Abstract
Colorectal cancer (CRC) liver metastasis accounts for the majority of fatalities associated with CRC. Early detection of metastasis is crucial for improving patient outcomes but can be delayed due to a lack of symptoms. In this research, we aimed to investigate CRC metastasis-related biomarkers by employing a machine learning (ML) approach and experimental validation. The gene expression profile of CRC patients with liver metastasis was obtained using the GSE41568 dataset, and the differentially expressed genes between primary and metastatic samples were screened. Subsequently, we carried out feature selection to identify the most relevant DEGs using LASSO and Penalized-SVM methods. DEGs commonly selected by these methods were selected for further analysis. Finally, the experimental validation was done through qRT-PCR. 11 genes were commonly selected by LASSO and P-SVM algorithms, among which seven had prognostic value in colorectal cancer. It was found that the expression of the MMP3 gene decreases in stage IV of colorectal cancer compared to other stages (P value < 0.01). Also, the expression level of the WNT11 gene was observed to increase significantly in this stage (P value < 0.001). It was also found that the expression of WNT5a, TNFSF11, and MMP3 is significantly lower, and the expression level of WNT11 is significantly higher in liver metastasis samples compared to primary tumors. In summary, this study has identified a set of potential biomarkers for CRC metastasis using ML algorithms. The findings of this research may provide new insights into identifying biomarkers for CRC metastasis and may potentially lay the groundwork for innovative therapeutic strategies for treatment of this disease.
Similar content being viewed by others
Introduction
Despite numerous research efforts aimed at identifying strategies for cancer prevention, colorectal cancer (CRC) is still the second cause of cancer mortality worldwide, with about one million deaths in 20201. An estimated 90% of all cancer-related deaths are caused by cancer metastasis, making it a significant obstacle to effective cancer care2. Nearly 20% of CRC patients present with metastatic disease at initial diagnosis, and the liver is the most general metastatic site for CRC, accounting for almost 50% of all cases3. Metastasis of CRC leads to a poor prognosis4. Despite receiving standard treatments such as surgical removal, radiation therapy, and systemic chemotherapy, many patients with CRC liver metastasis still experience high rates of recurrence and less favorable clinical outcomes2,5. Hence, early diagnosis of liver metastases of CRC is crucial for improving patients’ prognosis and clinical outcomes6. Imaging examinations and focal biopsies are necessary for diagnosing CRC liver metastasis. However, the sensitivity of imaging techniques for CRC liver metastasis is still insufficient to accomplish the benefit of early diagnosis7. Incorporating biomarkers alongside imaging methods can significantly enhance the accuracy of detecting CRC liver metastasis8. Various biomarkers are utilized for detecting CRC liver metastasis, including but not limited to CEA, CA19-9, CA125, and others9,10. On the other hand, while serum markers such as CEA are helpful in diagnosing CRC, their limitations in terms of sensitivity and specificity decrease their reliability in identifying hepatic metastases particularly6. Thus, it is imperative to explore novel biomarkers to improve patients' diagnostic accuracy and clinical outcomes. Several types of biomarkers are used for cancer screening, including DNA, protein, and RNA biomarkers11. Transcriptional biomarkers are a promising class of biomarkers reflecting changes in the levels of RNA molecules produced from DNA in cells, including mRNAs, micro RNAs, long non-coding RNAs, and circular RNAs. They are non-invasive and highly sensitive, making them a valuable tool for the early detection and monitoring of various cancers12. Cancer initiation, development, and metastasis are influenced by complex processes and alterations at the transcriptome levels13.
Healthcare digitalization and the development of high-throughput technologies, such as microarray and next-generation sequencing (NGS), have enabled the collection of large amounts of transcriptome data in comprehensive databases such as The Gene Expression Omnibus (GEO) and The Cancer Genome Atlas (TCGA) database, which can be used to understand the underlying mechanisms of diseases which in turn developing precision medicine14. The analysis of transcriptome data can be complex and time-consuming, requiring expertise in bioinformatics and statistics. Traditional methods for analyzing transcriptome data involve manual curation and interpretation, which can be error-prone and may not be beneficial in handling the large amounts of data generated by modern sequencing technologies15. Artificial Intelligence (AI) algorithms such as machine learning (ML) can identify patterns and relationships in the data that would be difficult or impossible to detect using manual methods16. ML is a branch of AI that applies statistical methods and algorithms to achieve data parsing, categorization, and pattern recognition. This obtained information later enables computers to learn from data processing experiences and make more accurate predictions17. It is anticipated that ML will have a preeminent impact in the therapeutic context for detecting and treating cancer in the near future. It is worth mentioning that ML has been progressively utilized for the screening, diagnosis, and therapy of CRC over the course of the past five years18.
