Abstract
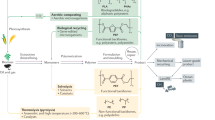
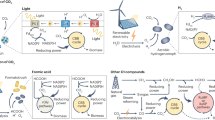
The increase in population-related and environmental issues has emphasized the need for more efficient and sustainable production strategies for foods and chemicals. Carbohydrates are macronutrients sourced from crops and undergone transformation into various products ranging from foods to chemicals. Continuous efforts have led to the identification of a promising hybrid system that couples the electrochemical reduction of CO2 to intermediates containing one to three carbons (C1–3) with the transformation of the intermediates using engineered microorganisms into valuable products. Here we use yeast to transform C1–3 substrates into glucose and structurally tailored glucose derivatives, such as the sugar alcohol myo-inositol, the amino monosaccharide glucosamine, the disaccharide sucrose and the polysaccharide starch. By metabolic rewiring and mitigation of glucose repression, the titre of glucose and sucrose reached dozens of grams per litre. These results provide directions for microbial sugar-derived foods and chemicals production from renewable reduced CO2-based feedstocks.

This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
Source data are provided with this paper.
References
Pastor, A. et al. The global nexus of food–trade–water sustaining environmental flows by 2050. Nat. Sustain. 2, 499–507 (2019).
Eibl, R. et al. Cellular agriculture: opportunities and challenges. Annu. Rev. Food Sci. 12, 51–73 (2021).
Cestellos-Blanco, S. et al. Molecular insights and future frontiers in cell photosensitization for solar-driven CO2 conversion. iScience 24, 102952 (2021).
Sellers, P. et al. Comparison of radiative and physiological effects of doubled atmospheric CO2 on climate. Science 271, 1402–1406 (1996).
Liu, Y. et al. Biofuels for a sustainable future. Cell 184, 1636–1647 (2021).
Roy, S., Cherevotan, A. & Peter, S. C. Thermochemical CO2 hydrogenation to single carbon products: scientific and technological challenges. ACS Energy Lett. 3, 1938–1966 (2018).
Ross, M. B. et al. Designing materials for electrochemical carbon dioxide recycling. Nat. Catal. 2, 648–658 (2019).
Jia, S. et al. Electrochemical transformation of CO2 to value-added chemicals and fuels. CCS Chem. 4, 3213–3229 (2022).
Grim, R. G. et al. Transforming the carbon economy: challenges and opportunities in the convergence of low-cost electricity and reductive CO2 utilization. Energy Environ. Sci. 13, 472–494 (2020).
Pachaiappan, R. et al. A review of recent progress on photocatalytic carbon dioxide reduction into sustainable energy products using carbon nitride. Chem. Eng. Res. Des. 177, 304–320 (2022).
Bierbaumer, S. et al. Enzymatic conversion of CO2: from natural to artificial utilization. Chem. Rev. 123, 5702–5754 (2023).
Yuan, L. et al. Coupling strategy for CO2 valorization integrated with organic synthesis by heterogeneous photocatalysis. Angew. Chem. Int. Ed. 60, 21150–21172 (2021).
Song, X. et al. Towards sustainable CO2 electrochemical transformation via coupling design strategy. Mater. Today Sustain. 19, 100179 (2022).
Davidson, E. A. Carbohydrate. Encyclopedia Britannica https://www.britannica.com/science/carbohydrate (2023).
Farrán, A. et al. Green solvents in carbohydrate chemistry: from raw materials to fine chemicals. Chem. Rev. 115, 6811–6853 (2015).
Martíneza, J. B. G. et al. Chemical synthesis of food from CO2 for space missions and food resilience. J. CO2 Util. 53, 101726 (2021).
Harbaugh, J. NASA awards $750,000 in competition to convert CO2 into sugar. https://www.nasa.gov/directorates/spacetech/centennial_challenges/75K-awarded-in-competition-to-convert-carbon-dioxide-into-sugar.html (2021).
Cestellos-Blanco, S. et al. Toward abiotic sugar synthesis from CO2 electrolysis. Joule 6, 2304–2323 (2022).
