Abstract
Obesity is a major public health burden worldwide and is characterized by chronic low-grade inflammation driven by the cooperation of the innate immune system and dysregulated metabolism in adipose tissue and other metabolic organs. Receptor-interacting serine/threonine-protein kinase 1 (RIPK1) is a central regulator of inflammatory cell function that coordinates inflammation, apoptosis and necroptosis in response to inflammatory stimuli. Here we show that genetic polymorphisms near the human RIPK1 locus associate with increased RIPK1 gene expression and obesity. We show that one of these single nucleotide polymorphisms is within a binding site for E4BP4 and increases RIPK1 promoter activity and RIPK1 gene expression in adipose tissue. Therapeutic silencing of RIPK1 in vivo in a mouse model of diet-induced obesity dramatically reduces fat mass, total body weight and improves insulin sensitivity, while simultaneously reducing macrophage and promoting invariant natural killer T cell accumulation in adipose tissue. These findings demonstrate that RIPK1 is genetically associated with obesity, and reducing RIPK1 expression is a potential therapeutic approach to target obesity and related diseases.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Data availability
Human and mouse scRNA-seq data have been deposited in the GEO database (https://www.ncbi.nlm.nih.gov/geo/) under accession number GSE151889. Other publicly available data used in this study are available at the GTEx database, or under GEO accession numbers GSE36032 and GSE63358 (mouse) and GSE115469 (human liver). All other data that support the findings of this study are available from the corresponding author upon request. Source data are provided with this paper.
Change history
08 October 2020
An amendment to this paper has been published and can be accessed via a link at the top of the paper.
References
Berg, A. H. & Scherer, P. E. Adipose tissue, inflammation and cardiovascular disease. Circ. Res. 96, 939–949 (2005).
Weisberg, S. P. et al. Obesity is associated with macrophage accumulation in adipose tissue. J. Clin. Invest. 112, 1796–1808 (2003).
Stunkard, A. J., Foch, T. T. & Hrubec, Z. A twin study of human obesity. JAMA 256, 51–54 (1986).
Frayling, T. M. et al. A common variant in the FTO gene is associated with body mass index and predisposes to childhood and adult obesity. Science 316, 889–894 (2007).
Gerken, T. et al. The obesity-associated FTO gene encodes a 2-oxoglutarate-dependent nucleic acid demethylase. Science 318, 1469–1472 (2007).
Scuteri, A. et al. Genome-wide association scan shows genetic variants in the FTO gene are associated with obesity-related traits. PLoS Genet. 3, e115 (2007).
Dina, C. et al. Variation in FTO contributes to childhood obesity and severe adult obesity. Nat. Genet. 39, 724–726 (2007).
Speakman, J. R., Rance, K. A. & Johnstone, A. M. Polymorphisms of the FTO gene are associated with variation in energy intake, but not energy expenditure. Obesity (Silver Spring) 16, 1961–1965 (2008).
Smemo, S. et al. Obesity-associated variants within FTO form long-range functional connections with IRX3. Nature 507, 371–375 (2014).
Loos, R. J. et al. Common variants near MC4R are associated with fat mass, weight and risk of obesity. Nat. Genet. 40, 768–775 (2008).
Thorleifsson, G. et al. Genome-wide association yields new sequence variants at seven loci that associate with measures of obesity. Nat. Genet. 41, 18–24 (2009).
Nikpay, M., Turner, A. W. & McPherson, R. Partitioning the pleiotropy between coronary artery disease and body mass index reveals the importance of low frequency variants and central nervous system-specific functional elements. Circ. Genom. Precis. Med. 11, e002050 (2018).
Karunakaran, D. et al. Targeting macrophage necroptosis for therapeutic and diagnostic interventions in atherosclerosis. Sci. Adv. 2, e1600224 (2016).
Silke, J., Rickard, J. A. & Gerlic, M. The diverse role of RIP kinases in necroptosis and inflammation. Nat. Immunol. 16, 689–697 (2015).
Kondylis, V., Kumari, S., Vlantis, K. & Pasparakis, M. The interplay of IKK, NFκB and RIPK1 signaling in the regulation of cell death, tissue homeostasis and inflammation. Immunol. Rev. 277, 113–127 (2017).