We used a machine learning-based feature selection technique to explore liver metastasis-related biomarkers in CRC. These methods are particularly beneficial for handling high-dimensional data and complex interactions between features and improving predictive models' performance which, results in identifying relevant features based on their predictive power. In the present study, features commonly selected by two feature selection algorithms were investigated through in-silico and experimental validation.
Materials and methods
Study design, data resources, and preprocessing
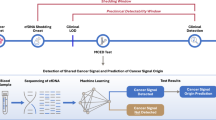
A schematic representation of the research process is illustrated in Fig. 1. Primary search was conducted using the terms "colorectal neoplasms" and metastasis in the Gene Expression Omnibus database (GEO, http://www.ncbi.nlm.nih.gov/geo) to identify any relevant available datasets based on these criteria: (1) datasets contain primary CRC and metastatic liver tumor samples; (2) comprise more than 20 samples in each group; (3) more than 10,000 genes in each dataset. Based on these considerations, three microarray datasets, GSE41568, GSE41258, and GSE68468, were included in the present study (Table 1). GSE41568 dataset (GPL570, Affymetrix Human Genome U133 Plus 2.0 Array) containing 80 liver metastases and 39 primary tumors was selected as the main dataset. GSE41258 (GPL96, Affymetrix Human Genome U133A Array) with 186 primary tumors and 67 liver metastases and GSE68468 (GPL96, Affymetrix Human Genome U133A Array) with 186 primary tumors and 47 liver metastases were considered as validation datasets. In addition, we used the TCGA database (https://portal.gdc.cancer.gov/) to obtain RNA-seq data and clinical details of 644 CRC patients (89 metastatic (M1) and 555 non-metastatic (M0) samples) as well as another validation set in our study.
GSE41568 dataset was analyzed for identifying differentially expressed mRNAs (DEGs) in primary CRC samples and liver metastases. Next, using ML-based feature selection methods, most relevant DEGs were selected. Further analyses such as survival analysis and their potential targeting drugs and related TFs were investigated. Experimental validation was also carried out as the last step of our study.
On each platform, raw data was retrieved for these three datasets. All datasets were normalized if needed in R software(version 3.6.0; https://www.r-project.org/)19. Raw data was also evaluated to contain logarithmic fold change values, and if necessary, logarithm 2 of values was obtained. The probe IDs were converted into gene expression symbols on each annotation platform. Multiple probes related to a single gene were averaged to give the gene expression value. Also, probes with a vacancy were removed.
Gene set enrichment analysis (GSEA)
GSEA is a popular method for identifying genes' biological significance by analyzing gene set expression patterns. This approach holds the potential to offer valuable insights into underlying biological processes that are associated with expressed genes. In this regard, we used GSEA software (version 4.1.0) to conduct this analysis to uncover the pathways most relevant to all expressed genes between primary tumors and liver metastases of CRC patients in the GSE41568 dataset by setting FDR criteria to < 0.05.
Screening of differentially expressed genes (DEGs)
To screen DEGs between primary tumors and liver metastasis samples in the GSE41568 dataset, we carried out differential gene expression analysis using the limma package20 in R. |log2Fold Change |≥ 1 and False Discovery Rate threshold (FDR) < 0.05 were considered as cut-off criteria.
Gene ontology (GO) and KEGG pathway enrichment analysis of DEGs
To investigate the biological properties of these DEGs, KEGG (Kyoto Encyclopedia of Genes and Genomes (http://www.Kegg.jp)) and GO enrichment analysis of selected DEGs was carried out using ClusterProfiler21, and GOplot22 packages in R. Statistical significance was assigned in terms of Benjamini < 0.05.
