Abstract
Sustained responses to transient environmental stimuli are important for survival. The mechanisms underlying long-term adaptations to temporary shifts in abiotic factors remain incompletely understood. Here, we find that transient cold exposure leads to sustained transcriptional and metabolic adaptations in brown adipose tissue, which improve thermogenic responses to secondary cold encounter. Primary thermogenic challenge triggers the delayed induction of a lipid biosynthesis programme even after cessation of the original stimulus, which protects from subsequent exposures. Single-nucleus RNA sequencing and spatial transcriptomics reveal that this response is driven by a lipogenic subpopulation of brown adipocytes localized along the perimeter of Ucp1hi adipocytes. This lipogenic programme is associated with the production of acylcarnitines, and supplementation of acylcarnitines is sufficient to recapitulate improved secondary cold responses. Overall, our data highlight the importance of heterogenous brown adipocyte populations for ‘thermogenic memory’, which may have therapeutic implications for leveraging short-term thermogenesis to counteract obesity.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
All snRNA-seq and spatial transcriptomics data reported in here have been deposited in the Gene Expression Omnibus under accession number GSE218711. RNA-seq data are available at the sequence read archive under BioProject number PRJNA866352. The snRNA-seq data of BAT from human come from a published study31 with accession number E-MTAB-8564 as described in the snRNA-seq section in the methods. Source data are provided with this paper. All other data are available in the main text or the Supplementary Information.
Code availability
Representative code to reproduce the spatial transcriptomics analysis are available at https://github.com/kpcoleman/BAT-SpaDecon/. Any additional information required to analyse the data reported in this paper is available from C.A.T. upon request.
References
Farber, D. L., Netea, M. G., Radbruch, A., Rajewsky, K. & Zinkernagel, R. M. Immunological memory: lessons from the past and a look to the future. Nat. Rev. Immunol. 16, 124–128 (2016).
Netea, M. G. et al. Defining trained immunity and its role in health and disease. Nat. Rev. Immunol. 20, 375–388 (2020).
Josselyn, S. A., Köhler, S. & Frankland, P. W. Finding the engram. Nat. Rev. Neurosci. 16, 521–534 (2015).
Kandel, E. R., Dudai, Y. & Mayford, M. R. The molecular and systems biology of memory. Cell 157, 163–186 (2014).
Naik, S. et al. Inflammatory memory sensitizes skin epithelial stem cells to tissue damage. Nature 550, 475–480 (2017).
Ordovas-Montanes, J., Beyaz, S., Rakoff-Nahoum, S. & Shalek, A. K. Distribution and storage of inflammatory memory in barrier tissues. Nat. Rev. Immunol. 20, 308–320 (2020).
Ordovas-Montanes, J. et al. Allergic inflammatory memory in human respiratory epithelial progenitor cells. Nature 560, 649–654 (2018).
Cannon, B. & Nedergaard, J. Brown adipose tissue: function and physiological significance. Physiol. Rev. 84, 277–359 (2004).
Chouchani, E. T., Kazak, L. & Spiegelman, B. M. New advances in adaptive thermogenesis: UCP1 and beyond. Cell Metab. 29, 27–37 (2019).
Cypess, A. M. et al. Identification and importance of brown adipose tissue in adult humans. N. Engl. J. Med. 360, 1509–1517 (2009).
Van Marken Lichtenbelt, W. D. et al. Cold-activated brown adipose tissue in healthy men. N. Engl. J. Med. 360, 1500–1508 (2009).
Becher, T. et al. Brown adipose tissue is associated with cardiometabolic health. Nat. Med. 27, 58–65 (2021).
Bartelt, A. et al. Brown adipose tissue activity controls triglyceride clearance. Nat. Med. 17, 200–205 (2011).
Stanford, K. I. et al. Brown adipose tissue regulates glucose homeostasis and insulin sensitivity. J. Clin. Invest. 123, 215–223 (2013).
Hanssen, M. J. W. et al. Short-term cold acclimation improves insulin sensitivity in patients with type 2 diabetes mellitus. Nat. Med. 21, 863–865 (2015).
