Abstract
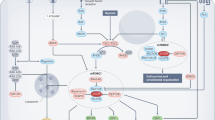
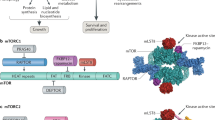
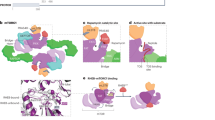
The protein kinase complex mechanistic target of rapamycin complex 1 (mTORC1) serves as a key conduit between growth signals and the metabolic processes underlying cell growth. The activation state of mTORC1 is controlled by intracellular nutrients and energy, as well as exogenous hormones and growth factors, thereby integrating local and systemic growth signals. Here we discuss the molecular logic of the mTORC1 signalling network and its importance in coupling growth signals to the control of cellular metabolism. After activation, mTORC1 promotes the conversion of available nutrients and energy into the major macromolecular species contributing to cellular mass, including proteins, nucleic acids and lipids, while suppressing the autophagic recycling of these macromolecules back into their nutrient constituents. Given that uncoupling of mTORC1 from its normal regulatory inputs contributes to many diseases—including cancer, genetic tumour syndromes, metabolic diseases, autoimmune diseases and neurological disorders—understanding the molecular logic of the mTORC1 network and how to modulate it may present therapeutic opportunities for treatment of a broad range of diseases and potentially even for the extension of lifespan.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Singh, K., Sun, S. & Vézina, C. Rapamycin (AY-22,989), a new antifungal antibiotic. IV. Mechanism of action. J. Antibiot. (Tokyo) 32, 630–645 (1979).
Düvel, K. et al. Activation of a metabolic gene regulatory network downstream of mTOR complex 1. Mol. Cell 39, 171–183 (2010). This study defines a set of transcripts encoding metabolic enzymes that are sensitive to mTORC1 activation and inhibition. Transcriptional profiling and metabolomics reveal that mTORC1 induces glycolysis, the oxidative pentose-phosphate pathway and de novo lipid synthesis through activation of the HIF1 and SREBP transcription factors.
Siddiqui, N. & Sonenberg, N. Signalling to eIF4E in cancer. Biochem. Soc. Trans. 43, 763–772 (2015).
Ma, X. M. & Blenis, J. Molecular mechanisms of mTOR-mediated translational control. Nat. Rev. Mol. Cell Biol. 10, 307–318 (2009).
Raught, B. et al. Phosphorylation of eucaryotic translation initiation factor 4B Ser422 is modulated by S6 kinases. EMBO J. 23, 1761–1769 (2004).
Holz, M. K., Ballif, B. A., Gygi, S. P. & Blenis, J. mTOR and S6K1 mediate assembly of the translation preinitiation complex through dynamic protein interchange and ordered phosphorylation events. Cell 123, 569–580 (2005).
Csibi, A. et al. The mTORC1/S6K1 pathway regulates glutamine metabolism through the eIF4B-dependent control of c-Myc translation. Curr. Biol. 24, 2274–2280 (2014).
JefferiesH. B.., ReinhardC.., KozmaS. C.. & ThomasG.. Rapamycin selectively represses translation of the "polypyrimidine tract" mRNA family. Proc. Natl Acad. Sci. USA 91, 4441–4445 (1994).
Hsieh, A. C. et al. The translational landscape of mTOR signalling steers cancer initiation and metastasis. Nature 485, 55–61 (2012).
Thoreen, C. C. et al. A unifying model for mTORC1-mediated regulation of mRNA translation. Nature 485, 109–113 (2012).
Hong, S. et al. LARP1 functions as a molecular switch for mTORC1-mediated translation of an essential class of mRNAs. eLife 6, e25237 (2017).
Lahr, R. M. et al. The La-related protein 1-specific domain repurposes HEAT-like repeats to directly bind a 5’TOP sequence. Nucleic Acids Res. 43, 8077–8088 (2015).
Fonseca, B. D. et al. La-related Protein 1 (LARP1) represses terminal oligopyrimidine (TOP) mRNA translation downstream of mTOR complex 1 (mTORC1). J. Biol. Chem. 290, 15996–16020 (2015).
Heintz, C. et al. Splicing factor 1 modulates dietary restriction and TORC1 pathway longevity in C. elegans. Nature 541, 102–106 (2017).
Ma, X. M., Yoon, S. O., Richardson, C. J., Jülich, K. & Blenis, J. SKAR links pre-mRNA splicing to mTOR/S6K1-mediated enhanced translation efficiency of spliced mRNAs. Cell 133, 303–313 (2008).
Lee, G. et al. Post-transcriptional regulation of de novo lipogenesis by mTORC1-S6K1-SRPK2 signaling. Cell 171, 1545–1558.e18 (2017).
