Abstract
The present study examined human N-myristoylated proteins that specifically localize to mitochondria among the 1,705 human genes listed in MitoProteome, a mitochondrial protein database. We herein employed a strategy utilizing cellular metabolic labeling with a bioorthogonal myristic acid analog in transfected COS-1 cells established in our previous studies. Four proteins, DMAC1, HCCS, NDUFB7, and PLGRKT, were identified as N-myristoylated proteins that specifically localize to mitochondria. Among these proteins, DMAC1 and NDUFB7 play critical roles in the assembly of complex I of the mitochondrial respiratory chain. DMAC1 functions as an assembly factor, and NDUFB7 is an accessory subunit of complex I. An analysis of the intracellular localization of non-myristoylatable G2A mutants revealed that protein N-myristoylation occurring on NDUFB7 was important for the mitochondrial localization of this protein. Furthermore, an analysis of the role of the CHCH domain in NDUFB7 using Cys to Ser mutants revealed that it was essential for the mitochondrial localization of NDUFB7. Therefore, the present results showed that NDUFB7, a vital component of human mitochondrial complex I, was N-myristoylated, and protein N-myrisotylation and the CHCH domain were both indispensable for the specific targeting and localization of NDUFB7 to mitochondria.
Similar content being viewed by others
Introduction
Protein N-myristoylation is an important fatty acylation of proteins in eukaryotes and viruses. In this modification, myristic acid is covalently bound to the α-amino group of the N-terminal Gly of proteins through an amido bond after the removal of the initiating Met by methionine aminopeptidase1,2,3,4. Therefore, protein N-myristoylation occurs cotranslationally on newly synthesized proteins. In addition to cotranslational N-myristoylation, posttranslational N-myristoylation was found to occur on many caspase-cleavage products on an internal Gly exposed by caspase cleavage during apoptosis5,6,7. N-myristoyltransferase, a member of the GCN5-related N-acetyltransferase superfamily of proteins8, catalyzes both cotranslational and posttranslational N-myristoylation. Many N-myristoylated proteins play essential roles in the regulation of a wide variety of signal transduction pathways in cells, and include guanine nucleotide-binding proteins, Ca2+-binding proteins, protein kinases and their substrates, phosphatases, E3-ubiquitin ligases, and apoptosis-related proteins. Besides these cellular signal transduction proteins, protein N-myristoylation is found on many disease-associated proteins9,10,11,12,13. In many cases, reversible membrane binding mediated by protein N-myristoylation regulates the functions of these proteins. Therefore, protein N-myristoylation is regarded as a lipid modification that mainly occurs on cytoplasmic soluble proteins; however, some N-myristoylated proteins that specifically localized to intracellular organelles were also found. We previously established a strategy to comprehensively identify human N-myristoylated proteins from human cDNA clones in human cDNA resources. In this strategy, N-myristoylated proteins were detected by metabolic labeling and mass spectrometric analyses of proteins obtained by an in vitro translation system14,15,16.
Using this strategy, we identified two mitochondrial membrane proteins, TOMM40 and SAMM50, as N-myristoylated membrane proteins17,18. Both of these proteins are β-barrel proteins that localize to the outer membrane of mitochondria. TOMM40 is a major component of the translocase of the mitochondrial outer membrane (TOM complex) that acts as a general import pore for most mitochondrial precursor proteins19,20. SAMM50 is a major component of the SAM complex required for the sorting and assembly of β-barrel proteins21. An analysis of the intracellular localization of non-myristoylatable G2A mutants revealed that protein N-myristoylation occurring on SAMM50, but not on TOMM40, plays a critical role in the mitochondrial targeting of this protein18. However, the function of protein N-myristoylation occurring on TOMM40 remains unclear. Based on these findings, protein N-myristoylation may mediate diverse roles in the intracellular localization and function of mitochondrial proteins. In the present study, we searched for human N-myristoylated proteins that specifically localize to mitochondria among the 1,705 human genes listed in MitoProteome, a mitochondrial protein database22. The results obtained revealed that four proteins, DMAC1, HCCS, NDUFB7, and PLGRKT, were N-myristoylated proteins that specifically localized to mitochondria. Among these proteins, DMAC1 and NDUFB7 were previously found to play critical roles in the assembly of complex I of the mitochondrial respiratory chain as an assembly factor and an accessory subunit, respectively23. An analysis of the intracellular localization of non-myristoylatable G2A mutants revealed that protein N-myristoylation occurring on NDUFB7 played a critical role in the mitochondrial localization of this protein. Furthermore, an analysis of the role of the CHCH domain (CHCHD) found in NDUFB7 using Cys to Ser mutants revealed that CHCHD was vital for the mitochondrial localization of NDUFB7. Therefore, the present results showed that NDUFB7, an essential component of human mitochondrial complex I, was N-myristoylated, and protein N-myristoylation and CHCHD were both indispensable for the specific targeting and localization of NDUFB7 to mitochondria.
Results
Identification of human N-myristoylated mitochondrial proteins from human genes listed in the MitoProteome protein database
We searched for human N-myristoylated mitochondrial proteins among the human genes listed in MitoProteome, a mitochondrial protein database, using a strategy established in our previous studies14,15,16,17,18. In this strategy, 131 genes with N-terminal Met-Gly motifs were extracted from the 1,705 human genes listed in the MitoProteome protein database22. After applying the N-terminal sequences of the proteins encoded by these genes to two protein N-myristoylation prediction programs, the MYR Predictor and Myristoylator24,25, 34 positively predicted genes were identified (Supplementary Table S1). Among these positively predicted genes, 12 candidate genes (BCAP31, CLN3, DMAC1/TMEM261, FLAD1, HCCS, LRRC10, MARC1/MTARC1, ME3, NDUFB7, NOL3, PLGRKT and TUSC3) for N-myristoylated mitochondrial proteins were selected and their susceptibility to protein N-myristoylation was evaluated using fusion proteins in which the 10 N-terminal amino acid residues were fused to a FLAG-tagged model protein, tGelsolin, as previously described14 (Fig. 1A,B). The information about the 13 genes analyzed in this study was summarized in Supplementary Table S2.
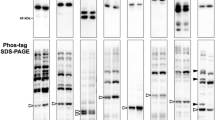
Identification of N-myristoylated mitochondrial proteins from human genes listed in the MitoProteome protein database. Twelve candidate genes (BCAP31, CLN3, DMAC1/TMEM261, FLAD1, HCCS, LRRC10, MARC1/MTARC1, ME3, NDUFB7, NOL3, PLGRKT and TUSC3) were selected as N-myristoylated mitochondrial proteins from the human genes listed in MitoProteome, a mitochondrial protein database, and their susceptibility to protein N-myristoylation was evaluated by in vivo metabolic labeling in transfected COS-1 cells using fusion proteins in which the 10 N-terminal amino acid residues were fused to the FLAG-tagged model protein, tGelsolin. (A) Strategy to identify N-myristoylated mitochondrial proteins from human genes listed in the MitoProteome protein database. (B) Schematic representation of the generation of X(N10)-tGelsolin-FLAG with 10 N-terminal amino acids of the candidate protein X at its N terminus. (C) Analysis of the protein N-myristoylation of X(N10)-tGelsolin-FLAGs by metabolic labeling in transfected COS-1 cells. cDNAs coding X(N10)-tGelsolin-FLAGs were transfected into COS-1 cells. The expression of proteins was evaluated by Western blotting using an anti-FLAG antibody (upper panels). Protein N-myristoylation was evaluated by metabolic labeling with a myristic acid analog followed by click chemistry, as described in the Methods (lower panels). Images obtained by Western blotting or metabolic labeling were cropped. Uncropped full-length raw image data are shown in Supplementary Figure S1.
