Abstract
Microbial biochemistry contributes to a dynamic environment in the gut. Yet, how bacterial metabolites such as hydrogen sulfide (H2S) mechanistically alter the gut chemical landscape is poorly understood. Here we show that microbially generated H2S drives the abiotic reduction of azo (R–N = N–R’) xenobiotics, which are commonly found in Western food dyes and drugs. This nonenzymatic reduction of azo compounds is demonstrated in Escherichia coli cultures, in human faecal microbial communities and in vivo in male mice. Changing dietary levels of the H2S xenobiotic redox partner Red 40 transiently decreases mouse faecal sulfide levels, demonstrating that a xenobiotic can attenuate sulfide concentration and alleviate H2S accumulation in vivo. Cryptic H2S redox chemistry thus can modulate sulfur homeostasis, alter the chemical landscape in the gut and contribute to azo food dye and drug metabolism. Interactions between chemicals derived from microbial communities may be a key feature shaping metabolism in the gut.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
Data supporting the findings of this study are available within the paper and Supplementary Information. No custom computer code was used to generate results reported in the paper. Correspondence and requests for materials may be addressed to L.K. Source data are provided with this paper.
References
Magee, E. A., Richardson, C. J., Hughes, R. & Cummings, J. H. Contribution of dietary protein to sulfide production in the large intestine: an in vitro and a controlled feeding study in humans. Am. J. Clin. Nutr. 72, 1488–1494 (2000).
Barton, L. L., Ritz, N. L., Fauque, G. D. & Lin, H. C. Sulfur cycling and the intestinal microbiome. Dig. Dis. Sci. 62, 2241–2257 (2017).
Hanson, B. T. et al. Sulfoquinovose is a select nutrient of prominent bacteria and a source of hydrogen sulfide in the human gut. ISME J. https://doi.org/10.1038/s41396-021-00968-0 (2021).
Guidotti, T. L. Hydrogen sulfide: advances in understanding human toxicity. Int J. Toxicol. 29, 569–581 (2010).
Shatalin, K., Shatalina, E., Mironov, A. & Nudler, E. H2S: a universal defense against antibiotics in bacteria. Science 334, 986–990 (2011).
Nguyen, L. H. et al. The sulfur microbial diet is associated with increased risk of early-onset colorectal cancer precursors. Gastroenterology 161, 1423–1432 (2021).
Wolf, P. G. et al. Diversity and distribution of sulfur metabolic genes in the human gut microbiome and their association with colorectal cancer. Microbiome 10, 64 (2022).
Motta, J.-P. et al. Hydrogen sulfide protects from colitis and restores intestinal microbiota biofilm and mucus production. Inflamm. Bowel Dis. 21, 1006–1017 (2015).
Hu, L.-F. et al. Neuroprotective effects of hydrogen sulfide on Parkinson’s disease rat models. Aging Cell 9, 135–146 (2010).
Stevens, L. J., Burgess, J. R., Stochelski, M. A. & Kuczek, T. Amounts of artificial food dyes and added sugars in foods and sweets commonly consumed by children. Clin. Pediatr. 53, 133–140 (2015).
Zou, L. et al. Bacterial metabolism rescues the inhibition of intestinal drug absorption by food and drug additives. Proc. Natl Acad. Sci. USA 117, 16009–16018 (2020).
He, Z. et al. Food colorants metabolized by commensal bacteria promote colitis in mice with dysregulated expression of interleukin-23. Cell Metab. 33, 1358–1371 (2021).
Braccia, D. J., Jiang, X., Pop, M. & Hall, A. B. The capacity to produce hydrogen sulfide (H2S) via cysteine degradation is ubiquitous in the human gut microbiome. Front. Microbiol. 12, 705583 (2021).
Lobel, L., Cao, Y. G., Fenn, K., Glickman, J. N. & Garrett, W. S. Diet posttranslationally modifies the mouse gut microbial proteome to modulate renal function. Science 369, 1518–1524 (2020).
Rey, F. E. et al. Metabolic niche of a prominent sulfate-reducing human gut bacterium. Proc. Natl Acad. Sci. USA 110, 13582–13587 (2013).
Pereira, F. C. & Berry, D. Microbial nutrient niches in the gut. Environ. Microbiol. 19, 1366–1378 (2017).
Blachier, F., Beaumont, M. & Kim, E. Cysteine-derived hydrogen sulfide and gut health: a matter of endogenous or bacterial origin. Curr. Opin. Clin. Nutr. Metab. Care 22, 68–75 (2019).
Shen, X. et al. Microbial regulation of host hydrogen sulfide bioavailability and metabolism. Free Radic. Biol. Med. 60, 195–200 (2013).
