Abstract
The daily activity in the brain is typically fine-tuned by the circadian clock in the local neurons as well as by the master circadian clock in the suprachiasmatic nucleus (SCN) of the hypothalamus. In the olfactory response, odor-evoked activity in the piriform cortex (PC) and olfactory behavior retain circadian rhythmicity in the absence of the SCN, yet how the circadian rhythm in the PC is achieved independently of the SCN remains elusive. Here, to define neurons regulating the circadian rhythm of the odor-evoked activity in the PC, we knocked out the clock gene Bmal1 in a host of specific neurons along the olfactory circuit. We discovered that Bmal1 knockout in the PC largely abolishes the circadian rhythm of the odor-evoked activity. We further showed that isolated PC exhibits sustained circadian rhythms of the clock gene Per2 expression. Quantitative PCR analysis revealed that expression patterns of multiple genes involved in neural activity and synaptic transmission exhibit circadian rhythm in the PC in a BMAL1-dependent manner. Our findings indicate that BMAL1 acts intrinsically in the PC to control the circadian rhythm of the odor-evoked activity in the PC, possibly through regulating expression patterns of multiple genes involved in neural activity and transmission.
Similar content being viewed by others
Introduction
Many organs including the brain exhibit circadian rhythms in their biological processes. Recent physiological and behavioral studies showed that the circadian rhythmicity is observed in multiple neural functions including memory & learning, emotional response, and sensory activity such as audition and olfaction1,2,3,4. These circadian rhythms of neural activities are typically generated and maintained through a series of transcription factors called clock genes. The transcription/translation levels of these clock genes oscillate in a 24-h cycle, via the negative transcription-translation feedback loop regulation5,6,7. Indeed, knocking out the core clock genes such as Bmal1 and Period1/2 (Per1/2) largely abolishes the circadian rhythms of neural activities8,9, underscoring their crucial roles in the circadian rhythms of neural activities.
In mammals, the suprachiasmatic nucleus (SCN) of the hypothalamus is widely believed to be the master circadian oscillator orchestrating the clock of the entire body10,11,12. Consistently, the core clock genes are abundantly expressed in the SCN13,14. On the other hand, a growing body of evidence suggests that the core clock genes are widely expressed in virtually all cells of the entire body, and that certain neural activities show circadian rhythms independent of the SCN15,16. One such example is the olfactory circuit, in which odor information is initially detected by the olfactory sensory neurons in the olfactory epithelium, then relayed to the mitral/tufted cells in the olfactory bulb, and eventually processed in the pyramidal neurons of the piriform cortex (PC)17,18. Odor-evoked activity in the PC shows a circadian rhythm with the highest response at night1, which is paralleled by the olfactory discrimination accuracy19,20. This circadian rhythmicity of the odor-evoked activity in the PC is unaffected by surgical removal of the SCN, although the oscillation phase tends to be affected1. These data suggest that the circadian rhythmicity of the odor-evoked activity in the PC is generated by circadian oscillators outside of the SCN. Consistent with this notion, surgical removal of the olfactory bulb in mice partially disturbs the circadian rhythm of the odor-evoked activity in the PC1. It is, however, still unknown what neural populations in the olfactory circuit contributes to the circadian rhythm of the odor-evoked activity in the PC, partially due to less accessibility of the surgical approach.
In this study, by combining multiple Cre recombinase-expressing mice and adeno-associated virus (AAV) transfection, we knocked out the clock gene Bmal1 in specific sets of neurons in the olfactory circuit. We show that BMAL1 in the PC is required for the circadian rhythm of the odor-evoked activity. Furthermore, the isolated PC ex vivo maintained the circadian oscillation of Per2 expression, another core clock gene. Interestingly, we found that expression levels of multiple genes related to neural activity and synaptic transmission exhibit circadian rhythms in the PC in a BMAL1-dependent manner. These findings suggest that the intrinsic molecular clock in the PC is critical for the circadian rhythm of the odor-evoked neural activity, and hint at the possibility that the intrinsic clock might exert such function through transcriptional regulation of the molecules related to neural activity and synaptic transmission.
Results
Odor-evoked neural activity in the PC exhibits a circadian rhythm
To investigate the circadian rhythm of the odor-evoked activity in the mouse brain, we presented cedar oil to the mouse as described previously1, and quantified the number of c-Fos positive cells21 in the PC for six timepoints throughout the day (Fig. 1a). Consistent with previous studies1,22, by comparing the number of c-Fos positive cells among six timepoints, we observed a peak at the subjective night CT16 (Fig. 1b, Supplementary Fig. 1a). Of note, the number of c-Fos positive cells showed no obvious correlation with other external conditions such as humidity and temperature (Supplementary Fig. 1b), suggesting that the circadian rhythm of odor-evoked PC activity is unrelated to external stimuli, but rather regulated through an internal mechanism. Additionally, mice exposed to another neutral odorant limonene(-) showed a significantly higher number of c-Fos positive cells in the PC at CT16 compared to CT4 (Supplementary Fig. 1c), comparable to the response to cedar oil. Thus, the circadian rhythm of the odor-evoked activity in the PC is not specific to cedar oil.
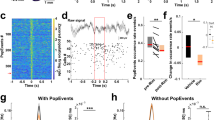
a Schematic illustration of the odor-presentation experiment for c-Fos quantification. 100 µL of cedar oil (1:1000 dilution in paraffin oil) was presented at six different timepoints for 5 min. b The number of c-Fos positive cells in the PC of mice exposed to the odor at six timepoints (CT0, 4, 8, 12, 16, 20). N = 4, 6, 3, 4, 8, 4 for CT0, CT4, CT8, CT12, CT16, CT20, respectively. p = 0.001185, one-way ANOVA. The line at the center of each boxplot depicts the median; the box depicts the third quartile and first quartile. c Representative whole-brain reconstruction of c-Fos positive cells in mice that were kept in home cages and then sacrificed at two timepoints (CT4, CT16). Red dots indicate individual c-Fos positive cells. d Quantification of the number of c-Fos positive cells in the whole brain. N = 4 and 2 for CT4 and CT16, respectively. e The number of c-Fos positive cells in five sensory cortical areas: visual cortex (V1), auditory cortex (A1), gustatory cortex, and PC in mice that were kept in home cages and then sacrificed at two timepoints (CT4, CT16). N = 4 and 2 for CT4 and CT16, respectively.
