Abstract
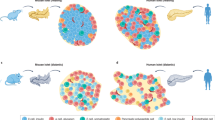
β-cell dysfunction is a hallmark of type 1 and type 2 diabetes. Type 2 diabetes is strongly associated with ageing-related β-cell abnormalities that arise through unknown mechanisms. Here we show better β-cell identity, less β-cell senescence, enhanced glucose-stimulated insulin secretion and improved glucose homeostasis in global microsomal prostaglandin E synthase-2 (mPGES-2)-deficient mice challenged with a high-fat diet or bred with a genetic model of type 2 diabetes (db/db mice). Furthermore, the function of mPGES-2 in β-cells is validated using mice with β-cell-specific mPGES-2 deficiency or overexpression. Mechanistically, the protective role of mPGES-2 deletion is induced by antagonizing β-cell senescence via interference of the PGE2–EP3–NR4A1 signalling axis. We also discover an inhibitor of mPGES-2, SZ0232, which protects against β-cell dysfunction and diabetes, similar to mPGES-2 deletion. We conclude that mPGES-2 contributes to ageing-associated β-cell senescence and dysfunction via the PGE2–EP3–NR4A1 signalling axis. Pharmacologic blockade of mPGES-2 might be effective for treating ageing-associated β-cell dysfunction and diabetes.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
Data availability
Source data are provided with this paper. All other data are available in the manuscript or the extended data. Additional source data are available from the corresponding authors upon reasonable request. RNA-seq data were deposited in NCBI Sequence Read Archive (SRA) under the accession number PRJNA795309.
References
Weir, G. C. & Bonner-Weir, S. Islet β cell mass in diabetes and how it relates to function, birth, and death. Ann. NY Acad. Sci. 1281, 92 (2013).
Aguayo-Mazzucato, C. et al. Acceleration of β cell aging determines diabetes and senolysis improves disease outcomes. Cell Metab. 30, 129–142 (2019).
White, M. G., Shaw, J. A. & Taylor, R. Type 2 diabetes: the pathologic basis of reversible β-cell dysfunction. Diabetes care 39, 2080–2088 (2016).
Irles, E. et al. Enhanced glucose-induced intracellular signaling promotes insulin hypersecretion: pancreatic beta-cell functional adaptations in a model of genetic obesity and prediabetes. Mol. Cell. Endocrinol. 404, 46–55 (2015).
Aguayo-Mazzucato, C. et al. β cell aging markers have heterogeneous distribution and are induced by insulin resistance. Cell Metab. 25, 898–910 (2017).
Helman, A. et al. Effects of ageing and senescence on pancreatic β-cell function. Diabetes Obes. Metab. 18, 58–62 (2016).
Tan, S. Y. et al. Type 1 and 2 diabetes mellitus: a review on current treatment approach and gene therapy as potential intervention. Diabetes Metab. Syndr. Clin. Res. Rev. 13, 364–372 (2019).
Amiel, S. A., Dixon, T., Mann, R. & Jameson, K. Hypoglycaemia in type 2 diabetes. Diabet. Med. 25, 245–254 (2008).
Cryer, P. E. Mechanisms of hypoglycemia-associated autonomic failure in diabetes. N. Engl. J. Med. 369, 362–372 (2013).
Hara, S. et al. Prostaglandin E synthases: understanding their pathophysiological roles through mouse genetic models. Biochimie 92, 651–659 (2010).
Consalvi, S., Biava, M. & Poce, G. COX inhibitors: a patent review (2011–2014). Expert Opin. Ther. Pat. 25, 1357–1371 (2015).
Corso, G. et al. A novel multi-step virtual screening for the identification of human and mouse mPGES-1 inhibitors. Mol. Inf. 35, 358–368 (2016).
Takusagawa, F. Microsomal prostaglandin E synthase type 2 (mPGES2) is a glutathione-dependent heme protein, and dithiothreitol dissociates the bound heme to produce active prostaglandin E2 synthase in vitro. J. Biol. Chem. 288, 10166–10175 (2013).