Feature selection using machine learning algorithms
ML methods are tools to develop and evaluate classification and prediction algorithms. Data collection, model selection, training the model, and testing the model are the four steps that make up the foundation of machine learning23. Large numbers of input characteristics are challenging for ML methods to manage. Consequently, data preparation is a necessary task for supporting the use of machine learning in real-world settings. Feature selection is among the most used data preparation techniques for screening outcome-related variables from a large pool of variables24. Obtaining the appropriate features or subsets of elements from the literature to fulfill their classification goals has become an essential part of the ML procedure. In addition to the benefits of feature selection processes to search for a subset of important features, they are also employed to prevent overfitting and produce more efficient models23. We used two feature selection algorithms in the present study to pick cancer-related genes that accurately discriminate metastatic samples from non-metastatic ones, including 1). Random Forest (RF), 2). Penalized Support Vector Machine (P-SVM) with two penalties of Smoothly Clipped Absolute Deviation (SCAD) and Least Absolute Shrinkage and Selection Operator (LASSO).
Random Forest
Random Forest, proposed by Breiman (2001), is a well-known technique that belongs to the ensemble algorithms used for classification and regression issues. This method forecasts an outcome by averaging the results of hundreds or more decision trees. RF is also employed as a variable selection strategy to identify informative variables25. The "randomForestSRC" R package was utilized in our investigation to determine the best features.
Penalized support vector machine
Support vector machine (SVM) classification is one of the most popular and effective classification approaches26. However, a significant shortcoming of this method is that it cannot perform automated gene selection. To handle this problem, the " P-SVM" method was introduced, including two wrapper feature selection techniques for SVM classification utilizing the penalty function27.
Let us consider \(\left\{\left({x}_{1},{y}_{1}\right),\dots ,\left({x}_{n},{y}_{n}\right)\right\},{x}_{i}\in {\mathfrak{R}}^{d},{y}_{i}\in \{-\mathrm{1,1}\}\) be the training data (x is the input data, and y is the binary outcome variable. The following linear boundary function separates the classes of the outcome variable in the linear SVM problem:
The "wj" is the regression coefficient of the obtained hyperplane, and "b" stands for its intercept.
Then the assignment rule for the test dataset to each class is given as: \({y}_{\text{test }}=\mathrm{sign}\left[f\left({x}_{\text{test}}\right)\right]\)
In the above problem, the finding of the optimal hyperplane is conducted by convex optimization. In the penalized version, maximizing the margins or optimization is achieved by the following penalized problem:
Here the \({\mathrm{pen}}_{\lambda }(w)\) is the LASSO (least absolute shrinkage and selection operator) and SCAD (Smoothly Clipped Absolute Deviation).
Penalized logistic regression
The logistic regression model considers a linear relationship between predictors (here, gene profiles) with a binary (dichotomous) outcome (here, having colorectal cancer liver metastasis or being a healthy control)28. So, the outcome, which variable takes y = 1 or y = 0, is considered to have a Bernoulli distribution with \(P(y=1)=\pi =\frac{\mathrm{exp}\left({\beta }^{T}X\right)}{1+\mathrm{exp}\left({\beta }^{T}X\right)}\). Here, \(\beta\) is the vector of regression coefficients, including an intercept term, and X is the data matrix of the gene expression profile of the patients and healthy control. Therefore, considering the logit transform, we have the following regression form:
Then, the parameter estimation is conducted by the maximum likelihood estimation by considering the following log-likelihood function:
Like the P-SVM, to handle the high dimension problem (having a greater number of predictors than the sample size), the penalized likelihood is used for variable selection. The penalty terms can have different forms. Here, we considered Lasso and SCAD (Smoothly Clipped Absolute Deviation).
LASSO and SCAD penalties
The LASSO approach imposes a constraint on the total of the absolute values of the model parameters; the sum must be smaller than a predetermined value29: (in the equation below, λ ≥ 0 is the tuning parameter).
In terms of variable selection, LASSO and SCAD have similar performances30. SCAD penalty, suggested by Fan and Li in 200131, is used to reduce the bias while estimating large regression coefficients32:
The tuning parameters in this equation are λ ≥ 0 and a > 2. In this study, we set the tuning parameter α's value at 3.7. However, the lambda parameter was tuned by the cross-validation method.