Hanssen, M. J. W. et al. Short-term cold acclimation recruits brown adipose tissue in obese humans. Diabetes 65, 1179–1189 (2015).
Roh, H. C. et al. Warming induces significant reprogramming of beige, but not brown, adipocyte cellular identity. Cell Metab. 27, 1121–1137 (2018).
Gerhart-Hines, Z. et al. The nuclear receptor Rev-erbα controls circadian thermogenic plasticity. Nature 503, 410–413 (2013).
Golozoubova, V. et al. Only UCP1 can mediate adaptive nonshivering thermogenesis in the cold. FASEB J. 15, 2048–2050 (2001).
Cannon, B. & Nedergaard, J. Nonshivering thermogenesis and its adequate measurement in metabolic studies. J. Exp. Biol. 214, 242–253 (2011).
Enerbäck, S. et al. Mice lacking mitochondrial uncoupling protein are cold-sensitive but not obese. Nature 387, 90–94 (1997).
Sanchez-Gurmaches, J. et al. Brown fat AKT2 is a cold-induced kinase that stimulates ChREBP-mediated de novo lipogenesis to optimize fuel storage and thermogenesis. Cell Metab. 27, 195–209 (2018).
Adlanmerini, M. et al. Circadian lipid synthesis in brown fat maintains murine body temperature during chronic cold. Proc. Natl Acad. Sci. USA 116, 18691–18699 (2019).
Mottillo, E. P. et al. Coupling of lipolysis and de novo lipogenesis in brown, beige, and white adipose tissues during chronic β3-adrenergic receptor activation. J. Lipid Res. 55, 2276–2286 (2014).
Yu, X. X., Lewin, D. A., Forrest, W. & Adams, S. H. Cold elicits the simultaneous induction of fatty acid synthesis and β-oxidation in murine brown adipose tissue: prediction from differential gene expression and confirmation in vivo. FASEB J. 16, 155–168 (2002).
Matsuda, M. et al. SREBP cleavage-activating protein (SCAP) is required for increased lipid synthesis in liver induced by cholesterol deprivation and insulin elevation. Genes Dev. 15, 1206–1216 (2001).
Moon, Y.-A. et al. The Scap/SREBP pathway is essential for developing diabetic fatty liver and carbohydrate-induced hypertriglyceridemia in animals. Cell Metab. 15, 240–246 (2012).
Habib, N. et al. Div-Seq: single-nucleus RNA-seq reveals dynamics of rare adult newborn neurons. Science 353, 925–928 (2016).
Habib, N. et al. Massively parallel single-nucleus RNA-seq with DroNc-seq. Nat. Methods 14, 955–958 (2017).
Lake, B. B. et al. Neuronal subtypes and diversity revealed by single-nucleus RNA sequencing of the human brain. Science 352, 1586–1590 (2016).
Sun, W. et al. snRNA-seq reveals a subpopulation of adipocytes that regulates thermogenesis. Nature 587, 98–102 (2020).
Paulo, E. et al. Sympathetic inputs regulate adaptive thermogenesis in brown adipose tissue through cAMP-Salt inducible kinase axis. Sci. Rep. 8, 11001 (2018).
Coleman, K., Hu, J., Schroeder, A., Lee, E. B. & Li, M. SpaDecon: cell-type deconvolution in spatial transcriptomics with semi-supervised learning. Commun. Biol. 6, 378 (2023).
Simcox, J. et al. Global analysis of plasma lipids Identifies liver-derived acylcarnitines as a fuel source for brown fat thermogenesis. Cell Metab. 26, 509–522 (2017).
McCormack, J. G. & Denton, R. M. Evidence that fatty acid synthesis in the interscapular brown adipose tissue of cold-adapted rats is increased in vivo by insulin by mechanisms involving parallel activation of pyruvate dehydrogenase and acetyl-coenzyme A carboxylase. Biochem. J. 166, 627–630 (1977).
Shimazu, T. & Takahashi, A. Stimulation of hypothalamic nuclei has differential effects on lipid synthesis in brown and white adipose tissue. Nature 284, 62–63 (1980).
