Abstract
Barth syndrome (BTHS) is a life-threatening genetic disorder with unknown pathogenicity caused by mutations in TAFAZZIN (TAZ) that affect remodeling of mitochondrial cardiolipin (CL). TAZ deficiency leads to accumulation of mono-lyso-CL (MLCL), which forms a peroxidase complex with cytochrome c (cyt c) capable of oxidizing polyunsaturated fatty acid-containing lipids. We hypothesized that accumulation of MLCL facilitates formation of anomalous MLCL–cyt c peroxidase complexes and peroxidation of polyunsaturated fatty acid phospholipids as the primary BTHS pathogenic mechanism. Using genetic, biochemical/biophysical, redox lipidomic and computational approaches, we reveal mechanisms of peroxidase-competent MLCL–cyt c complexation and increased phospholipid peroxidation in different TAZ-deficient cells and animal models and in pre-transplant biopsies from hearts of patients with BTHS. A specific mitochondria-targeted anti-peroxidase agent inhibited MLCL–cyt c peroxidase activity, prevented phospholipid peroxidation, improved mitochondrial respiration of TAZ-deficient C2C12 myoblasts and restored exercise endurance in a BTHS Drosophila model. Targeting MLCL–cyt c peroxidase offers therapeutic approaches to BTHS treatment.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
Data availability
Source data for Figs. 1–7 and Extended Data Figs. 4–8 are provided with the manuscript. Raw flow cytometry data can be accessed from the FlowRepository database (repository ID FR-FCM-Z6XG). MS data are available upon request. Source data are provided with this paper.
Code availability
No custom code was generated for this study.
References
Reid Thompson, W. et al. A phase 2/3 randomized clinical trial followed by an open-label extension to evaluate the effectiveness of elamipretide in Barth syndrome, a genetic disorder of mitochondrial cardiolipin metabolism. Genet. Med. 23, 471–478 (2021).
Dabner, L. et al. Treatment of Barth syndrome by cardiolipin manipulation (CARDIOMAN) with bezafibrate: protocol for a randomized placebo-controlled pilot trial conducted in the nationally commissioned Barth syndrome service. JMIR Res. Protoc. 10, e22533 (2021).
Kagan, V. E., Chu, C. T., Tyurina, Y. Y., Cheikhi, A. & Bayir, H. Cardiolipin asymmetry, oxidation and signaling. Chem. Phys. Lipids 179, 64–69 (2014).
Oemer, G. et al. Molecular structural diversity of mitochondrial cardiolipins. Proc. Natl Acad. Sci. USA 115, 4158–4163 (2018).
Maguire, J. J. et al. Known unknowns of cardiolipin signaling: the best is yet to come. Biochim. Biophys. Acta Mol. Cell Biol. Lipids 1862, 8–24 (2017).
Ren, M., Miller, P. C., Schlame, M. & Phoon, C. K. L. A critical appraisal of the tafazzin knockdown mouse model of Barth syndrome: what have we learned about pathogenesis and potential treatments? Am. J. Physiol. Heart Circ. Physiol. 317, H1183–H1193 (2019).
Schlame, M. & Greenberg, M. L. Biosynthesis, remodeling and turnover of mitochondrial cardiolipin. Biochim. Biophys. Acta Mol. Cell Biol. Lipids 1862, 3–7 (2017).
Claypool, S. M. & Koehler, C. M. The complexity of cardiolipin in health and disease. Trends Biochem. Sci. 37, 32–41 (2012).
Beranek, A. et al. Identification of a cardiolipin-specific phospholipase encoded by the gene CLD1 (YGR110W) in yeast. J. Biol. Chem. 284, 11572–11578 (2009).
Sparagna, G. C. & Lesnefsky, E. J. Cardiolipin remodeling in the heart. J. Cardiovasc. Pharmacol. 53, 290–301 (2009).
Ji, J. & Greenberg, M. L. Cardiolipin function in the yeast S. cerevisiae and the lessons learned for Barth syndrome. J. Inherit. Metab. Dis. 45, 60–71 (2022).
Li, M. et al. Activation of cytochrome C peroxidase function through coordinated foldon loop dynamics upon interaction with anionic lipids. J. Mol. Biol. 433, 167057 (2021).
Li, M. et al. Surface-binding to cardiolipin nanodomains triggers cytochrome c pro-apoptotic peroxidase activity via localized dynamics. Structure 27, 806–815 (2019).
Baile, M. G., Whited, K. & Claypool, S. M. Deacylation on the matrix side of the mitochondrial inner membrane regulates cardiolipin remodeling. Mol. Biol. Cell 24, 2008–2020 (2013).