Ryan, A. Azoreductases in drug metabolism. Br. J. Pharmacol. 174, 2161–2173 (2017).
van der Zee, F. P. et al. The contribution of biotic and abiotic processes during azo dye reduction in anaerobic sludge. Water Res. 37, 3098–3109 (2003).
Blachier, F. et al. Luminal sulfide and large intestine mucosa: friend or foe? Amino Acids 39, 335–347 (2010).
Costa, M. C., Mota, F. S. B., Santos, A. B. D., Mendonça, G. L. F. & Nascimento, R. Fdo Effect of dye structure and redox mediators on anaerobic azo and anthraquinone dye reduction. Quím. Nova 35, 482–486 (2012).
van der Zee, F. P., Bouwman, R. H. M., Strik, D. P. B. T. B., Lettinga, G. & Field, J. A. Application of redox mediators to accelerate the transformation of reactive azo dyes in anaerobic bioreactors. Biotechnol. Bioeng. 75, 691–701 (2001).
Awano, N., Wada, M., Mori, H., Nakamori, S. & Takagi, H. Identification and functional analysis of Escherichia coli cysteine desulfhydrases. Appl. Environ. Microbiol. 71, 4149–4152 (2005).
Canfield, D. E. et al. A cryptic sulfur cycle in oxygen-minimum-zone waters off the Chilean coast. Science 330, 1375–1378 (2010).
Nayfach, S., Fischbach, M. A. & Pollard, K. S. MetaQuery: a web server for rapid annotation and quantitative analysis of specific genes in the human gut microbiome. Bioinformatics 31, 3368–3370 (2015).
Loddeke, M. et al. Anaerobic cysteine degradation and potential metabolic coordination in Salmonella enterica and Escherichia coli. J. Bacteriol. 199, e00117-17 (2017).
Bastie, C. C. et al. Dietary cholecalciferol and calcium levels in a Western-style defined rodent diet alter energy metabolism and inflammatory responses in mice. J. Nutr. 142, 859–865 (2012).
Miller, R. A. et al. Methionine-deficient diet extends mouse lifespan, slows immune and lens aging, alters glucose, T4, IGF-I and insulin levels, and increases hepatocyte MIF levels and stress resistance. Aging Cell 4, 119–125 (2005).
Reese, A. T. et al. Antibiotic-induced changes in the microbiota disrupt redox dynamics in the gut. eLife 7, e35987 (2018).
Shatalin, K. et al. Inhibitors of bacterial H2S biogenesis targeting antibiotic resistance and tolerance. Science 372, 1169–1175 (2021).
Kondo, K. et al. H2S protects against pressure overload-induced heart failure via upregulation of endothelial nitric oxide synthase. Circulation 127, 1116–1127 (2013).
Yang, G. et al. H2S as a physiologic vasorelaxant: hypertension in mice with deletion of cystathionine γ-lyase. Science 322, 587–590 (2008).
Wolfe, R. S. Techniques for cultivating methanogens. Methods Enzymol. 494, 1–22 (2011).
A. Webster, T., J. Sismaet, H., J. Chan, I.-ping & D. Goluch, E. Electrochemically monitoring the antibiotic susceptibility of Pseudomonas aeruginosa biofilms. Analyst 140, 7195–7201 (2015).
R Core Team. R: a language and environment for statistical computing. R Foundation for Statistical Computing. https://www.R-project.org/ (2020).
Wickham, H. ggplot2: Elegant Graphics for Data Analysis (Springer-Verlag, 2016).
Cline, J. D. Spectrophotometric determination of hydrogen sulfide in natural waters. Limnol. Oceanogr. 14, 454–458 (1969).
Strocchi, A., Furne, J. K. & Levitt, M. D. A modification of the methylene blue method to measure bacterial production in feces. J. Microbiol. Methods 15, 75–82 (1992).
Phelps, C. D. & Young, L. Y. Anaerobic biodegradation of BTEX and gasoline in various aquatic sediments. Biodegradation 10, 15–25 (1999).
Baker, F. D., Papiska, H. R. & Campbell, L. L. Choline fermentation by Desulfovibrio desulfuricans. J. Bacteriol. 84, 973–978 (1962).
Benson, D. A. et al. GenBank. Nucleic Acids Res. 41, D36–D42 (2013).
Kanehisa, M. & Goto, S. KEGG: Kyoto Encyclopedia of Genes and Genomes. Nucleic Acids Res. 28, 27–30 (2000).
Huerta-Cepas, J. et al. eggNOG 4.5: a hierarchical orthology framework with improved functional annotations for eukaryotic, prokaryotic and viral sequences. Nucleic Acids Res. 44, D286–D293 (2016).