Cai, T. et al. Cell-free chemoenzymatic starch synthesis from carbon dioxide. Science 373, 1523–1527 (2021).
Zheng, T. et al. Upcycling CO2 into energy-rich long-chain compounds via electrochemical and metabolic engineering. Nat. Catal. 5, 388–396 (2022).
Parapouli, M. et al. Saccharomyces cerevisiae and its industrial applications. AIMS Microbiol 6, 1–31 (2020).
Spohner, S. C. et al. Expression of enzymes for the usage in food and feed industry with Pichia pastoris. J. Biotechnol. 202, 118–134 (2015).
Kayikci, Ö. & Nielsen, J. Glucose repression in Saccharomyces cerevisiae. FEMS Yeast Res. 15, fov068 (2015).
Zhang, B. & Sun, L. Artificial photosynthesis: opportunities and challenges of molecular catalysts. Chem. Soc. Rev. 48, 2216–2264 (2019).
Peng, C. et al. Double sulfur vacancies by lithium tuning enhance CO2 electroreduction to n-propanol. Nat. Commun. 12, 1580 (2021).
Pronk, J. T. et al. Propionate metabolism in Saccharomyces cerevisiae: implications for the metabolon hypothesis. Microbiology 140, 717–722 (1994).
Carnegie, D. & Ramsay, J. A. Anaerobic ethylene glycol degradation by microorganisms in poplar and willow rhizospheres. Biodegradation 20, 551–558 (2009).
Pandit, A. V., Harrison, E. & Mahadevan, R. Engineering Escherichia coli for the utilization of ethylene glycol. Microb. Cell. Fact. 20, 22 (2021).
Snoeckx, R. & Bogaerts, A. Plasma technology—a novel solution for CO2 conversion? Chem. Soc. Rev. 46, 5805–5863 (2017).
Lum, Y. & Ager, J. W. Evidence for product-specific active sites on oxide-derived Cu catalysts for electrochemical CO2 reduction. Nat. Catal. 2, 86–93 (2019).
Lin, Y. et al. Tunable CO2 electroreduction to ethanol and ethylene with controllable interfacial wettability. Nat. Commun. 14, 3575 (2023).
Brown, M. E., Mukhopadhyay, A. & Keasling, J. D. Engineering bacteria to catabolize the carbonaceous component of sarin: teaching E. coli to eat isopropanol. ACS Synth. Biol. 5, 1485–1496 (2016).
Hausinger, R. P. New insights into acetone metabolism. J. Bacteriol. 189, 671–673 (2007).
Pfeiffer, M., Wildberger, P. & Nidetzky, B. Yihx-encoded haloacid dehalogenase-like phosphatase HAD4 from Escherichia coli is a specific α-d-glucose 1-phosphate hydrolase useful for substrate-selective sugar phosphate transformations. J. Mol. Catal. B 110, 39–46 (2014).
Somoza-Tornos, A. et al. Process modeling, techno-economic assessment, and life cycle assessment of the electrochemical reduction of CO2: a review. iScience 24, 102813 (2021).
Jouny, M. et al. General techno-economic analysis of CO2 electrolysis systems. Ind. Eng. Chem. Res. 57, 2165–2177 (2018).
Li, F. et al. Cooperative CO2-to-ethanol conversion via enriched intermediates at molecule–metal catalyst interfaces. Nat. Catal. 3, 75–82 (2020).
Zhou, Y. et al. Dopant-induced electron localization drives CO2 reduction to C2 hydrocarbons. Nat. Chem. 10, 974–980 (2018).
Arán-Ais, R. M. et al. The role of in situ generated morphological motifs and Cu(I) species in C2+ product selectivity during CO2 pulsed electroreduction. Nat. Energy 5, 317–325 (2020).
Birdja, Y. Y. et al. Advances and challenges in understanding the electrocatalytic conversion of carbon dioxide to fuels. Nat. Energy 4, 732–745 (2019).