Tuominen, E. K., Wallace, C. J. & Kinnunen, P. K. Phospholipid-cytochrome c interaction: evidence for the extended lipid anchorage. J. Biol. Chem. 277, 8822–8826 (2002).
Nantes, I. L., Zucchi, M. R., Nascimento, O. R. & Faljoni-Alario, A. Effect of heme iron valence state on the conformation of cytochrome c and its association with membrane interfaces. A CD and EPR investigation. J. Biol. Chem. 276, 153–158 (2001).
Kagan, V. E. et al. Cytochrome c acts as a cardiolipin oxygenase required for release of proapoptotic factors. Nat. Chem. Biol. 1, 223–232 (2005).
Battistuzzi, G. et al. Role of Met80 and Tyr67 in the low-pH conformational equilibria of cytochrome c. Biochemistry 51, 5967–5978 (2012).
Li, Y. et al. Cardiolipin-induced activation of pyruvate dehydrogenase links mitochondrial lipid biosynthesis to TCA cycle function. J. Biol. Chem. 294, 11568–11578 (2019).
Imai, M. et al. Investigation of the redox-dependent modulation of structure and dynamics in human cytochrome c. Biochem. Biophys. Res. Commun. 469, 978–984 (2016).
Matlahov, I. & van der Wel, P. C. A. Hidden motions and motion-induced invisibility: dynamics-based spectral editing in solid-state NMR. Methods 148, 123–135 (2018).
Andronesi, O. C. et al. Determination of membrane protein structure and dynamics by magic-angle-spinning solid-state NMR spectroscopy. J. Am. Chem. Soc. 127, 12965–12974 (2005).
Powell, G. L. & Marsh, D. Polymorphic phase behavior of cardiolipin derivatives studied by 31P NMR and X-ray diffraction. Biochemistry 24, 2902–2908 (1985).
Lou, W. et al. Genetic re-engineering of polyunsaturated phospholipid profile of Saccharomyces cerevisiae identifies a novel role for Cld1 in mitigating the effects of cardiolipin peroxidation. Biochim. Biophys. Acta Mol. Cell Biol. Lipids 1863, 1354–1368 (2018).
Horvath, S. E. & Daum, G. Lipids of mitochondria. Prog. Lipid Res. 52, 590–614 (2013).
Lou, W. et al. Loss of tafazzin results in decreased myoblast differentiation in C2C12 cells: a myoblast model of Barth syndrome and cardiolipin deficiency. Biochim. Biophys. Acta Mol. Cell Biol. Lipids 1863, 857–865 (2018).
Acehan, D. et al. Cardiac and skeletal muscle defects in a mouse model of human Barth syndrome. J. Biol. Chem. 286, 899–908 (2011).
Kobayashi, T. et al. A lipid associated with the antiphospholipid syndrome regulates endosome structure and function. Nature 392, 193–197 (1998).
Showalter, M. R. et al. The emerging and diverse roles of bis(monoacylglycero) phosphate lipids in cellular physiology and disease. Int. J. Mol. Sci. 21, 8067 (2020).
Atkinson, J. et al. A mitochondria-targeted inhibitor of cytochrome c peroxidase mitigates radiation-induced death. Nat. Commun. 2, 497 (2011).
Xu, Y. et al. A Drosophila model of Barth syndrome. Proc. Natl Acad. Sci. USA 103, 11584–11588 (2006).
Malhotra, A. et al. Role of calcium-independent phospholipase A2 in the pathogenesis of Barth syndrome. Proc. Natl Acad. Sci. USA 106, 2337–2341 (2009).
Creed, S. & McKenzie, M. Measurement of mitochondrial membrane potential with the fluorescent dye tetramethylrhodamine methyl ester (TMRM). Methods Mol. Biol. 1928, 69–76 (2019).
Pang, J., Bao, Y., Mitchell-Silbaugh, K., Veevers, J. & Fang, X. Barth syndrome cardiomyopathy: an update. Genes 13, 656 (2022).
Hornby, B. et al. Natural history comparison study to assess the efficacy of elamipretide in patients with Barth syndrome. Orphanet. J. Rare Dis. 17, 336 (2022).
Scott, R. A. & Mauk, A. G. Cytochrome c: A Multidisciplinary Approach. (University Science Books, 1996).
Alvarez-Paggi, D. et al. Multifunctional cytochrome c: learning new tricks from an old dog. Chem. Rev. 117, 13382–13460 (2017).
Kagan, V. E. et al. Cytochrome c/cardiolipin relations in mitochondria: a kiss of death. Free Radic. Biol. Med 46, 1439–1453 (2009).