Yamada, T. & Takusagawa, F. PGH2 degradation pathway catalyzed by GSH-heme complex bound microsomal prostaglandin E2 synthase type 2: the first example of a dual-function enzyme. Biochemistry 46, 8414–8424 (2007).
Sun, Y. et al. mPGES-2 deletion remarkably enhances liver injury in streptozotocin-treated mice via induction of GLUT2. J. Hepatol. 61, 1328–1336 (2014).
Jania, L. A. et al. Microsomal prostaglandin E synthase-2 is not essential for in vivo prostaglandin E2 biosynthesis. Prostaglandins Other Lipid Mediat. 88, 73–81 (2009).
Wang, H. et al. Microsomal prostaglandin E synthase 2 deficiency is resistant to acetaminophen-induced liver injury. Arch. Toxicol. 93, 2863–2878 (2019).
Fisher, E. et al. Candidate gene association study of type 2 diabetes in a nested case-control study of the EPIC-Potsdam cohort–role of fat assimilation. Mol. Nutr. Food Res. 51, 185–191 (2007).
Lindner, I. et al. Prostaglandin E synthase 2 (PTGES2) Arg298His polymorphism and parameters of the metabolic syndrome. Mol. Nutr. Food Res. 51, 1447–1451 (2007).
Fischer, A. et al. Association analysis between the prostaglandin E synthase 2 R298H polymorphism and body mass index in 8079 participants of the KORA study cohort. Genet. Test. Mol. Biomark. 13, 223–226 (2009).
Stott, N. L. & Marino, J. S. High fat rodent models of type 2 diabetes: from rodent to human. Nutrients 12, 3650 (2020).
Parazzoli, S. et al. Cyclooxygenase-2, not microsomal prostaglandin E synthase-1, is the mechanism for interleukin-1β-induced prostaglandin E2 production and inhibition of insulin secretion in pancreatic islets. J. Biol. Chem. 287, 32246–32253 (2012).
Granot, Z. et al. LKB1 regulates pancreatic β cell size, polarity, and function. Cell Metab. 10, 296–308 (2009).
Zhan, Y. Y. et al. The orphan nuclear receptor Nur77 regulates LKB1 localization and activates AMPK. Nat. Chem. Biol. 8, 897–904 (2012).
Kovalovsky, D. et al. Activation and induction of NUR77/NURR1 in corticotrophs by CRH/cAMP: involvement of calcium, protein kinase A, and MAPK pathways. Mol. Endocrinol. 16, 1638–1651 (2002).
Maxwell, M. A. et al. Nur77 regulates lipolysis in skeletal muscle cells evidence for cross-talk between the β-adrenergic and an orphan nuclear hormone receptor pathway. J. Biol. Chem. 280, 12573–12584 (2005).
Pei, L. et al. NR4A orphan nuclear receptors are transcriptional regulators of hepatic glucose metabolism. Nat. Med. 12, 1048–1055 (2006).
Kahn, S. E., Cooper, M. E. & Del Prato, S. Pathophysiology and treatment of type 2 diabetes: perspectives on the past, present, and future. Lancet 383, 1068–1083 (2014).
Eguchi, K. & Nagai, R. Islet inflammation in type 2 diabetes and physiology. J. Clin. Invest. 127, 14–23 (2017).
Ji, Y. et al. Toll-like receptors TLR2 and TLR4 block the replication of pancreatic β cells in diet-induced obesity. Nat. Immunol. 20, 677–686 (2019).
Wang, D. & DuBois, R. N. Role of prostanoids in gastrointestinal cancer. J. Clin. Invest. 128, 2732–2742 (2018).
Nasrallah, R., Hassouneh, R. & Hebert, R. L. PGE2, kidney disease, and cardiovascular risk: beyond hypertension and diabetes. J. Am. Soc. Nephrol. 27, 666–676 (2016).
Kimple, M. E. et al. Prostaglandin E2 receptor, EP3, is induced in diabetic islets and negatively regulates glucose-and hormone-stimulated insulin secretion. Diabetes 62, 1904–1912 (2013).