De Luna, P. et al. What would it take for renewably powered electrosynthesis to displace petrochemical processes? Science 364, eaav3506 (2019).
Gonzalez-Uarquin, F., Rodehutscord, M. & Huber, K. Myo-inositol: its metabolism and potential implications for poultry nutrition—a review. Poult. Sci. 99, 893–905 (2020).
Li, Y. et al. Production of myo-inositol: recent advance and prospective. Biotechnol. Appl. Biochem. 69, 1101–1111 (2022).
Tang, E. et al. Synergetic utilization of glucose and glycerol for efficient myo-inositol biosynthesis. Biotechnol. Bioeng. 117, 1247–1252 (2020).
Zhang, Q. et al. Metabolic engineering of Pichia pastoris for myo-inositol production by dynamic regulation of central metabolism. Microb. Cell. Fact. 21, 112 (2022).
Liu, L. et al. Microbial production of glucosamine and N-acetylglucosamine: advances and perspectives. Appl. Microbiol. Biotechnol. 97, 6149–6158 (2013).
Shintani, T. Food industrial production of monosaccharides using microbial, enzymatic, and chemical methods. Fermentation 5, 47 (2019).
Yin, W. et al. Metabolic engineering of E. coli for xylose production from glucose as the sole carbon source. ACS Synth. Biol. 10, 2266–2275 (2021).
Godinho, L. M. & de Sá-Nogueira, I. Characterization and regulation of a bacterial sugar phosphatase of the haloalkanoate dehalogenase superfamily, AraL, from Bacillus subtilis. FEBS J. 278, 2511–2524 (2011).
Marques, W. L. et al. Elimination of sucrose transport and hydrolysis in Saccharomyces cerevisiae: a platform strain for engineering sucrose metabolism. FEMS Yeast Res. 17, fox006 (2017).
Sakulsingharoj, C. et al. Engineering starch biosynthesis for increasing rice seed weight: the role of the cytoplasmic ADP-glucose pyrophosphorylase. Plant Sci. 167, 1323–1333 (2004).
Zhou, Y., Grof, C. P. L. & Patrick, J. W. Proof of concept for a novel functional screening system for plant sucrose effluxers. J. Microbiol. Methods 1, e5 (2014).
Jiang, T., Duan, Q., Zhu, J., Liu, H. & Yu, L. Starch-based biodegradable materials: challenges and opportunities. Ind. Eng. Chem. Res. 3, 8–18 (2020).
Keeling, P. L. & Myers, A. M. Biochemistry and genetics of starch synthesis. Annu Rev. Food Sci. Technol. 1, 271–303 (2010).
Pfister, B. et al. Recreating the synthesis of starch granules in yeast. eLife 5, e15552 (2016).
Liu, X. et al. Fusion of cellobiose phosphorylase and potato alpha-glucan phosphorylase facilitates substrate channeling for enzymatic conversion of cellobiose to starch. Prep. Biochem. Biotechnol. 52, 611–617 (2021).
You, C. et al. Enzymatic transformation of nonfood biomass to starch. Proc. Natl Acad. Sci. USA 110, 7182–7187 (2013).
Peng, B., Wood, R. J., Nielsen, L. K. & Vickers, C. E. An expanded heterologous GAL promoter collection for diauxie-inducible expression in Saccharomyces cerevisiae. ACS Synth. Biol. 7, 748–751 (2018).
Westfall, P. J. et al. Production of amorphadiene in yeast, and its conversion to dihydroartemisinic acid, precursor to the antimalarial agent artemisinin. Proc. Natl Acad. Sci. USA 110, 7182–7187 (2013).
Hers, H. G. & Hue, L. Gluconeogenesis and related aspects of glycolysis. Annu. Rev. Biochem. 52, 617–653 (1983).
Flores, C.-L. & Gancedo, C. Construction and characterization of a Saccharomyces cerevisiae strain able to grow on glucosamine as sole carbon and nitrogen source. Sci. Rep. 8, 16949 (2018).