Evaluating the performance of feature selection techniques
In order to assess the efficiency of genes selected by each method in differentiating primary samples from metastatic tumors, we applied the Artificial Neural Network (ANN) method using the Multilayer Perceptron procedure in SPSS 24.0. Algorithms with an area under the ROC curve (AUC) > 0.9 were opted, and commonly selected features by these algorithms were considered as the main DEGs for further analysis.
Establishment of the SVM model
SVM is a supervised ML algorithm that is mainly used for data categorization33. This algorithm distinguishes sample type by estimating the degree of a sample that belongs to a specific class34. As a part of this study, We constructed an SVM classifier based on selected features for the GSE41568 training set using the "e1071" package in R35. The SVM classifier's efficacy was assessed on the training and three independent validation sets (GSE68468, GSE41258, and TCGA COAD-READ).
Survival analysis to identify genes with prognostic value
Disease-free survival (DFS) and overall survival (OS) analyses were carried out to identify genes with prognostic significance. In this regard, for OS analysis, 644 TCGA COAD-READ samples were categorized into high-expression and low-expression groups based on the optimal cut-off points determined by the "survminer" and "maxstat" R packages. Using the "survival" package and P value < 0.05, we conducted a Kaplan–Meier survival analysis with a log-rank test to determine which genes are associated with overall survival.
Transcription factor-DEGs network construction
Toward identifying the transcription factors (TFs) of the key genes, we utilized the NetworkAnalyst online tool36. NetworkAnalyst is a web-based application for comprehensive gene expression profiling and meta-analysis using network-based visual analytics36. To construct the TF-gene network, the final DEGs were submitted to NetworkAnalyst to collect information on TF-gene and microRNA-gene interactions, and the resultant list of datasets was exported to Cytoscape software (version 3.7.1) for additional analysis.
Drug–DEGs interaction network
The Drug Gene Interaction Database (DGIdb) (https://www.dgidb.org/) was utilized to find the potential drugs that target the final genes. This database is linked to 22 different databases. To find drug–DEG interactions in the current investigation, only empirically verified interactions were examined.
Sample collection
40 CRC samples, including 16 stage IV CRC samples and 24 CRC samples from other stages, were obtained from Iranian patients who underwent surgery in Mortaz Hospital in Yazd, Iran. All tumor samples were preserved at – 80 °C until the RNA extraction process. (Table 2). Also, five liver metastasis paraffin-embedded samples were received from the archive of the Cancer Institute of Imam Khomeini Hospital. The Study procedure was authorized by The Hamadan University of Medical Science Ethics Committee (ethical code: IR.UMSHA.REC.1400.530). Additionally, informed consent was acquired from all participating patients in this study. The procedures were carried out in compliance with the Helsinki Declaration's laws and recommendations.
Experimental validation: real-time PCR assay
We extracted total cellular RNA from both fresh frozen and paraffin-embedded tissues using the RNX kit (RNX, Cina Gene Company, Iran). Then Yekta Tajhiz cDNA Synthesis Kit (Yekta Tajhiz Azama, Iran) was used to convert the extracted RNA into cDNA. The quantitative reverse transcription PCR (RT-qPCR) was carried out on each sample in duplicate using SYBR Green Master Mix Kit (©Ampliqon, Herlev, Denmark) in a LightCycler 96 Real-Time PCR detection system (Roche, United States) in accordance with the manufacturer’s guidelines. The primer sequence of four evaluated genes in this study is presented in Table 3. Among 11 feature genes, we selected four with the highest AUC in the SVM model (Supplementary Figure 1) and prognostic value. In this analysis, GAPDH was used as the reference gene. Finally, we calculated the relative expression levels of the genes using the 2−△△Ct method37. The gene expression levels were assessed and compared among three distinct groups: Stage IVCRC samples, CRC samples from stages I to III, and liver metastasis samples.
Statistical analysis
The data were analyzed utilizing the R programming language, SPSS 24.0, and the GraphPad Prism 9.0 software. The differences between expression values of the three sample groups were evaluated through a one-way ANOVA test. Statistical significance was determined using the thresholds of *P value < 0.05, **P value < 0.01, and ***P value < 0.001 for all statistical tests.
Ethical approval
Ethical issues (Including plagiarism, informed consent, misconduct, data fabrication and/or falsification, double publication and/or submission, redundancy, etc.) have been completely observed by the authors.