Civelek, M. et al. Genetic regulation of adipose gene expression and cardio-metabolic traits. Am. J. Hum. Genet. 100, 428–443 (2017).
Laakso, M. et al. The Metabolic Syndrome in Men study: a resource for studies of metabolic and cardiovascular diseases. J. Lipid. Res. 58, 481–493 (2017).
The GTEx Consortium. The Genotype-Tissue Expression (GTEx) pilot analysis: multitissue gene regulation in humans. Science 348, 648–660 (2015).
Davies, R. W. et al. A 680-kb duplication at the FTO locus in a kindred with obesity and a distinct body fat distribution. Eur. J. Hum. Genet. 21, 1417–1422 (2013).
van Rheenen, W., Peyrot, W. J., Schork, A. J., Lee, S. H. & Wray, N. R. Genetic correlations of polygenic disease traits: from theory to practice. Nat. Rev. Genet. 20, 567–581 (2019).
Pingault, J. B. et al. Using genetic data to strengthen causal inference in observational research. Nat. Rev. Genet. 19, 566–580 (2018).
Zhu, Z. et al. Causal associations between risk factors and common diseases inferred from GWAS summary data. Nat. Commun. 9, 224 (2018).
Westra, H. J. et al. Systematic identification of trans eQTLs as putative drivers of known disease associations. Nat. Genet. 45, 1238–1243 (2013).
van Nas, A. et al. The systems genetics resource: a web application to mine global data for complex disease traits. Front. Genet. 4, 84 (2013).
Ward, L. D. & Kellis, M. HaploReg: a resource for exploring chromatin states, conservation, and regulatory motif alterations within sets of genetically linked variants. Nucleic Acids Res. 40, D930–D934 (2011).
Cowell, I. G. E4BP4/NFIL3, a PAR-related bZIP factor with many roles. Bioessays 24, 1023–1029 (2002).
Lynch, L. et al. Regulatory iNKT cells lack expression of the transcription factor PLZF and control the homeostasis of Treg cells and macrophages in adipose tissue. Nat. Immunol. 16, 85–95 (2015).
Roy, S. et al. Redefining the transcriptional regulatory dynamics of classically and alternatively activated macrophages by deepCAGE transcriptomics. Nucleic Acids Res. 43, 6969–6982 (2015).
Takahashi, S. et al. A promoter in the novel exon of hPPARγ directs the circadian expression of PPARγ. J. Atheroscler. Thromb. 17, 73–83 (2010).
Lynch, L. et al. Adipose tissue iNKT cells protect against diet-induced obesity and metabolic disorder through regulatory cytokine production. Immunity 37, 574–587 (2012).
Ji, Y. et al. Activation of natural killer T cells promotes M2 macrophage polarization in adipose tissue and improves systemic glucose tolerance via IL-4/STAT6 protein signaling axis in obesity. J. Biol. Chem. 287, 13561–13571 (2012).
Jaitin, D. A. et al. Lipid-associated macrophages control metabolic homeostasis in a Trem2-dependent manner. Cell 178, 686–698(2019).
MacParland, S. A. et al. Single cell RNA sequencing of human liver reveals distinct intrahepatic macrophage populations. Nat. Commun. 9, 4383 (2018).
Kenna, T. et al. NKT cells from normal and tumor-bearing human livers are phenotypically and functionally distinct from murine NKT cells. J. Immunol. 171, 1775–1779 (2003).
Fengler, V. H. et al. Susceptibility of different mouse wild-type strains to develop diet-induced NAFLD/AFLD-associated liver disease. PLoS ONE 11, e0155163 (2016).
Berger, S. B. et al. Cutting edge: RIP1 kinase activity is dispensable for normal development but is a key regulator of inflammation in SHARPIN-deficient mice. J. Immunol. 192, 5476–5480 (2014).
Woo, Y. D. C., Kim, H. Y. & Chung, D. H. Invariant NKT cells recruit CD103+ dendritic cells via XCL1–XCR1 axis, thereby promoting allergic airway resistance. J. Immunol. 198, 1 (2017).
van der Klaauw, A. A. & Farooqi, I. S. The hunger genes: pathways to obesity. Cell 161, 119–132 (2015).