In this experiment, N-myristoylatable tGelsolin-FLAG, its non-N-myristoylatable mutant (tGelsolin-G2A-FLAG), and the tGelsolin fusion protein of MIC19 (CHCHD3), a known N-myristoylated mitochondrial protein, were used as control proteins. To establish whether these fusion proteins are N-myristoylated, cellular metabolic labeling experiments on COS-1 cells using Alk-Myr, a bioorthogonal myristic acid analog, were performed16,17,18. In these experiments, after labeling cells transfected with cDNA coding C-terminally FLAG-tagged tGelsolin-fusion proteins with Alk-Myr, the N-myristoylated protein was detected with click chemistry. As shown in the upper panels of Fig. 1C, efficient protein expression in COS-1 cells was equally observed for all the fusion proteins tested, including 3 control proteins, according to a Western blotting analysis. Metabolic labeling experiments revealed the efficient incorporation of Alk-Myr into the fusion proteins of 10 candidate genes (BCAP31, CLN3, DMAC1/TMEM261, HCCS, LRRC10, MARC1/MTARC1, ME3, NDUFB7, NOL3, and PLGRKT), similar to two N-myristoylatable control proteins (tGelsolin-FLAG and MIC19(N10)-tGelsolin-FLAG); however, this incorporation was not observed in the 2 fusion proteins of 2 candidate genes (FLAD1 and TUSC3), as was the case for the non-N-myristoylatable tGelsolin-G2A mutant (Fig. 1C, lower panels).
To clarify whether the full-length protein was N-myristoylated, we selected 7 genes (CLN3, DMAC1/TMEM261, HCCS, MARC1/MTARC1, NDUFB7, NOL3, and PLGRKT) and their susceptibility to protein N-myristoylation was evaluated using the same strategy with the full-length cDNA coding for the C-terminally FLAG-tagged protein. The results obtained are shown in Fig. 2A.
DMAC1, HCCS, NDUFB7, and PLGRKT are N-myristoylated mitochondrial proteins. Seven proteins (CLN3, DMAC1, HCCS, MARC1, NDUFB7, NOL3, and PLGRKT) were selected from proteins in which protein N-myristoylation was detected in X(N10)-tGelsolin-FLAG, and their protein N-myristoylation and intracellular localization were evaluated using C-terminally FLAG-tagged full-length proteins. (A) Analysis of the protein N-myristoylation of CLN3, DMAC1, HCCS, MARC1, NDUFB7, NOL3, and PLGRKT expressed in transfected COS-1 cells by metabolic labeling. cDNAs coding C-terminally FLAG-tagged full-length proteins were transfected into COS-1 cells and cells were then labeled with a myristic acid analog. The expression of proteins was evaluated by Western blotting using an anti-FLAG antibody (left panel). Protein N-myristoylation was evaluated by metabolic labeling followed by click chemistry, as described in the Methods (right panel). Arrows indicate the position of the expressed proteins. The images obtained by Western blotting or metabolic labeling were cropped. Uncropped full-length raw image data are shown in Supplementary Figure S2. (B) Analysis of the intracellular localization of CLN3, DMAC1, HCCS, MARC1, NDUFB7, NOL3, and PLGRKT by an immunofluorescence microscopic analysis. The intracellular localization of seven proteins was assessed by an immunofluorescence analysis of COS-1 cells transfected with cDNA coding C-terminally FLAG-tagged full-length proteins using an anti-FLAG antibody. MIC19-FLAG was used as a control mitochondrial N-myristoylated protein. Hoechst and MitoTracker Red were used as organelle markers for the nucleus and mitochondria, respectively. Experiments were repeated 3 times and similar results were obtained. Representative data are shown. Abbreviation used: PC, phase contrast image. The results of the line profile analysis are shown. In the line profile analysis, corresponding line-scan graphs of the relative fluorescence intensities of green anti-FLAG fluorescence (green line) and red MitoTracker red fluorescence (red line) along the white line indicated in the merged images are shown.
In the left panel, all proteins, including three control proteins (tGelsolin-FLAG, tGelsolin-G2A-FLAG, and MIC19-FLAG), were expressed based on a Western blotting analysis using an anti-FLAG antibody. Metabolic labeling experiments revealed the incorporation of Alk-Myr in all 7 proteins encoded by the 7 tested genes, similar to the two N-myristoylatable control proteins (tGelsolin-FLAG and MIC19-FLAG), as shown in Fig. 2A right panel, indicating that the 7 proteins encoded by the 7 tested genes (CLN3, DMAC1/TMEM261, HCCS, MARC1/MTARC1, NDUFB7, NOL3, and PLGRKT) were N-myristoylated proteins. The characteristics of these proteins are summarized in Table 1.
DMAC1, HCCS, NDUFB7, and PLGRKT are N-myristoylated mitochondrial proteins
To assess the intracellular localization of the 7 proteins, an immunofluorescence analysis of COS-1 cells transfected with cDNA coding for the C-terminally FLAG-tagged protein was performed. In this analysis, MitoTracker Red and Hoechst 33342 were used as organelle markers for mitochondria and the nucleus, respectively. As shown in Fig. 2B, left panel and Supplementary Fig. S3, specific localization to mitochondria was observed with DMAC1, HCCS, NDUFB7, and PLGRKT, similar to MIC19, a known N-myristoylated mitochondrial protein. In contrast, selective localization to mitochondria was not observed with CLN3, MARC1, or NOL3. A line profile analysis of the images obtained further confirmed the specific localization of DMAC1, HCCS, NDUFB7, and PLGRKT to mitochondria (Fig. 2B, right panel).
Protein N-myristoylation plays a critical role in the mitochondrial targeting of NDUFB7
As shown in Fig. 3A, the high conservation of the N-myristoylation motif was observed among vertebrates by the interspecies alignment of the N-terminal sequences of DMAC1, HCCS, NDUFB7, and PLGRKT.
Protein N-myristoylation plays a critical role in the mitochondrial localization of NDUFB7. To clarify the role of protein N-myristoylation in the mitochondrial localization of DMAC1, HCCS, NDUFB7, and PLGRKT, non-myristoylatable G2A mutants were generated and their susceptibility to protein N-myristoylation and intracellular localization were evaluated. (A) Interspecies alignment of the N-terminal sequences of DMAC1, HCCS, NDUFB7, and PLGRKT. N-myristoylation motifs are shown in purple in N-terminal sequences. (B) Analysis of protein N-myristoylation of the WT and non-myristoylatable G2A mutants of DMAC1, HCCS, NDUFB7, and PLGRKT. cDNAs coding the WT and non-myristoylatable G2A mutants of four proteins were transfected into COS-1 cells. The expression of proteins was evaluated by Western blotting using an anti-FLAG antibody (upper panel). Protein N-myristoylation was evaluated by metabolic labeling with a myristic acid analog followed by click chemistry, as described in the Methods (lower panel). Arrows indicate the position of the expressed proteins. The images obtained by Western blotting or metabolic labeling were cropped. Uncropped full-length raw image data are shown in Supplementary Figure S4. (C) Analysis of the intracellular localization of the non-myristoylatable G2A mutant of DMAC1, HCCS, NDUFB7, and PLGRKT. The intracellular localization of the G2A mutant of the four proteins was assessed by an immunofluorescence analysis of COS-1 cells transfected with cDNA coding the C-terminally FLAG-tagged G2A mutant using an anti-FLAG antibody. MIC19-G2A-FLAG was used as a control non-myristoylatable G2A mutant. Hoechst and MitoTracker Red were used as organelle markers for the nucleus and mitochondria, respectively. Experiments were repeated 3 times and similar results were obtained. Representative data are shown. Abbreviation used: PC, phase contrast image. The results of the line profile analysis are shown. In the line profile analysis, corresponding line-scan graphs of the relative fluorescence intensities of green anti-FLAG fluorescence (green line) and red MitoTracker red fluorescence (red line) along the white line indicated in the merged images are shown.