We next assessed whether odor-evoked neural activity might specifically fluctuate in the PC. To this end, we collected brain samples from subjective day CT4 and subjective night CT16 and quantified the number of c-Fos positive cells in serial sections spanning the whole brain. We found that the total number of c-Fos positive cells was higher at CT16 compared to CT4 (Fig. 1c, d). We further quantified the number of c-Fos positive cells in each sensory cortex by mapping the c-Fos positive cells onto a reference mouse brain of the Allen Brain institute three-dimensionally23. As a result, while the PC showed a higher number of c-Fos positive cells at CT16 (Fig. 1e), the number of c-Fos positive cells was scarce both at CT4 and CT16 in the primary visual cortex (V1), primary auditory cortex (A1) and gustatory cortex (Fig. 1e). The circadian fluctuation of neural activity is thus most obvious in the PC among sensory cortices. Interestingly, we found multiple brain areas in which c-Fos-positive cells were significantly higher at CT16 compared to CT4 including the somatomotor cortex, cortical subplate, basolateral amygdala, and hypothalamus (Supplementary Fig. 1d).
Circadian rhythmicity of the odor-evoked activity in the PC are intrinsically maintained independently of the SCN
Circadian rhythms of neural activities are widely thought to be generated by the SCN10,11,12 and the clock genes such as Bmal18,9. While a previous report has suggested that the circadian rhythm of the odor-evoked activity in the PC operates independent of the SCN1, this report solely relied on surgical removal of the SCN, which could be confounded by inevitable damages to other brain structures. We thus sought to test whether BMAL1 expression in the SCN contributes to the circadian rhythm of the odor-evoked activity in the PC with less invasive methodologies. To this end, we took advantage of the evidence that the SCN is mostly comprised of GABAergic neurons, and that GABAergic neurons are required for the circadian rhythm of SCN activity24. Specifically, we conditionally knocked out the clock gene Bmal1 in GABAergic neurons by crossing Vgat-Cre mice25 against Bmal1-floxed mice in which loxP sequences are inserted in the coding region of the Bmal1 gene26. As expected, the number of BMAL1-positive cells in the SCN was significantly reduced in these conditional KO mice (“Vgat-Bmal1 KO mice”) compared to control mice (Fig. 2a). While control mice showed a clear circadian rhythm in locomotor activity both under the LD condition (light condition is indicated by yellow-shaded area) and the following DD condition (Fig. 2b, left), Vgat-Bmal1 KO mice failed to show a circadian rhythm under the DD condition (Fig. 2b, right), suggesting that the circadian rhythm in the SCN was effectively disrupted in Vgat-Bmal1 KO mice. By contrast, the number of c-Fos positive cells in the PC following cedar oil presentation retained circadian rhythmicity (Fig. 2c), suggesting that BMAL1 expression in GABAergic SCN neurons is dispensable for the circadian rhythmicity of the odor-evoked activity in the PC. We note that the number of c-Fos positive cells peaked at CT4 in Vgat-Bmal1 KO mice while this number peaked at CT16 in control mice (Fig. 1d), consistent with a previous study in which SCN was surgically removed1. These data suggests that BMAL1 in GABAergic SCN neurons is dispensable for rhythm generation but might contribute to phase control of the circadian rhythm of the odor-evoked activity in the PC.
a Representative images of the suprachiasmatic nucleus (SCN) of Bmal1-control mice and Vgat-Bmal1 KO mice, immunohistochemically stained with BMAL1 antibody. The ratios of BMAL1 + cells over DAPI + cells were quantified in the SCN. b Representative locomotor activity patterns of Bmal1-control mice and Vgat-Bmal1 KO mice. Yellow shades and gray shades indicate light ON and OFF respectively. c Quantification of c-Fos positive cells in the PC of Vgat-Bmal1 KO mice which underwent odorant exposure identical to Fig. 5A. N = 3, 2, 4, 2 for CT4, CT8, CT16, CT20, respectively. p = 0.03224, one-way ANOVA. The line at the center of each boxplot depicts the median; the box depicts the third quartile and first quartile.
Bmal1 KO in neurons containing GCs and PC disrupts the circadian rhythm of the odor-evoked activity in the PC
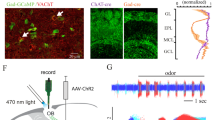
Since we failed to detect contribution of BMAL1 in the SCN to the circadian rhythmicity of the odor-evoked activity of the PC, we next asked whether BMAL1 functions inside the olfactory circuit. To this end, against Bmal1-floxed mice, we crossed multiple transgenic mice expressing Cre recombinase known to label specific neuronal populations along the olfactory circuit, including Omp-Cre mice27, Pcdh21-CreER mice28, and Emx1-Cre mice29. To check precisely where and how efficiently Bmal1 was knocked out in these conditional KO mice (“Omp-Bmal1 KO mice”, “Pcdh21-Bmal1 KO mice”, “Emx1-Bmal1 KO mice”), we first conducted immunohistochemical analyses in the olfactory circuit including mitral/tufted cells, granule cells of the olfactory bulb, and the PC. In the mitral/tufted cell layer, we analyzed the ratio of BMAL1-positive cells against Tbx21-positive mitral/tufted cells and found that 97.6% of Tbx21-positive cells were BMAL1-positive in Bmal1-control mice while this ratio dropped to 64.9 ± 2.29% and 7.02 ± 5.85% in Pcdh21-Bmal1 KO mice and Emx1-Bmal1 KO mice, respectively (Fig. 3a). In the granule cell layer of the olfactory bulb, we analyzed the ratio of BMAL1-positive cells in five different depths since different granule cell types likely reside in the different depths30,31. As a result, while >90% of Nissl-positive neurons was BMAL1-positive in Bmal1-control mice and Pcdh21-Bmal1 KO mice, this ratio was reduced in all depths of the Emx1-Bmal1 KO mice (Fig. 3b). In the PC, while the ratio of BMAL1-positive cells was 98.6 ± 0.69% in Bmal1-control mice, this ratio was reduced to 90.2 ± 3.78% and 25.3 ± 5.40% in Pcdh21-Bmal1 KO mice and Emx1-Bmal1 KO mice, respectively (Fig. 3c). Thus, by utilizing the three Cre-lines, we successfully managed to knock out Bmal1 in different neuronal subsets of the olfactory circuit (Fig. 3d).