Akiyama, M. et al. Genome-wide association study identifies 112 new loci for body mass index in the Japanese population. Nat. Genet. 49, 1458–1467 (2017).
Welsh, P. et al. Unraveling the directional link between adiposity and inflammation: a bidirectional Mendelian randomization approach. J. Clin. Endocrinol. Metab. 95, 93–99 (2010).
Vucic, D., Dixit, V. M. & Wertz, I. E. Ubiquitylation in apoptosis: a post-translational modification at the edge of life and death. Nat. Rev. Mol. Cell. Biol. 12, 439–452 (2011).
Chan, F. K., Luz, N. F. & Moriwaki, K. Programmed necrosis in the cross talk of cell death and inflammation. Annu. Rev. Immunol. 33, 79–106 (2015).
Peltzer, N., Darding, M. & Walczak, H. Holding RIPK1 on the ubiquitin leash in TNFR1 signaling. Trends Cell Biol. 26, 445–461 (2016).
Roderick, J. E. et al. Hematopoietic RIPK1 deficiency results in bone marrow failure caused by apoptosis and RIPK3-mediated necroptosis. Proc. Natl Acad. Sci. USA 111, 14436–14441 (2014).
Gautheron, J. et al. The necroptosis-inducing kinase RIPK3 dampens adipose tissue inflammation and glucose intolerance. Nat. Commun. 7, 11869 (2016).
Dannappel, M. et al. RIPK1 maintains epithelial homeostasis by inhibiting apoptosis and necroptosis. Nature 513, 90–94 (2014).
Berzins, S. P., Smyth, M. J. & Baxter, A. G. Presumed guilty: natural killer T cell defects and human disease. Nat. Rev. Immunol. 11, 131–142 (2011).
Lynch, L. Adipose invariant natural killer T cells. Immunology 142, 337–346 (2014).
Huh, J. Y. et al. Deletion of CD1d in adipocytes aggravates adipose tissue inflammation and insulin resistance in obesity. Diabetes 66, 835–847 (2017).
Vieth, J. A. et al. TCRα–TCRβ pairing controls recognition of CD1d and directs the development of adipose NKT cells. Nat. Immunol. 18, 36–44 (2017).
Mookerjee-Basu, J. & Kappes, D. J. iNKT cells do a fat lot of good. Nat. Immunol. 18, 10–12 (2016).
Lynch, L. et al. Invariant NKT cells and CD1d+ cells amass in human omentum and are depleted in patients with cancer and obesity. Eur. J. Immunol. 39, 1893–1901 (2009).
Engel, I. et al. Innate-like functions of natural killer T cell subsets result from highly divergent gene programs. Nat. Immunol. 17, 728–739 (2016).
Plenge, R. M. Priority index for human genetics and drug discovery. Nat. Genet. 51, 1073–1075 (2019).
Fang, H. et al. A genetics-led approach defines the drug target landscape of 30 immune-related traits. Nat. Genet. 51, 1082–1091 (2019).
Nelson, M. R. et al. The support of human genetic evidence for approved drug indications. Nat. Genet. 47, 856–860 (2015).
Plenge, R. M., Scolnick, E. M. & Altshuler, D. Validating therapeutic targets through human genetics. Nat. Rev. Drug Discov. 12, 581–594 (2013).
Paik, J. & Duggan, S. Volanesorsen: first global approval. Drugs 79, 1349–1354 (2019).
Witztum, J. L. et al. Volanesorsen and triglyceride levels in familial chylomicronemia syndrome. N. Engl. J. Med. 381, 531–542 (2019).
Fitzgerald, K. et al. A highly durable RNAi therapeutic inhibitor of PCSK9. N. Engl. J. Med. 376, 41–51 (2017).
Duell, P. B. et al. Long-term mipomersen treatment is associated with a reduction in cardiovascular events in patients with familial hypercholesterolemia. J. Clin. Lipidol. 10, 1011–1021 (2016).
Adams, D. et al. Patisiran, an RNAi therapeutic, for hereditary transthyretin amyloidosis. N. Engl. J. Med. 379, 11–21 (2018).
Stein, C. A. & Castanotto, D. FDA-approved oligonucleotide therapies in 2017. Mol. Ther. 25, 1069–1075 (2017).