Pinheiro, T. J. & Watts, A. Lipid specificity in the interaction of cytochrome c with anionic phospholipid bilayers revealed by solid-state 31P NMR. Biochemistry 33, 2451–2458 (1994).
Cassina, A. M. et al. Cytochrome c nitration by peroxynitrite. J. Biol. Chem. 275, 21409–21415 (2000).
Hannibal, L. et al. Alternative conformations of cytochrome c: structure, function, and detection. Biochemistry 55, 407–428 (2016).
Schweitzer-Stenner, R. Relating the multi-functionality of cytochrome c to membrane binding and structural conversion. Biophys. Rev. 10, 1151–1185 (2018).
Droghetti, E., Oellerich, S., Hildebrandt, P. & Smulevich, G. Heme coordination states of unfolded ferrous cytochrome C. Biophys. J. 91, 3022–3031 (2006).
Dunford, H. B. & Stillman, J. S. On the function and mechanism of action of peroxidases. Coord. Chem. Rev. 19, 187–251 (1976).
Basova, L. V. et al. Cardiolipin switch in mitochondria: shutting off the reduction of cytochrome c and turning on the peroxidase activity. Biochemistry 46, 3423–3434 (2007).
Belikova, N. A. et al. Peroxidase activity and structural transitions of cytochrome c bound to cardiolipin-containing membranes. Biochemistry 45, 4998–5009 (2006).
Barr, D. P. & Mason, R. P. Mechanism of radical production from the reaction of cytochrome c with organic hydroperoxides. An ESR spin trapping investigation. J. Biol. Chem. 270, 12709–12716 (1995).
Diederix, R. E. et al. Kinetic stability of the peroxidase activity of unfolded cytochrome c: heme degradation and catalyst inactivation by hydrogen peroxide. Inorg. Chem. 42, 7249–7257 (2003).
Chicco, A. J. & Sparagna, G. C. Role of cardiolipin alterations in mitochondrial dysfunction and disease. Am. J. Physiol. Cell Physiol. 292, C33–C44 (2007).
Amoscato, A. A. et al. Formation of protein adducts with hydroperoxy-PE electrophilic cleavage products during ferroptosis. Redox Biol. 63, 102758 (2023).
Claypool, S. M., McCaffery, J. M. & Koehler, C. M. Mitochondrial mislocalization and altered assembly of a cluster of Barth syndrome mutant tafazzins. J. Cell Biol. 174, 379–390 (2006).
Schlame, M. Cardiolipin synthesis for the assembly of bacterial and mitochondrial membranes. J. Lipid Res. 49, 1607–1620 (2008).
Ge, Y., Boopathy, S., Nguyen, T. H., Lugo, C. M. & Chao, L. H. Absence of cardiolipin from the outer leaflet of a mitochondrial inner membrane mimic restricts Opa1-mediated fusion. Front. Mol. Biosci. 8, 769135 (2021).
Duncan, A. L. Monolysocardiolipin (MLCL) interactions with mitochondrial membrane proteins. Biochem. Soc. Trans. 48, 993–1004 (2020).
Xu, Y. et al. Loss of protein association causes cardiolipin degradation in Barth syndrome. Nat. Chem. Biol. 12, 641–647 (2016).
Musatov, A. Contribution of peroxidized cardiolipin to inactivation of bovine heart cytochrome c oxidase. Free Radic. Biol. Med 41, 238–246 (2006).
Petrosillo, G., Casanova, G., Matera, M., Ruggiero, F. M. & Paradies, G. Interaction of peroxidized cardiolipin with rat-heart mitochondrial membranes: induction of permeability transition and cytochrome c release. FEBS Lett. 580, 6311–6316 (2006).
Clarke, S. L. et al. Barth syndrome. Orphanet. J. Rare Dis. 8, 23 (2013).
Gonzalvez, F. et al. Barth syndrome: cellular compensation of mitochondrial dysfunction and apoptosis inhibition due to changes in cardiolipin remodeling linked to tafazzin (TAZ) gene mutation. Biochim. Biophys. Acta 1832, 1194–1206 (2013).
Chu, C. T. et al. Cardiolipin externalization to the outer mitochondrial membrane acts as an elimination signal for mitophagy in neuronal cells. Nat. Cell Biol. 15, 1197–1205 (2013).
Zhang, J., Liu, X., Nie, J. & Shi, Y. Restoration of mitophagy ameliorates cardiomyopathy in Barth syndrome. Autophagy 18, 2134–2149 (2022).
Hsu, P. et al. Cardiolipin remodeling by TAZ/tafazzin is selectively required for the initiation of mitophagy. Autophagy 11, 643–652 (2015).