Trayhurn, P. Fatty acid synthesis in vivo in brown adipose tissue, liver and white adipose tissue of the cold-acclimated rat. FEBS Lett. 104, 13–16 (1979).
Guilherme, A. et al. Control of adipocyte thermogenesis and lipogenesis through β3-adrenergic and thyroid hormone signal integration. Cell Rep. 31, 107598 (2020).
Broeders, E. P. M. et al. Thyroid hormone activates brown adipose tissue and increases non-shivering thermogenesis-a cohort study in a group of thyroid carcinoma patients. PLoS ONE 11, e0145049 (2016).
Weiner, J. et al. Thyroid hormone status defines brown adipose tissue activity and browning of white adipose tissues in mice. Sci. Rep. 6, 38124 (2016).
Yeh, W. J., Leahy, P. & Freake, H. C. Regulation of brown adipose tissue lipogenesis by thyroid hormone and the sympathetic nervous system. Am. J. Physiol. Metab. 265, E252–E258 (1993).
Tan, C. Y. et al. Brown adipose tissue thermogenic capacity is regulated by Elovl6. Cell Rep. 13, 2039–2047 (2015).
Westerberg, R. et al. ELOVL3 is an important component for early onset of lipid recruitment in brown adipose tissue. J. Biol. Chem. 281, 4958–4968 (2006).
Guilherme, A. et al. Neuronal modulation of brown adipose activity through perturbation of white adipocyte lipogenesis. Mol. Metab. 16, 116–125 (2018).
Guilherme, A. et al. Adipocyte lipid synthesis coupled to neuronal control of thermogenic programming. Mol. Metab. 6, 781–796 (2017).
Lodhi, I. J. et al. Inhibiting adipose tissue lipogenesis reprograms thermogenesis and PPARγ activation to decrease diet-induced obesity. Cell Metab. 16, 189–201 (2012).
Lee, S.-H. et al. Lack of stearoyl-CoA desaturase 1 upregulates basal thermogenesis but causes hypothermia in a cold environment. J. Lipid Res. 45, 1674–1682 (2004).
Foster, D. W. Malonyl-CoA: the regulator of fatty acid synthesis and oxidation. J. Clin. Invest. 122, 1958–1959 (2012).
McGarry, J. D., Leatherman, G. F. & Foster, D. W. Carnitine palmitoyltransferase I. The site of inhibition of hepatic fatty acid oxidation by malonyl-CoA. J. Biol. Chem. 253, 4128–4136 (1978).
Jung, S. M. et al. In vivo isotope tracing reveals the versatility of glucose as a brown adipose tissue substrate. Cell Rep. 36, 109459 (2021).
Benador, I. Y. et al. Mitochondria bound to lipid droplets have unique bioenergetics, composition, and dynamics that support lipid droplet expansion. Cell Metab. 27, 869–885 (2018).
Yu, J. et al. Lipid droplet remodeling and interaction with mitochondria in mouse brown adipose tissue during cold treatment. Biochim. Biophys. Acta 1853, 918–928 (2015).
Benador, I. Y., Veliova, M., Liesa, M. & Shirihai, O. S. Mitochondria bound to lipid droplets: where mitochondrial dynamics regulate lipid storage and utilization. Cell Metab. 29, 827–835 (2019).
Fedorenko, A., Lishko, P. V. & Kirichok, Y. Mechanism of fatty-acid-dependent UCP1 uncoupling in brown fat mitochondria. Cell 151, 400–413 (2012).
Sárvári, A. K. et al. Plasticity of epididymal adipose tissue in response to diet-induced obesity at single-nucleus resolution. Cell Metab. 33, 437–453 (2021).
Lee, Y. -H., Kim, S. -N., Kwon, H. -J. & Granneman, J. G. Metabolic heterogeneity of activated beige/brite adipocytes in inguinal adipose tissue. Sci. Rep. 7, 39794 (2017).
Rosenwald, M., Perdikari, A., Rülicke, T. & Wolfrum, C. Bi-directional interconversion of brite and white adipocytes. Nat. Cell Biol. 15, 659–667 (2013).