Pelletier, J., Thomas, G. & Volarević, S. Ribosome biogenesis in cancer: new players and therapeutic avenues. Nat. Rev. Cancer 18, 51–63 (2018).
Iadevaia, V., Liu, R. & Proud, C. G. mTORC1 signaling controls multiple steps in ribosome biogenesis. Semin. Cell Dev. Biol. 36, 113–120 (2014).
Hannan, K. M. et al. mTOR-dependent regulation of ribosomal gene transcription requires S6K1 and is mediated by phosphorylation of the carboxy-terminal activation domain of the nucleolar transcription factor UBF. Mol. Cell. Biol. 23, 8862–8877 (2003).
Mayer, C., Zhao, J., Yuan, X. & Grummt, I. mTOR-dependent activation of the transcription factor TIF-IA links rRNA synthesis to nutrient availability. Genes Dev. 18, 423–434 (2004).
Lee, D. et al. SHPRH regulates rRNA transcription by recognizing the histone code in an mTOR-dependent manner. Proc. Natl Acad. Sci. USA 114, E3424–E3433 (2017).
Liu, Y. et al. PWP1 mediates nutrient-dependent growth control through nucleolar regulation of ribosomal gene expression. Dev. Cell 43, 240–252.e5 (2017).
Michels, A. A. et al. mTORC1 directly phosphorylates and regulates human MAF1. Mol. Cell. Biol. 30, 3749–3757 (2010).
Shor, B. et al. Requirement of the mTOR kinase for the regulation of Maf1 phosphorylation and control of RNA polymerase III-dependent transcription in cancer cells. J. Biol. Chem. 285, 15380–15392 (2010).
Iadevaia, V., Zhang, Z., Jan, E. & Proud, C. G. mTOR signaling regulates the processing of pre-rRNA in human cells. Nucleic Acids Res. 40, 2527–2539 (2012).
Blobel, G. & Potter, V. R. Studies on free and membrane-bound ribosomes in rat liver. I. Distribution as related to total cellular RNA. J. Mol. Biol. 26, 279–292 (1967).
Ben-Sahra, I., Howell, J. J., Asara, J. M. & Manning, B. D. Stimulation of de novo pyrimidine synthesis by growth signaling through mTOR and S6K1. Science 339, 1323–1328 (2013).
Ben-Sahra, I., Hoxhaj, G., Ricoult, S. J. H., Asara, J. M. & Manning, B. D. mTORC1 induces purine synthesis through control of the mitochondrial tetrahydrofolate cycle. Science 351, 728–733 (2016).
Robitaille, A. M. et al. Quantitative phosphoproteomics reveal mTORC1 activates de novo pyrimidine synthesis. Science 339, 1320–1323 (2013). The studies in refs. 27–29 demonstrate that mTORC1 stimulates de novo nucleotide synthesis: ref. 27 and ref. 29 show that metabolic flux through the pyrimidine-nucleotide-synthesis pathway is acutely stimulated through S6K1-mediated phosphorylation of the first enzyme in the pathway, CAD; ref. 28 shows that mTORC1 activation promotes purine-nucleotide synthesis through transcriptional mechanisms including stress-independent activation of ATF4, which in turn induces expression of the mitochondrial tetrahydrofolate-cycle enzyme MTHFD2.
French, J. B. et al. Spatial colocalization and functional link of purinosomes with mitochondria. Science 351, 733–737 (2016).
Cunningham, J. T., Moreno, M. V., Lodi, A., Ronen, S. M. & Ruggero, D. Protein and nucleotide biosynthesis are coupled by a single rate-limiting enzyme, PRPS2, to drive cancer. Cell 157, 1088–1103 (2014).
Porstmann, T. et al. SREBP activity is regulated by mTORC1 and contributes to Akt-dependent cell growth. Cell Metab. 8, 224–236 (2008).
Li, S., Brown, M. S. & Goldstein, J. L. Bifurcation of insulin signaling pathway in rat liver: mTORC1 required for stimulation of lipogenesis, but not inhibition of gluconeogenesis. Proc. Natl Acad. Sci. USA 107, 3441–3446 (2010). Together with ref. 2 , the studies in refs. 32 and 33 reveal that mTORC1 induces de novo lipid synthesis through activation of the SREBP family of transcription factors.
Horton, J. D., Goldstein, J. L. & Brown, M. S. SREBPs: activators of the complete program of cholesterol and fatty acid synthesis in the liver. J. Clin. Invest. 109, 1125–1131 (2002).
Owen, J. L. et al. Insulin stimulation of SREBP-1c processing in transgenic rat hepatocytes requires p70 S6-kinase. Proc. Natl Acad. Sci. USA 109, 16184–16189 (2012).