Hullin-Matsuda, F. et al. De novo biosynthesis of the late endosome lipid, bis(monoacylglycero)phosphate. J. Lipid Res. 48, 1997–2008 (2007).
Sohn, J. et al. A new murine model of Barth syndrome neutropenia links TAFAZZIN deficiency to increased ER stress-induced apoptosis. Blood Adv. 6, 2557–2577 (2022).
Esposti, M. D., Cristea, I. M., Gaskell, S. J., Nakao, Y. & Dive, C. Proapoptotic Bid binds to monolysocardiolipin, a new molecular connection between mitochondrial membranes and cell death. Cell Death Differ. 10, 1300–1309 (2003).
Human Protein Atlas https://www.proteinatlas.org/ENSG00000102125-TAZ/tissue
Houtkooper, R. H. et al. Cardiolipin and monolysocardiolipin analysis in fibroblasts, lymphocytes, and tissues using high-performance liquid chromatography-mass spectrometry as a diagnostic test for Barth syndrome. Anal. Biochem. 387, 230–237 (2009).
Cole, L. K. et al. Aberrant cardiolipin metabolism is associated with cognitive deficiency and hippocampal alteration in tafazzin knockdown mice. Biochim. Biophys. Acta Mol. Basis Dis. 1864, 3353–3367 (2018).
Olivar-Villanueva, M., Ren, M. & Phoon, C. K. L. Neurological & psychological aspects of Barth syndrome: clinical manifestations and potential pathogenic mechanisms. Mitochondrion 61, 188–195 (2021).
Reynolds, S. Successful management of Barth syndrome: a systematic review highlighting the importance of a flexible and multidisciplinary approach. J. Multidiscip. Health. 8, 345–358 (2015).
Raches, D. & Mazzocco, M. M. Emergence and nature of mathematical difficulties in young children with Barth syndrome. J. Dev. Behav. Pediatr. 33, 328–335 (2012).
Mazzocco, M. M., Henry, A. E. & Kelly, R. I. Barth syndrome is associated with a cognitive phenotype. J. Dev. Behav. Pediatr. 28, 22–30 (2007).
Damschroder, D., Reynolds, C. & Wessells, R. Drosophila tafazzin mutants have impaired exercise capacity. Physiol. Rep. 6, e13604 (2018).
Damschroder, D. et al. Stimulating the sir2–spargel axis rescues exercise capacity and mitochondrial respiration in a Drosophila model of Barth syndrome. Dis. Model Mech. 15, dmm049279 (2022).
Schlame, M., Xu, Y. & Ren, M. The basis for acyl specificity in the tafazzin reaction. J. Biol. Chem. 292, 5499–5506 (2017).
Taylor, W. A. & Hatch, G. M. Identification of the human mitochondrial linoleoyl-coenzyme A monolysocardiolipin acyltransferase (MLCL AT-1). J. Biol. Chem. 284, 30360–30371 (2009).
Rieger, B., Krajcova, A., Duwe, P. & Busch, K. B. ALCAT1 overexpression affects supercomplex formation and increases ROS in respiring mitochondria. Oxid. Med. Cell Longev. 2019, 9186469 (2019).
Miklas, J. W. et al. TFPa/HADHA is required for fatty acid β-oxidation and cardiolipin re-modeling in human cardiomyocytes. Nat. Commun. 10, 4671 (2019).
Li, J. et al. Cardiolipin remodeling by ALCAT1 links oxidative stress and mitochondrial dysfunction to obesity. Cell Metab. 12, 154–165 (2010).
Price, T. R. et al. Lipidomic QTL in Diversity Outbred mice identifies a novel function for α/β hydrolase domain 2 (Abhd2) as an enzyme that metabolizes phosphatidylcholine and cardiolipin. PLoS Genet. 19, e1010713 (2023).
Maddalena, L. A., Ghelfi, M., Atkinson, J. & Stuart, J. A. The mitochondria-targeted imidazole substituted oleic acid ‘TPP-IOA’ affects mitochondrial bioenergetics and its protective efficacy in cells is influenced by cellular dependence on aerobic metabolism. Biochim. Biophys. Acta Bioenerg. 1858, 73–85 (2017).
Reily, C. et al. Mitochondrially targeted compounds and their impact on cellular bioenergetics. Redox Biol. 1, 86–93 (2013).
Trnka, J., Elkalaf, M. & Andel, M. Lipophilic triphenylphosphonium cations inhibit mitochondrial electron transport chain and induce mitochondrial proton leak. PLoS ONE 10, e0121837 (2015).