Carboneau, B. A., Breyer, R. M. & Gannon, M. Regulation of pancreatic β-cell function and mass dynamics by prostaglandin signaling. J. Cell Commun. Signal. 11, 105–116 (2017).
Neuman, J. C. et al. Enriching islet phospholipids with eicosapentaenoic acid reduces prostaglandin E2 signaling and enhances diabetic β-cell function. Diabetes 66, 1572–1585 (2017).
Robertson, R. P. The COX-2/PGE2/EP3/Gi/o/cAMP/GSIS pathway in the islet: the beat goes on. Diabetes 66, 1464–1466 (2017).
Amior, L., Srivastava, R., Nano, R., Bertuzzi, F. & Melloul, D. The role of Cox‐2 and prostaglandin E2 receptor EP3 in pancreatic β-cell death. FASEB J. 33, 4975–4986 (2019).
Chao, L. C. et al. Nur77 coordinately regulates expression of genes linked to glucose metabolism in skeletal muscle. Mol. Endocrinol. 21, 2152–2163 (2007).
Kanzleiter, T. et al. Overexpression of the orphan receptor Nur77 alters glucose metabolism in rat muscle cells and rat muscle in vivo. Diabetologia 53, 1174–1183 (2010).
Kone, M. et al. LKB1 and AMPK differentially regulate pancreatic β-cell identity. FASEB J. 28, 4972–4985 (2014).
Liu, J. et al. SIRT1 promotes proliferation and prevents senescence through targeting LKB1 in primary porcine aortic endothelial cells. Circ. Res. 106, 1384–1393 (2010).
Baker, D. J. et al. Clearance of p16 Ink4a-positive senescent cells delays ageing-associated disorders. Nature 479, 232–236 (2011).
Krishnamurthy, J. et al. p16 INK4a induces an age-dependent decline in islet regenerative potential. Nature 443, 453–457 (2006).
Uchida, T. et al. Deletion of Cdkn1b ameliorates hyperglycemia by maintaining compensatory hyperinsulinemia in diabetic mice. Nat. Med. 11, 175–182 (2005).
Xiao, X. et al. Pancreatic cell tracing, lineage tagging and targeted genetic manipulations in multiple cell types using pancreatic ductal infusion of adeno-associated viral vectors and/or cell-tagging dyes. Nat. Protoc. 9, 2719–2724 (2014).
Meng, Z. et al. Activation of liver X receptors inhibits pancreatic islet beta cell proliferation through cell cycle arrest. Diabetologia 52, 125 (2009).
Waterhouse, A. et al. SWISS-MODEL: homology modelling of protein structures and complexes. Nucleic Acids Res. 46, W296–W303 (2018).
Studer, G. et al. QMEANDisCo—distance constraints applied on model quality estimation. Bioinformatics 36, 1765–1771 (2020).
Guex, N., Peitsch, M. C. & Schwede, T. Automated comparative protein structure modeling with SWISS-MODEL and Swiss-PdbViewer: a historical perspective. Electrophoresis 30, S162–S173 (2009).
Bienert, S. et al. The SWISS-MODEL Repository—new features and functionality. Nucleic Acids Res. 45, D313–D319 (2017).
Bertoni, M., Kiefer, F., Biasini, M., Bordoli, L. & Schwede, T. Modeling protein quaternary structure of homo- and hetero-oligomers beyond binary interactions by homology. Sci. Rep. 7, 10480 (2017).
Liu, L. et al. Identification of protein arginine methyltransferase 7 (PRMT7) inhibitor by virtual screening and biological evaluation in vitro. Med. Chem. Res. 28, 125–132 (2019).