Santangelo, G. M. Glucose signaling in Saccharomyces cerevisiae. Microbiol. Mol. Biol. Rev. 70, 253–282 (2006).
Leech, A., Nath, N., McCartney, R. R. & Schmidt, M. C. Isolation of mutations in the catalytic domain of the snf1 kinase that render its activity independent of the snf4 subunit. Eukaryot. Cell 2, 265–273 (2003).
Dombek, K. M., Kacherovsky, N. & Young, E. T. The Reg1-interacting proteins, Bmh1, Bmh2, Ssb1, and Ssb2, have roles in maintaining glucose repression in Saccharomyces cerevisiae. J. Biol. Chem. 279, 39165–39174 (2004).
Rubenstein, E. M. et al. Access denied: Snf1 activation loop phosphorylation is controlled by availability of the phosphorylated threonine 210 to the PP1 phosphatase. J. Biol. Chem. 283, 222–230 (2008).
Palomino, A., Herrero, P. & Moreno, F. Tpk3 and Snf1 protein kinases regulate Rgt1 association with Saccharomyces cerevisiae HXK2 promoter. Nucleic Acids Res. 34, 1427–1438 (2006).
Rolland, F., Winderickx, J. & Thevelein, J. M. Glucose-sensing and -signalling mechanisms in yeast. FEMS Yeast Res. 2, 183–201 (2002).
Nishimura, A. et al. The Cdc25/Ras/cAMP-dependent protein kinase A signaling pathway regulates proline utilization in wine yeast Saccharomyces cerevisiae under a wine fermentation model. Biosci. Biotechnol. Biochem. 86, 1318–1326 (2022).
Wang, Y. et al. Ras and Gpa2 mediate one branch of a redundant glucose signaling pathway in yeast. PLoS Biol. 2, e128 (2004).
Ma, P., Wera, S., Van Dijck, P. & Thevelein, J. M. The PDE1-encoded low-affinity phosphodiesterase in the yeast Saccharomyces cerevisiae has a specific function in controlling agonist-induced cAMP signaling. Mol. Cell. Biol. 10, 91–104 (1999).
Deng, M. D. et al. Metabolic engineering of Escherichia coli for industrial production of glucosamine and N-acetylglucosamine. Metab. Eng. 7, 201–214 (2005).
Yin, Z., Hatton, L. & Brown, A. J. Differential post-transcriptional regulation of yeast mRNAs in response to high and low glucose concentrations. Mol. Microbiol. 35, 553–565 (2000).
Wang, F. et al. Technologies and perspectives for achieving carbon neutrality. Innovation 2, 100180 (2021).
Molitor, B., Mishra, A. & Angenent, L. T. Power-to-protein: converting renewable electric power and carbon dioxide into single cell protein with a two-stage bioprocess. Energy Environ. Sci. 12, 3515–3521 (2019).
Bertels, L. K., Fernández Murillo, L. & Heinisch, J. J. The pentose phosphate pathway in yeasts-more than a poor cousin of glycolysis. Biomolecules 11, 725 (2021).
Wong, D. W. S. Food Enzymes: Structure and Mechanism Ch. 13 (Springer, 1995).
Sarthy, A. V. et al. Expression of the Escherichia coli xylose isomerase gene in Saccharomyces cerevisiae. Appl. Environ. 53, 1996–2000 (1987).
Gancedo, J. M. & Gancedo, C. Catabolite repression mutants of yeast. FEMS Microbiol. Rev. 1, 179–187 (1986).
Verduyn, C., Postma, E., Scheffers, W. A. & Van Dijken, J. P. Effect of benzoic acid on metabolic fluxes in yeasts: a continuous-culture study on the regulation of respiration and alcoholic fermentation. Yeast 8, 501–517 (1992).
Yu, T. et al. Metabolic engineering of Saccharomyces cerevisiae for production of very long chain fatty acid-derived chemicals. Nat. Commun. 8, 15587 (2017).
Mans, R. et al. CRISPR/Cas9: a molecular Swiss army knife for simultaneous introduction of multiple genetic modifications in Saccharomyces cerevisiae. FEMS Yeast Res. 15, fov004 (2015).