To clarify the role of protein N-myristoylation in the mitochondrial localization of these four proteins, non-myristoylatable G2A mutants were generated and their susceptibility to protein N-myristoylation and intracellular localization were evaluated. As shown in Fig. 3B upper panel, the efficient expression of proteins was observed in the wild types and G2A mutants of four C-terminally FLAG-tagged proteins, except for the G2A mutant of DMAC1-FLAG. Metabolic labeling experiments revealed the efficient incorporation of Alk-Myr in wild-type proteins, whereas it was abolished in the G2A mutant (Fig. 3B, lower panel). When the intracellular localization of the G2A mutants of these four proteins was evaluated by an immunofluorescence analysis using an anti-FLAG antibody, marked differences were observed among the four proteins, as shown in Fig. 3C, left panel and Supplementary Fig. S5. Specific localization to mitochondria was observed for the G2A mutants of DMAC1, HCCS, and PLGRKT, similar to the wild-type proteins. The G2A mutant of NDUFB7 localized exclusively to the nucleus (Fig. 3C, left panel, Supplementary Fig. S5, and Supplementary Fig. S7A, upper panel). These results were further confirmed by the line profile analysis of the images obtained, as shown in Fig. 3C, right panel, and Supplementary Fig. S7A, lower panel.
However, in the case of MIC19, in which protein N-myristoylation was shown to be required for mitochondrial targeting, the G2A mutant localized to the cytoplasm (Fig. 3C, left panel, Supplementary Fig. S5, and Supplementary Fig. S8B, upper panel). This result was further confirmed by the line profile analysis of the images obtained, as shown in Fig. 3C, right panel. Therefore, protein N-myristoylation occurring on NDUFB7 played a critical role in mitochondrial localization, whereas that of DMAC1, HCCS, and PLGRKT did not affect the mitochondrial localization of these proteins.
Structure of NDUFB7
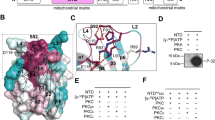
Similar to MIC19, NDUFB7 is a member of the CHCHD-containing protein family31. The structures of NDUFB7 and MIC19 are shown in Fig. 4.
Structure of NDUFB7. Schematic representation of the structures of MIC19 and NDUFB7. Interspecies alignments of the N-myristoylation motif, CHCH domain, and nuclear localization signal-like (NLSL) sequence of NDUFB7 are shown. N-myristoylation motifs were boxed. Four conserved Cys residues in the CHCH domain were indicated by the blue characters in the amino acid sequences. Conserved RKKRR sequences in the NLSL sequence were indicated by the red characters in the amino acid sequences.
In addition to the N-terminal N-myristoylation motif, NDUFB7 contains CHCHD at positions 56 to 98. CHCHD contains two Cx9C motifs at positions 59 to 69 and 80 to 90 that are predicted to form a helix-coil-helix structure, permitting the formation of intramolecular disulfide bonds32. Since the non-myristoylatable G2A mutant of NDUFB7 specifically localized to the nucleus, NDUFB7 was predicted to contain a nuclear localization signal-like sequence (NLSL). When the two public WWW server-based prediction programs for nuclear localization signals, cNLS Mapper (http://nls-mapper.iab.keio.ac.jp/cgi-bin/NLS_Mapper_form.cgi)33 and NucPred (http://www.sbc.su.se/~maccallr/nucpred)34, were applied to the amino acid sequence of NDUFB7, potential nuclear localization signals were predicted at positions 109 to 118 (LQRKKRREKK) and 111 to 115 (RKKRR), respectively. As shown in Fig. 4, the high conservation of the N-myristoylation motif, CHCHD, and NLSL was observed among vertebrates by the interspecies alignment of the protein sequence of NDUFB7.
Positive charge cluster localized to the C-terminal region functions as a nuclear localization signal of the non-myristoylatable G2A mutant of NDUFB7
Two public WWW server-based prediction programs for nuclear localization signals predicted a possible nuclear localization signal and the predicted nuclear localization signal overlapped at positions 111 to 115 (RKKRR). Therefore, we attempted to clarify whether the positive charge cluster at this position functioned as a nuclear localization signal. Two mutants in which amino acids at positions 111 to 115 of NDUFB7-FLAG and NDUFB7-G2A-FLAG were changed to Ala (NDUFB7-5KRtoA-FLAG and NDUFB7-G2A-5KRtoA-FLAG, respectively) were generated (Fig. 5A) and their susceptibility to protein N-myristoylation and their intracellular localization were evaluated.
Positive charge cluster localized in the C-terminal region functions as a nuclear localization signal of the non-myristoylatable G2A mutant of NDUFB7. To establish whether the positive charge cluster at positions 111 to 115 of NDUFB7 function as a nuclear localization signal, two mutants in which amino acids at this positions of NDUFB7-FLAG and NDUFB7-G2A-FLAG were changed to Ala (NDUFB7-5KRtoA-FLAG and NDUFB7-G2A-5KRtoA-FLAG, respectively) were generated (Fig. 5A), and their susceptibility to protein N-myristoylation and their intracellular localization were evaluated. The results obtained on protein expression and protein N-myristoylation from a Western blotting analysis and metabolic labeling experiments, respectively, are shown in Supplementary Figure S6. (A) Structure of NDUFB7 mutants lacking the C-terminal positive charge cluster. (B) Analysis of the intracellular localization of NDUFB7 mutants lacking the C-terminal positive charge cluster. The intracellular localization of NDUFB7-FLAG, NDUFB7-5KRtoA-FLAG, NDUFB7-G2A-FLAG, and NDUFB7-G2A-5KRtoA-FLAG was assessed by an immunofluorescence analysis of COS-1 cells transfected with cDNA coding these four proteins using an anti-FLAG antibody. Hoechst and MitoTracker Red were used as organelle markers for the nucleus and mitochondria, respectively. Experiments were repeated 3 times and similar results were obtained. Representative data are shown. Abbreviation used: PC, phase contrast image. The results of the line profile analysis are shown. In the line profile analysis, corresponding line-scan graphs of the relative fluorescence intensities of green anti-FLAG fluorescence (green line) and red MitoTracker red fluorescence (red line) along the white line indicated in the merged images are shown.