Peterson, T. R. et al. mTOR complex 1 regulates lipin 1 localization to control the SREBP pathway. Cell 146, 408–420 (2011).
Quinn, W. J. III et al. mTORC1 stimulates phosphatidylcholine synthesis to promote triglyceride secretion. J. Clin. Invest. 127, 4207–4215 (2017).
Arsham, A. M., Plas, D. R., Thompson, C. B. & Simon, M. C. Phosphatidylinositol 3-kinase/Akt signaling is neither required for hypoxic stabilization of HIF-1 alpha nor sufficient for HIF-1-dependent target gene transcription. J. Biol. Chem. 277, 15162–15170 (2002).
He, L. et al. mTORC1 promotes metabolic reprogramming by the suppression of GSK3-dependent Foxk1 phosphorylation. Mol. Cell 70, 949–960.e4 (2018).
DeBerardinis, R. J. & Chandel, N. S. Fundamentals of cancer metabolism. Sci. Adv. 2, e1600200 (2016).
Fan, J. et al. Quantitative flux analysis reveals folate-dependent NADPH production. Nature 510, 298–302 (2014).
Zabala-Letona, A. et al. mTORC1-dependent AMD1 regulation sustains polyamine metabolism in prostate cancer. Nature 547, 109–113 (2017).
Dever, T. E. & Ivanov, I. P. Roles of polyamines in translation. J. Biol. Chem. 293, 18719–18729 (2018).
Mandal, S., Mandal, A., Johansson, H. E., Orjalo, A. V. & Park, M. H. Depletion of cellular polyamines, spermidine and spermine, causes a total arrest in translation and growth in mammalian cells. Proc. Natl Acad. Sci. USA 110, 2169–2174 (2013).
Arif, A. et al. EPRS is a critical mTORC1-S6K1 effector that influences adiposity in mice. Nature 542, 357–361 (2017).
Egan, D., Kim, J., Shaw, R. J. & Guan, K. L. The autophagy initiating kinase ULK1 is regulated via opposing phosphorylation by AMPK and mTOR. Autophagy 7, 643–644 (2011).
Kim, J., Kundu, M., Viollet, B. & Guan, K. L. AMPK and mTOR regulate autophagy through direct phosphorylation of Ulk1. Nat. Cell Biol. 13, 132–141 (2011).
Shang, L. et al. Nutrient starvation elicits an acute autophagic response mediated by Ulk1 dephosphorylation and its subsequent dissociation from AMPK. Proc. Natl Acad. Sci. USA 108, 4788–4793 (2011).
Dunlop, E. A. & Tee, A. R. mTOR and autophagy: a dynamic relationship governed by nutrients and energy. Semin. Cell Dev. Biol. 36, 121–129 (2014).
Sardiello, M. et al. A gene network regulating lysosomal biogenesis and function. Science 325, 473–477 (2009).
Settembre, C. et al. TFEB links autophagy to lysosomal biogenesis. Science 332, 1429–1433 (2011).
Peña-Llopis, S. et al. Regulation of TFEB and V-ATPases by mTORC1. EMBO J. 30, 3242–3258 (2011).
Settembre, C. et al. A lysosome-to-nucleus signalling mechanism senses and regulates the lysosome via mTOR and TFEB. EMBO J. 31, 1095–1108 (2012).
Martina, J. A., Chen, Y., Gucek, M. & Puertollano, R. MTORC1 functions as a transcriptional regulator of autophagy by preventing nuclear transport of TFEB. Autophagy 8, 903–914 (2012).
Roczniak-Ferguson, A. et al. The transcription factor TFEB links mTORC1 signaling to transcriptional control of lysosome homeostasis. Sci. Signal. 5, ra42 (2012).
Guo, J. Y. et al. Autophagy provides metabolic substrates to maintain energy charge and nucleotide pools in Ras-driven lung cancer cells. Genes Dev. 30, 1704–1717 (2016).
Frankel, L. B., Lubas, M. & Lund, A. H. Emerging connections between RNA and autophagy. Autophagy 13, 3–23 (2017).
An, H. & Harper, J. W. Systematic analysis of ribophagy in human cells reveals bystander flux during selective autophagy. Nat. Cell Biol. 20, 135–143 (2018).
Wyant, G. A. et al. NUFIP1 is a ribosome receptor for starvation-induced ribophagy. Science 360, 751–758 (2018). The studies in refs. 58 and 59 reveal that mammalian cells degrade ribosomes through either bulk autophagy (ref. 58 ) or selective autophagy (ref. 59 ), referred to as ribophagy.
Abu-Remaileh, M. et al. Lysosomal metabolomics reveals V-ATPase- and mTOR-dependent regulation of amino acid efflux from lysosomes. Science 358, 807–813 (2017).