Di Paola, M. & Lorusso, M. Interaction of free fatty acids with mitochondria: coupling, uncoupling and permeability transition. Biochim. Biophys. Acta 1757, 1330–1337 (2006).
Severin, F. F. et al. Penetrating cation/fatty acid anion pair as a mitochondria-targeted protonophore. Proc. Natl Acad. Sci. USA 107, 663–668 (2010).
Davis, B. H. et al. Determination of optimal replicate number for validation of imprecision using fluorescence cell-based assays: proposed practical method. Cytom. B Clin. Cytom. 84, 329–337 (2013).
Olteanu, A. et al. Stability and apoptotic activity of recombinant human cytochrome c. Biochem. Biophys. Res. Commun. 312, 733–740 (2003).
Mandal, A. et al. Structural changes and proapoptotic peroxidase activity of cardiolipin-bound mitochondrial cytochrome c. Biophys. J. 109, 1873–1884 (2015).
Mandal, A., Boatz, J. C., Wheeler, T. B. & van der Wel, P. C. A. On the use of ultracentrifugal devices for routine sample preparation in biomolecular magic-angle-spinning NMR. J. Biomol. NMR 67, 165–178 (2017).
Delaglio, F. et al. NMRPipe: a multidimensional spectral processing system based on UNIX pipes. J. Biomol. NMR 6, 277–293 (1995).
Pettersen, E. F. et al. UCSF Chimera-a visualization system for exploratory research and analysis. J. Comput. Chem. 25, 1605–1612 (2004).
Bushnell, G. W., Louie, G. V. & Brayer, G. D. High-resolution three-dimensional structure of horse heart cytochrome c. J. Mol. Biol. 214, 585–595 (1990).
Jo, S., Kim, T., Iyer, V. G. & Im, W. CHARMM-GUI: a web-based graphical user interface for CHARMM. J. Comp. Chem. 29, 1859–1865 (2008).
Autenrieth, F., Tajkhorshid, E., Baudry, J. & Luthey‐Schulten, Z. Classical force field parameters for the heme prosthetic group of cytochrome c. J. Comput. Chem. 25, 1613–1622 (2004).
Phillips, J. et al. Scalable molecular dynamics with NAMD. J. Comput. Chem. 26, 1781–1802 (2005).
Koes, D. R., Baumgartner, M. P. & Camacho, C. J. Lessons learned in empirical scoring with smina from the CSAR 2011 benchmarking exercise. J. Chem. Inf. Mod. 53, 1893–1904 (2013).
Zoete, V., Cuendet, M. A., Grosdidier, A. & Michielin, O. SwissParam: a fast force field generation tool for small organic molecules. J. Comp. Chem. 32, 2359–2368 (2011).
Bakan, A., Meireles, L. M. & Bahar, I. ProDy: protein dynamics inferred from theory and experiments. Bioinformatics 27, 1575–1577 (2011).
Acknowledgements
The study was supported by the National Institutes of Health grant GM134715 (to V.E.K. and M.L.G.), HL117880 (M.L.G.), NS076511 (V.E.K. and H.B.) and AG059683 (R.W.), the Polish National Science Centre (2019/35/D/ST4/02203 to K.M.-R) and the Natural Sciences and Engineering Research Council of Canada (RGPIN/5368-2019 to G.M.H). G.M.H. is the Canada Research Chair in Molecular Cardiolipin Metabolism.
Author information
Authors and Affiliations
Contributions
Conceptualization was the responsibility of V.E.K., H.B. and M.L.G. Methodology was the responsibility of V.E.K., Y.Y.T., V.A.T., A.A.K., A.A.A., J.P., J.J., R.W., M.L.G., P.C.A.W., A.L., G.M.H. and E.V.N. Software was the responsibility of Y.Y.T., K.M.-R. and P.C.A.W. Validation was carried out by V.E.K. and M.L.G. Formal analysis was conducted by Y.Y.T., V.A.T., A.A.K., A.A.A., S.N.S., M.A.A., G.K.V., E.-K.B., D.D., J.J. and A.R. Computational modeling was carried out by K.M.-R. and I.B. Investigation was conducted by Y.Y.T., V.A.T., A.A.K., M.A.A., S.N.S., H.H.D., A.B.S., A.K., Z.L., D.D., J.J., A.R., P.L., A.L., P.R., L.K.C., E.V.N. and K.K. Resources were the responsibility of K.M.-R., M.W.S., B.K., G.M.H., A.N., H.B., J.A. and J.V. Data curation was carried out by Y.Y.T., K.M.-R. and P.C.A.W. Writing of the original draft was carried out by V.E.K., H.B. and M.L.G. Review and editing was carried out by Y.Y.T., K.M.-R., P.C.A.W., M.W.S., I.B., G.M.H. and E.V.N. Visualization was the responsibility of Y.Y.T., K.M.-R., A.R. and A.L. Supervision was the responsibility of V.E.K., R.W., M.L.G., P.C.A.W. and J.V. Project administration was the responsibility of V.E.K. and M.L.G. Funding acquisition was the responsibility of V.E.K. and M.L.G.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Metabolism thanks Steven M. Claypool, Rafael Radi and the other, anonymous, reviewers for their contribution to the peer review of this work. Primary Handling Editor: Yanina-Yasmin Pesch, in collaboration with the Nature Metabolism team.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 SSNMR analysis of structural and dynamic rearrangements in the cyt c peroxidase complex with MLCL.