Acknowledgements
We thank X. Han (Nanjing Medical University) for guidance with islet isolation and sharing the β-cell line, H.G. for language advice, F. Dong (Public Experimental Research Center) and Z. Xu for supporting in the laser scanning confocal microscopy and quality control of compound. This work was supported by the National Natural Science Foundation of China (grant no. 81603179 to Y.S. and 82000683 to D.Z.), the China Postdoctoral Science Foundation (grant no. 2016M601893 to Y.S.), the China Postdoctoral Science Special Foundation (grant no. 2018T110555 to Y.S.), the Key Project of Natural Science Research in Jiangsu Universities (21KJA350002 to Y.S.), Jiangsu Research Innovation Program for College Graduates (KYCX20_2504 to J.C.), the Natural Science Research Project of Jiangsu Universities (20KJB310023 to Z.D.), Jiangsu Province Innovative and Entrepreneurial Team Program (RC7042012) and the Xuzhou Science and Technology Plan Project (KC20150 to Z.D.).
Author information
Authors and Affiliations
Contributions
Y.S. and Z.J. provided the study concept and design. D.Z., Y.S., Z.W., J.C., L.Q. and H.G. contributed to data acquisition and drafting of the manuscript. D.Z., Z.W., T.T., R.Z., C.F., M.W., D.G. and H.Z. participated in critical data analysis and discussion of the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Metabolism thanks Manuel Collado and the other, anonymous, reviewers for their contribution to the peer review of this work. Primary Handling Editor: Isabella Samuelson.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 The expression of mPGES-2 in islet of mice and human.
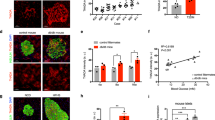
a, Genotyping of mPGES-2 WT and KO mice. Genotyping was repeated more than twice, independently, with similar results. b, Expression of mPGES-2 in different tissues in mPGES-2 WT and KO mice. The experiment was repeated more than twice, independently, with similar results. c, Body weight of mPGES-2 WT and KO mice fed an NCD at different weeks (n = 10 per point except n = 9 for 52 w WT). d, qRT–PCR analysis of Ptges, Ptges2 and Ptges3 expression in WT islets at 8 and 32 weeks (n = 4). e, Labeling for insulin and mPGES-2, or glucagon and mPGES-2 on the same sections in mouse islets; scale bars = 20 μm. The experiment was repeated more than twice, independently, with similar results. f, Labeling for insulin and mPGES-2, or glucagon and mPGES-2 on the same sections in human islets; scale bars = 20 μm. The experiment was repeated more than twice, independently, with similar results. g, Body weight of mPGES-2 WT and KO mice fed an HFD at different weeks (n = 6). Data are expressed as mean ± SEM. Statistical significance was assessed using a two-tailed unpaired Student’s t-test. Exact P values are indicated.
Extended Data Fig. 2 Phenotype of mPGES-1-deficient mice.
a, Genotyping of mPGES-1 WT (Ptges+/+) and KO (Ptges−/−) mice. Genotyping was repeated more than twice, independently, with similar results. b, Body weight of mPGES-1 WT and KO mice fed an NCD at different weeks (n = 5 per group). c, Body weight of mPGES-1 WT and KO mice fed an HFD at different weeks (n = 6 per group). d, IPGTT and AUC of mPGES-1 WT or KO mice fed an HFD at 16 weeks (n = 6). e, IPGTT and AUC of mPGES-1 WT or KO mice fed an NCD at 32 weeks (n = 5). f, IPGTT and AUC of mPGES-1 WT or KO mice fed an NCD at 52 weeks (n = 5). g, Insulin secretion from mouse islets isolated from 16-week-old WT and mPGES-1 KO mice under conditions of low or high glucose (n = 3). h, PGE2 levels in islets isolated from mPGES-1 WT or KO mice (n = 3). Data are expressed as mean ± SEM. Statistical significance was assessed using and a two-tailed unpaired Student’s t-test.