Edgar, R. C. Search and clustering orders of magnitude faster than BLAST. Bioinformatics 26, 2460–2461 (2010).
Edgar, R. C. MUSCLE: multiple sequence alignment with high accuracy and high throughput. Nucleic Acids Res. 32, 1792–1797 (2004).
Eddy, S. R. Accelerated Profile HMM Searches. PLOS Comput. Biol. 7, e1002195 (2011).
Pasolli, E. et al. Accessible, curated metagenomic data through ExperimentHub. Nat. Methods 14, 1023–1024 (2017).
Segata, N. et al. Metagenomic microbial community profiling using unique clade-specific marker genes. Nat. Methods 9, 811–814 (2012).
F, M. et al. The EMBL-EBI search and sequence analysis tools APIs in 2019. Nucleic Acids Res. 47, W636–W641 (2019).
Trifinopoulos, J., Nguyen, L.-T., von Haeseler, A. & Minh, B. Q. W-IQ-TREE: a fast online phylogenetic tool for maximum likelihood analysis. Nucleic Acids Res. 44, W232–W235 (2016).
Letunic, I. & Bork, P. Interactive Tree Of Life (iTOL) v5: an online tool for phylogenetic tree display and annotation. Nucleic Acids Res. https://doi.org/10.1093/nar/gkab301 (2021).
Li, W. et al. The nutritional environment determines which and how intestinal stem cells contribute to homeostasis and tumorigenesis. Carcinogenesis 40, 937–946 (2019).
Acknowledgements
The authors thank A. Ryan (Northumbria University) for sharing knowledge of azoreductase enzymes, and the laboratories of S. Almo and T. Grove (Albert Einstein College of Medicine) for experimental and intellectual support in the related work. L.K. was supported in part by a US Department of Defense Cancer Research Program Career Development Award (CA171019) and the National Institutes of Health NHLBI (R01HL069438-21). R.H. was supported by the Albert Einstein College of Medicine PhD in Clinical Investigation training grant (TL1TR001072). S.K. was supported by the Einstein Medical Scientist Training Program (2T32GM007288) and a National Institutes of Health T32 Fellowship in Geographic Medicine and Emerging Infectious Diseases (2T32AI070117). L.A. was supported by National Cancer Institute Department of Cancer Prevention Nutrition grants (1R01CA214625 and 1R01CA229216). Support was also received by the Albert Einstein Cancer Center core support grant (P30CA013330).
Author information
Authors and Affiliations
Contributions
S.J.W., R.H., L.K. and L.A. drafted the manuscript. S.J.W. performed E. coli experiments, inactivated faecal experiments, metagenomic analysis and mouse faecal sulfide determination. R.H. performed abiotic and faecal slurry experiments, as well as MS. K.P. performed all mouse experimentation and maintained the mouse colony. L.A. guided mouse experiments and diet formulation. T.Y. performed cyclic voltammetry of azo compounds. E.D.G. guided electrochemistry experiments. M.M. and Z.C. assisted with faecal slurry experiments. S.K. assisted with metagenomic analysis. All authors edited the manuscript and contributed intellectually.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Primary Handling Editor: Yanina-Yasmin Pesch, in collaboration with the Nature Metabolism team.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Azoreduction with and without FMN.
Normalized MS intensities of azo compounds and their proposed azoreduced metabolites (mean ± sd, n = 3 independent incubations per compound). Dashed lines indicate conditions without FMN as an electron shuttle, solid lines indicate conditions supplemented with FMN as an electron shuttle.
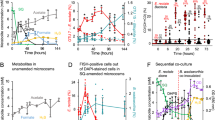
Extended Data Fig. 2 Dietary cysteine and faecal sulfide.
Mice fed 8 g/kg cysteine (High Cys) could not be distinguished from mice fed 4 g/kg cysteine (Control Cys) after 1 week. After 2 weeks, fecal sulfide is lower in 0 Cysteine diet than either Control or High Cysteine diets. (n = 30, 10/diet). Boxes show the upper and lower quartiles, and whiskers depict range excluding outliers. Outliers are defined as points > 1.5 times the inter-quartile range. Statistical difference was determined by one-way ANOVA, Tukey’s test.
Extended Data Fig. 3 Initial rate of azoreduction in active faecal microcosm.
Rate of initial azoreduction in faecal microcosms (as in Fig. 3a, n = 3 biologically independent incubations per condition) (mean ± SEM). Conditions azoreduce at different rates (one-way ANOVA, p = 0.0004), Mucin azoreduces faster than the Enzymatic Control (Dunnett’s multiple comparisons test, p = 0.047).