Levin, A. A. Treating disease at the RNA level with oligonucleotides. N. Engl. J. Med. 380, 57–70 (2019).
Nikpay, M. et al. A comprehensive 1000 Genomes-based genome-wide association meta-analysis of coronary artery disease. Nat. Genet. 47, 1121–1130 (2015).
Ramkhelawon, B. et al. Netrin-1 promotes adipose tissue macrophage retention and insulin resistance in obesity. Nat. Med. 20, 377–384 (2014).
Karunakaran, D. et al. Therapeutic inhibition of miR-33 promotes fatty acid oxidation but does not ameliorate metabolic dysfunction in diet-induced obesity. Arterioscler. Thromb. Vasc. Biol. 35, 2536–2543 (2015).
Butler, A., Hoffman, P., Smibert, P., Papalexi, E. & Satija, R. Integrating single-cell transcriptomic data across different conditions, technologies and species. Nat. Biotechnol. 36, 411–420 (2018).
Hafemeister, C. & Satija, R. Normalization and variance stabilization of single-cell RNA-seq data using regularized negative binomial regression. Genome Biol. 20, 296 (2019).
Becht, E. et al. Dimensionality reduction for visualizing single-cell data using UMAP. Nat. Biotechnol. 37, 38–44 (2018).
McInnes, L., Healy, J., Saul, N. & Großberger, L. UMAP: uniform manifold approximation and projection. J. Open Source Softw. 3, 861 (2018).
Frankish, A. et al. GENCODE reference annotation for the human and mouse genomes. Nucleic Acids Res. 47, D766–D773 (2019).
Stuart, T. et al. Comprehensive integration of single cell data. Cell 177, 1888–1902.e21 (2019).
Carvalho, B. S. & Irizarry, R. A. A framework for oligonucleotide microarray preprocessing. Bioinformatics 26, 2363–2367 (2010).
Ritchie, M. E. et al. Limma powers differential expression analyses for RNA-sequencing and microarray studies. Nucleic Acids Res. 43, e47 (2015).
MacDonald, J. W. mogene11sttranscriptcluster.db: Affymetrix mogene11 annotation data (chip mogene11sttranscriptcluster). R package version 8.7.0. (2017).
Riveros-McKay, F. et al. Genetic architecture of human thinness compared to severe obesity. PLoS Genet. 15, e1007603 (2019).
Lloyd-Jones, L. R. et al. The genetic architecture of gene expression in peripheral blood. Am. J. Hum. Genet. 100, 228–237 (2017).
Acknowledgements
The authors thank X. Zhao, K. Sheikheleslamy and G. Palidwor for their technical and statistical support. We thank J. Kaprio and A. Rissanen for their contributions and the individuals who participated in the Finnish Twin study. We also thank the University of Ottawa StemCore Facility and the UNGC Sequencing Core at UCLA for performing RNA sequencing. This study was supported with funding from the following organizations: Canadian Institutes of Health Research (K.J.R., R.M., M.E.H., B.C.V. and D.P.C.), NIH (R01 HL119047 to K.J.R.; DK117850 and HL147883 to A.J.L., HL095056, HL28481 and U01 DK105561 to P.P.), Foundation Leduc (A.J.L.), National Psoriasis Foundation (D.K.), Academy of Finland (M.L.; 272376, 266286, 314383 and 315035 to K.H.P.), European Union (M.L.) and Sigrid Juselius Foundation (M.L. and K.H.P.); Finnish Medical Foundation (K.H.P.) and the Finnish Diabetes Research Foundation, Novo Nordisk Foundation, Gyllenberg Foundation, Helsinki University Hospital Research Funds and University of Helsinki (K.H.P.); University of Ottawa Heart Institute Cardiac Endowment Fellowship (A.C.D.); and HHMI Gilliam Fellowship (M.A.). The funders had no role in study design, data collection and analysis, decision to publish or preparation of the article.