a Representative images of the mitral/tufted cell layer in the olfactory bulb of Bmal1-floxedfl/fl mice (Bmal1-control), Pcdh21-CreERTg/+; Bmal1-floxedfl/fl mice (Pcdh21-Bmal1 KO) and Emx1-CreTg/+; Bmal1-floxedfl/fl mice (Emx1-Bmal1 KO) immunohistochemically stained with BMAL1 antibody and mitral/tufted cell marker Tbx21 antibody. Arrowheads indicate BMAL1-; Tbx21+ cells. The ratios of BMAL1 + cells over Tbx21+ cells were 97.6%, 64.9 ± 2.29%, 7.1 ± 5.85% (mean ± standard deviation), in Bmal1-control mice, Pcdh21-Bmal1 KO mice, and Emx1-Bmal1 KO mice, respectively. The line at the center of each boxplot depicts the median; the box depicts the third quartile and first quartile. b Representative images of the granule cell layer in the olfactory bulb of Bmal1-control mice, Pcdh21-Bmal1 KO mice and Emx1-Bmal1 KO mice immunohistochemically stained with BMAL1 antibody and neuron marker Nissl. The ratio of BMAL1 + cells over Nissl+ cells were quantified in granule cell layers divided into five layers. N = 3 for each genotype, 91.1 ± 5.24, 99.0 ± 1.75, 53.5 ± 13.6%; 100.0 ± 0.00, 98.1 ± 0.09, 64.1 ± 5.53%; 97.7 ± 4.03, 98.6 ± 1.23, 73.9 ± 6.52%; 98.6 ± 1.18, 97.8 ± 2.68, 84.6 ±;8.55%; 96.9 ± 2.73, 98.3 ± 1.63, 82.1 ± 6.72% (mean ± standard deviation), in layers 1~5 of Bmal1-control mice, Pcdh21-Bmal1 KO mice, and Emx1-Bmal1 KO mice, respectively. The line at the center of each boxplot depicts the median; the box depicts the third quartile and first quartile. c Representative images of the PC in Bmal1-control mice, Pcdh21-Bmal1 KO mice, and Emx1-Bmal1 KO mice immunohistochemically stained with BMAL1 antibody and DAPI. The ratios of BMAL1 + cells over DAPI + cells were 98.6;± 0.69%, 90.2 ± 3.78%, 25.3 ± 5.43% (mean ± standard deviation), in Bmal1-control mice, Pcdh21-Bmal1 KO mice, and Emx1-Bmal1 KO mice, respectively. N = 4, 2, 4 for Bmal1-control mice, Pcdh21-Bmal1 KO mice, and Emx1-Bmal1 KO mice, respectively. The line at the center of each boxplot depicts the median; the box depicts the third quartile and first quartile. d Summary of the regions where Bmal1 was knocked-out in each Cre-line. In Omp-Bmal1KO mice, Bmal1 was knocked out in olfactory sensory neurons. In Pcdh21-Bmal1 KO mice, Bmal1 was knocked out in mitral/tufted cells. In Emx1-Bmal1 KO mice, Bmal1 was knocked out in mitral/tufted cells, granule cells, and PC neurons. e The number of c-Fos positive cells in the PC of Bmal1-control mice, Omp-Bmal1 KO mice, Pcdh21-Bmal1 KO mice, Emx1-Bmal1 KO mice exposed to an odorant. For Bmal1-control mice, N = 13 each for CT4 and CT16. For Omp-Bmal1 KO mice, N = 11 and 9 for CT4 and CT16, respectively. For Pcdh21-Bmal1 KO mice, N = 11 and 8 for CT4 and CT16, respectively. For Emx1-Bmal1 KO mice, N = 9 each for CT4 and CT16. p = 0.005072, 0.006549, 0.02052, 0.1615 for Bmal1-control mice, Omp-Bmal1 KO mice, Pcdh21-Bmal1 KO mice, Emx1-Bmal1 KO mice, respectively. Wilcoxon rank-sum test with Bonferroni correction. The line at the center of each boxplot depicts the median; the box depicts the third quartile and first quartile.
We next presented cedar oil to these mice and quantified the number of c-Fos positive cells in the PC. First, in the control group where Cre recombinase is not expressed (Bmal1-control mice), the number of c-Fos positive cells in the PC was significantly higher at CT16 compared to CT4 (Fig. 3e). Similarly, the number of c-Fos positive cells in the PC was significantly higher at CT16 compared to CT4 both in Omp-Bmal1 KO mice and in Pcdh21-Bmal1 KO mice (Fig. 3e). By contrast, we observed no significant difference in the number of c-Fos positive cells in the PC between CT4 and CT16 in Emx1-Bmal1 KO mice (Fig. 3e). Further, Pcdh21-Bmal1 KO mice and Emx1-Bmal1 KO mice both showed rhythmic circadian locomotor activity in a constant dark condition (Supplementary Fig. 2), suggesting that the SCN-dependent circadian regulation was maintained in both KO mice. Given that Bmal1 is knocked out in the mitral/tufted cells, granule cells, and the PC in Emx1-Bmal KO mice and that Pcdh21-Bmal1 KO mice in which Bmal1 is knocked out in the mitral/tufted cells retained circadian rhythm of the odor-evoked activity in the PC, our data suggest that BMAL1 in the granule cells and/or the PC is required for the circadian rhythm of the odor-evoked activity in the PC.
AAV-mediated PC-specific Bmal1 KO abolishes the circadian rhythm of the odor-evoked activity in the PC
To further define the neural population required for the circadian rhythm of the odor-evoked activity in the PC, we limited Bmal1 KO into a specific population of the olfactory circuit by expressing Cre recombinase via adeno-associated virus (AAV) injection. Specifically, we generated AAV where Cre recombinase and red fluorescent protein mCherry are simultaneously expressed downstream of the CaMKIIα promoter. CaMKIIα is known to be expressed both in the granule cells of the olfactory bulb and in the pyramidal neurons of the PC32,33,34. When we injected this AAV to the granule cell layers of the olfactory bulb (Fig. 4a), mCherry expression was largely observed in the deeper layers of the granule cell, and correspondingly BMAL1 expression was especially weak in the deeper layers (Fig. 4b, c, Supplementary Fig. 3a, b). We validated that mCherry is not expressed in cells outside of the granule cell layers, i.e. mitral/tufted cells (Supplementary Fig. 3a). When we injected the AAV to the PC (Fig. 4d), 55.5 ± 6.14% of Nissl-positive cells expressed mCherry (Supplementary Fig. 3c, d), and only 13.5 ± 8.59% of Nissl-positive cells expressed BMAL1 (Fig. 4e, f).
