Abstract
Glutamine is a critical metabolite for rapidly proliferating cells as it is used for the synthesis of key metabolites necessary for cell growth and proliferation. Glutamine metabolism has been proposed as a therapeutic target in cancer and several chemical inhibitors are in development or in clinical trials. How cells subsist when glutamine is limiting is poorly understood. Here, using an unbiased screen, we identify ALDH18A1, which encodes P5CS, the rate-limiting enzyme in the proline biosynthetic pathway, as a gene that cells can downregulate in response to glutamine starvation. Notably, P5CS downregulation promotes de novo glutamine synthesis, highlighting a previously unrecognized metabolic plasticity of cancer cells. The glutamate conserved from reducing proline synthesis allows cells to produce the key metabolites necessary for cell survival and proliferation under glutamine-restricted conditions. Our findings reveal an adaptive pathway that cancer cells acquire under nutrient stress, identifying proline biosynthesis as a previously unrecognized major consumer of glutamate, a pathway that could be exploited for developing effective metabolism-driven anticancer therapies.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Data availability
The datasets for the shRNA screening have been deposited in the NCBI Gene Expression Omnibus under accession no. GSE162314. Other data have been provided as source data with the article or will be available upon request. Source data are provided with this paper.
References
Altman, A. B. J., Stine, Z. E. & Dang, C. V. From Krebs to clinic: glutamine metabolism to cancer therapy. Nat. Rev. Cancer 16, 619–634 (2016).
Zhang, J., Pavlova, N. N. & Thompson, C. B. Cancer cell metabolism: the essential role of the nonessential amino acid, glutamine. EMBO J. 36, 1302–1315 (2017).
Krebs, H. A. Metabolism of amino-acids: the synthesis of glutamine from glutamic acid and ammonia, and the enzymic hydrolysis of glutamine in animal tissues. Biochem. J. 29, 1951–1969 (1935).
Eagle, H. The minimum vitamin requirements of the L and HeLa cells in tissue culture, the production of specific vitamin deficiencies, and their cure. J. Exp. Med. 102, 595–600 (1955).
Bott, J. et al. Oncogenic Myc induces expression of glutamine synthetase through promoter demethylation. Cell Metab. 22, 1068–1077 (2015).
Tardito, S. et al. Glutamine synthetase activity fuels nucleotide biosynthesis and supports growth of glutamine-restricted glioblastoma. Nat. Cell Biol. 17, 1556–1568 (2015).
Pavlova, N. N. et al. As extracellular glutamine levels decline, asparagine becomes an essential amino acid. Cell Metab. 27, 428–438 (2018).
Bott, J. et al. Glutamine anabolism plays a critical role in pancreatic cancer by coupling carbon and nitrogen metabolism. Cell Rep. 29, 1287–1298 (2019).
Wise, D. R. et al. Myc regulates a transcriptional program that stimulates mitochondrial glutaminolysis and leads to glutamine addiction. Proc. Natl Acad. Sci. USA 105, 18782–18787 (2008).
Mayers, J. R. & Heiden, M. G. V. Famine versus feast: understanding the metabolism of tumors in vivo. Trends Biochem. Sci. 40, 130–140 (2015).
Roberts, E., Simonsen, D. G., Tanaka, K. K. & Tanaka, T. Free amino acids in growing and regressing ascites cell tumors: host resistance and chemical agents. Cancer Res. 16, 970–978 (1956).
Rivera, S., Azcón-Bieto, J., López-Soriano, F. J., Miralpeix, M. & Argilés, J. M. Amino acid metabolism in tumour-bearing mice. Biochem. J. 249, 443–449 (1988).
Márquez, J., Sánchez-Jiménez, F., Medina, M. A., Quesada, A. R. & de Castro, I. N. Nitrogen metabolism in tumor bearing mice. Arch. Biochem. Biophys. 268, 667–675 (1989).
Pan, M. et al. Regional glutamine deficiency in tumours promotes dedifferentiation through inhibition of histone demethylation. Nat. Cell Biol. 18, 1090–1101 (2016).
Lee, S.-W. et al. EGFR-Pak signaling selectively regulates glutamine deprivation-induced macropinocytosis. Dev. Cell 50, 381–392 (2019).