Author information
Authors and Affiliations
Contributions
K.J.R. and D.K. conceived the project and wrote the manuscript with input from all authors. A.W.T., S.S. and P. Lau performed and analysed the chromatin immunoprecipitation experiments in human adipose tissue, and A.W.T. performed the promoter luciferase experiments. A.C.D. and D.S. conducted the flow cytometry analysis of human adipose tissue and liver. D.K. and A.R. conducted and analysed the mouse obesity experiments. J.W.K., M.G., M.A.N. and H.W. performed biochemical analyses on mouse tissue. M.E.H. provided assistance with the indirect calorimetry experiments in mice. K.J.R., A.R. and D.P.C. conducted and analysed the single-cell sequencing of human and mouse adipose tissue. M.N. performed the MR and analyses of the OBLE cohort. C.P., M.L. and A.J.L. conducted and analysed data associated with the METSIM cohort and the HMDP. L.B. and B.R. collected and analysed qPCR analyses of human adipose tissue for RIPK1 expression. M.A., K.H.P. and P.P. performed the experiments and analyses of the human adipose single-nuclei data. R.L. provided the ASOs used in the mouse studies. S.B.B., P.J.G. and J.B. developed and provided the Ripk1 mutant mice. B.C.V., P. Liu, A.J.L. and R.M. provided supervision and guidance throughout the study.
Corresponding authors
Ethics declarations
Competing interests
R.L. is an employee of Ionis Pharmaceuticals, S.B.B. is an employee of Janssen Pharmaceuticals, P.J.G. is an employee of Inzen Therapeutics and J.B. is an employee of GlaxoSmithKline. The other authors declare no competing interests.
Additional information
Peer review information Primary Handling Editor: Pooja Jha.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 SNPs in the RIPK1 locus associate with obesity in humans.
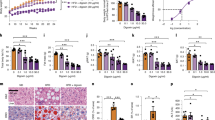
a, eQTL analysis of SNPs rs2272990 and rs2064310 in the RIPK1 locus and levels of RIPK1 mRNA in adipose tissue from the METSIM cohort. b, Regional association plots of eQTLs for RIPK1 in subcutaneous adipose from GTEX (probe ENSG00000137275.9). SNPs are coloured based on their linkage disequilibrium (r2) with the labeled top eQTL (rs2064310), which has the smallest P value in the region. Linkage disequilibrium and recombination rate calculations are based on the European population from 1000 Genomes reference panel (Phase I; release 3). c, d, Individual SNPs from the GTEx database and their association with RIPK1 expression in visceral (c) and subcutaneous (d) adipose tissue. Number of subjects homozygous or heterozygous for the minor (Ref) or major (Alt) alleles are indicated. p-values as calculated by the GTEx Consortium detailed at: https://www.gtexportal.org. Box plot shows ranked normalized gene expression in median, 1st and 3rd quartiles, 1.5 interquartile range (IQR) of 1st and 3rd quartiles. e, Odds ratio for the index SNP rs6907943 and LD SNPs in the Ottawa OBLE cohort and comparison to rs9939609 and LD SNPs in the FTO locus. f–i, Mendelian Randomization analysis of SNPs that are independently (linkage disequilibrium r2<0.2) associated with expression of RIPK1 and significantly (P<0.05) associated with obesity metrics. The beta and standard error of the effect sizes of RIPK1 expression (eQTL, x-axis) and the effect sizes on obesity metrics (y-axis) are plotted. f- UKHLS& GTEX (Adipose tissue) g- UKHLS & METSIM (Adipose tissue) h- UK Biobank & GTEX (Adipose tissue), i- UK Biobank & Llyod-Jones, 2017 (Whole Blood).
Extended Data Fig. 2 Single nuclei sequencing of human adipose tissue and HMDP data from mice.
a, Single-nuclei RNA sequencing on human adipose tissue: UMAP visual representation of 10 cell types derived from single-nucleus RNA-seq of human abdominal subcutaneous adipose tissue. Clusters are identified using the Seurat pipeline and cell types are assigned by biological function of marker genes. Vertical bar plot of average cell type-specific expression of RIPK1 across the 10 cell types identified in human abdominal subcutaneous adipose tissue. Expression was normalized using SCTransform. b, Data from the Hybrid Mouse Diversity Panel (HMDP) showing correlations of Ripk1 mRNA expression and NMR, visceral fat mass at sacrifice, serum insulin and HOMA-IR index in males from approximately 100 inbred strains of mice19. c, Luciferase reporter activity containing the variant sequence plus the human RIPK1 promoter upstream of the firefly luciferase gene (as in Fig. 2b). Data is shown as mean ± SEM of n=4 independent experiments, d–f. Single cell RNA sequencing of mouse CD45+ immune cells (from GSE128518) after 6, 12 and 18 weeks high fat diet feeding. d, Horizontal bar plot of cell clusters in normal chow (NC) and high fat diet (HFD). e, UMAP plot of Xcl1 cluster (cluster 9, representing NK/NKT cells) and (f) the corresponding expression of Nfil3 and Ripk1 in cluster 9.