Gassler, T., Heistinger, L., Mattanovich, D., Gasser, B. & Prielhofer, R. CRISPR/Cas9-mediated homology-directed genome editing in Pichia pastoris. Methods Mol. Biol. 1923, 211–225 (2019).
Lin-Cereghino, J. et al. Condensed protocol for competent cell preparation and transformation of the methylotrophic yeast Pichia pastoris. BioTechniques 38, 44 (2005). 46, 48.
de Lima, P. B. A. et al. Novel homologous lactate transporter improves l-lactic acid production from glycerol in recombinant strains of Pichia pastoris. Microb. Cell Fact. 15, 158 (2016).
Du, W., Liang, F., Duan, Y., Tan, X. & Lu, X. Exploring the photosynthetic production capacity of sucrose by cyanobacteria. Metab. Eng. 19, 17–25 (2013).
Acknowledgements
This work was financially supported by the National Key Research and Development Program of China (grant no. 2020YFA0907800 to T.Y.), the National Natural Science Foundation of China (grant no. 32071416 to T.Y. and 22308369 to W.C.), the Shenzhen Institute of Synthetic Biology Scientific Research Program (grant no. JCHZ20200003 to T.Y.), Shenzhen Key Laboratory for the Intelligent Microbial Manufacturing of Medicines (grant no. ZDSYS20210623091810032 to T.Y.), Key-Area Research and Development Program of Guangdong Province (grant no. 2022B1111080005 to T.Y.), the Strategic Priority Research Program of the Chinese Academy of Sciences (grant no. XDB0480000 to T.Y.), the National Key Research and Development Program of China (grant no. 2021YFA0911000 to T.Y.), CAS Project for Young Scientists in Basic Research (YSBR-051 to J.Z.), the China Postdoctoral Science Foundation (grant no. 2020M682973 to S.G.) and Guangdong Basic and Applied Basic Research Foundation (grant no. 2020A1515110927 to S.G.). We acknowledge the related fundings supported by China Merchants Research Institute of Advanced Technology Company Limited and China BlueChemical Ltd. We acknowledge the Shenzhen Infrastructure for Synthetic Biology for instrument support and technical assistance with plasmid construction. We thank X. Zhang and L. Xia for critical discussion, and the SIAT Mass Spectrometry Infrastructure for assistance with metabolite analysis.
Author information
Authors and Affiliations
Contributions
T.Y. and J.D.K. conceived this study. H.T., L.W. and S.G. designed and performed most of the experiments, analysed the data and drafted the manuscript. W.M., X.W. and J.S. assisted with the experiments and products detection. W.C. assisted with data analysis and interpretation. M.W., Q.Z., X.L. and J.Z. contributed to the manuscript revision. T.Y., J.D.K. and M.H. revised the manuscript. All authors revised and approved the manuscript.
Corresponding authors
Ethics declarations
Competing interests
J.D.K. has a financial interest in Amyris, Lygos, Demetrix, Napigen, Maple Bio, Apertor Labs, Zero Acre Farms, Berkeley Yeast and Ansa Biotechnology. X.L. has a financial interest in Demetrix and Synceres. All other authors declare no competing interests.
Peer review
Peer review information
Nature Catalysis thanks Yong-Cheol Park, Rodrigo Ledesma-Amaro and Tianwei Tan for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Information
Supplementary Figs. 1–22, Tables 1-–5, note and references.
Source data
Source Data Fig. 2
Statistical source data.
Source Data Fig. 3
Statistical source data.
Source Data Fig. 4
Statistical source data.
Source Data Fig. 5
Statistical source data.
Source Data Fig. 6
Statistical source data.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Tang, H., Wu, L., Guo, S. et al. Metabolic engineering of yeast for the production of carbohydrate-derived foods and chemicals from C1–3 molecules. Nat Catal 7, 21–34 (2024). https://doi.org/10.1038/s41929-023-01063-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41929-023-01063-7