Kong, X. et al. IRF4 is a key thermogenic transcriptional partner of PGC-1α. Cell 158, 69–83 (2014).
Alwarawrah, Y. et al. Fasnall, a selective FASN inhibitor, shows potent anti-tumor activity in the MMTV-Neu model of HER2+ breast cancer. Cell Chem. Biol. 23, 678–688 (2016).
Rothwell, N. J. & Stock, M. J. Surgical removal of brown fat results in rapid and complete compensation by other depots. Am. J. Physiol. Integr. Comp. Physiol. 257, R253–R258 (1989).
Kong, X. et al. Brown adipose tissue controls skeletal muscle function via the secretion of myostatin. Cell Metab. 28, 631–643 (2018).
Qing, H. et al. Origin and function of stress-induced IL-6 in murine models. Cell 182, 372–387 (2020).
Emmett, M. J. et al. Histone deacetylase 3 prepares brown adipose tissue for acute thermogenic challenge. Nature 546, 544–548 (2017).
Virtue, S. & Vidal-Puig, A. GTTs and ITTs in mice: simple tests, complex answers. Nat. Metab. 3, 883–886 (2021).
Bray, N. L., Pimentel, H., Melsted, P. & Pachter, L. Near-optimal probabilistic RNA-seq quantification. Nat. Biotechnol. 34, 525–527 (2016).
R Core Team. R: A language and environment for statistical computing. R Foundation for Statistical Computing http://www.R-project.org/ (2013).
Huber, W. et al. Orchestrating high-throughput genomic analysis with Bioconductor. Nat. Methods 12, 115–121 (2015).
Soneson, C., Love, M. I. & Robinson, M. D. Differential analyses for RNA-seq: transcript-level estimates improve gene-level inferences. F1000Res 4, 1521 (2015).
Robinson, M. D., McCarthy, D. J. & Smyth, G. K. edgeR: a Bioconductor package for differential expression analysis of digital gene expression data. Bioinformatics 26, 139–140 (2010).
Ritchie, M. E. et al. limma powers differential expression analyses for RNA-sequencing and microarray studies. Nucleic Acids Res. 43, e47 (2015).
Chen, E. Y. et al. Enrichr: interactive and collaborative HTML5 gene list enrichment analysis tool. BMC Bioinformatics 14, 128 (2013).
Kuleshov, M. V. et al. Enrichr: a comprehensive gene-set enrichment analysis web server 2016 update. Nucleic Acids Res. 44, W90–W97 (2016).
Xie, Z. et al. Gene set knowledge discovery with Enrichr. Curr. Protoc. 1, e90 (2021).
Stuart, T. et al. Comprehensive integration of single-cell data. Cell 177, 1888–1902 (2019).
Hafemeister, C. & Satija, R. Normalization and variance stabilization of single-cell RNA-seq data using regularized negative binomial regression. Genome Biol. 20, 296 (2019).
Finak, G. et al. MAST: a flexible statistical framework for assessing transcriptional changes and characterizing heterogeneity in single-cell RNA sequencing data. Genome Biol. 16, 278 (2015).
Linderman, G. C. et al. Zero-preserving imputation of single-cell RNA-seq data. Nat. Commun. 13, 192 (2022).