Extended Data Fig. 3 Analysis of flow cytometry and single cell RNA sequencing of human adipose.
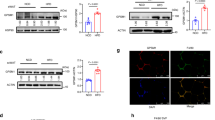
a, Mean fluorescence intensity (MFI) of RIPK1 and E4BP4 in iNKT cells from human omental adipose tissue (left) and liver (right) from obese patients (n= 12 patients total). iNKT were identified as CD19-CD3+αGalCer-loaded CD1d-tetramer+ population and based on negative control (isotype IgG) staining. Outliers were removed according to ROUT method with a specified FDR of 1% (adipose, two E4BP4 samples; liver one E4BP4 and one RIPK1 sample). Data is mean ± SD, ***p≤0.001 by paired two-tailed Student’s t-test. b, UMAP embeddings of scRNA-seq data from human omentum samples. The colour maps represent the log2-based gene expression values of various T/NK cell markers (CD3D, XCL1, NKG7, CD8A) and myeloid markers (LYZ, CD14). The bottom panels show RIPK1 and NFIL3 expression across these populations. Data deposited in the GEO database (accession number GSE151889).
Extended Data Fig. 5 Metabolic analyses of RIPK1 ASO and cont ASO treated mice.
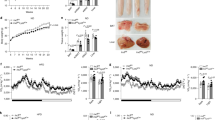
a, C57BL6 mice were treated with 50mg/kg control ASO, RIPK1 ASO-A or ASO-B over 20 weeks while fed a normal chow diet, and body weight was measured weekly. Data depicts mean ± SEM of n=3 mice per group. Two-way ANOVA using Holm-Sidak’s multiple comparison test. *p<0.05 control ASO vs. RIPK1 ASO-A and ASO-B. b, Western blot of RIPK1 (top) and GAPDH (bottom) from adipose tissue of ASO treated mice after 5 weeks of treatment. Quantification of RIPK1 band density normalized to GAPDH band density was performed using Image J. Data representative of mean ± SD of n=7-8 mice/group, p=0.09 RIPK1 ASO-A vs cont ASO, p=0.02 RIPK1 ASO-B vs. cont ASO by two-tailed Student’s t-test. One outlier was removed according to ROUT method with a specified FDR of 1%. c–f, Mice were caged in metabolic cages with indirect calorimetric and activity monitoring. VO2 (c) and respiratory exchange rations (RER) (d) and beam breaks were recorded as a measure of physical activity (e). Total food consumption (f) was measured over a 48h period. Data representative of mean ± SEM of n=8 mice/group, *p<0.05, **p<0.01, ***p<0.001, ****p<0.0001, one-way ANOVA using Tukey’s multiple comparison test. g, Plasma levels of alanine transaminase (ALT) and aspartate transaminase (AST) were determined after 20 weeks of ASO treatment. Data representative of mean ± SEM of n=8 mice/group analyzed by one-way ANOVA using Tukey’s multiple comparison test. *p<0.05 vs. control ASO and RIPK1 ASO-B.
Extended Data Fig. 6 Analysis of diet-induced obesity in RIPK1 kinase dead mice.
WT or RIP1K45A mice (male, n=8/group) were fed a high fat diet for 20 weeks. a, Body weight. b, Epididymal adipose tissue, liver and spleen weights at termination of study. c, d, Fasting glucose tolerance test (GTT, c) or insulin tolerance test (ITT, d) and area under the curve. Data representative of mean ± SEM and analyzed by Two-way ANOVA (body weight, GTT and ITT) or two-tailed Student’s t-test (tissues weights).
Extended Data Fig. 7 Serum cytokine analysis of cont ASO and RIPK1 ASO treated mice.