As shown in Supplementary Fig. S6, efficient protein expression and protein N-myristoylation were equally observed on NDUFB7-FLAG and NDUFB7-5KRtoA-FLAG. In contrast, the level of protein expression of NDUFB7-G2A-5KRtoA-FLAG was lower than that of NDUFB7-G2A-FLAG. As expected, protein N-myristoylation was not observed on these two G2A mutants. As shown in Fig. 5B upper left panel, NDUFB7-5KRtoA-FLAG specifically localized to mitochondria, similar to wild-type NDUFB7-FLAG. In contrast, NDUFB7-G2A-5KRtoA-FLAG localized to the cytoplasm, whereas NDUFB7-G2A-FLAG specifically localized to the nucleus (Fig. 5B lower left panel and Supplementary Fig. S7, A, B upper panels). These results were further confirmed by the line profile analysis of the images obtained, as shown in Fig. 5B, right panels and Supplementary Fig. S7, A B, lower panels.
CHCHD plays a critical role in the mitochondrial localization of NDUFB7
The Mia40/CHCHD4 import pathway has been shown to mediate the import of (CX9C)2 motif-carrying substrates into the intermembrane space (IMS) through outer membrane-localized TOM translocase35. The formation of two disulfide bonds among the (CX9C)2 motif in the substrates catalyzed by the oxidase Mia40/CHCHD4 plays a critical role in this transport pathway31. Regarding NDUFB7, two intra-molecular disulfide bridges between two Cys pairs (Cys59/Cys90 and Cys69/Cys80) were proposed to be formed by this system32. Therefore, we attempted to elucidate the role of CHCHD in NDUFB7 in the mitochondrial localization of this protein. To achieve this, two mutants in which Cys residues at positions 59 and 69 in the wild type and G2A mutant of NDUFB7 were changed to Ser (NDUFB7-CtoS-FLAG and NDUFB7-G2A-CtoS-FLAG, respectively) were generated (Fig. 6A and Supplementary Fig. S8A) and their susceptibility to protein N-myristoylation and their intracellular localization were evaluated.
The CHCH domain plays a critical role in the mitochondrial localization of NDUFB7. To elucidate the role of the CHCH domain in NDUFB7 in the mitochondrial localization of this protein, two mutants in which Cys residues at positions 59 and 69 in the wild type and G2A mutant of NDUFB7 were changed to Ser (NDUFB7-CtoS-FLAG and NDUFB7-G2A-CtoS-FLAG, respectively) were generated (Fig. 6A and Supplementary Fig. S8A) and their susceptibility to protein N-myristoylation and their intracellular localization were evaluated. As controls for this analysis, two MIC19 mutants in which the corresponding Cys residues at 183 and 193 in the wild type and G2A mutant of MIC19-FLAG were changed to Ser (MIC19-CtoS-FLAG and MIC19-G2A-CtoS-FLAG, respectively) were generated and the same experiments were performed. The results obtained on protein expression and protein N-myristoylation from a Western blotting analysis and metabolic labeling experiments, respectively, are shown in Supplementary Figure S9. (A) Structures of MIC19-FLAG, MIC19-CtoS-FLAG, NDUFB7-FLAG, and NDUFB7-CtoS-FLAG. (B) Analysis of the intracellular localization of MIC19-FLAG, MIC19-CtoS-FLAG, NDUFB7-FLAG, and NDUFB7-CtoS-FLAG. The intracellular localization of MIC19-FLAG, MIC19-CtoS-FLAG, NDUFB7-FLAG, and NDUFB7-CtoS-FLAG was assessed by an immunofluorescence analysis of COS-1 cells transfected with cDNA coding these four proteins using an anti-FLAG antibody. Hoechst, MitoTracker Red, and TGN-46-EGFP were used as organelle markers for the nucleus, mitochondria, and TGN, respectively. Experiments were repeated 3 times and similar results were obtained. Representative data are shown. Abbreviation used: PC, phase contrast image. (C) A line profile analysis of the intracellular localization of NDUFB7-FLAG and NDUFB7-CtoS-FLAG. The results of a line profile analysis are shown. In the line profile analysis, corresponding line-scan graphs of the relative fluorescence intensities of green anti-FLAG or TGN46-EGFP fluorescence (green line) and red MitoTracker red or anti-FLAG fluorescence (red line) along the white line indicated in the merged images are shown.
As controls for this analysis, two MIC19 mutants in which the corresponding Cys residues at positions 183 and 193 in the wild type and G2A mutant of MIC19-FLAG were changed to Ser (MIC19-CtoS-FLAG and MIC19-G2A-CtoS-FLAG, respectively) were generated. As shown in Supplementary Fig. S9, efficient protein expression and effective protein N-myristoylation were equally observed on NDUFB7-FLAG and NDUFB7-CtoS-FLAG. In contrast, the protein expression level of NDUFB7-G2A-CtoS-FLAG was significantly lower than that of NDUFB7-G2A-FLAG. Protein N-myristoylation was not observed on NDUFB7-G2A-FLAG as expected. As shown in Fig. 6B, the CtoS mutant of NDUFB7-FLAG (NDUFG7-CtoS-FLAG) did not localize to mitochondria, but specifically localized to the Golgi apparatus (TGN), as was the case with MIC19-CtoS-FLAG. These results were further confirmed by the line profile analysis of the images obtained, as shown in Fig. 6C. In contrast, the CtoS mutant derived from the G2A mutant of NDUFB7 (NDUFB7-G2A-CtoS-FLAG) localized to the nucleus, as was the case with NDUFB7-G2A-FLAG (Supplementary Fig. S8B). Regarding the CtoS mutant of MIC19-G2A-FLAG (MIC19-G2A-CtoS-FLAG), a cytosolic distribution was observed, similar to MIC19-G2A-FLAG, as shown in Supplementary Fig. S8B upper panels. These results indicated that CHCHD played a critical role in the mitochondrial localization of NDUFB7, as was the case for MIC19. Furthermore, protein N-myristoylation and CHCHD were both required for the proper mitochondrial targeting and localization of NDUFB7.
Discussion
In an attempt to identify novel human N-myristoylated proteins from human cDNA clones in human cDNA resources, we previously found that two mitochondrial outer membrane proteins, SAMM50 and TOMM40, were N-myristoylated integral membrane proteins18. The protein N-myristoylation of SAMM50, but not TOMM40, was also shown to be required for mitochondrial targeting. In the present study, we searched for human N-myristoylated mitochondrial proteins in which protein N-myristoylation was required for their specific localization to mitochondria. Among the 1,705 human genes listed in MitoProteome, 12 candidate genes for N-myristoylated mitochondrial proteins were selected and their susceptibility to protein N-myristoylation and their intracellular localization were evaluated, as previously described14,15,16,17,18,36. As a result, 7 proteins (CLN3, DMAC1, HCCS, MARC1, NDUFB7, NOL3, and PLGRKT) were found to be N-myristoylated proteins. The endogenous protein N-myristoylation occurring on these 7 proteins expressed in HeLa cells have been reported in recent reports 37,38.
Among them, four proteins, DMAC1, HCCS, NDUFB7, and PLGRKT, were identified as N-myristoylated proteins that specifically localized to mitochondria (Fig. 2). None of these proteins contain predicted cleavable presequences that direct mitochondrial localization. Among these proteins, DMAC1, HCCS, and NDUFB7 have been shown to be mitochondrial proteins. DMAC1 and NDUFB7 were found to play important roles in the assembly of complex I of the mitochondrial respiratory chain as an assembly factor and an accessory subunit, respectively23. Both DMAC1 and NDUFB7 localize to the inner membrane and associate with complex I of the mitochondrial respiratory chain23,39. The import pathway for NDUFB7 has been characterized. NDUFB7 was first synthesized by cytosolic ribosomes in the cytoplasm, and imported into intermembrane space through outer membrane-localized TOM complex in an unfolded reduced form, then folded and oxidized by the disulfide relay-dependent Mia40/CHCHD4 import machinery39,40. The import pathway for DMAC1 has not been characterized so far.