Palm, W. et al. The utilization of extracellular proteins as nutrients is suppressed by mTORC1. Cell 162, 259–270 (2015).
Nofal, M., Zhang, K., Han, S. & Rabinowitz, J. D. mTOR inhibition restores amino acid balance in cells dependent on catabolism of extracellular protein. Mol. Cell 67, 936–946.e5 (2017).
Hara, K. et al. Amino acid sufficiency and mTOR regulate p70 S6 kinase and eIF-4E BP1 through a common effector mechanism. J. Biol. Chem. 273, 14484–14494 (1998).
Hoxhaj, G. et al. The mTORC1 signaling network senses changes in cellular purine nucleotide levels. Cell Rep. 21, 1331–1346 (2017).
Emmanuel, N. et al. Purine nucleotide availability regulates mTORC1 activity through the Rheb GTPase. Cell Rep. 19, 2665–2680 (2017).
Patel, J., Wang, X. & Proud, C. G. Glucose exerts a permissive effect on the regulation of the initiation factor 4E binding protein 4E-BP1. Biochem. J. 358, 497–503 (2001).
Dennis, P. B. et al. Mammalian TOR: a homeostatic ATP sensor. Science 294, 1102–1105 (2001).
Inoki, K., Zhu, T. & Guan, K. L. TSC2 mediates cellular energy response to control cell growth and survival. Cell 115, 577–590 (2003).
Sancak, Y. et al. The Rag GTPases bind raptor and mediate amino acid signaling to mTORC1. Science 320, 1496–1501 (2008). This is the first study indicating that mTORC1 translocates to a specific subcellular compartment, now established to be the lysosome, in response to amino acids, where it encounters and is activated by Rheb.
Menon, S. et al. Spatial control of the TSC complex integrates insulin and nutrient regulation of mTORC1 at the lysosome. Cell 156, 771–785 (2014). This study shows that the TSC complex interacts with Rheb at the lysosome in the absence of growth factors and that insulin acutely stimulates release of the TSC complex from this location via Akt-mediated phosphorylation of TSC2. This regulation allows Rheb to subsequently activate mTORC1, which is brought independently to the lysosome in response to amino acids, thus suggesting mTORC1 regulation via a spatial AND gate.
Kim, E., Goraksha-Hicks, P., Li, L., Neufeld, T. P. & Guan, K. L. Regulation of TORC1 by Rag GTPases in nutrient response. Nat. Cell Biol. 10, 935–945 (2008). Together with ref. 69 , this study identifies Rag GTPase heterodimers as direct binding partners of mTORC1 required for amino acid sensing.
Inoki, K., Li, Y., Xu, T. & Guan, K.-L. Rheb GTPase is a direct target of TSC2 GAP activity and regulates mTOR signaling. Genes Dev. 17, 1829–1834 (2003).
Tee, A. R., Manning, B. D., Roux, P. P., Cantley, L. C. & Blenis, J. Tuberous sclerosis complex gene products, Tuberin and Hamartin, control mTOR signaling by acting as a GTPase-activating protein complex toward Rheb. Curr. Biol. 13, 1259–1268 (2003).
Sancak, Y. et al. PRAS40 is an insulin-regulated inhibitor of the mTORC1 protein kinase. Mol. Cell 25, 903–915 (2007).
Long, X., Lin, Y., Ortiz-Vega, S., Yonezawa, K. & Avruch, J. Rheb binds and regulates the mTOR kinase. Curr. Biol. 15, 702–713 (2005).
Yang, H. et al. Mechanisms of mTORC1 activation by RHEB and inhibition by PRAS40. Nature 552, 368–373 (2017). This study provides important structural insights into the direct interactions and activation of mTOR within mTORC1 by Rheb–GTP.
Saxton, R. A. & Sabatini, D. M. mTOR signaling in growth, metabolism, and disease. Cell 169, 361–371 (2017).
Dibble, C. C. & Manning, B. D. Signal integration by mTORC1 coordinates nutrient input with biosynthetic output. Nat. Cell Biol. 15, 555–564 (2013).
Sancak, Y. et al. Ragulator-Rag complex targets mTORC1 to the lysosomal surface and is necessary for its activation by amino acids. Cell 141, 290–303 (2010).
Zoncu, R. et al. mTORC1 senses lysosomal amino acids through an inside-out mechanism that requires the vacuolar H+-ATPase. Science 334, 678–683 (2011).
Sekiguchi, T., Hirose, E., Nakashima, N., Ii, M. & Nishimoto, T. Novel G proteins, Rag C and Rag D, interact with GTP-binding proteins, Rag A and Rag B. J. Biol. Chem. 276, 7246–7257 (2001).