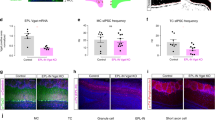
a AAVDJ-CaMKII-iCre-T2A-mCherry injection to the olfactory bulb (GC-Bmal1 KO). b Representative images of the granule cell layer in the olfactory bulb of GC-Bmal1 KO mice, immunohistochemically stained with BMAL1 antibody. c The ratio of BMAL1 + cells over Nissl+ cells were quantified in granule cell layers divided into five layers. For Bmal1-control mice: 91.1 ± 5.24%, 100.0 ± 0.00%, 97.7 ± 4.03%, 98.6 ± 1.18%, 96.9 ± 2.73%, (mean ± standard deviation), in layers 1~5, respectively. For GC-Bmal1 KO mice: 92.5 ± 7.81%, 96.8 ± 2.80%, 69.3 ± 15.8%, 17.1 ± 6.82%, 36.6 ± 8.85%, (mean ± standard deviation), in layers 1~5, respectively. N = 3 and 4 for Bmal1-control mice and GC-Bmal1 KO mice, respectively. The line at the center of each boxplot depicts the median; the box depicts the third quartile and first quartile. d AAVDJ-CaMKII-iCre-T2A-mCherry injection to the PC (PC-Bmal1 KO). e Representative images of the PC of PC-Bmal1 KO mice, immunohistochemically stained with BMAL1 antibody. f The ratios of BMAL1 + cells over Nissl+ cells were quantified in the PC. 98.6 ± 0.69%, 13.5 ± 8.59% (mean ± standard deviation) for Bmal1-control mice and PC-Bmal1 KO mice, respectively. N = 4 and 7 for Bmal-control mice and PC-Bmal1 KO mice, respectively. The line at the center of each boxplot depicts the median; the box depicts the third quartile and first quartile. g The number of c-Fos positive cells in the PC of olfactory bulb-injected granule cell conditional KO mice exposed to an odorant. N = 9 each for CT4 and CT16. p = 0.02394, Wilcoxon rank-sum test. The line at the center of each boxplot depicts the median; the box depicts the third quartile and first quartile. h The number of c-Fos positive cells in the PC of PC-injected with AAVDJ-CaMKII-mCherry (PC Control) and PC conditional KO mice (PC-Bmal1 KO) exposed to an odorant. For PC Control, N = 4 and 6 for CT4 and CT16, respectively. p = 0.07323, one-way ANOVA. For PC-Bmal1 KO, N = 7, 2, 7, 3 for CT4, CT10, CT16, CT22, respectively. p = 0.9506, one-way ANOVA. The line at the center of each boxplot depicts the median; the box depicts the third quartile and first quartile.
We next presented cedar oil to these mice and quantified the number of c-Fos positive cells in the PC. In mice where Bmal1 was conditionally knocked out in granule cells (“GC-Bmal1 KO mice”), the number of c-Fos positive cells was significantly higher at CT16 compared to CT4 (Fig. 4g). On the other hand, in mice where Bmal1 was knocked out in the PC (“PC-Bmal1 KO mice”), there was no significant difference in the number of c-Fos positive cells in the PC across four timepoints (Fig. 4h). Since the number of c-Fos positive cells was significantly higher at CT16 compared to CT4 in control group in which AAV-CaMKII-mCherry was injected to the PC (Fig. 4h), AAV injection alone has no significant effect on odor-evoked activity in the PC. These data suggest that the circadian rhythm of the odor-evoked activity in the PC was abolished in these mice. PC-Bmal1 KO mice showed rhythmic locomotor activity in a constant dark condition (Supplementary Fig. 3e), suggesting that the molecular clock in the SCN is functionally intact in PC-Bmal1 KO mice. These results indicate that BMAL1 expression in the CaMKIIα-positive pyramidal neurons of the PC is required for the circadian rhythm of the odor-evoked activity in the PC.
PC can maintain circadian rhythm of clock gene expression in an autonomous manner
Given that BMAL1 expression in the CaMKIIα-positive pyramidal neurons of the PC is responsible for the circadian rhythm of the odor-evoked activity in the PC, we reasoned that PC may be capable of self-sustaining the circadian rhythm of its clock gene expression. To test this possibility, we next conducted an ex vivo measurement of the circadian rhythm of the intrinsic clock using isolated tissues from PER2::LUC knock-in mice, in which the firefly luciferase cDNA is inserted immediately downstream of the Per2 gene35. In brief, we generated acute slice sections of the PC and measured the luminescence level for up to 1 week in a luciferin-containing medium (Fig. 5a). As a positive control, we first confirmed that the SCN showed a sustained circadian rhythm of luminescence with a period of 24.4 ± 1.10 h (Fig. 5b) as reported previously35. Importantly, we observed sustained circadian rhythm in the PC with a period of 24.73 ± 0.43 h (Fig. 5b). There was no significant difference in the observed period between the SCN and the PC (Fig. 5c). To investigate whether action potential-based neural activity is necessary for the oscillation of clock gene expression in the PC, we applied tetrodotoxin (TTX) to the culture medium and found that the circadian rhythm damped out in the PC as well as the SCN and that the rhythm was re-activated by media change safter TTX treatment. These data suggest that neural activity is necessary for circadian oscillation of clock gene expression in the PC (Fig. 5d). Our findings thus indicate that the PC can intrinsically maintain the circadian rhythm of the clock gene expression in the absence of other brain regions.
a Schematic illustration of luciferase assay. The PC and suprachiasmatic nucleus were cut out from PER2::LUC mice and incubated in a luciferin-containing medium for up to 1 week. b Luminescence levels of the suprachiasmatic nucleus and the PC. Lines indicate the mean luminescence levels, and the shade indicates the mean ± standard error. c Mean period of the suprachiasmatic nucleus and PC. Periods were calculated by the Lomb-Scargle periodogram analysis. N = 4, 24.4 ± 1.10 h; N = 13, 24.73 ± 0.43 h, for the SCN and PC, respectively. p = 0.5331, Wilcoxon rank-sum test. The line at the center of each boxplot depicts the median; the box depicts the third quartile and first quartile. d Representative graphs of the luminescence levels of the SCN and the PC with TTX application.
Genes related to neural activity show circadian rhythms in the PC in a BMAL1-dependent manner
Given that clock genes including Bmal1 and Per2 encode transcriptional regulators, the molecular clock in the PC is likely to regulate neural activity in a circadian manner through downstream gene expression. A previous study36 identified a set of genes acting downstream of the clock genes in the SCN, and we reasoned that at least some of these genes are likely to operate similarly in the PC. We thus extracted nine candidates from this set of genes deemed related to neural function through our gene ontology analyses and measured the mRNA levels of these genes in the PC at six timepoints throughout the day (Fig. 6a, Supplementary Table 1). We found that seven out of nine genes—Avpr1a, Cacna2d3, Chrnb2, Clcn4, Gad1, Snap25, and SynI—showed circadian rhythms of expression levels, and all seven peaked at CT8 (Fig. 6b). Next, we investigated whether the mRNA levels of these seven genes are regulated through BMAL1 transcription regulation. To this end, we collected the PC from Emx1-Bmal1 KO mice and quantified the mRNA levels of these genes. First, we confirmed that the mRNA levels of Bmal1 were significantly suppressed compared to wild-type mice (Supplementary Fig. 4). Importantly, we failed to detect circadian rhythms in the seven genes that showed circadian rhythm in the wild-type mice (Fig. 6c), suggesting that the expression of these genes is transcriptionally regulated through BMAL1 in the PC. Taken together, our data suggest that BMAL1 in the PC regulates the expression patterns of multiple target genes related to neural function in a circadian manner, which could contribute to intrinsic control of the odor-evoked neural activity of the PC.