HCCS localizes to the intermembrane space and functions as a lyase that catalyzes the covalent linking of the heme group to the cytochrome C apoprotein to produce mature functional cytochrome C27.
The localization and import pathway of HCCS has been well characterized in yeast41. HCCS was synthesized in the cytosol and directly imported to intermembrane space through outer membrane-localized TOM complex, then localized to the outer surface of the inner membrane.
Different from DMAC1, NDUFB7 and HCCS, PLGRKT has been shown to be a cell surface receptor for plasminogen that regulates cell surface plasminogen activation30. The presence and biological function of PLGRKT in mitochondria have not yet been reported. Thus, intramitochondrial localization and import pathway for PLGRKT have not been characterized yet.
An analysis of the intracellular localization of non-myristoylatable G2A mutants indicated that protein N-myristoylation occurring on NDUFB7, an accessory subunit of human mitochondrial respiratory chain complex I, played a vital role in the mitochondrial localization of this protein (Fig. 3). NDUFB7 is a member of the CHCHD-containing family of proteins (CHCHD proteins), which are imported into the mitochondrial IMS, and it has been proposed to stabilize assembled mitochondrial respiratory chain complex I by binding to its surface32. Mitochondrial respiratory chain complex I is the largest complex in the oxidative phosphorylation system. It comprises 14 central subunits, 31 accessory subunits, and at least 15 assembly factors40. Significant advances have recently been made in elucidating the structure and assembly steps of complex I. NDUFB7 is an accessory subunit of module PD-b. A critical role has been suggested for NDUFB7 in the assembly of complex I in the yeast Yarrowia lipolytica42. Furthermore, from in vitro experiments, the absence of any of the accessory subunits of the PD-b module may critically affect the assembly and function of human complex I23,42,43.
In fully assembled complex I, the module PD-b is composed of the central subunit ND5 and accessory subunits NDUFB2, NDUFB3, NDUFB7, NDUFB8, and NDUFB940,43. Deleterious changes in proteins of the PD-b module have been described and associated with complex I disorders44,45,46. Regarding NDUFB7, a recent study reported that an intronic mutation leading to a cryptic splice site in the NDUFB7 gene caused a mitochondrial disorder47. In this case, an intronic mutation in NDUFB7 caused severe congenital lactic acidosis and hypertrophic cardiomyopathy. The detected mutation resulted in a significant reduction in the NDUFB7 protein and reduced complex I activity. It was also revealed that complementation studies with the expression of wild-type NDUFB7 in patient fibroblasts normalized complex I function. Therefore, NDUFB7 plays an important role in the formation and function of complex I.
The CHCHD-containing family of proteins are proteins that localize to the mitochondrial IMS and contain (CX3C)2 or (CX9C)2 motifs31. These CHCHD proteins are specifically transported into mitochondria by the disulfide relay-dependent Mia40/CHCHD4 import machinery35. They are involved in mitochondrial biogenesis, bioenergetics, dynamics, and quality control in the IMS31. Similar to other CHCHD proteins, CHCHD is required for the mitochondrial localization of NDUFB7; however, the present study revealed that NDUFB7 required protein N-myristoylation in addition to CHCHD. G2A mutant in which protein N-myristoylation was inhibited did not localize to mitochondria, but specifically localized to the nucleus (Figs. 3, 5). The nuclear localization of the G2A mutant of NDUFB7 was directed by the NLSL RKKRR, located at positions 111–115 (Fig. 5). This result indicated that protein N-myristoylation-mediated mitochondrial localization is preferential over nuclear localization mediated by the NLSL RKKRR.
A known CHCHD protein that requires protein N-myristoylation for mitochondrial localization is MIC19 (CHCHD3), an N-myristoylated mitochondrial inner membrane-binding protein that interacts with the luminal surface of the MICOS complex and is involved in the formation of cristae18,48. In addition to MIC19, protein N-myristoylation has been proposed to occur on another CHCHD protein, MIC25 (CHCHD6), which localizes to the mitochondrial IMS. A phylogenic analysis indicated that human MIC19 and MIC25 resulted from a gene duplication at the root of vertebrates49. Therefore, they are both orthologous to MIC19 in yeast and likely the result of whole genome duplication50. Regarding the mitochondrial localization of MIC19 and MIC25, neither a canonical cleavable N-terminal mitochondrial targeting signal (presequence) nor another specific mitochondrial targeting signal has been identified. However, as for MIC19, protein N-myristoylation and CHCHD were both essential for the proper mitochondrial localization of this protein48.
MIC25 was predicted to be N-myristoylated; however, direct biochemical evidence for protein N-myristoylation has not yet been reported. We previously showed that exogenously and endogenously expressed MIC19 and MIC25 were efficiently N-myristoylated18. Overexpressed MIC19 specifically localized to mitochondria, whereas the non-myristoylatable G2A mutant of MIC19 localized to the cytoplasm, confirming the requirement of protein N-myristoylation for the mitochondrial localization of MIC19. However, the majority of overexpressed N-myristoylated MIC25 was detected in the cytoplasm, but not in mitochondria. A similar cytosolic localization was observed for the non-myristoylatable G2A mutant of MIC25. Therefore, protein N-myristoylation did not appear to play a critical role in the mitochondrial localization of MIC25. The mechanisms responsible for differences in the role of protein N-myristoylation in the mitochondrial localization of MIC19 and MIC25 remains unclear.
A mechanism for the protein N-myristoylation-dependent mitochondrial localization of Mic19 was recently proposed using yeast Mic1951. This study demonstrated that the CHCHD-mediated mitochondrial translocation of Mic19 was inhibited by an unfolded domain in its N-terminal region, called the DAF domain. Protein N-myristoylation circumvented this inhibitory effect by mediating the binding of MIC19 to Tom20 in the TOM complex and to the mitochondrial outer membrane, thereby achieving the efficient mitochondrial localization of MIC19. It remains unclear whether protein N-myristoylation plays a similar role with respect to NDUFB7. However, since there is no DUF domain-like unfolded domain in the N-terminal region of NDUFB7, it is unlikely that the protein N-myristoylation of NDUFB7 circumvented the inhibitory effects of the DUF domain-like unfolded domain on mitochondrial localization. In addition, the non-myristoylatable G2A mutant of NDUFB7 was shown to localize to the nucleus rather than to the cytoplasm, different from the case for MIC19 (Fig. 5B lower panel, Supplementary Fig. S5, Supplementary Fig. S7A). Furthermore, the nuclear localization of the non-myristoylatable G2A mutant of NDUFB7 was mediated by the NLSL RKKRR, located at positions 111 ~ 115 (Fig. 5B lower panel, Supplementary Fig. S7). Therefore, protein N-myristoylation in NDUFB7 appears to function in the inhibition of nuclear localization mediated by the NLSL RKKRR rather than circumventing the suppressive effects of the DUF domain on mitochondrial localization, as observed for Mic19. It currently remains unclear why a positive charge amino acid cluster, such as RKKRR, is present in NDUFB7. In some proteins, positively charged amino acids play a role in protein–protein interactions and the assembly of protein complexes. For example, electrostatic interactions between positively and negatively charged residues play critical roles in the structure and function of mammalian respiratory complex I39,52. Therefore, the amino acid sequence RKKRR in NDUFB7 may be involved in the function of NDUFB7 as an accessory subunit of respiratory chain complex I.
