liao, C. Y., Rikke, B. A., Johnson, T. E., Diaz, V. & Nelson, J. F. Genetic variation in the murine lifespan response to dietary restriction: from life extension to life shortening. Aging Cell 9, 92–95 (2010).
Rando, T. A. & Chang, H. Y. Aging, rejuvenation, and epigenetic reprogramming: resetting the aging clock. Cell 148, 46–57 (2012).
Villeda, S. A. et al. Young blood reverses age-related impairments in cognitive function and synaptic plasticity in mice. Nat. Med. 20, 659–663 (2014).
Wyss-Coray, T. Ageing, neurodegeneration and brain rejuvenation. Nature 539, 180–186 (2016).
Ocampo, A. et al. In vivo amelioration of age-associated hallmarks by partial reprogramming. Cell 167, 1719–1733 (2016).
Sarkar, T. J. et al. Transient non-integrative expression of nuclear reprogramming factors promotes multifaceted amelioration of aging in human cells. Nat. Commun. 11, 1545 (2020).
Dobzhansky, T. Nothing in biology makes sense except in the light of evolution. Am. Biol. Teacher 35, 125–129 (1973).
Zhang, R., Chen, H. Z. & Liu, D. P. The four layers of aging. Cell Syst. 1, 180–186 (2015).
Acknowledgements
This work was supported by the Glenn Foundation for Medical Research, Nan Fung Life Sciences and the NOMIS Foundation.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information Nature Aging thanks Steven Austad, Vadim Gladyshev and Jing-Dong Jackie Han for their contribution to the peer review of this work.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Rando, T.A., Wyss-Coray, T. Asynchronous, contagious and digital aging. Nat Aging 1, 29–35 (2021). https://doi.org/10.1038/s43587-020-00015-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s43587-020-00015-1
This article is cited by
-
Tissue-specific profiling of age-dependent miRNAomic changes in Caenorhabditis elegans
Nature Communications (2024)
-
LKRSDH-dependent histone modifications of insulin-like peptide sites contribute to age-related circadian rhythm changes
Nature Communications (2024)
-
Nonlinear DNA methylation trajectories in aging male mice
Nature Communications (2024)
-
Review: myogenic and muscle toxicity targets of environmental methylmercury exposure
Archives of Toxicology (2024)
-
Emerging roles of mitochondrial functions and epigenetic changes in the modulation of stem cell fate
Cellular and Molecular Life Sciences (2024)