a, Region from the 2D CP-DARR spectrum of 15N,13C-labeled cyt c bound to DOPC:MLCL(L)3 (1:1; blue). Overlaid red contours show simulated peak patterns constructed from the solution NMR shifts of the N- and C-terminal α-helices (BMRB ID 25908). b, 2D 15N-13C NCA ssNMR spectrum of the same sample, along with a corresponding simulated spectrum for the N- and C-terminal α-helices. The simulated peaks from the blue foldon coincide with strong peaks in the experimental ssNMR spectra. c, 2D (CP-based) 15N-13C NCA ssNMR spectrum of membrane-bound cyt c, alongside the analogous simulated spectrum (d) for cyt c in solution (BMRB ID 25908). Key resonances missing in the experimental spectrum (c) are indicated with green circles; these mostly belong to the Ω-loop D (for example Met80 and Ile81). These 2D NCA ssNMR spectra were acquired at 253 K and 10 kHz MAS. e, 1D 13C INEPT, CP, and direct excitation spectra of 15N,13C-labeled cyt c bound to DOPC:MLCL(L)3 (1:1), measured at 278K. These ssNMR spectra detect flexible (INEPT), rigid (CP), or both (direct excitation) sample components. The liquid crystalline lipids contribute many peaks to the INEPT spectrum (marked with blue arrows), while the labeled protein dominates the other two spectra. f, Overlay of 2D INEPT-TOBSY spectra for 15N,13C-labeled cyt c bound to MLCL/DOPC (blue) and CL/DOPC (red) vesicles. Cross-peaks stem from the labeled protein and must reflect flexible residues. Many more (and stronger) peaks are seen for MLCL-bound cyt c.
Extended Data Fig. 2 31P ssNMR analysis of MLCL-induced structural and dynamic changes in the lipid membrane.
a, 1D 31P ssNMR static spectra of DOPC:MLCL (1:1) (blue) and DOPC:CL (1:1) (red) vesicles with bound cyt c. b, Analogous 31P NMR spectra in absence of cyt c. Panels (a,b) show that the presence of MLCL results in a narrower extra component, that is neither isotropic nor reflects a typical non-bilayer phase. As previously discussed23, these signals reflect MLCL in the liquid crystalline bilayer, undergoing distinct, increased dynamics (of the phosphate groups) compared to normal CL. c, 1D 31P ssNMR MAS spectra of DOPC:MLCL (1:1) at 10 kHz (blue curve) and 1.8 kHz (light blue curve) MAS rates. d, Zoomed-in region from (C) showing the isotropic peak (left) and first sideband (at 1.8 kHz MAS). Peak assignments are indicated, based on prior publications13,23. Note that in the side bands, the PC signals are strong, one MLCL signal is partly retained and one MLCL signal is attenuated or missing. This is consistent with the shown assignments and cited literature.
Extended Data Fig. 3 Final shapshots from simulations of cyt c – membrane interactions and superposition of the conformations of cyt c reached after 100 ns runs.
a, Final shapshots from simulations of cyt c – membrane interactions, and residues making frequent contacts with MLCL. Final conformations of cyt c reached after 100 ns in twenty independent MD trajectories. Red labels denote the systems which were further extended to 300 ns. The colors of the components of the membrane are following: DOPC – in orange, DOPE – in green, MLCL – in transparent blue. b, Superposition of the conformations of cyt c reached after 100 ns runs, in a system without membrane which contained Fe-S bond (sharp structure) and two independent runs where the bond was drastically weakened (transparent structures). All final conformations of heme group are displayed as grey shadow balls-sticks. c, Final conformations of cyt c after association with the membrane, observed in four runs MD1-MD4 (labeled). The white arrow in the upper left panel shows the residue M80 which inserted deep into the membrane, being trapped by several MLCL molecules, also shown in Fig. 2b.