In MIC19, protein N-myristoylation played a critical role not only in mitochondrial localization, but also in interactions with SAMM50 localized to the mitochondrial outer membrane48. In the case of NDUFB7, protein N-myristoylation was first reported on bovine NDUFB7, and this modification was proposed to function as a membrane anchor to the mitochondrial inner membrane53. Concerning the protein–protein interactions mediated by protein N-myristoylation of NDUFB7, recent study on the structure of mammalian mitochondrial complex I indicated that the N-terminal myristoyl group of NDUFB7 is bound in a groove on ND5 formed by TMH12ND5, TMH13ND5 and TMH15ND539. Thus, it is quite probable that the protein–protein interactions mediated by protein N-myristoylation are involved in the function of NDUFB7 as an accessory subunit of respiratory chain complex I.
Schematic representation of possible roles of protein N-myristoylation and the CHCH domain in the mitochondrial targeting and localization of NDUFB7 is shown in Fig. 7.
Schematic representation of possible roles of protein N-myristoylation and the CHCH domain in the mitochondrial targeting and localization of NDUFB7. NDUFB7 was synthesized by cytosolic ribosomes in the cytoplasm, targeted to the mitochondrial outer membrane, and then translocated across the outer membrane through outer membrane-localized TOM translocase (TOM complex) by a protein N-myristoylation-dependent mechanism. The Mia40/CHCHD4 import machinery then mediated the oxidation-dependent retention of NDUFB7 in the intermembrane space. NDUFB7 functioned as an accessory subunit of complex I of the mitochondrial respiratory chain (Complex I). In this mechanism, protein N-myristoylation and CHCH-domain were both required for the proper mitochondrial localization of NDUFB7.
NDUFB7 was first synthesized by cytosolic ribosomes in the cytoplasm, targeted to the mitochondrial outer membrane, and then translocated across the outer membrane through outer membrane-localized TOM translocase (TOM complex) by a protein N-myristoylation-dependent mechanism. The Mia40/CHCHD4 import machinery then mediated the oxidation-dependent retention of NDUFB7 in the intermembrane space. The NDUFB7 localized in the intermembrane space functioned as an accessory subunit of complex I of the mitochondrial respiratory chain (Complex I). In this mechanism, protein N-myristoylation and CHCH-domain were both required for the proper mitochondrial localization of NDUFB7. However, the relative roles of protein N-myristoylation and CHCH-domain in the intracellular localization and function of NDUFB7 remained to be elucidated. Further analyses are required to fully elucidate the role of protein N-myristoylation in the intracellular localization and function of NDUFB7.
Methods
Materials
cDNA coding human MIC19 was purchased from Promega Corp. (Madison, WI, USA). Other human cDNAs were obtained from the RIKEN BioResource Research Center (RIKEN BRC, Tsukuba, Japan) through the National Bio-Resource Project of the MEXT, Japan54,55,56,57. ECL prime Western blotting detection reagents were from GE Healthcare (Buckinghamshire, UK). The dye terminator cycle sequencing kit, Lipofectamine LTX and Plus reagents, MitoTracker Red CMXRos, and Hoechst 33342 were from Life Technologies Corporation (Carlsbad, CA, USA). An anti-FLAG monoclonal antibody and anti-mouse IgG-FITC antibody were from Sigma (St. Louis, MO, USA). An anti-mouse IgG-ALEXA594 antibody was from Thermo Scientific (Rockford, IL, USA). 13-Tetradecynoic acid (Alk-Myr) was from Cayman (Ann Arbor, MI, USA). Azide TAMRA (Az-TAMRA) was from Click Chemistry Tools (Scottsdale, AZ, USA). Tris(2-carboxyethyl)phosphine hydrochloride (TCEP) and tris[(1-benzyl-1H-1,2,3-triazol-4-yl)methyl]amine (TBTA) were from Sigma (St. Louis, MO, USA). Protein G-HRP conjugate was from Bio-Rad (Hercules, CA, USA). The other reagents used were from Wako Pure Chemical (Osaka, Japan) or Daiichi Pure Chemicals (Tokyo, Japan) and were of analytical or DNA grade.
Prediction of protein N-myristoylation using prediction programs
Two public WWW server-based prediction programs for protein N-myristoylation, MYR Predictor (http://mendel.imp.ac.at/myristate/SUPLpredictor.htm)24 and Myristoylator (http://www.expasy.org/tools/myristoylator/)25, were used to predict protein N-myristoylation. The entire amino acid sequences deduced from the nucleotide sequences of the ORFs were used as the query.
Plasmid construction
The nucleotide sequences of oligonucleotides used for plasmid construction are summarized in Supplementary Tables S3, S4, and S5. The plasmids pcDNA3-tGelsolin-FLAG, pcDNA3-FLAG, and pcDNA3-TGN46-EGFP were constructed as previously described58,59. pcDNA3 plasmids containing the cDNAs coding tGelsolin fusion proteins with the 10 N-terminal amino acids of the proteins listed in the MitoProteome protein database at the N terminus (pcDNA3-X(N10)-tGelsolin-FLAG) were constructed using the oligonucleotides listed in Table S3 and the plasmids pcDNA3-tGelsolin-FLAG and pcDNA3-FLAG, as described in Table S6. pcDNA3 plasmids, including cDNA coding C-terminally FLAG-tagged full-length proteins and their mutants, were constructed by PCR or site-directed mutagenesis, as summarized in Supplementary Tables S6 and S7.
Transfection of cells
COS-1 (a simian virus 40-transformed African green monkey kidney cell line, American Type Culture Collection) cells were maintained in Dulbecco’s modified Eagle’s medium (DMEM; Gibco BRL [Palo Alto, CA, USA]) supplemented with 10% fetal calf serum (FCS; Gibco BRL). Cells (2 × 105) were plated onto 35-mm dishes 1 day before transfection. pcDNA3 constructs (2 μg) containing cDNA coding FLAG-tagged or EGFP-fusion proteins were transfected into cells on each plate using 2.5 μL of Lipofectamine LTX and 2 μL of Plus reagent in 1 mL of serum-free medium18. After an incubation at 37 °C for 5 h, serum-containing medium was added and cells were incubated again at 37 °C for the appropriate periods.
Metabolic labeling of cells
The metabolic labeling of cells with the myristic acid analog (Alk-Myr) was performed as previously described18. Cells (2 × 105) were transfected with pcDNA3 constructs (2 μg) containing cDNA as described above and incubated at 37 °C for 12 h. They were then washed once with 1 mL of serum-free DMEM and incubated at 37 °C for 10 h in 1 mL of DMEM (+ 2% FCS) containing 25 μM Alk-Myr. Cells were then washed three times with Dulbecco’s phosphate-buffered saline (DPBS), harvested, and lysed with 200 μL of RIPA buffer (50 mM Tris–HCl (pH 7.5), 150 mM NaCl, 1% Nonidet P-40, 0.5% sodium deoxycholate, 0.1% SDS, and protease inhibitors) on ice for 20 min. Samples labeled with Alk-Myr were reacted with Az-TAMRA by click chemistry and protein N-myristoylation was analyzed by an in-gel fluorescence analysis36.