Acknowledgements
We thank all members of the laboratory of C.A.T. for valuable discussions and input, as well as members of the P.V.S. laboratory (J. Ishibashi and A. Angueira), the J. Wherry laboratory (Z. Chen) and D. Allman laboratory (B. Gaudette) at UPenn for their scientific and technical advice. We thank the Penn Metabolomics Core in the Penn Cardiovascular Institute for LC–MS quantification of acylcarnitines. We acknowledge L. Micha (UPenn) for excellent mouse husbandry. We further thank M. Lazar (UPenn) and C. Wolfrum (ETH Zurich) for Scapfl/fl Ucp1-CreER mice. We thank C. Semenkovich (WashU) for sharing Fasnfl/fl mice. We gratefully acknowledge M. Lazar, J. Henao-Mejia, R. Faryabi and N. Betley (all at UPenn) for scientific advice throughout this study. C.A.T. is a Pew Biomedical Scholar and a Kathryn W. Davis Aging Brain Scholar, and is supported by a National Institutes of Health Director’s New Innovator Award (DP2AG067492), the Edward Mallinckrodt, Jr. Foundation, the Global Probiotics Council, the Mouse Microbiome Metabolic Research Program of the National Mouse Metabolic Phenotyping Centers, and grants by the IDSA Foundation, the Thyssen Foundation, the PennCHOP Microbiome Program, the Penn Institute for Immunology, the Penn Center for Molecular Studies in Digestive and Liver Diseases (P30-DK-050306), the Penn Skin Biology and Diseases Resource-based Center (P30-AR-069589), the Penn Diabetes Research Center (P30-DK-019525) and the Penn Institute on Aging. This work was further supported by a National Institutes of Health Training Grant T32AI141393 (to P.L.), Fellowship Grant F31HL160065 (to P.L.), Medical Scientist Training Program T32 GM07170 (to P.V.S. and L.L.), Training Grant in Computational Biology 5-T32-HG-000046-21 (to L.L.), Boehringer Ingelheim Fonds MD Fellowship (to S.K.), and University of Pennsylvania Center for Undergraduate Research Fellowship (to J.C.). Graphical images in Figs. 1a and Fig. 2a and Extended Data Fig. 3a were created with BioRender.com.
Author information
Authors and Affiliations
Contributions
P.L. conceived the study, designed and performed the experiments, interpreted the results, and wrote the manuscript. P.V.S., L.D., G.T.U., S.K., H.C.D., C.D., J.C., K. Chellappa, T.O.C., Y.H., S.R.P., C.S. and C.P. performed experiments. K. Coleman, L.L., P.V.S., K.B. and M. Li performed computational and statistical analyses. P.S., M. Levy, M. Li, K.E.W., N.W.S., J.A.B., A.R. and O.S. provided essential tools and insights. C.A.T. conceived the study, designed the experiments, interpreted the results and wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Metabolism thanks the anonymous reviewers for their contribution to the peer review of this work. Primary Handling Editor: Isabella Samuelson, in collaboration with the Nature Metabolism team.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Whole-body energy expenditure is increased in a secondary cold challenge 4 days after a primary cold challenge.
a, Whole-body energy expenditure over time in the primary cold challenge compared to secondary cold challenge of the same mice over time (n = 6 independent animals). b, Area under the curve (AUC) analysis of energy expenditure over time in primary and secondary cold challenge (9am-5pm). Error bars indicate means ± s.e.m. *, P < 0.05. Exact P values are presented in the source data file for Extended Data Fig. 1.
Extended Data Fig. 2 The transcriptional induction of lipid biosynthesis following a primary thermogenic response is specific to BAT.
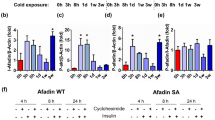
a, Relative gene expression of Ucp1 in iWAT and BAT following transient cold exposure (n = 5 independent animals per condition, except iWAT Day 1 n = 4, iWAT Day 2 n = 2, and iWAT Day 4 n = 4). b, Relative gene expression of Fasn in iWAT and BAT following transient cold exposure (n = 5 independent animals per condition, except iWAT Day 1 n = 4, iWAT Day 2 n = 3, and iWAT Day 4 n = 4). c, Relative gene expression of Fasn in BAT, liver, and muscle from mice in their primary thermogenic response (1cyc) and secondary thermogenic response (2cyc) (n = 5 independent animals per condition). d, Western blot of FASN protein in BAT from cold-naïve mice compared to cold-experienced mice, and relative expression analysis compared to housekeeping gene (Vinculin) (n = 4 independent animals per condition). e, Western blot of UCP1 protein in BAT from cold-naïve mice compared to cold-experienced mice, and relative expression analysis compared to housekeeping gene (Tubulin) (n = 4 independent animals per condition). f, Representative H&E histology images of BAT from cold-naïve mice and cold-experienced mice 4 days after a transient cold exposure (n = 4 independent animals per condition), scale bar = 50μm. Error bars indicate means ± s.e.m. ns, not significant; *, P < 0.05; **, P < 0.01; ***, P < 0.001; ****, P < 0.0001. Exact P values are presented in the source data file for Extended Data Fig. 2.