The ethical protocol of this study was approved by the Ethics Committee of Hamadan University of Medical Sciences. (Ethical code: IR.UMSHA.REC.1400.530.) and written informed consent was obtained from all patients to participate in the study.
Results
Identification of DEGs related to CRC liver metastasis in the datasets
Details of selected databases are summarized in Table 1. The gene expression levels of selected samples are shown in Fig. 2. Samples in the GSE41568 dataset were divided into two groups: primary tumor and metastatic tumor, and gene expression analysis was carried out (cut-off criteria: |logFC|≥ 1 and FDR < 0.05). This analysis identified 496 DEGs containing 393 upregulated and 103 downregulated genes, comparing metastatic and non-metastatic samples. To investigate the biological functions of all expressed genes between primary and metastatic CRC patients in the GSE41568 dataset, the GSEA method was employed using the ‘hallmarks’ gene set. As shown in Fig. 3, these genes were enriched in pathways including “Hedgehog signaling”, “Hypoxia”, “ Complement”, “ Epithelial-Mesenchymal Transition”, “ KRAS signaling Down”, and “ Angiogenesis” pathways (nominal P value < 0.05).
Gene set enrichment analysis of all expressed genes between primary and metastatic CRC patients in the GSE41568. This analysis demonstrated that these genes are enriched in (A) Hedgehog signaling, (B) Hypoxia, (C) Complement, (D) Epithelial-Mesenchymal Transition, (E) KRAS signaling Down, and (F) Angiogenesis pathways with nominal P value < 0.05.
GO, KEGG pathway analysis
GO categorizes genes according to their molecular function (MF), biological process (BP), and cellular component (CC)38. The DEGs were most strongly related to the BP of “humoral immune response” (GO:0006959), CC of “blood microparticle” (GO:0072562), and MF of “peptidase regulatory activity” (GO:0052547), according to the analysis of the GO terms. Remarkably, most of the top enriched gene ontology BP, CC, and MF terms were related to a lipid metabolic process, including the “fatty acid metabolic process” and “peptidase activity”. Moreover, the screened DEGs were considerably enriched in the “complement and coagulation cascade pathway” (Fig. 4).
Selecting the features
Machine learning algorithms of RF, P-SVM, and logistic regressions with LASSO and SCAD penalties were employed for selecting features associated with metastasis among 496 screened DEGs. The LASSO method applied by the “glmnet”, P-SVM method by the “penalizedSVM” package, and SCAD by “grpreg” package in R selected 20, 32, and 6 features, respectively, as the most relevant features. Also, 43 features were selected by random forest algorithm. Genes selected by each algorithm are listed in supplementary table 1.
Using ANN to compare variable selection methods
A multi-layer perceptron artificial neural network was utilized to compare the accuracy of feature selection methods. In this regard, ANN was trained with 70% of the data based on features selected by each algorithm, a hold-out validation technique was applied in SPSS, and the performance of the algorithms was evaluated with respect to the area under the receiver operating characteristic (ROC) curve. The AUC results revealed that P-SVM and LASSO are the most accurate models, with an AUC of 0.94 and 0.9, respectively. Roc curves of different algorithms are presented in Fig. 5. Features picked by both LASSO and P-SVM algorithms were considered as the main features for this study which were MMP3, TNFSF11, WNT5A, EPHA3, WNT11, CXCR4, MAP2, MAB21L2, FOXC1, TMEM158, PDE4D. These 11 genes may have the potential to be considered as a diagnostic panel for colorectal cancer metastasis.
Development and verification of predictive SVM model based on feature genes
The 11 mentioned (previous paragraph) genes from the GSE41568 training dataset were used to construct an SVM predictive classification model. The C-Classification SVM method was applied with Radial Based Function (RBF) kernel and tenfold cross-validation. The AUC in the GSE41568 was 1, standing for sensitivity and specificity of 100% (Fig. 6). We also used three external datasets (GSE68468, GSE41258, and TCGA COAD-READ) to evaluate the model. In the GSE68468 and the GSE41258 validation sets, the AUC was 0.75 and 0.77, respectively. TCGA COAD-READ dataset was used as the other validation set. We divided the samples into two groups (M0 and M1) based on the TNM staging characteristics of each sample. The AUC for this validation dataset was 0.75.