a Schematic illustration of real-time quantitative PCR. Mice were kept in 12-h Light/Dark condition for at least 3 weeks. Prior to sacrifice, mice underwent a constant dark condition, and PC and SCN were dissected for and RNA extraction, purification for real-time quantitative PCR. b Relative mRNA expression of Avpr1a, Cacna2d3, Calb2, Chrnb2, Clcn4, Gad1, Snap25, Sst, Syn1 mRNA levels in the PC of wild-type mice. N = 4, 3, 4, 4, 3, 4 for CT0, CT4, CT8, CT12, CT16, CT20, respectively. One-way ANOVA with Bonferroni correction. Error bars indicate standard error. c Relative mRNA expression of Avpr1a, Cacna2d3, Calb2, Chrnb2, Clcn4, Gad1, Snap25, Sst, Syn1 mRNA levels in the PC of Emx1-Bmal1 KO mice. N = 2, 3, 3, 2, 2, 2 for CT0, CT4, CT8, CT12, CT16, CT20, respectively.
Discussion
In this study, we focused on the mouse olfactory system and showed that BMAL1 in the pyramidal neurons of the PC intrinsically regulates the circadian rhythm of the odor-evoked neural activity. This notion is supported by the following lines of evidence. Firstly, the circadian rhythm of the odor-evoked neural activity was maintained in Vgat-Bmal1 KO mice in which BMAL1 expression was largely diminished in GABAergic neurons including over 90% of SCN neurons (Fig. 2), suggesting that the clock genes in GABAergic neurons in the SCN are dispensable for the odor-evoked neural activity in the PC. Secondly, Bmal1 KO exclusively in the CaMKIIα-positive pyramidal neurons in the PC largely abolished the circadian rhythm of the odor-evoked neural activity in the PC (Figs. 3, 4). Lastly, isolated brain slices of the PC maintained the circadian rhythm of Per2 expression at least for 3 days and this rhythm damped out upon TTX application (Fig. 5), suggesting that action potential-based neural activity likely contributes to intrinsic control of the clock gene expressions in the PC.
The present study showed an indispensable role of BMAL1 expression in the PC for the circadian rhythm of the odor-evoked activity in the PC. However, we could not clearly test the role of BMAL1 expression in the olfactory bulb because Pcdh21-Bmal1 KO removed BMAL1 from only ~30% of mitral/tufted cells (Fig. 3). Notably, a previous report suggested that the olfactory bulb functions as an independent circadian system regulating circadian rhythms of the odor-evoked neural activity, as surgical removal of the olfactory bulb abolished the circadian rhythms of the odor-evoked response in the PC1. Since the PC receives the vast majority of afferent fibers from the olfactory bulb37,38, it would be important in the future to understand how the olfactory bulb and the PC interact with each other to shape the circadian rhythms of the odor-evoked activity in the PC. Given that the mitral/tufted cells in the olfactory bulb project to the PC37,38, one potential scenario is that neural activity might contribute to synchronization between the olfactory bulb project and the PC.
The present study identified seven genes whose mRNA expressions showed circadian rhythms under the control of BMAL1 (Fig. 6). Of the seven genes identified, Avpr1a encodes a G-protein coupled arginine vasopressin receptor, and it increases intracellular Ca2+ levels to activate ERK/CREB39. Chrnb2 encodes a subunit of a nicotinic acetylcholine receptor that promotes cation influx to depolarize the membrane potential40. Clcn4 encodes a voltage-gated chloride channel. Hence the circadian rhythms of expressions of these genes may directly shape the circadian rhythm of the odor-evoked activity in the PC. Unexpectedly, other genes are associated with presynaptic, rather than postsynaptic, functions. Cacna2d3 encodes a subunit of the voltage-gated calcium channel Cav2.241 which functions at presynaptic terminals42,43,44. Cav2.2 channels are known to interact with the active zone proteins including SNAP2545,46,47,48. SynI encodes Synapsin I which contributes to the size control of the synaptic vesicle pool within the presynapses49. One possible scenario, therefore, is that BMAL1 in the PC regulates the expression of Cav2.2, SNAP25, and Synapsin I to regulate the circadian rhythm of the odor-evoked output from the PC. Consistently, we found that action potential-based neural activity is required for in intrinsic circadian oscillation of clock gene Per2 in the isolated SCN as well as the PC (Fig. 5d). However, we have to take into account that our experiments were limited to c-Fos induction, which does not directly measure neural activity. We would need to conduct other experiments such as calcium imaging and electrophysiological analyses to measure neural activity in a less indirect approach. For example, further investigations using electrophysiology and calcium imaging will be needed to understand how the local circadian clock indeed impacts on neural activity and synapse transmission in the PC.
Overall, we show in our current analyses that the molecular clockwork in the pyramidal neurons of the PC intrinsically regulates the circadian rhythm of the odor-evoked neural activity. We further show candidate genes that might contribute to the circadian rhythm of the odor-evoked neural activity under the control of the molecular clock. Since the molecular clock appears to target distinct genes in different tissues50,51, it is highly possible that additional genes might be transcriptionally regulated by BMAL1 and contribute to the circadian rhythms of the olfactory response in the PC. It will be thus important to understand the comprehensive sets of BMAL1-targeting genes in the PC and how the gene products could contribute to the intrinsic control of the circadian rhythm of neural activity.
Methods
Animals
All animal experiments had received approval from the Animal Care Committee at the University of Tokyo. C57BL/6 J wild-type mice were purchased from Japan SLC, Inc. Bmal1-floxed mice, Emx1-Cre mice, Vgat-Cre mice, and PER2::LUC mice were provided as a kind gift from Dr. Fukada, Omp-Cre mice, Pcdh21-CreER mice, and tdTomato-flox(Ai9) mice were purchased from The Jackson Laboratory. Animals were group-housed, up to four animals per cage, in a temperature (23 ± 1˚C) and humidity (50 ± 20%) controlled environment, in a 12 h light, 12 h dark cycle with ad libitum access to food and water, light on at 7:00 AM. Male mice were used in all experiments.
Odor presentation paradigm
Adult male mice (2–3 months of age) were isolated to a new cage (Japan CLEA, CL-0113-1) 1 week prior to odor presentation with ad libitum access to food and water. Two days prior to odor presentation, mice were transferred to a new cage and underwent a constant dark condition for >24 . On the day of the experiment, 100 µL of odorant (cedar oil, limonene(-)) was applied to a cotton swab and placed on the cage lid for 5 min. 50 min after odor presentation, mice were perfused for further histological experiments.