Rebsamen, M. et al. SLC38A9 is a component of the lysosomal amino acid sensing machinery that controls mTORC1. Nature 519, 477–481 (2015).
Wang, S. et al. Lysosomal amino acid transporter SLC38A9 signals arginine sufficiency to mTORC1. Science 347, 188–194 (2015). The studies in ref. 82 and ref. 83 identify SLC38A9 as a lysosomal amino acid transporter that engages Rag heterodimers and regulates mTORC1 in response to amino acids.
Bar-Peled, L., Schweitzer, L. D., Zoncu, R. & Sabatini, D. M. Ragulator is a GEF for the rag GTPases that signal amino acid levels to mTORC1. Cell 150, 1196–1208 (2012).
Martina, J. A. & Puertollano, R. Rag GTPases mediate amino acid-dependent recruitment of TFEB and MITF to lysosomes. J. Cell Biol. 200, 475–491 (2013).
Lawrence, R. E. et al. A nutrient-induced affinity switch controls mTORC1 activation by its Rag GTPase-Ragulator lysosomal scaffold. Nat. Cell Biol. 20, 1052–1063 (2018).
Zhang, Y. et al. Rheb is a direct target of the tuberous sclerosis tumour suppressor proteins. Nat. Cell Biol. 5, 578–581 (2003).
Garami, A. et al. Insulin activation of Rheb, a mediator of mTOR/S6K/4E-BP signaling, is inhibited by TSC1 and 2. Mol. Cell 11, 1457–1466 (2003).
Dibble, C. C. et al. TBC1D7 is a third subunit of the TSC1-TSC2 complex upstream of mTORC1. Mol. Cell 47, 535–546 (2012).
Manning, B. D., Tee, A. R., Logsdon, M. N., Blenis, J. & Cantley, L. C. Identification of the tuberous sclerosis complex-2 tumor suppressor gene product tuberin as a target of the phosphoinositide 3-kinase/akt pathway. Mol. Cell 10, 151–162 (2002).
Inoki, K., Li, Y., Zhu, T., Wu, J. & Guan, K. L. TSC2 is phosphorylated and inhibited by Akt and suppresses mTOR signalling. Nat. Cell Biol. 4, 648–657 (2002).
Roux, P. P., Ballif, B. A., Anjum, R., Gygi, S. P. & Blenis, J. Tumor-promoting phorbol esters and activated Ras inactivate the tuberous sclerosis tumor suppressor complex via p90 ribosomal S6 kinase. Proc. Natl Acad. Sci. USA 101, 13489–13494 (2004).
Ma, L., Chen, Z., Erdjument-Bromage, H., Tempst, P. & Pandolfi, P. P. Phosphorylation and functional inactivation of TSC2 by Erk implications for tuberous sclerosis and cancer pathogenesis. Cell 121, 179–193 (2005).
Inoki, K. et al. TSC2 integrates Wnt and energy signals via a coordinated phosphorylation by AMPK and GSK3 to regulate cell growth. Cell 126, 955–968 (2006).
Shaw, R. J. et al. The LKB1 tumor suppressor negatively regulates mTOR signaling. Cancer Cell 6, 91–99 (2004).
Howell, J. J. et al. Metformin inhibits hepatic mTORC1 signaling via dose-dependent mechanisms involving AMPK and the TSC complex. Cell Metab. 25, 463–471 (2017).
Brugarolas, J. et al. Regulation of mTOR function in response to hypoxia by REDD1 and the TSC1/TSC2 tumor suppressor complex. Genes Dev. 18, 2893–2904 (2004).
Yecies, J. L. et al. Akt stimulates hepatic SREBP1c and lipogenesis through parallel mTORC1-dependent and independent pathways. Cell Metab. 14, 21–32 (2011).
Sengupta, S., Peterson, T. R., Laplante, M., Oh, S. & Sabatini, D. M. mTORC1 controls fasting-induced ketogenesis and its modulation by ageing. Nature 468, 1100–1104 (2010).
Sabatini, D. M. Twenty-five years of mTOR: uncovering the link from nutrients to growth. Proc. Natl Acad. Sci. USA 114, 11818–11825 (2017).
Bar-Peled, L. et al. A tumor suppressor complex with GAP activity for the Rag GTPases that signal amino acid sufficiency to mTORC1. Science 340, 1100–1106 (2013).
Panchaud, N., Péli-Gulli, M. P. & De Virgilio, C. Amino acid deprivation inhibits TORC1 through a GTPase-activating protein complex for the Rag family GTPase Gtr1. Sci. Signal. 6, ra42 (2013). The studies in ref. 101 and ref. 102 identify evolutionarily conserved protein complexes (GATOR1 and GATOR2 in mammalian cells) as essential upstream regulators of mTORC1 activity by amino acid signals that influence the Rag GTPases, and show that GATOR1 serves as a RagA/B GAP.