Cu(I)-catalyzed azide-alkyne cycloaddition (CuAAC)
Cell lysates labeled with Alk-Myr (46 µL) were reacted with 4 µL of a freshly premixed click chemistry reaction cocktail (1 µL Az-TAMRA [5 mM], 1 µL TCEP [50 mM], 1 µL TBTA [5 mM], and 1 µL CuSO4·5H2O [50 mM]) in a total reaction volume of 50 µL at room temperature for 1 h. After CuAAC, 500 µL of MeOH was added to the samples and they were stored at − 80 °C overnight. After centrifugation at 15,000 rpm at 4 °C for 30 min, the supernatant was removed. Thereafter, the pellet was washed with 500 µL of MeOH and air dried18. Samples were denatured by sonication in SDS-sample buffer and then subjected to SDS-PAGE. An in-gel fluorescence analysis of SDS-PAGE gels was performed using Typhoon FLA9500 (GE Healthcare Bio-Sciences AB, Uppsala, Sweden)60.
Western blotting
Proteins were resolved by 12.5% SDS-PAGE and then transferred to an Immobilon-P transfer membrane. After blocking with non-fat milk, the membrane was probed with a primary antibody, as previously described61. Immunoreactive proteins were specifically detected by an incubation with a protein G-HRP conjugate. The membrane was developed using ECL Prime Western blotting detection reagent and bands were detected using a MicroChemi Chemiluminescence Imaging System (Berthold Technologies, Bad Wildbad, Germany).
Immunofluorescence analysis and fluorescence microscopy
An immunofluorescence analysis of transfected cells was performed 24 h after transfection62. After staining with Hoechst 33342 and MitoTracker Red, cells were washed with DPBS, fixed in 4% paraformaldehyde in DPBS for 15 min, and permeabilized with 0.1% Triton X-100 in DPBS for 10 min at room temperature, followed by washing with 0.1% gelatin in DPBS36. Permeabilized cells were incubated with a specific antibody in DPBS at room temperature for 1 h. After washing with 0.1% gelatin in DPBS, cells were incubated with the anti-IgG-FITC or anti-IgG-ALEXA594 antibody at room temperature for 1 h. After washing with 0.1% gelatin in DPBS, cells were observed using a Leica AF7000 fluorescence microscope (Leica, Solmser, Germany)36. A line profile analysis was performed using the Line Profile Tool of Leica LAS AF software (Leica, Solmser, Germany). In this analysis, white lines in merged panels were converted to line profiles using the Line Profile Tool.
Data availability
All data generated or analyzed during this study are included in this published article and its supplementary information files.
References
Meinnel, T., Dian, C. & Giglione, C. Myristoylation, an ancient protein modification mirroring eukaryogenesis and evolution. Trends Biochem. Sci. 45, 619–632 (2020).
Wang, B. et al. Protein N-myristoylation: Functions and mechanisms in control of innate immunity. Cell. Mol. Immunol. 18, 878–888 (2021).
Jiang, H. et al. Protein lipidation: Occurrence, mechanisms, biological functions, and enabling technologies. Chem. Rev. 118, 919–988 (2018).
Udenwobele, D. I. et al. Myristoylation: An important protein modification in the immune response. Front Immunol. 8, 751 (2017).
Zha, J. et al. Posttranslational N-myristoylation of BID as a molecular switch for targeting mitochondria and apoptosis. Science 290, 1761–1765 (2000).
Utsumi, T., Sakurai, N., Nakano, K. & Ishisaka, R. C-terminal 15 kDa fragment of cytoskeletal actin is posttranslationally N-myristoylated upon caspase-mediated cleavage and targeted to mitochondria. FEBS Lett. 539, 37–44 (2003).
Martin, D. D., Beauchamp, E. & Berthiaume, L. G. Post-translational myristoylation: Fat matters in cellular life and death. Biochimie 93, 18–31 (2011).
Dyda, F., Klein, D. C. & Hickman, A. B. GCN5-related N-acetyltransferases: a structural overview. Annu. Rev. Biophys. Biomol. Struct. 29, 81–103 (2000).
Fhu, C. W. & Ali, A. Protein lipidation by palmitoylation and myristoylation in cancer. Front. Cell Dev. Biol. 9, 673647 (2021).
Wright, M. H., Heal, W. P., Mann, D. J. & Tate, E. W. Protein myristoylation in health and disease. J. Chem. Biol. 3, 19–35 (2010).
Resh, M. D. Targeting protein lipidation in disease. Trends Mol. Med. 18, 206–214 (2012).
Burnaevskiy, N. et al. Proteolytic elimination of N-myristoyl modifications by the Shigella virulence factor IpaJ. Nature 496, 106–109 (2013).
Timothy, P. H. et al. Asciminib in chronic myeloid leukemia after ABL kinase inhibitor failure. N. Engl. J. Med. 381, 2315–2326 (2019).
Suzuki, T. et al. Strategy for comprehensive identification of human N-myristoylated proteins using an insect cell-free protein synthesis system. Proteomics 10, 1780–1793 (2010).
Yamauchi, S. et al. The consensus motif for N-myristoylation of plant proteins in a wheat germ cell-free translation system. FEBS J. 277, 3596–3607 (2010).
Takamitsu, E. et al. Cell-free identification of novel N-myristoylated proteins from complementary DNA resources using bioorthogonal myristic acid analogues. Anal. Biochem. 464, 83–93 (2014).
Takamitsu, E. et al. Identification of human N-myristoylated proteins from human complementary DNA resources by cell-free and cellular metabolic labeling analyses. PLoS ONE 10, e0136360 (2015).
Utsumi, T. et al. Identification and characterization of protein N-myristoylation occurring on four human mitochondrial proteins, SAMM50, TOMM40, MIC19, and MIC25. PLoS ONE 13, e0206355 (2018).
Chacinska, A. et al. Importing mitochondrial proteins: machineries and mechanisms. Cell 138, 628–644 (2009).
Rapaport, D. & Neupert, W. Biogenesis of Tom40, core component of the TOM complex of mitochondria. J Cell Biol. 146, 321–331 (1999).
Kozjak, V. et al. An essential role of Sam50 in the protein sorting and assembly machinery of the mitochondrial outer membrane. J. Biol. Chem. 278, 48520–48523 (2003).
Cotter, D., Guda, P., Fahy, E. & Subramaniam, S. MitoProteome: Mitochondrial protein sequence database and annotation system. Nucleic Acids Res. 32, D463–D467 (2004).
Stroud, D. A. et al. Accessory subunits are integral for assembly and function of human mitochondrial complex I. Nature 538, 123–126 (2016).
Stroh, S. M., Eisenhaber, B. & Eisenhaber, F. N-terminal N-myristoylation of proteins: Prediction of substrate proteins from amino acid sequence. J. Mol. Biol. 317, 541–557 (2002).
Bologna, G., Yvon, C., Duvaud, S. & Veuthey, A.-L. N-Terminal myristoylation predictions by ensembles of neural networks. Proteomics 4, 1626–1632 (2004).
Luiro, K. et al. Interconnections of CLN3, Hook1 and Rab proteins link Batten disease to defects in the endocytic pathway. Hum. Mol. Genet. 13, 3017–3027 (2004).
San Francisco, B., Bretsnyder, E. C. & Kranz, R. G. Human mitochondrial holocytochrome c synthase’s heme binding, maturation determinants, and complex formation with cytochrome c. Proc. Natl. Acad. Sci. USA 110, E788-E797 (2013).