Immunohistochemistry
Mice were anesthetized with Isoflurane (Pfizer), then transcardially perfused with ice-cold 10 mL 0.1 M phosphate buffer (pH 7.4), followed by ice-cold 25 mL fixative (4% paraformaldehyde in 0.1 M phosphate buffer). Whole brains were dissected, immersed in ice-cold fixative for 6 h. Next, brains were immersed in 0.1 M PBS (pH 7.4) containing 20% sucrose for 6 h, then transferred to 0.1 M PBS (pH 7.4) containing 30% sucrose for 6 h. Forty µm-thick coronal sections were cut with a sliding microtome (Yamato Kohki). Cut sections were stored in an Anti-freezing buffer (25% glycerol (Nakalai Tesque), 30% ethylene glycol (Nakalai Tesque), 0.1 M PBS) at −30 °C.
Brain sections were incubated in blocking buffer (3% Normal Donkey Serum (Jackson Immuno Research Laboratories), 0.05% TritonX-100, PBS) for 2 h at room temperature. Next, sections were incubated in primary antibody (Anti-c-Fos Rabbit pAb, Calbiochem, 1:20,000 dilution in blocking buffer; Anti-BMAL1 Rabbit Polyclonal, Novus Biologicals, 1:1000 dilution in blocking buffer; Anti-Tbr1 Rabbit Polyclonal, Abcam, 1:1000 dilution in blocking buffer; anti-Tbx21 guinea pig, a gift from Y Yoshihara, 1:10,000 dilution in blocking buffer) for 3 days at 4 °C. After primary antibody incubation, sections were washed with PBST for 10 min, five times at room temperature. Next, sections were incubated with a second antibody (Alexa 488-conjugated donkey anti-rabbit, Invitrogen, 1:500 in blocking buffer). Afterward, sections were washed with PBST for 10 min, five times at room temperature. Finally, for nuclear staining, sections were incubated with Nissl (Invitrogen, 1:200 in PBS) or DAPI (Sigma-Aldrich, 1:2000 in PBS) for 1 h at room temperature. Sections were mounted with Prolong Diamond (Thermo Fisher Scientific) antifade.
Microscopy and cell counting
For whole-brain 3D Mapping, brain sections were imaged by KEYENCE.
For other imaging, brain slices were imaged by confocal microscopy (Leica, SP8).
Brain sections immunostained with c-Fos antibody were quantified with the following procedures using ImageJ. First, the background signal intensity was calculated by averaging ten areas of the brain section without any cells. The background intensity was subtracted and despeckled. The signal threshold was set to 40, and the signals >30 µm were counted as c-Fos positive cells. For other imaging, brain slices were imaged by confocal microscopy (Leica, SP8).
Locomotor activity analysis
Mice were housed individually, and their spontaneous locomotor activities were recorded using an area sensor (Elekit) with an infrared detector. Locomotor activity was collected every minute and analyzed by ClockLab software (Actimetrics).
Adeno-associated virus production
AAV293 cells (Agilent Technologies) were cultured in 10 cm culture dish (1.25~1.5 × 106 cells/dish) with 10 mL culture medium (DMEM (Sigma-Aldrich, D5796-500ML), 10% FBS (GE Life Sciences, HyClone SH30396.03), Penicillin Streptomycin (gibco, 15140-122), GlutaMax (gibco, 35050-061)). After 2~3 days, cells were passaged into T-150 flasks (2.5~3.0 × 106 cells/flask) with 35 mL of culture medium. After 48~72 h, when cells reached 60~70% confluency, 25 mL of culture medium was removed, and 9 mL of fresh culture medium was added. Subsequently, plasmids were transfected via calcium phosphate co-precipitation. Briefly, three plasmids—pAAV-DJ, pHelper, plasmid including genes of interest—were added to 6 mL of 0.3 M CaCl2. Next, 6 mL of 2x HBS (280 mM NaCl, 1.5 mM Na2HPO4, 50 mM Hepes) was added with thorough mixing. 3 mL of the mixed compound was applied to each flask and further cultured for 6~12 h. Next, the culture medium was replaced with 35 mL of medium containing trichostatin A (DMEM, 2% FBS, Penicillin, Glutamax, 100 nM trichostatin A (Wako)), and further cultured for 48~96 h. Subsequently, the medium containing scraped cells was collected and underwent freeze-thaw three times. Finally, 1/10,000 volume of 250 U/µL TurboNuclease (Accelagen) was added and incubated for 30 min at 37 °C. The AAVs were purified with ViraPur Kit (Virapur).
Adeno-associated virus injection
We performed AAVs injection as previously described52,53. In brief, mice 6–8 weeks of age were stereotaxically injected. Animals were anesthetized with isoflurane (1–2%) and placed in a stereotaxic frame. The adeno-associated virus AAVDJ-CaMKII-iCre-T2A-mCherry (3.55~7.87 ×1011 vg/mL) or AAVDJ-CAMKII-mCherry (7.83 × 1012 vg/mL) were injected to the olfactory bulb or the PC bilaterally. For AAV transduction to the granule cell layer of the olfactory bulb, we injected 300 nL of AAV to each olfactory bulb bilaterally. For AAV transduction to the PC, we injected AAV to three coordinates to either side of the brain, with a total of six injection coordinates bilaterally with 300 nL injection volume per injection coordinate. For injection, we used a pulled glass capillary (Drummond) and injected through nanoliter pressure injection (Nanoject3, Drummond). Stereotactic injection coordinates to target the olfactory bulb or PC were obtained from the Paxinos and Franklin atlas (for olfactory bulb AP:4.2 mm ML:1.0 mm DV:1.0 mm from brain surface; for PC AP:1.8, 0.5, −0.5 mm ML:2.7, 3.5, 3.9 mm, DV:3.8, 3.8, 4.2 mm from brain surface). Animals were allowed to recover for at least 4 weeks before being used in experiments.
Luciferase assay
PER2::LUC mice 4 weeks of age were used for luciferase assay. Mice were sacrificed, and dissected brains were immediately cooled in ice-cold Hank’s Balanced Sodium Saline. The piriform cortex and the SCN were further coronally dissected in 150μm-thickness with vibratome. Sectioned brain slices were placed on Milli Cell immersed in culture medium (5% MEM, 18% HBSS, 15 mM NaHCO3, 2 mM HEPES, 10 mM D-glucose, 1 mM L-glutamine, 175 mg/L Ascorbic Acid, 1 mg/mL insulin in 0.01 M HCl, 25% Horse Serum; pH adjusted to 7.0~7.4) with Luciferin, and incubated at 37 °C, 5% CO2. Luminescence was detected by Kronos Dio (Atto) with 1-min exposure, 10-min intervals. For tetrodotoxin application, Tetrodotoxin (Fujifilm) was applied to the culture medium with a final concentration of 20 µM.