Extended Data Fig. 3 The effect of mPGES-2 on β-cell function under physiological conditions.
a, qRT–PCR analysis of Mafa, Slc2a2, Nkx6-1 and Pdx1 expression in WT and KO islets at 32 weeks (n = 5 per group). b, Representative IF for Glut2 (green) and insulin (red) in islets of WT and KO mice fed an NCD at 16, 52 weeks and an HFD at 52 weeks; scale bars = 20 μm. c, Statistical quantification of IF for Glut2 (n = 4 per group). d, H&E staining of pancreas in db/db Ptges2+/+ and db/db Ptges2−/− mice (n = 4 per group); scale bars = 20 μm. e, Body weights of db/db Ptges2+/+ and db/db Ptges2−/− mice at different weeks (n = 10 per group). f, Representative IF and quantification for Glut2 (green) and insulin (red) in islets of db/db Ptges2+/+ and db/db Ptges2−/− mice; scale bars = 20 μm (n = 5 per group). H&E staining and IF were repeated more than twice, independently and similar results were obtained. Data are expressed as mean ± SEM. Statistical significance was assessed using a two-tailed unpaired Student’s t-test. Exact P values are indicated.
Extended Data Fig. 4 β-cell-specific mPGES-2 deficiency improves β-cell senescence and glucose metabolism.
Ptges2flox/flox mice (six-week-old) were injected with AAV8-MIP-control or AAV8-MIP-cre via pancreatic intraductal viral infusion. a, Pancreatic tissues were collected from mice infused after 2 weeks. Representative images of insulin, visualized by staining of β-cell-specific mPGES-2-knockout islets with an anti-insulin antibody (red), GFP by direct fluorescence (green) and DAPI (blue); scale bars = 20 μm. The experiment was repeated more than twice, independently, with similar results. b-g, Control mice (β-WT) and β-cell-specific mPGES-2-knockout mice (β-KO) were fed an HFD for 10 weeks after viral infusion. b, Western blotting and quantification of mPGES-2 expression in islets from control and β-cell-specific mPGES-2-knockout mice (n = 3 per group). c, Results of an IPGTT (n = 5 per group). d, Measurement of plasma insulin levels (n = 5 per group). e, IF analysis of MafA, Pdx1, p16, IGF1R (red) and GFP (green); scale bars = 20 μm. f, Statistical quantification of IF intensities for MafA, Pdx1, p16, and IGF1R (n = 4 per group). g, Representative images of β-Gal staining and quantification (n = 4 per group); scale bars = 20 μm. Data are expressed as mean ± SEM. Statistical significance was assessed using a two-tailed unpaired Student’s t-tests. Exact P values are indicated.
Extended Data Fig. 5 Assessment of mPGES-2 function in β-cells.
a, PGE2 levels were determined in tissues of liver, kidney, or brown fat in mPGES-2 WT and mPGES-2 KO mice (n = 5 per group). b, Islets were isolated from mPGES-2 WT and KO mice to assess the expression levels of different PGE2 synthases. Levels of Ptges, Ptges2 and Ptges3 mRNA expression (n = 4 WT, n = 5 KO). c, Immunoblotting and quantification of mPGES-2 in INS-1 cells (n = 4). d, NR4A1 expression induced by 10 μM 8-Br-cAMP at different times (n = 5), the samples derive from the same experiment and that gels/blots were processed in parallel. e, The expression of p21 in NR4A1-overexpressing INS-1 cells (n = 6). f, The expression of p21 and p16 in INS-1 cells with mPGES-2 overexpression or NR4A1 overexpression or dual overexpression (n = 4). g, The effect of mPGES-2 knockdown on NR4A1 signaling pathway, senescence markers and quantification of western blotting in INS-1 cells (n = 4). NC, negative control; KD, knockdown. h, The influence of mPGES-2 knockdown on insulin secretion under low glucose or high glucose (n = 3). Data are expressed as mean ± SEM. Statistical significance was assessed using one-way ANOVA with Tukey’s test (d and f) and a two-tailed unpaired Student’s t-test (a, b, c, e, g and h). Exact P values are indicated.