Yuneva, M., Zamboni, N., Oefner, P., Sachidanandam, R. & Lazebnik, Y. Deficiency in glutamine but not glucose induces MYC-dependent apoptosis in human cells. J. Cell Biol. 178, 93–105 (2007).
Zhang, J. et al. Asparagine plays a critical role in regulating cellular adaptation to glutamine depletion. Mol. Cell. 56, 205–218 (2014).
Krall, S., Xu, S., Graeber, T. G., Braas, D. & Christofk, H. R. Asparagine promotes cancer cell proliferation through use as an amino acid exchange factor. Nat. Commun. 7, 11457 (2016).
Yang, L. et al. Targeting stromal glutamine synthetase in tumors disrupts tumor microenvironment-regulated cancer cell growth. Cell Metab. 24, 685–700 (2016).
Linares, J. F. et al. ATF4-induced metabolic reprograming is a synthetic vulnerability of the p62-deficient tumor stroma. Cell Metab. 26, 817–829 (2017).
Mishra, R. et al. Stromal epigenetic alterations drive metabolic and neuroendocrine prostate cancer reprogramming. J. Clin. Invest. 128, 4472–4484 (2018).
Tajan, M. et al. A role for p53 in the adaptation to glutamine starvation through the expression of SLC1A3. Cell Metab. 28, 721–736 (2018).
Alkan, H. F. et al. Cytosolic aspartate availability determines cell survival when glutamine is limiting. Cell Metab. 28, 706–720 (2018).
Lowman, X. H. et al. p53 promotes cancer cell adaptation to glutamine deprivation by upregulating Slc7a3 to increase arginine uptake. Cell Rep. 26, 3051–3060 (2019).
Jiang, J., Srivastava, S. & Zhang, J. Starve cancer cells of glutamine: break the spell or make a hungry monster? Cancers 11, 804 (2019).
Hu, C. A., Lin, W. W., Obie, C. & Valle, D. Molecular enzymology of mammalian δ1-pyrroline-5-carboxylate synthase. Alternative splice donor utilization generates isoforms with different sensitivity to ornithine inhibition. J. Biol. Chem. 274, 6754–6762 (1999).
Olivares, O. et al. Collagen-derived proline promotes pancreatic ductal adenocarcinoma cell survival under nutrient limited conditions. Nat. Commun. 8, 16031 (2017).
Zhu, J. et al. Mitochondrial NADP(H) generation is essential for proline biosynthesis. Science 372, 968–972 (2021).
Tran, D. H. et al. Mitochondrial NADP+ is essential for proline biosynthesis during cell growth. Nat. Metab. 3, 571–585 (2021).
Windmueller, H. G. & Spaeth, A. E. Uptake and metabolism of plasma glutamine by the small intestine. J. Biol. Chem. 249, 5070–5070 (1974).
Liu, W. et al. Reprogramming of proline and glutamine metabolism contributes to the proliferative and metabolic responses regulated by oncogenic transcription factor c-MYC. Proc. Natl Acad. Sci. USA 109, 8983–8988 (2012).
Liu, W., Hancock, C. N., Fischer, J. W., Harman, M. & Phang, J. M. Proline biosynthesis augments tumor cell growth and aerobic glycolysis: involvement of pyridine nucleotides. Sci. Rep. 5, 17206 (2015).
Burke, L. et al. The Janus-like role of proline metabolism in cancer. Cell Death Discov. 6, 104 (2020).
Kardos, G. R., Wastyk, H. C. & Robertson, G. P. Disruption of proline synthesis in melanoma inhibits protein production mediated by the GCN2 pathway. Mol. Cancer Res. 13, 1408–1420 (2015).
Loayza-Puch, F. et al. Tumour-specific proline vulnerability uncovered by differential ribosome codon reading. Nature 530, 490–494 (2016).
Sahu, N. et al. Proline starvation induces unresolved ER stress and hinders mTORC1-dependent tumorigenesis. Cell Metab. 24, 753–761 (2016).
Grinde, M. T. et al. Glutamine to proline conversion is associated with response to glutaminase inhibition in breast cancer. Breast Cancer Res. 21, 61 (2019).