Wolfson, R. L. et al. KICSTOR recruits GATOR1 to the lysosome and is necessary for nutrients to regulate mTORC1. Nature 543, 438–442 (2017).
Peng, M., Yin, N. & Li, M. O. SZT2 dictates GATOR control of mTORC1 signalling. Nature 543, 433–437 (2017).
Chantranupong, L. et al. The CASTOR proteins are arginine sensors for the mTORC1 pathway. Cell 165, 153–164 (2016).
Wolfson, R. L. et al. Sestrin2 is a leucine sensor for the mTORC1 pathway. Science 351, 43–48 (2016).
Saxton, R. A., Chantranupong, L., Knockenhauer, K. E., Schwartz, T. U. & Sabatini, D. M. Mechanism of arginine sensing by CASTOR1 upstream of mTORC1. Nature 536, 229–233 (2016).
Saxton, R. A. et al. Structural basis for leucine sensing by the Sestrin2-mTORC1 pathway. Science 351, 53–58 (2016).
Gu, X. et al. SAMTOR is an S-adenosylmethionine sensor for the mTORC1 pathway. Science 358, 813–818 (2017). Collectively, the studies in refs. 105–109 identify and molecularly characterize direct amino acid sensors that act upstream of the GATOR complexes and regulate mTORC1.
Shen, K. & Sabatini, D. M. Ragulator and SLC38A9 activate the Rag GTPases through noncanonical GEF mechanisms. Proc. Natl Acad. Sci. USA 115, 9545–9550 (2018).
Wyant, G. A. et al. mTORC1 activator SLC38A9 is required to efflux essential amino acids from lysosomes and use protein as a nutrient. Cell 171, 642–654.e12 (2017). The studies in ref. 110 and ref. 111 show that SLC38A9 serves as both a GEF for RagA/B (ref. 110 ) and an amino acid transporter performing efflux of neutral amino acids out of the lysosome in response to arginine binding.
Lei, H. T., Ma, J., Sanchez Martinez, S. & Gonen, T. Crystal structure of arginine-bound lysosomal transporter SLC38A9 in the cytosol-open state. Nat. Struct. Mol. Biol. 25, 522–527 (2018).
Castellano, B. M. et al. Lysosomal cholesterol activates mTORC1 via an SLC38A9-Niemann-Pick C1 signaling complex. Science 355, 1306–1311 (2017).
Petit, C. S., Roczniak-Ferguson, A. & Ferguson, S. M. Recruitment of folliculin to lysosomes supports the amino acid-dependent activation of Rag GTPases. J. Cell Biol. 202, 1107–1122 (2013).
Tsun, Z. Y. et al. The folliculin tumor suppressor is a GAP for the RagC/D GTPases that signal amino acid levels to mTORC1. Mol. Cell 52, 495–505 (2013).
Shen, K., Choe, A. & Sabatini, D. M. Intersubunit crosstalk in the Rag GTPase heterodimeR Enables mTORC1 to respond rapidly to amino acid availability. Mol. Cell 68, 552–565.e8 (2017).
Choo, A. Y. et al. Glucose addiction of TSC null cells is caused by failed mTORC1-dependent balancing of metabolic demand with supply. Mol. Cell 38, 487–499 (2010).
Efeyan, A. et al. Regulation of mTORC1 by the Rag GTPases is necessary for neonatal autophagy and survival. Nature 493, 679–683 (2013).
Long, X., Ortiz-Vega, S., Lin, Y. & Avruch, J. Rheb binding to mTOR is regulated by amino acid sufficiency. J. Biol. Chem. 280, 23433–23436 (2005).
Smith, E. M., Finn, S. G., Tee, A. R., Browne, G. J. & Proud, C. G. The tuberous sclerosis protein TSC2 is not required for the regulation of the mammalian target of rapamycin by amino acids and certain cellular stresses. J. Biol. Chem. 280, 18717–18727 (2005).
Nobukuni, T. et al. Amino acids mediate mTOR/raptor signaling through activation of class 3 phosphatidylinositol 3OH-kinase. Proc. Natl Acad. Sci. USA 102, 14238–14243 (2005).
Demetriades, C., Doumpas, N. & Teleman, A. A. Regulation of TORC1 in response to amino acid starvation via lysosomal recruitment of TSC2. Cell 156, 786–799 (2014).
Carroll, B. et al. Control of TSC2-Rheb signaling axis by arginine regulates mTORC1 activity. eLife 5, e11058 (2016).