Extended Data Fig. 4 Peroxidase activity of cyt c–MLCL complex causes phospholipid peroxidation in vitro and induces changes in lipidome and oxy-lipidome of genetically manipulated yeast.
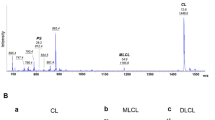
MS/MS spectra of (a) HOO-MLCL(L)3 (left panel) and oxidatively truncated ONA-MLCL(L)3 (right panel), (b) PC(18:0/18:2-OOH) (left panel) and PE(18:0/18:2-OOH) (right panel), (c) PC(18:0/ONA) (left panel) and PE(18:0/ONA) (right panel). Possible structures of oxidation products are inserted. ONA- 9-oxo nonanoic acid. d, Typical MS spectra of CL (upper panels) and MLCL (lower panels) obtained from yeast cells. e, Effect of D12desaturase on composition of CL (left panel) and MLCL (right panel) in WT and taz1D yeasts. SFA – saturated fatty acids, MUFA – monounsaturated fatty acids, PUFA – polyunsaturated fatty acids. f, Content of PC (upper panels) and PE (lower panels) molecular species in WT and D12/taz1D cells. Data are presented as mean values ± SD. Each data point represents a biologically independent sample. Upper panels: left ****P < 0.0001, middle ****P < 0.0001, ***P = 0.0004, right *P = 0.0202, ***P = 0.0001. ****P < 0.0001. One-way ANOVA, Tukey’s multiple comparison test. Lower panels; left ****P < 0.0001, middle **P = 0.0115. ***P = 0.0006, unpaired two-tailed t-test, ***P < 0.0001, ****P < 0.0001, One-way ANOVA, Tukey’s multiple comparison test. right ***P = 0.0056, unpaired two-tailed t-test, ****P < 0.0001, One-way ANOVA, Tukey’s multiple comparison test. g, OPLS-DA analysis showing the differences in oxy-lipidomes of D12 and D12/taz1D yeast cells; h, Pie plots showing the number of oxidatively modified phospholipid species. In total, 60 oxygenated phospholipid species were detected in yeast cells.
Extended Data Fig. 5 TAZ deficiency induces the changes in phospholipidome of mouse myoblasts and human lymphoblasts in vitro and mouse and human heart in vivo.
a, Table: Human heart biopsy sample description. Typical spectra of MLCL (left panels) and CL (right panels) obtained from WT and TAZ-KO C2C12 cells (b), WT and TAZ-KD mice (c), human lymphoblasts (d) and heart biopsy samples from control, non-BTHS-associated heart failure (NBHF) and BTHS-associated heart failure (BTHS) patients (e). TAZ deficiency in cells and mice and TAZ mutation in human results in a decrease of MLCL molecular species with C18:2 and appearance of MLCL molecular species containing C20:4 (shown in red); (f) Pie plots showing the total number of MLCL species in C2C12 cells, mouse heart, human lymphoblasts and heart samples from BTHD patients. (g) The score plots of OPLS-DA analysis show the differences in oxy-lipidomes in WT and TAZ-KO cells, WT and TAZ-KD mice, human control and BTHS lymphoblasts and control heart samples and BTHS heart samples.
Extended Data Fig. 6 Content of PE and PC species in cells, mouse heart, human lymphoblasts and human heart biopsy samples.
C2C12 cells (a-d): a, Total content of diacyl-PC (left panel) and its molecular species (right panel). b, Total content of plasmalogen-PC (left panel) and its molecular species (right panel). c, Total content of diacyl-PE (left panel) and its molecular species (right panel). d, Total content of plasmalogen-PE (left panel) and its molecular species (right panel). Mouse heart (e-h): e, Total content of diacyl-PC (left panel) and its molecular species (right panel). f, Total content of plasmalogen-PC (left panel) and its molecular species (right panel). g, Total content of diacyl-PE (left panel) and its molecular species (right panel). h, Total content of plasmalogen-PE (left panel) and its molecular species (right panel). Human lymphocytes (i-l): i, Total content of diacyl-PC (left panel) and its molecular species (right panel). j, Total content of plasmalogen-PC (left panel) and its molecular species (right panel). k, Total content of diacyl-PE (left panel) and its molecular species (right panel). l, Total content of plasmalogen-PE (left panel) and its molecular species (right panel). Human heart biopsy samples (m-p): m, Total content of diacyl-PC (left panel) and its molecular species (right panel). n, Total content of plasmalogen-PC (left panel) and its molecular species (right panel). o, Total content of diacyl-PE (left panel) and its molecular species (right panel). p, Total content of plasmalogen-PE (left panel) and its molecular species (right panel). *P < 0.05, **P < 0.01. DB - double bond number.