Rubin, H. Deprivation of glutamine in cell culture reveals its potential for treating cancer. Proc. Natl Acad. Sci. USA 116, 6964–6968 (2019).
Sullivan, K. D. et al. Trisomy 21 consistently activates the interferon response. eLife 5, e16220 (2016).
Langmead, B. & Salzberg, S. L. Fast gapped-read alignment with Bowtie 2. Nat. Methods 9, 357–359 (2012).
Mathew, R., Degenhardt, K., Haramaty, L., Karp, C. M. & White, E. Immortalized mouse epithelial cell models to study the role of apoptosis in cancer. Methods Enzymol. 446, 77–106 (2008).
Zhong, L. et al. The histone deacetylase Sirt6 regulates glucose homeostasis via Hif1α. Cell 140, 280–293 (2010).
Schindelin, J. et al. Fiji: an open-source platform for biological-image analysis. Nat. Methods 9, 676–682 (2012).
Schneider, C. A., Rasband, W. S. & Eliceiri, K. W. NIH Image to ImageJ: 25 years of image analysis. Nat. Methods 9, 671–675 (2012).
Buescher, J. M. et al. A roadmap for interpreting (13)C metabolite labeling patterns from cells. Curr. Opin. Biotech. 34, 189–201 (2015).
Mackay, G. M., Zheng, L., van den Broek, N. J. F. & Gottlieb, E. Analysis of cell metabolism using LC–MS and isotope tracers. Methods Enzymol. 561, 171–196 (2015).
Boon, R. et al. Amino acid levels determine metabolism and CYP450 function of hepatocytes and hepatoma cell lines. Nat. Commun. 11, 1393 (2020).
Bernasocchi, T. et al. Dual functions of SPOP and ERG dictate androgen therapy responses in prostate cancer. Nat. Commun. 12, 734 (2021).
Acknowledgements
We thank all members of the Mostoslavsky laboratory for helpful discussions and critical reading of the paper. We thank R. DeBerardinis (University of Texas Southwestern Medical Center) for providing the MycER MEF cell line used in the screen and staff at the Metabolite Profiling Core (Whitehead Institute) for their helpful discussions and technical expertise. S.J.L. is the recipient of a National Institutes of health (NIH) F31 Ruth L. Kirschstein Predoctoral fellowship (F31CA210310). T.B. is a recipient of an EMBO Postdoctoral Fellowship, ALTF 359-2022 . R. Mostoslavsky is the Laurel Schwartz Endowed Chair in Oncology. This work is partially supported by NIH grants R33ES025638 and R01GM128448 and a Massachusetts Life Sciences Center Bits to Bytes award to R. Mostoslavsky, NIH grants R01CA117907, R01GM120109 and P30CA046934, NSF grant MCB-1817582 and grants from the Wings of Hope and Golfers Against Cancer foundations to J.M.E. B.R.R. is funded by the Nile Albright Research Foundation and Vincent Memorial Hospital Foundation.
Author information
Authors and Affiliations
Contributions
S.J.L., T.B., B.M.-P., C.M.F., R.B., C.V. and H.M.C. performed the experiments. R. Massri and E.G. generated and analyzed the P5CS CRISPR KO cells. K.D.S., M.D.G. and J.M.E. provided the shRNA library and performed the analysis of screening data. C.A.L. performed and analyzed the metabolomics assays. K.N.R. analyzed the TCGA data. S. Schroff provided the patient-derived tumor tissue slides. J.P.O.-C., G.G.S. and S. Stott imaged and analyzed the IHC of the tissue slides. Y.M., E.K. and B.R.R. provided de-identified formalin-fixed paraffin-embedded sections representing endometrial cancer and endometrial cancer cell lysates from the Vincent Center for Reproductive Biology Gynecologic Tissue Repository for analysis. S.J.L., T.B. and R. Mostoslavsky conceptualized and designed the study, analyzed results and wrote the paper. R. Mostoslavsky and T.B. supervised the project.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Metabolism thanks Jiyeon Kim and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Primary Handling Editor: Alfredo Giménez-Cassina, in collaboration with the Nature Metabolism team.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Quality controls of the genome-wide shRNA-library, GLN deprivation screen.