Demetriades, C., Plescher, M. & Teleman, A. A. Lysosomal recruitment of TSC2 is a universal response to cellular stress. Nat. Commun. 7, 10662 (2016).
Gwinn, D. M. et al. AMPK phosphorylation of raptor mediates a metabolic checkpoint. Mol. Cell 30, 214–226 (2008).
Carrière, A. et al. Oncogenic MAPK signaling stimulates mTORC1 activity by promoting RSK-mediated raptor phosphorylation. Curr. Biol. 18, 1269–1277 (2008).
Carriere, A. et al. ERK1/2 phosphorylate Raptor to promote Ras-dependent activation of mTOR complex 1 (mTORC1). J. Biol. Chem. 286, 567–577 (2011).
Foster, K. G. et al. Regulation of mTOR complex 1 (mTORC1) by raptor Ser863 and multisite phosphorylation. J. Biol. Chem. 285, 80–94 (2010).
Kovacina, K. S. et al. Identification of a proline-rich Akt substrate as a 14-3-3 binding partner. J. Biol. Chem. 278, 10189–10194 (2003).
Vander Haar, E., Lee, S. I., Bandhakavi, S., Griffin, T. J. & Kim, D. H. Insulin signalling to mTOR mediated by the Akt/PKB substrate PRAS40. Nat. Cell Biol. 9, 316–323 (2007).
Jewell, J. L. et al. Differential regulation of mTORC1 by leucine and glutamine. Science 347, 194–198 (2015).
Ye, J. et al. GCN2 sustains mTORC1 suppression upon amino acid deprivation by inducing Sestrin2. Genes Dev. 29, 2331–2336 (2015).
Wolfson, R. L. & Sabatini, D. M. The dawn of the age of amino acid sensors for the mTORC1 pathway. Cell Metab. 26, 301–309 (2017).
Parmigiani, A. et al. Sestrins inhibit mTORC1 kinase activation through the GATOR complex. Cell Rep. 9, 1281–1291 (2014).
Kim, J. S. et al. Sestrin2 inhibits mTORC1 through modulation of GATOR complexes. Sci. Rep. 5, 9502 (2015).
Fruman, D. A. et al. The PI3K pathway in human disease. Cell 170, 605–635 (2017).
Kang, S. A. et al. mTORC1 phosphorylation sites encode their sensitivity to starvation and rapamycin. Science 341, 1236566 (2013).
Thoreen, C. C. et al. An ATP-competitive mammalian target of rapamycin inhibitor reveals rapamycin-resistant functions of mTORC1. J. Biol. Chem. 284, 8023–8032 (2009).
Nyfeler, B. et al. Relieving autophagy and 4EBP1 from rapamycin resistance. Mol. Cell. Biol. 31, 2867–2876 (2011).
Valvezan, A. J. et al. mTORC1 couples nucleotide synthesis to nucleotide demand resulting in a targetable metabolic vulnerability. Cancer Cell 32, 624–638.e5 (2017). This study on the concept of anabolic balance downstream of mTORC1 has demonstrated that uncoupling mTORC1-stimulated nucleotide synthesis, via IMPDH inhibitors, from its induction of rRNA synthesis, induces rapid nucleotide depletion, replication stress and cell death in cell and tumour models of tuberous sclerosis complex .
Young, R. M. et al. Dysregulated mTORC1 renders cells critically dependent on desaturated lipids for survival under tumor-like stress. Genes Dev. 27, 1115–1131 (2013).
Zhang, Y. & Manning, B. D. mTORC1 signaling activates NRF1 to increase cellular proteasome levels. Cell Cycle 14, 2011–2017 (2015).
Zhao, J., Zhai, B., Gygi, S. P. & Goldberg, A. L. mTOR inhibition activates overall protein degradation by the ubiquitin proteasome system as well as by autophagy. Proc. Natl Acad. Sci. USA 112, 15790–15797 (2015).
Rousseau, A. & Bertolotti, A. An evolutionarily conserved pathway controls proteasome homeostasis. Nature 536, 184–189 (2016).
Griffiths, B. et al. Sterol regulatory element binding protein-dependent regulation of lipid synthesis supports cell survival and tumor growth. Cancer Metab. 1, 3 (2013).
Zhang, Y. et al. Coordinated regulation of protein synthesis and degradation by mTORC1. Nature 513, 440–443 (2014).
Fok, W. C. et al. Mice fed rapamycin have an increase in lifespan associated with major changes in the liver transcriptome. PLoS One 9, e83988 (2014).
Zhang, Y. et al. Rapamycin extends life and health in C57BL/6 mice. J. Gerontol. A Biol. Sci. Med. Sci. 69, 119–130 (2014).