Extended Data Fig. 3 Workflow and validation of nuclei isolation from BAT.
a, Workflow schematic. b, Nuclei sorting strategy using DAPI positivity, including a re-sort validation experiment. c, Inspection and counting of nuclei quality on haemocytometer.
Extended Data Fig. 4 Single-nucleus RNA-sequencing of brown adipose tissue resolves cell types and adipocyte subpopulations.
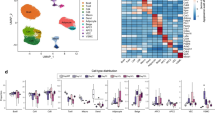
a, Expression of canonical marker genes of cell types identified by BAT snRNAseq. b-c, UMAP plots of cell types separated by cold-naïve BAT (b) and cold-experienced BAT (4 days following transient acute cold) (c). d, Cell type distributions separated by condition. e, Feature plots for Fasn, Ucp1, and Cpt2 expression within the adipocyte population. f, Dot plot for Slc7a10, Ucp1, and Fasn expression across five identified adipocyte subpopulations (A1-A5).
Extended Data Fig. 5 Cell type distributions in spatial transcriptomics data and snRNAseq data.
a, b, Cell type distributions quantified in spatial transcriptomics data (a) and snRNAseq data (b). c, Violin plots for the expression of indicated genes across all spots in each condition from BAT spatial transcriptomics data.
Extended Data Fig. 6 Total free fatty acids, glycerolipids, or phospholipids & sphingolipids in brown adipose tissue following a primary thermogenic response.
a-d, Relative abundance and distribution of free fatty acids (a), glycerolipids (b), and phospholipid & sphingolipid (c) species in BAT in different experimental conditions. Data presented in panels a-d is based on metabolomics from n = 5 independent animals per condition. Each data point represents the average value across replicates for each species. d, Comparison of indicated lipid metabolite species in BAT between TN, day 4, and day 4 with Fasn inhibition (Fasni). Each data point represents the average value across replicates for each species. Error bars indicate means ± s.e.m. ns, not significant; *, P < 0.05; ***, P < 0.001; ****, P < 0.0001. Exact P values are presented in the source data file for Extended Data Fig. 6.
Extended Data Fig. 7 Scap knockout in brown adipocytes does not affect the expression of genes involved in acylcarnitine transport or fatty acid oxidation.
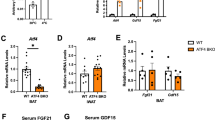
a-c, Relative gene expression of Cact (a), Lcad (b), and Mcad (c) in brown adipose tissue from ScapΔUcp1 (n = 7 independent animals) and Scapflox mice (n = 7 independent animals) 4 days after primary cold exposure. Error bars indicate means ± s.e.m. ns, not significant. Exact P values are presented in the source data file for Extended Data Fig. 7.
Supplementary information
Supplementary Information
Supplementary Table 1
Source data
Source Data Fig. 1
Source data.
Source Data Fig. 2
Source data.
Source Data Fig. 3
Source data.
Source Data Fig. 4
Source data.
Source Data Fig. 5
Source data.
Source Data Fig. 6
Source data.
Source Data Extended Data Fig. 1
Source data.
Source Data Extended Data Fig. 2
Source data.
Source Data Extended Data Fig. 4
Source data.
Source Data Extended Data Fig. 5
Source data.
Source Data Extended Data Fig. 6
Source data.
Source Data Extended Data Fig. 7
Source data.
Source Data Extended Data Fig. 2
Unprocessed western blot gels.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Lundgren, P., Sharma, P.V., Dohnalová, L. et al. A subpopulation of lipogenic brown adipocytes drives thermogenic memory. Nat Metab 5, 1691–1705 (2023). https://doi.org/10.1038/s42255-023-00893-w
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s42255-023-00893-w
This article is cited by
-
Futile lipid cycling: from biochemistry to physiology
Nature Metabolism (2024)
-
Fuelling the fire: de novo lipogenesis primes thermogenesis
Nature Metabolism (2023)