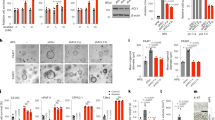
A) Western blot of MycER and Myc transcriptional target Ldha in whole cell lysate (WCL) and chromatin isolated from MycER MEFs exposed to 200 nM 4-OHT for 24 and 48 hours, (n = 3) B) Phase contrast microscopy of MycER cells + 200 nM 4-OHT grown in normal media for 3 days or without GLN for 3 and 11 days, (n = 3). C) Heatmap showing pairwise sample-sample Euclidean distances, arranged by treatment group. Dendrograms show hierarchical clustering to highlight similarities between samples. D) MA plot of log2(mean counts-per-million) against log2(median fold change), with points colored by density to highlight data trend(s). E) Levels of Glutamine (GLN), Glutamate (GLU), and Proline (PRO) in Fetal bovine serum. Data are represented as mean ± SD of three independent experiments F) WT and two clonal P5CS CRISPR KO mouse kidney epithelial cell lines grown in the presence of 2 mM GLN for 1 week, or absence of GLN for 1 month, with corresponding quantification of growth. Statistically significant differences were evaluated using a paired, one-tailed T-Test. Data are presented as mean values ± SD. G) Growth curves of control and Aldh18a1 knockdown cells under +GLN and -GLN conditions. Data are presented as mean values ± SD of three independent experiments. Statistical significance was determined using two-way ANOVA. N = 3 biological independent samples. H) Repeats of western blot against P5CS in short-term (S) or long-term (L) culture of shScram and shAldh18a1 MycER MEFs in the presence or absence of 2 mM glutamine, (n = 3).
Extended Data Fig. 2 Proline depletion upon P5CS/Aldh18a1 downregulation.
A) Relative abundance of intracellular PRO in WT mouse kidney epithelial cells from two different thaws (WT-1 and WT-2) and two P5CS KO subclones (C2 and C9) infected either with empty vector ( + Vec) or reconstituted with P5CS ( + P5CS). Cells were grown for 24 hours in proline-free high-glucose DMEM medium with 0.65 mM GLN and 2.5%FBS, and metabolite levels normalized by total protein (n = 3) Statistical significance was determined using an unpaired, Two-tailed T-Test. Data are presented as mean values ± SD. B) Metabolomic profiling of amino acid consumption and secretion. This panel of plots illustrates the relative consumption and secretion rates of various amino acids by control and P5CS KD cells, under low-glutamine conditions. The lower bar graphs show the consumption rates of amino acids from the media, while the upper ones depict the secretion rates of amino acids into the media. Data are represented as mean ± SD of three independent experiments. The statistical analysis was performed using two-way ANOVA.
Extended Data Fig. 3 Evaluation of Cell Death in Response to Glutamine Deprivation.
A) Gating strategy for sorting MycER control cells and shALDH18A1 cells under different conditions: normal medium, glutamine-deprived medium, and medium supplemented with 7 mM alpha-ketoglutarate (aKG). B) Propidium iodide staining of NUGC2 control and shALDH18A1 cells quantified by Flow cytometry. Data are presented as mean ± SD from three independent experiments. Statistical analysis was performed using two-way ANOVA. On the right, the gating strategy for sorting in control and shALDH18A1 NUGC2 cells under different conditions: normal medium, glutamine-deprived medium, and medium supplemented with 7 mM alpha-ketoglutarate (aKG).
Extended Data Fig. 4 Proline is not a major source of carbon and/or nitrogen in GLN-starved cells.