Yun, Y. S. et al. mTORC1 coordinates protein synthesis and immunoproteasome formation via PRAS40 to prevent accumulation of protein stress. Mol. Cell 61, 625–639 (2016).
Choi, J. H. et al. mTORC1 accelerates retinal development via the immunoproteasome. Nat. Commun. 9, 2502 (2018).
Radhakrishnan, S. K. et al. Transcription factor Nrf1 mediates the proteasome recovery pathway after proteasome inhibition in mammalian cells. Mol. Cell 38, 17–28 (2010).
Steffen, J., Seeger, M., Koch, A. & Krüger, E. Proteasomal degradation is transcriptionally controlled by TCF11 via an ERAD-dependent feedback loop. Mol. Cell 40, 147–158 (2010).
Torrence, M. E. & Manning, B. D. Nutrient sensing in cancer. Annu Rev. Cancer Biol. 2, 251–269 (2018).
Perl, A. metabolic control of immune system activation in rheumatic diseases. Arthritis Rheumatol. 69, 2259–2270 (2017).
Cummings, N. E. & Lamming, D. W. Regulation of metabolic health and aging by nutrient-sensitive signaling pathways. Mol. Cell. Endocrinol. 455, 13–22 (2017).
Johnson, S. C., Rabinovitch, P. S. & Kaeberlein, M. mTOR is a key modulator of ageing and age-related disease. Nature 493, 338–345 (2013).
Ilagan, E. & Manning, B. D. Emerging role of mTOR in the response to cancer therapeutics. Trends Cancer 2, 241–251 (2016).
Nickerson, M. L. et al. Mutations in a novel gene lead to kidney tumors, lung wall defects, and benign tumors of the hair follicle in patients with the Birt-Hogg-Dubé syndrome. Cancer Cell 2, 157–164 (2002).
Grabiner, B. C. et al. A diverse array of cancer-associated MTOR mutations are hyperactivating and can predict rapamycin sensitivity. Cancer Discov. 4, 554–563 (2014).
Wagle, N. et al. Activating mTOR mutations in a patient with an extraordinary response on a phase I trial of everolimus and pazopanib. Cancer Discov. 4, 546–553 (2014).
Okosun, J. et al. Recurrent mTORC1-activating RRAGC mutations in follicular lymphoma. Nat. Genet. 48, 183–188 (2016).
Ben-Sahra, I. & Manning, B. D. mTORC1 signaling and the metabolic control of cell growth. Curr. Opin. Cell Biol. 45, 72–82 (2017).
Henske, E. P., Jóźwiak, S., Kingswood, J. C., Sampson, J. R. & Thiele, E. A. Tuberous sclerosis complex. Nat. Rev. Dis. Prim. 2, 16035 (2016).
French, J. A. et al. Adjunctive everolimus therapy for treatment-resistant focal-onset seizures associated with tuberous sclerosis (EXIST-3): a phase 3, randomised, double-blind, placebo-controlled study. Lancet 388, 2153–2163 (2016).
Marsan, E. & Baulac, S. Mechanistic target of rapamycin (mTOR) pathway, focal cortical dysplasia and epilepsy. Neuropathol. Appl. Neurobiol. 44, 6–17 (2018).
Curatolo, P., Moavero, R. & de Vries, P. J. Neurological and neuropsychiatric aspects of tuberous sclerosis complex. Lancet Neurol. 14, 733–745 (2015).
Frake, R. A., Ricketts, T., Menzies, F. M. & Rubinsztein, D. C. Autophagy and neurodegeneration. J. Clin. Invest. 125, 65–74 (2015).
Duman, R. S. Ketamine and rapid-acting antidepressants: a new era in the battle against depression and suicide. F1000Res. 7, F1000 (2018). Faculty Rev-659.
Um, S. H., D’Alessio, D. & Thomas, G. Nutrient overload, insulin resistance, and ribosomal protein S6 kinase 1, S6K1. Cell Metab. 3, 393–402 (2006).
Acknowledgements
Studies in the laboratory of B.D.M. related to this subject were supported by American Cancer Society postdoctoral fellowship 127106-PF-14-254-01-TBE to A.J.V., NIH grants R35-CA197459 and P01-CA120964 to B.D.M., and a Rothberg Courage Award from the Tuberous Sclerosis Alliance to B.D.M.
Author information
Authors and Affiliations
Contributions
A.J.V. and B.D.M. conceived, researched and wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
B.D.M. is a shareholder and scientific advisory board member of Navitor Pharmaceuticals and LAM Therapeutics.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Valvezan, A.J., Manning, B.D. Molecular logic of mTORC1 signalling as a metabolic rheostat. Nat Metab 1, 321–333 (2019). https://doi.org/10.1038/s42255-019-0038-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s42255-019-0038-7