Extended Data Fig. 7 Effect of TPP-IOA on structure of cyt c–MLCL peroxidase complex, with MLCL, lipid oxidation and the endurance of Drosophila melanogaster.
Effect of TPP-imidazole-oleic acid (TPP-IOA) on MLCL(L)3-dependent formation of heme iron high-spin forms assessed by absorbance at 620 nm (a) and absorbace at 695 nm (b). Right: the differential absorption spectra (a) and representative absorption spectra (b) of MLCL–cyt c with or without TPP-IOA. Data are presented as mean values ± SD. Each data point represents a biologically independent sample. ****p < 0.0001, One-way ANOVA, Tukey’s multiple comparison test. (a) N = 8 (control), N = 6 (TPP-IOA–cyt c = 2/1), N = 6 (TPP-IOA–cyt c = 4/1). (b) N = 8 (control), N = 10 (TPP-IOA–cyt c = 2/1), N = 9 (TPP-IOA–cyt c = 4/1).(c) TPP-IOA inhibits accumulation of HOO-PE (left panel) and oxidatively truncated PE (right panel) species formed in MLCL(L)3–cyt c-driven reaction. Data are presented as mean values ± SD. N = 3. Each data point represents a biologically independent sample. Right panel: *p = 0.0287, ****p < 0.0001, Left panel: ***p = 0.0007, ****p < 0.0001. One-way ANOVA, Tukey’s multiple comparison test. (d) Normalized peak intensity values from 13C-13C ssNMR spectra of cyt c bound to DOPC:MLCL(L)3 (1:1; blue), or DOPC:MLCL(L)3 (2:1) LUVs in the absence (black) and presence (orange) of a four-fold excess of IOA with respect to protein (1:40 P/L ratio). Shown are the Cα/Cβ (open) and Cβ/Cα (closed) peaks for Thr residues. Each residue shows two data points: one for either side of the diagonal: see Fig. 5e). Increasing the ratio of MLCL (over PC) caused a net decrease in peak intensities, signifying increased protein motion. Addition of the IOA inhibitor had a much more modest effect on the Thr peak volumes.
Extended Data Fig. 8 TAZ deficiency induces changes in Drosophila melanogaster lipidome.
Typical mass spectra of MLCL (a) and CL (b) obtained from control (W1118) and (TAZ889) mutant flies. Content MLCL containing C18:2 (c) and C18:3 (d) in control W1118 and TAZ889 deficient flies. Lipidomics analysis was performed using 6 vials (n = 20 fly torsos per vial). (c) *P = 0.0401, **P < 0.0015, ***P = 0.0005 unpaired two-tailed t-test. (d) *P = 0.0378, **P < 0.0019, ****p < 0.0001 unpaired two-tailed t-test. (e) TPP-IOA did not significantly affect the endurance of control flies. Endurance was measure using 8 vials (N = 20 flies) and significance was determined by log-rank analysis., ns = not significant. (f) Content of CL (left panel) and MLCL (right panel) in control W1118 flies after feeding with TPP-IOA. Lipidomics analysis was performed using 6 vials (n = 20 fly torsos per vial). For all violin plots presented, individual points including maximal and minimal are shown as black circles. Dashed black line indicates median and doted lines indicate quartiles.
Supplementary information
Supplementary Information
Supplementary Fig. 1 and Tables 1 and 2.
Supplementary Video 1
Video 1.
Supplementary Video 2
Video 2.
Supplementary Video 3
Video 3.
Source data
Source Data Fig. 1
Statistical source data.
Source Data Fig. 2
Statistical source data.
Source Data Fig. 3
Statistical source data.
Source Data Fig. 4
Statistical source data.
Source Data Fig. 5
Statistical source data.
Source Data Fig. 6
Statistical source data.
Source Data Fig. 7
Statistical source data.
Source Data Extended Data Fig. 4
Statistical source data.
Source Data Extended Data Fig. 5
Statistical source data.
Source Data Extended Data Fig. 6
Statistical source data.
Source Data Extended Data Fig. 7
Statistical source data.
Source Data Extended Data Fig. 8
Statistical source data.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Kagan, V.E., Tyurina, Y.Y., Mikulska-Ruminska, K. et al. Anomalous peroxidase activity of cytochrome c is the primary pathogenic target in Barth syndrome. Nat Metab 5, 2184–2205 (2023). https://doi.org/10.1038/s42255-023-00926-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s42255-023-00926-4