A) Graphical representation of the 13C5 glutamate labeling pathway, illustrating its incorporation into intermediates of the TCA cycle, amino acids, and pyrimidines. B) Tracer experiment using 13C5-labeled glutamate in Control and P5CS Knockdown Cells. The bar chart illustrates the proportion of proline derived from glutamine in both control and P5CS knockdown cells, as well as the corresponding increase in the labeling of amino acids, nucleosides, and TCA cycle intermediates. Data are represented as mean ± SD of three independent experiments. The statistical analysis was performed using two-way ANOVA. C) Crystal violet staining and corresponding quantification of MycER shScram cells cultured in the presence or absence of GLN for 5 days. Cells were cultured in DMEM supplemented with 7 mM dimethyl-α-ketoglutarate (α-KG), 2 mM ASN, 15 mM GlcNAc, or 250 μM each cytidine (C), thymidine (T), uridine (U), adenosine (A), and guanosine (G) in indicated combinations. Statistical significance was determined using an unpaired, Two-tailed, T-Test. Data are presented as mean values ± SD. N = 3 biological independent samples. D) Crystal violet staining and corresponding quantification of MycER shScram cells cultured in the presence or absence of GLN for 5 days. Cells were cultured in DMEM supplemented with 7 mM dimethyl-α-ketoglutarate (α-KG), 2 mM ASN, 15 mM GlcNAc, or 250 μM each cytidine (C), thymidine (T), uridine (U), adenosine (A), and guanosine (G) in presence or absence of L-Methionine sulfoximine (MSO). Statistical significance was determined using ordinary one-way ANOVA followed by Tukey’s post hoc test for multiple comparisons. Data are presented as mean values ± SD. N = 6 biological independent samples. E) Cell proliferation of Aldh18a1 KD cells transfected with either siScram or two independent siP5CDH grown in the absence of GLN for 5 days (n = 3) (Top). Corresponding western blot verifying P5CS and P5CDH knockdown (Bottom). Statistical significance was determined using ordinary one-way ANOVA followed by Tukey’s post hoc test for multiple comparisons. Data are presented as mean values ± SD. N = 3 biological independent samples. F) Crystal violet staining and corresponding quantification of MycER shScram cells cultured in the presence or absence of GLN for 5 days. Cells were cultured in DMEM supplemented with 0.5 mM Proline, 2 mM ASN, or 250uM each C, T, U, A, and G, in indicated combinations. Data are presented as mean values ± SD. Gray square: Presence, white square: Absence.
Extended Data Fig. 5 Downregulation of P5CS maintains NADPH/ NADP+ ratio in low GLN.
A) Crystal violet staining and corresponding quantification of MycER cells cultured in the absence of GLN or in the presence of aKG (7 mM) + ASN (2 mM) + C,T,U,A, and G (250uM each) or Ornithine/arginine 2 mM or Ornithine/Proline 2 mM or Arginine/Proline 2 mM, or Ornithine/proline/Arginine 2 mM. Each condition with or without N-Acetyl Cysteine (NAC) 5 days. Statistical significance was determined using ordinary two-way ANOVA followed by Dunnett’s post hoc test for multiple comparisons. Data are presented as mean values ± SD. N = 3 biological independent samples. B) Percentage of incorporation of 13C5-labeled glutamine into L-Proline in MycER shScram, shAldh18a1 #1, and shAldh18a1 #2 cells. These cells were cultured in DMEM supplemented with dialyzed FBS and 2 mM each of labeled Glutamine (13C5), Arginine, Proline, and Ornithine (n = 3). The fraction that was not labeled was designated as Proline M + 0. Corresponding western blot verifying P5CS knockdown (right side). Statistical significance was determined using ordinary one-way ANOVA followed by Dunnett’s post hoc test for multiple comparisons. Error bars represent standard deviation (SD). C and D) Left panel: Growth curves of control and Aldh18a1 knockdown cells under -GLN and +GLN conditions, in the presence of 0.5 mM Proline. Data are represented as mean ± SD of three independent experiments. Statistical significance was determined using a two-way ANOVA. Right panel: Growth curves of control and Aldh18a1 knockdown cells under +Proline and -Proline conditions, in the context of +GLN. Data are represented as mean ± SD of three independent experiments. The statistical analysis was performed using a two-way ANOVA. E) bar graph representing the NADPH/NADP+ ratio in MycER cells, comparing the control group with two separate shALDH18A1 knockdown groups, under low glutamine conditions for 48 hours. Data are represented as mean ± SD of three independent experiments. The statistical analysis was performed using two-way ANOVA.
Extended Data Fig. 6 Sensitivity of human cancer cell lines to GLN restriction.