Extended Data Fig. 4 Active faecal microcosm azoreduction with and without sulfate amendment.
Red 40 azoreduction does not differ between Enzymatic Control and cultures amended with sulfate (unpaired, two-sided Wilcoxon rank-sum test, p > 0.05 all timepoints) (mean ± sd; Cx/C0, ratio of remaining Red 40 to 500 µM starting amendment; n = 3 biological replicates per condition). Minimal hydrogen sulfide accumulates in the 6.5 hour experimental window regardless of Red 40 presence.
Extended Data Fig. 5 Dissimilatory sulfate reduction in active faecal microcosm during extended incubation.
The healthy human faecal microcosm was incubated for ~3.5 days (90.5 hours) to allow the sulfate reducing bacterial community to acclimate. No additional thiol or sulfur sources were provided in the chemically defined media. Hydrogen sulfide accumulation, beginning after 18.5 hours, indicated dissimilatory sulfate reduction activity, and cultures were autoclaved following a 90.5 hour incubation for the heat inactivated fecal SRB experiment (Fig. 3c). H2S does not differ between the two conditions incubated with sulfate allocated for the heat inactivated faecal SRB experiment (mean ± sd; sulfate/sulfate control, unpaired, two-sided Wilcoxon rank-sum test, p > 0.05 all timepoints; n = 3 biological replicates per condition).
Supplementary information
Supplementary Tables 1 and 2
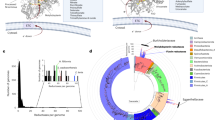
Supplementary Table 1 Profile of sulfidogenic cysteine-degrading genes in human gut symbiont genomes (Fig. 3d). Presence (1) or absence (0) of nine genes (CBS (K01697), CSE (K01758), cysK (K01738), cysM (K12339), cyuA (COG3681), malY (K14155), metC (K01760), sseA (K01011) and tnaA (K01667)) encoding sulfidogenic cysteine-degrading proteins in genomes of common, non-pathogenic human gut microorganisms. The subset of gut symbionts was identified from 8,548 metagenomic samples of participants from 51 studies using the R package curatedMetagenomicData48 (March 2021). To identify each gene in a genome, a reference database for each gene was aligned and HMMs were constructed. The nine HMMs were searched against bacterial genomes using hmmsearch with a cutoff of 1 × 10−10. One hit in a genome indicated the presence of the gene in a particular genome; multiple hits were ignored. Supplementary Table 2 Profile of sulfidogenic cysteine-degrading genes in bacteria with completely sequenced genomes. Presence (1) or absence (0) of nine genes ((CBS (K01697), CSE (K01758), cysK (K01738), cysM (K12339), cyuA (COG3681), malY (K14155); metC (K01760); sseA (K01011) and tnaA (K01667)) encoding sulfidogenic cysteine-degrading proteins in 24,758 NCBI bacterial genomes (GenBank, April 2021)42. To identify each gene in a genome, a reference database for each gene was aligned and HMMs were constructed. The nine HMMs were searched against bacterial genomes using hmmsearch with a cutoff of 1 × 10−10. One hit in a genome indicated the presence of the gene in a particular genome; multiple hits were ignored.
Source data
Source Data Fig. 1
MS and half-life data plotted in Fig. 1.
Source Data Fig. 2
Red 40 and sulfide concentrations plotted in Fig. 2.
Source Data Fig. 3
Red 40 and sulfide concentrations plotted in Fig. 3.
Source Data Fig. 4
Mouse cohort and sulfide concentrations plotted in Fig. 4.
Source Data Extended Data Fig. 1
MS data plotted in Extended Data Fig. 1.
Source Data Extended Data Fig. 2
Mouse cohort and sulfide concentrations plotted in Extended Data Fig. 2.
Source Data Extended Data Fig. 3
Red 40 loss rates as plotted in Extended Data Fig. 3.
Source Data Extended Data Fig. 4
Red 40 loss ratio as plotted in Extended Data Fig. 4.
Source Data Extended Data Fig. 5
Sulfide concentrations as plotted in Extended Data Fig. 5.
Rights and permissions
Springer Nature or its licensor holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Wolfson, S.J., Hitchings, R., Peregrina, K. et al. Bacterial hydrogen sulfide drives cryptic redox chemistry in gut microbial communities. Nat Metab 4, 1260–1270 (2022). https://doi.org/10.1038/s42255-022-00656-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s42255-022-00656-z
This article is cited by
-
Delayed gut microbiota maturation in the first year of life is a hallmark of pediatric allergic disease
Nature Communications (2023)
-
Gut reaction: it’s not all about enzymes
Nature Metabolism (2022)