Serum cytokines from mice treated with cont ASO or RIPK1 ASO-B for 24 weeks. Cytokine analysis was done using the Bioplex Pro Cytokine 23-plex assay. No significant differences between cytokines IL-1α, TNFα, IL-4 or IL-10 were observed. Data representative of mean ± SEM of n=8 mice/group and was analysed by one-way ANOVA using Dunnett’s multiple comparison test.
Extended Data Fig. 8 Analysis of single cell RNA sequencing of mouse adipose tissue.
a, Clustered UMAP embedding of scRNA-seq data from mouse adipose tissue. The colours and labels represent each of the 13 distinct clusters. b, The same embeddings as in (a), but separately coloring the cells derived from either Control ASO or RIPK1 ASO-B samples. c, A graph showing the percentage of cells from each sample that correspond to each cell type. d, UMAP embedding of scRNA-seq data from T cell populations of mouse adipose tissue. T cells were defined as CD3+ clusters. Plots show the log-transformed gene expression of various markers defining NKT (Xcl1), CD4+ T (Cd4), CD8+ T (Cd8a), Memory CD4+ T (Ccr7), Naive CD4+ T (S100a4), Regulatory T (Foxp3), and Gamma Delta T (Cd163l1). Data deposited in the GEO database (accession number GSE151889).
Extended Data Fig. 9 Flow cytometry of adipose and circulating immune cells from cont ASO and RIPK1 ASO treated mice.
a, Representative flow cytometry plots and NKT % (expressed as % of F4/80- cells that are αGalCer-loaded CD1d-tetramer+CD3e+) in spleen from mice treated for 5 weeks with cont ASO or RIPK1 ASO (n=8/group). Data representative of mean ± SEM of n=8 mice/group, analyzed by two-tailed Student’s t-test. b, c, Flow cytometry analysis of circulating leukocytes from mice treated with control ASO or RIP1 ASO-B at 5 weeks: Ly6Clow monocytes *p<0.05 (p=0.0452) Ly6Chigh monocytes *p<0.05 (p=0.0108), and CD3e+ T cells. (c) Flow cytometry analysis of circulating leukocytes from mice treated with control ASO or RIP1 ASO-B at 22 weeks: Ly6Clow monocytes, Ly6Chigh monocytes and CD3e+ T cells (p=0.0132). Data in b, c representative of mean ± SEM of n=8 mice/group, analyzed by two-tailed Student’s t-test.
Extended Data Fig. 10 Flow cytometry of adipose immune cells from cont ASO and RIPK1 ASO treated mice.
a, Regulatory T-cell (CD25+Foxp3+) and NK (NK1.1+) cells analysis by flow cytometry of isolated SVF of adipose tissue from mice treated with control ASO or RIP1 ASO-B for 5 weeks while fed a high fat diet (n=7-8/group). b, Gating strategy for selecting CD45+, CD3e+, CD4+ cells then those that were positive for CD25 and Foxp3 (to represent T regulatory cells) or NK1.1 (to represent NK cells). Data depicts mean ± SEM of n=8 mice/group.
Supplementary information
Supplementary Information
Supplementary Figs. 1–5 and Table 1
Source data
Source Data Fig. 1
Unprocessed western blots
Rights and permissions
About this article
Cite this article
Karunakaran, D., Turner, A.W., Duchez, AC. et al. RIPK1 gene variants associate with obesity in humans and can be therapeutically silenced to reduce obesity in mice. Nat Metab 2, 1113–1125 (2020). https://doi.org/10.1038/s42255-020-00279-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s42255-020-00279-2
This article is cited by
-
Mediators of necroptosis: from cell death to metabolic regulation
EMBO Molecular Medicine (2024)
-
Weight cycling based on altered immune microenvironment as a result of metaflammation
Nutrition & Metabolism (2023)
-
Polarity protein AF6 functions as a modulator of necroptosis by regulating ubiquitination of RIPK1 in liver diseases
Cell Death & Disease (2023)
-
Cell death and inflammation during obesity: “Know my methods, WAT(son)”
Cell Death & Differentiation (2023)
-
A common human MLKL polymorphism confers resistance to negative regulation by phosphorylation
Nature Communications (2023)