A) Kaplan–Meier survival curves analyzing the The Cancer Genome Atlas (TCGA) human uterine/endometrial or stomach cancer patient samples based on Aldh18a1 mRNA expression divided in quartiles. B) Bar plot illustrating the distribution of Uterine Endometrioid Carcinoma, Uterine Mixed Endometrial Carcinoma, and Uterine Serous Carcinoma cases, classified by low or high expression levels of ALDH18A1. (TCGA) C) Scatter plot demonstrating the ALDH18A1 mRNA expression levels across Uterine Endometrioid Carcinoma and Uterine Serous Carcinoma cases, based on TCGA data. Statistical analysis was conducted using an unpaired two-tailed t-test, and data are presented as mean ± SD. D) Bar plot demonstrating the distribution of Uterine cancer cases by histological grade, categorized by low or high ALDH18A1 RNA expression, as per TCGA data. n = 4 biological independent samples. E) Representative images and quantification of sensitivity to glutamine depletion of human cancer cell lines grown in the presence or absence of GLN for 4 days, represented by percent survival. Survival was calculated by dividing the average number of cells counted on Day 4 in wells treated with no GLN by the average number of cells in wells with GLN (n = 3 wells). F) Western blot analysis of NUGC2 and PC3 cell lines using scramble shRNA or two independent shRNAs targeting the 3’UTR of ALDH18A1, (n = 3). G) Western blot analysis of MFE-280 cell line using scramble shRNA or an shRNA for ALDH18A1 (n = 3).
Extended Data Fig. 7 Variation in P5CS Expression Levels among Human Cancer Cell Lines under Glutamine Deprivation.
A) Bar plot illustrating the weights of subcutaneous xenograft tumors from NUGC2 cells (PLKO, shALDH18A1, and rescue with dox-inducible NUGC2). Statistical analysis was conducted using a one-way ANOVA followed by Tukey’s multiple comparison test. Data are presented as mean values ± SD. N = 8 independent animals. B) 15N-labeled ammonia (15NH4 + ) in NUGC2 gastric cancer cells with genetically downregulated P5CS. Data are represented as mean ± SD of three independent experiments. Statistical significance was determined using two-way ANOVA. C) Western blot of P5CS in human cancer cell lines that do not endogenously downregulate P5CS upon acute GLN starvation. Cells were cultured in the presence or absence of 2 mM GLN for 24 hrs, (n = 3). D) Comparison of P5CS levels in xenograft tumors derived from MKN45 cells. On the left, a western blot illustrates the relative P5CS protein levels in tumor extracts, while on the right, a bar graph represents the quantification of these levels. Statistical significance was determined using a one-tailed unpaired t-test with Welch’s correction for unequal variances. Data are presented as mean values ± SD. E) Additional representative immunofluorescence (IF) of a tumor section of uterine serous carcinoma, depicting staining by IF for P5CS (red), GS (green), and Dapi (blue). On the right, a selected area of the tumors demonstrates an inverse correlation between P5CS and GS expression, (n = 3). F) Total number of cells with partner P5CSlow/GShigh or vice-versa detected in five tumor sections of uterine serous carcinoma (USC). Statistical analysis was performed using an unpaired two-tailed Mann-Whitney test. Data are presented as mean values ± SD.
Supplementary information
Supplementary Information
Supplementary Table 1.
Source data
Source Data Fig. 1
Statistical source data.
Source Data Fig. 2
Statistical source data.
Source Data Fig. 3
Statistical source data.
Source Data Fig. 4
Statistical source data.
Source Data Fig. 5
Statistical source data.
Source Data Extended Data Fig. 1
Statistical source data.
Source Data Extended Data Fig. 2
Statistical source data.
Source Data Extended Data Fig. 3
Statistical source data.
Source Data Extended Data Fig. 4
Statistical source data.
Source Data Extended Data Fig. 5
Statistical source data.
Source Data Extended Data Fig. 6
Statistical source data.
Source Data Extended Data Fig. 7
Statistical source data.
Source Data Supporting Blots
Source data blots.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Linder, S.J., Bernasocchi, T., Martínez-Pastor, B. et al. Inhibition of the proline metabolism rate-limiting enzyme P5CS allows proliferation of glutamine-restricted cancer cells. Nat Metab 5, 2131–2147 (2023). https://doi.org/10.1038/s42255-023-00919-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s42255-023-00919-3