Extended Data Fig. 6 Effects of SZ0232 on PGE2 synthase inhibitory rates and insulin secretion in vitro.
a, Cell viability of INS-1 cells at the indicated concentrations of SZ0232 (n = 6). b, Inhibition of COX-2 enzyme activity by SZ0232 (n = 3). c, Inhibition of mPGES-1 enzyme activity by SZ0232 (n = 3). d, Effect of SZ0232 on insulin secretion in islets isolated from WT mice at the indicated weeks (n = 3). e, Effects of SZ0232 on insulin secretion in islets isolated from mPGES-2 WT and KO mice under low glucose conditions (n = 3). f, Effects of SZ0232 on insulin secretion in islets isolated from mPGES-2 WT and KO mice under high glucose conditions (n = 3). g, Effects of SZ0232 on insulin secretion in islets isolated from β-WT and β-KO mice under high glucose conditions (n = 4 for control group, n = 3 for SZ0232 group). h, Representative images and quantification of β-Gal staining of INS-1 cells with or without mPGES-2 overexpression or OE cells treated with SZ0232 (n = 4); scale bars = 20 μm. β-Gal staining was repeated more than twice, independently, with similar results. Data are expressed as mean ± SEM. Statistical significance was assessed using a two-tailed unpaired Student’s t-test (d, e, f and g) and one-way ANOVA with Tukey’s test (h). Exact P values are indicated.
Extended Data Fig. 7 The effect of SZ0232 on db/db mice.
a-c, db/db mice were administered with saline or SZ0232 daily for 8 weeks and then sacrificed. a, Body weight (n = 5). b, Serum biochemistry parameters (n = 5). c, Representative pictures of H&E staining in different tissues (n = 4); scale bars = 20 μm. H&E staining was repeated more than twice, independently, and similar results were obtained. Data are expressed as mean ± SEM. Statistical significance was assessed using a two-tailed unpaired Student’s t-test.
Supplementary information
Supplementary Information
Supplementary Figs. 1–6 and Supplementary Tables 1 and 2.
Supplementary Data 1
All data presented in the graphs for the supplementary figures.
Supplementary Data 2
Unprocessed gel and western blot images for the supplementary figures.
Source data
Source Data Fig. 1
Statistical source data.
Source Data Fig. 2
Statistical source data.
Source Data Fig. 3
Statistical source data.
Source Data Fig. 4
Statistical source data.
Source Data Fig. 4
Unprocessed western blots.
Source Data Fig. 5
Statistical source data.
Source Data Fig. 5
Unprocessed western blots.
Source Data Fig. 6
Statistical source data.
Source Data Fig. 7
Statistical source data.
Source Data Extended Data Fig. 1
Statistical source data.
Source Data Extended Data Fig. 1
Unprocessed gel and western blots.
Source Data Extended Data Fig. 2
Statistical source data.
Source Data Extended Data Fig. 2
Unprocessed gel.
Source Data Extended Data Fig. 3
Statistical source data.
Source Data Extended Data Fig. 4
Statistical source data.
Source Data Extended Data Fig. 4
Unprocessed western blots.
Source Data Extended Data Fig. 5
Statistical source data.
Source Data Extended Data Fig. 5
Unprocessed western blots.
Source Data Extended Data Fig. 6
Statistical source data.
Source Data Extended Data Fig. 7
Statistical source data.
Rights and permissions
About this article
Cite this article
Zhong, D., Wan, Z., Cai, J. et al. mPGES-2 blockade antagonizes β-cell senescence to ameliorate diabetes by acting on NR4A1. Nat Metab 4, 269–283 (2022). https://doi.org/10.1038/s42255-022-00536-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s42255-022-00536-6
This article is cited by
-
Cellular rejuvenation: molecular mechanisms and potential therapeutic interventions for diseases
Signal Transduction and Targeted Therapy (2023)
-
Targeting mPGES-2 to protect against acute kidney injury via inhibition of ferroptosis dependent on p53
Cell Death & Disease (2023)
-
Caspase 6/NR4A1/SOX9 signaling axis regulates hepatic inflammation and pyroptosis in ischemia-stressed fatty liver
Cell Death Discovery (2023)
-
New drug target identified for β-cell dysfunction
Nature Reviews Endocrinology (2022)