Gruenewald, S. et al. The fourth molybdenum containing enzyme mARC: cloning and involvement in the activation of N-hydroxylated prodrugs. J. Med. Chem. 51, 8173–8177 (2008).
Gustafsson, A. B. et al. Apoptosis repressor with caspase recruitment domain protects against cell death by interfering with Bax activation. J. Biol. Chem. 279, 21233–21238 (2004).
Lighvani, S. et al. Regulation of macrophage migration by a novel plasminogen receptor Plg-RKT. Blood 118, 5622–5630 (2011).
Modjtahedi, N. et al. Mitochondrial proteins containing coiled-coil-helix-coiled-coil-helix (CHCH) domains in health and disease. Trends. Biochem. Sci. 41, 245–260 (2016).
Szklarczyk, R. et al. NDUFB7 and NDUFA8 are located at the intermembrane surface of complex I. FEBS Let. 585, 737–743 (2011).
Kosugi, S. et al. Systematic identification of cell cycle-dependent yeast nucleocytoplasmic shuttling proteins by prediction of composite motifs. Proc. Natl. Acad. Sci. USA 106, 10171–10176 (2009).
Brameier, M., Krings, A. & MacCallum, R. M. NucPred—Predicting nuclear localization of proteins. Bioinformatics 23, 1159–1160 (2007).
Al-Habib, H. & Ashcroft, M. CHCHD4(MIA40) and the mitochondrial disulfide relay system. Biochem. Soc. Trans. 49, 17–27 (2021).
Utsumi, T. et al. ANKRD22 is an N-myristoylated hairpin-like monotopic membrane protein specifically localized to lipid droplets. Sci. Rep. 11, 19233 (2021).
Thinon, E. et al. Global profiling of co- and post-translationally N-myristoylated proteomes in human cells. Nat. Commun. 5, 4919 (2014).
Tsumagari, K. et al. Application of liquid-liquid extraction for N-terminal myristoylation proteomics. Mol. Cell. Proteomics, in press.
Padavannil, A. et al. The mysterious multitude: Structural perspective on the accessory subunits on respiratory complex I. Front. Mol. Biosci. 8, 798353 (2022).
Formosa, L. E. et al. Building a complex complex: Assembly of mitochondrial respiratory chain complex I. Semin. Cell Dev. Biol. 76, 154–162 (2018).
Steiner, H. et al. Biogenesis of mitochondrial heme lyases in yeast. Import and folding in the intermembrane space. J. Biol. Chem. 270, 22842–22849 (1995).
Dröse, S. et al. Functional dissection of the proton pumping modules of mitochondrial complex I. PLoS Biol. 9, e1001128 (2011).
Formosa, L. E. et al. Dissecting the roles of mitochondrial complex I intermediate assembly complex factors in the biogenesis of complex I. Cell Rep. 31, 107541 (2020).
Haack, T. B. et al. Mutation screening of 75 candidate genes in 152 complex I deficiency cases identifies pathogenic variants in 16 genes including NDUFB9. J. Med. Genet. 49, 83–89 (2012).
Danhelovska, T. et al. Multisystem mitochondrial diseases due to mutations in mtDNA-encoded subunits of complex I. BMC Pediatr. 20, 1–13 (2020).
Piekutowska-Abramczuk, D. et al. NDUFB8 mutations cause mitochondrial complex I deficiency in individuals with Leigh-like encephalomyopathy. Am. J. Hum. Genet. 102, 460–467 (2018).
Correia, S. P. et al. Sever congenital lactic acidosis and hypertrophic cardiomyopathy caused by an intronic variant in NDUFB7. Hum. Mutat. 42, 378–384 (2021).
Darshi, M. et al. Targeting and import of coiled-coil helix coiled-coil helix domain-containing protein 3 (ChChd3) into the mitochondrial intermembrane space. J. Biol. Chem. 287, 39489–39491 (2012).
Schreiber, F. et al. TreeFam v9: A new website, more species and orthology-on-the-fly. Nucleic Acids Res. 42, D922–D925 (2014).
Szklarczyk, R. & Huynen, M. A. Expansion of the human mitochondrial proteome by intra- and inter-compartmental protein duplication. Genome Biol. 10, R135 (2009).
Ueda, E. et al. Myristoyl group-aided protein import into the mitochondrial intermembrane space. Sci. Rep. 9, 1185 (2019).
Kampjut, D. & Sazanov LA. The coupling mechanism of mammalian respiratory complex I. Science 370, eabc4209 (2020).
Carroll, J. et al. The post-translational modification of the nuclear encoded subunits of complex I from bovine heart mitochondria. Mol. Cell. Proteomics 4, 693–699 (2005).
Ota, T. et al. Complete sequencing and characterization of 21,243 full-length human cDNAs. Nat. Genet. 36, 40–45 (2004).
Otsuki, T. et al. Signal sequence and keyword trap in silico for selection of full-length human cDNAs coding secretion or membrane proteins from oligo-capped cDNA libraries. DNA Res. 12, 117–126 (2005).
Kimura, K. et al. Diversification of transcriptional modulation: Large-scale identification and characterization of putative alternative promoters of human genes. Genome Res. 16, 55–65 (2006).
Itoh, M. et al. Constructing ORFeome resources with removable termination codons. Biotechniques 41, 44–48 (2006).
Moriya, K. et al. Protein N-myristoylation plays a critical role in the endoplasmic reticulum morphological change induced by overexpression of protein Lunapark, an integral membrane protein of the endoplasmic reticulum. PLoS ONE 8, e78235 (2013).
Kinoshita-Kikuta, E. et al. Protein-N-myristoylation-dependent phosphorylation of serine 13 of tyrosine kinase Lyn by casein kinase 1γ at the Golgi during intracellular protein traffic. Sci. Rep. 10, 16273 (2020).
Kinoshita-Kikuta, E. et al. A strategy to identify protein-N-myristoylation-dependent phosphorylation reactions of cellular proteins by using Phos-tag SDS-PAGE. PLoS ONE 14, e0225510 (2019).
Utsumi, T. et al. Amino acid residue penultimate to the amino-terminal Gly residue strongly affects two cotranslational protein modifications, N-myristoylation and N-acetylation. J. Biol. Chem. 276, 10505–10513 (2001).
Sakurai, N. & Utsumi, T. Posttranslational N-myristoylation is required for the anti-apoptotic activity of human tGelsolin, the C-terminal caspase cleavage product of human gelsolin. J. Biol. Chem. 281, 14288–14295 (2006).
Acknowledgements
This study was supported in part by KAKENHI Grants 17K07758 and 20K05827 to T.U.
Author information
Authors and Affiliations
Contributions
T.U.: Project administration, Conceptualization methodology, resources, investigation, writing of the draft manuscript, review and editing of the draft manuscript, and funding acquisition. H.H.: Investigation, writing of the draft manuscript, and review and editing of the draft manuscript. K.M.: Investigation, review and editing of the draft manuscript. H.K.: Review and editing of the draft manuscript. N.I.: Conceptualization methodology, review and editing of the draft manuscript. All authors approved the final version of the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Harada, H., Moriya, K., Kobuchi, H. et al. Protein N-myristoylation plays a critical role in the mitochondrial localization of human mitochondrial complex I accessory subunit NDUFB7. Sci Rep 13, 22991 (2023). https://doi.org/10.1038/s41598-023-50390-z
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-023-50390-z
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.