In silico data analysis
From the list of genes reported to show circadian fluctuations of mRNA levels in the SCN36, we manually searched the function of each gene via UniProt (https://www.uniprot.org/) and identified genes that were documented to be involved in neuronal processes.
Real-time quantitative PCR
For RNA extraction, 8–10 weeks-old mice underwent a constant dark condition for 24–48 h. Brains were dissected, washed with HBSS, and immediately frozen with liquid nitrogen. RNA was extracted with RNeasy kit (Promega), and the cDNA library was created by first-strand reverse transcription with ReverTraAce Kit (Toyobo). For real-time quantitative PCR, cDNA samples were mixed with THUNDERBIRD SYBR qPCR Mix (Toyobo) and primers and underwent two-step PCR (95 ˚C 15 s→ 60 ˚C 60 s, 40 cycles).
Primer sequences for real-time quantitative PCR are explained in Supplementary Table 1.
Statistics and reproducibility
The statistical significance between values were determined by one-way ANOVA, one-way ANOVA with Bonferroni correction, Wilcoxon rank-sum test, Wilcoxon rank-sum test with Bonferroni correction. All experiments were independently repeated two or more times.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
References
Granados-Fuentes, D., Tseng, A. & Herzog, E. D. A circadian clock in the olfactory bulb controls olfactory responsivity. J. Neurosci. 26, 12219–12225 (2006).
Nakano, J. J., Shimizu, K., Shimba, S. & Fukada, Y. SCOP/PHLPP1β in the basolateral amygdala regulates circadian expression of mouse anxiety-like behavior. Sci. Rep. 6, 33500 (2016).
Park, J. et al. Identification of a circadian clock in the inferior colliculus and its dysregulation by noise exposure. J. Neurosci. 36, 5509–5519 (2016).
Shimizu, K. et al. SCOP/PHLPP1β mediates circadian regulation of long-term recognition memory. Nat. Commun. 7, 12926 (2016).
Honma, S. The mammalian circadian system: a hierarchical multi-oscillator structure for generating circadian rhythm. J. Physiol. Sci. 68, 207–219 (2018).
Ko, C. H. & Takahashi, J. S. Molecular components of the mammalian circadian clock. Hum. Mol. Genet. 15, R271–R277 (2006).
Takahashi, J. S. Transcriptional architecture of the mammalian circadian clock. Nat. Rev. Genet. 18, 164–179 (2017).
Bunger, M. K. et al. Mop3 is an essential component of the master circadian pacemaker in mammals. Cell 103, 1009–1017 (2000).
Zheng, B. et al. Nonredundant roles of the mPer1 and mPer2 genes in the mammalian circadian clock. Cell 105, 683–694 (2001).
Abe, M. et al. Circadian rhythms in isolated brain regions. J. Neurosci. 22, 350–356 (2002).
Dibner, C., Schibler, U. & Albrecht, U. The mammalian circadian timing system: organization and coordination of central and peripheral clocks. Annu. Rev. Physiol. 72, 517–549 (2010).
Reppert, S. M. & Weaver, D. R. Coordination of circadian timing in mammals. Nature 418, 935–941 (2002).
Hastings, M. H., Maywood, E. S. & Brancaccio, M. Generation of circadian rhythms in the suprachiasmatic nucleus. Nat. Rev. Neurosci. 19, 453–469 (2018).
Reppert, S. M. & Weaver, D. R. Molecular analysis of mammalian circadian rhythms. Annu. Rev. Physiol. 63, 647–676 (2001).
Granados-Fuentes, D., Saxena, M. T., Prolo, L. M., Aton, S. J. & Herzog, E. D. Olfactory bulb neurons express functional, entrainable circadian rhythms. Eur. J. Neurosci. 19, 898–906 (2004).
Ono, D., Honma, S. & Honma, K. Circadian PER2::LUC rhythms in the olfactory bulb of freely moving mice depend on the suprachiasmatic nucleus but not on behaviour rhythms. Eur. J. Neurosci. 42, 3128–3137 (2015).
Leinwand, S. G. & Chalasani, S. H. Olfactory networks: from sensation to perception. Curr. Opin. Genet. Dev. 21, 806–811 (2011).
Mombaerts, P. Axonal wiring in the mouse olfactory system. Annu. Rev. Cell Dev. Biol. 22, 713–737 (2006).
Herz, R. S. et al. The influence of circadian timing on olfactory sensitivity. Chem. Senses 43, 45–51 (2018).
Miller, J. K. et al. Vasoactive intestinal polypeptide mediates circadian rhythms in mammalian olfactory bulb and olfaction. J. Neurosci. 34, 6040–6046 (2014).
Krukoff, T. L. c-fos Expression as a Marker of Functional Activity in the Brain. in Cell Neurobiology Techniques (eds. Boulton, A. A., Baker, G. B. & Bateson, A. N.) 213–230 (Humana Press, 1999).
Amir, S., Cain, S., Sullivan, J., Robinson, B. & Stewart, J. In rats, odor-induced Fos in the olfactory pathways depends on the phase of the circadian clock. Neurosci. Lett. 272, 175–178 (1999).
Song, J. H. et al. Precise mapping of single neurons by calibrated 3D reconstruction of brain slices reveals topographic projection in mouse visual cortex. Cell Rep. 31, 107682 (2020).
Ono, D., Honma, K., Yanagawa, Y., Yamanaka, A. & Honma, S. GABA in the suprachiasmatic nucleus refines circadian output rhythms in mice. Commun. Biol. 2, 1–12 (2019).
Vong, L. et al. Leptin action on GABAergic neurons prevents obesity and reduces inhibitory tone to POMC neurons. Neuron 71, 142–154 (2011).
Shimba, S. et al. Deficient of a clock gene, brain and muscle Arnt-like protein-1 (BMAL1), induces dyslipidemia and ectopic fat formation. PLoS ONE 6, e25231 (2011).
Li, J., Ishii, T., Feinstein, P. & Mombaerts, P. Odorant receptor gene choice is reset by nuclear transfer from mouse olfactory sensory neurons. Nature 428, 393–399 (2004).
Yonekura, J. & Yokoi, M. Conditional genetic labeling of mitral cells of the mouse accessory olfactory bulb to visualize the organization of their apical dendritic tufts. Mol. Cell. Neurosci. 37, 708–718 (2008).
Iwasato, T. et al. Cortex-restricted disruption of NMDAR1 impairs neuronal patterns in the barrel cortex. Nature 406, 726–731 (2000).
Nagayama, S., Homma, R. & Imamura, F. Neuronal organization of olfactory bulb circuits. Front. Neural Circuits 8, 98 (2014).
Takahashi, H., Yoshihara, S. & Tsuboi, A. The functional role of olfactory bulb granule cell subtypes derived from embryonic and postnatal neurogenesis. Front. Mol. Neurosci. 11, 229 (2018).
Benson, D. L., Isackson, P. J., Gall, C. M. & Jones, E. G. Contrasting patterns in the localization of glutamic acid decarboxylase and Ca2+ /calmodulin protein kinase gene expression in the rat centrat nervous system. Neuroscience 46, 825–849 (1992).
Liu, N. Regional distribution of protein kinases in normal and odor-deprived mouse olfactory bulbs. Chem. Senses 25, 401–406 (2000).
Zou, D.-J., Greer, C. A. & Firestein, S. Expression pattern of αCaMKII in the mouse main olfactory bulb. J. Comp. Neurol. 443, 226–236 (2002).
Yoo, S.-H. et al. PERIOD2::LUCIFERASE real-time reporting of circadian dynamics reveals persistent circadian oscillations in mouse peripheral tissues. Proc. Natl Acad. Sci. 101, 5339–5346 (2004).
Panda, S. et al. Coordinated transcription of key pathways in the mouse by the circadian clock. Cell 109, 307–320 (2002).
Brunjes, P. C., Illig, K. R. & Meyer, E. A. A field guide to the anterior olfactory nucleus (cortex). Brain Res. Rev. 50, 305–335 (2005).
Wilson, D. A. & Sullivan, R. M. Cortical processing of odor objects. Neuron 72, 506–519 (2011).
Zhao, N. et al. Arginine vasopressin receptor 1a is a therapeutic target for castration-resistant prostate cancer. Sci. Transl. Med. 11, eaaw4636 (2019).
Hurst, R., Rollema, H. & Bertrand, D. Nicotinic acetylcholine receptors: from basic science to therapeutics. Pharmacol. Ther. 137, 22–54 (2013).
Neely, G. G. et al. A genome-wide Drosophila screen for heat nociception identifies α2δ3 as an evolutionarily conserved pain gene. Cell 143, 628–638 (2010).
Hoppa, M. B., Lana, B., Margas, W., Dolphin, A. C. & Ryan, T. A. α2δ expression sets presynaptic calcium channel abundance and release probability. Nature 486, 122–125 (2012).
Scott, M. B. & Kammermeier, P. J. CaV2 channel subtype expression in rat sympathetic neurons is selectively regulated by α2δ subunits. Channels 11, 555–573 (2017).
Stephani, F. et al. Deletion of the Ca2+ channel subunit α2δ3 differentially affects Cav2.1 and Cav2.2 currents in cultured spiral ganglion neurons before and after the onset of hearing. Front. Cell. Neurosci. 13, 278 (2019).
Kim, D. K. & Catterall, W. A. Ca2+-dependent and—independent interactions of the isoforms of the α1A subunit of brain Ca2+ channels with presynaptic SNARE proteins. Proc. Natl Acad. Sci. USA 94, 14782–14786 (1997).
Rettig, J. et al. Isoform-specific interaction of the alpha1A subunits of brain Ca2+ channels with the presynaptic proteins syntaxin and SNAP-25. Proc. Natl Acad. Sci. USA 93, 7363–7368 (1996).
Yokoyama, C. T. et al. Mechanism of SNARE protein binding and regulation of Cav2 channels by phosphorylation of the synaptic protein interaction site. Mol. Cell. Neurosci. 28, 1–17 (2005).
Yokoyama, C. T., Sheng, Z.-H. & Catterall, W. A. Phosphorylation of the synaptic protein interaction site on N-type calcium channels inhibits interactions with SNARE proteins. J. Neurosci. 17, 6929–6938 (1997).
Gitler, D. & Augustine, G. J. in Encyclopedia of Neuroscience (ed. Squire, L. R.) 709–717 (Academic Press, 2009).
Noya, S. B. et al. The forebrain synaptic transcriptome is organized by clocks but its proteome is driven by sleep. Science 366, eaav2642 (2019).
Pembroke, W. G., Babbs, A., Davies, K. E., Ponting, C. P. & Oliver, P. L. Temporal transcriptomics suggest that twin-peaking genes reset the clock. eLife 4, e10518 (2015).
Togashi, K. et al. Adeno associated virus-mediated single cell labeling of mitral cells in the mouse olfactory bulb: Insights into the developmental dynamics of dendrite remodeling. Front. Cell. Neurosci. 14, 572256 (2020).
Nakahama, R. et al. The tyrosine capsid mutations on retrograde adeno-associated virus accelerates gene transduction efficiency. Mol. Brain 15, 70 (2022).
Acknowledgements
We thank T. Imai for providing Omp-Cre transgenic mice, Atsu Aiba (Graduate School of Medicine, the University of Tokyo) for Emx1-Cre transgenic mice, and Shigeki Shimba (School of Pharmacology, Nihon University) for Bmal1-floxed mice; Y Yoshihara for Tbx21 antibody; S Sato, M Maetani, M Sakurai, A Nishino for technical assistance; the members of Emoto Lab for critical comments and discussion. This work is supported by the Graduate Program for Leaders in Life Innovation (GPLLI), World-Leading Innovative Graduate Study Program for Life Science and Technology (WINGS-LST), and Grant-in-Aid for JSPS Fellows to S.T.; MEXT Grants-in-Aid for Scientific Research on Innovative Areas “Dynamic regulation of brain function by Scrap and Build system” (KAKENHI 16H06456), JSPS (KAKENHI 16H02504), WPI-IRCN, AMED-CREST (JP21gm1310010), JST-CREST, Toray Foundation, Naito Foundation, Takeda Science Foundation, and Uehara Memorial Foundation to K.E.; and a Grant-in-Aid for Specially Promoted Research JSPS (KAKENHI 17H06096) to Y.F.
Author information
Authors and Affiliations
Contributions
S.T. performed all the experiments and data analyses with help of K.S. S.T. and K.E. are responsible for study conception, experimental design, and data interpretation. S.T. and K.E. wrote the manuscript with revisions and proofreading by K.S. and Y.F.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Communications Biology thanks Hugh Piggins, Christopher S. Colwell and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Primary Handling Editors: Daniel Bendor and George Inglis.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Takeuchi, S., Shimizu, K., Fukada, Y. et al. The circadian clock in the piriform cortex intrinsically tunes daily changes of odor-evoked neural activity. Commun Biol 6, 332 (2023). https://doi.org/10.1038/s42003-023-04691-8
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s42003-023-04691-8
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.