Identification of genes with prognostic value
Survival analysis was conducted by the optimal cut-off value. The results showed that seven of eleven feature genes were significantly related to the poor prognosis of CRC patients. In this case, WNT5a (P value < 0.0001), TNFSF11 (P value = 0.0015), MMP3 (P value = 0.0018), and MAP2 (P value = 0.0038) were the most significant genes predicting poor OS in CRC patients. The findings show that OS is lower in patients with low expression of MMP3, WNT5a, and TNFSF11. Also, patients with high expression of WNT11 had lower OS. The association between the expression of these genes with OS of CRC patients is presented in Fig. 7.
The association between expression of (A) EPHA3 (P value = 0.049), (B) FOXC1 (P value = 0.0045), (C) MAP2 (P value = 0.0038), (D) MMP3 (P value = 0.0018), (E) TNFSF11 (P value = 0.0015), (F) WNT5a (P value < 0.0001), and (G) WNT11 (P value = 0.0077) with overall survival of all patients in the TCGA COAD-READ dataset. The red line indicates high expression groups and the blue line represents the low expression group.
Transcription factors modulating feature genes
To uncover other underlying TFs regulating the selected genes, the TF–DEGs network was constructed using the networkanalyst tool and ENCODE database. According to this database, a total of 65 TFs were found to be related to the feature genes. The constructed network was visualized by Cytoscape and is depicted in Fig. 8. Seven genes had known interacting transcription factors. The resultant network shows that EZH2, the most interacting transcription factor in this network, regulates five feature genes.
The TF–DEGs interaction network. According to this network,65 TFs were found to be interacting with seven of selected genes. Among them EZH2 interacts with five genes and could be considered as the most important TF in this network. TFs are shown in different colors standing for their importance and interaction (this figure was drawn in the Cytoscape v.3.8.2 software).
Drug-DEGs network
The list of selected genes was imported into the DGIdb database to investigate any FDA-approved drugs related to these genes. Among feature genes MMP3, EPHA3, MAP2, TNFSF11, CXCR4 and PDE4D had approved targeting drugs. A total of 30 drugs were found, nine of which were antineoplastic, including Lenalidomide, Anastrozole, Letrozole, Colchicine, Plerixafor, Bevacizumab, Cisplatin, and Vandetanib. The drug-gene network was illustrated using Cytoscape and is represented in Fig. 9.
Illustration of the drug–gene interaction network. Totally, 27 candidate drugs were identified as modulators of the selected genes using DGIdb database. The red and green circles represent drugs and blue hexagon shapes represent genes. Red circle represents antineoplastic drugs. (this figure was drawn in the Cytoscape43 v.3.8.2 software).
Experimental validation using qRT-PCR:
In the final section of our study, we investigated the gene expression levels of MMP3, WNT5a, WNT11, and TNFSF11 genes. Analysis between three research groups revealed that expression of the MMP3 gene was significantly lower in the liver metastasis group compared to other groups. The expression of this gene was also lower in the stage IV CRC group compared to CRC samples from other stages. WNT11 was the other gene that showed a significant expression alternation in different groups. The results showed that this gene overexpressed significantly in liver metastases compared to stage 4 and stages 1,2,3 samples. This gene was also expressed in higher levels in stage 4 CRC samples compared to stages 1,2,3 CRC group. The gene expression levels of WNT5a and TNFSF11 were also significantly lower in liver metastases compared to stage 4 and stages 1,2,3 CRC groups but showed no significant expression alternations between stage 4 and stages1,2,3 CRC samples (Fig. 10).
Discussion
Ninety percent of cancer-related fatalities are due to metastatic spread39. Current cancer therapies are ineffectual for metastatic cancer because standard imaging tools cannot detect the disease in its early stages. In addition, the capacity to forecast cancer's ability to metastasize in advance will help to improve patient prognosis40. Therefore, it becomes crucial to investigate possible biomarkers with prognostic significance for metastatic CRC.
In the present study, we identified 11 distant metastasis-related genes using machine learning algorithms, seven of which substantially correlated with the survival of CRC patients. These genes were also utilized to construct an SVM predictive classification model with AUC = 1. In summary, 496 DEGs were screened by comparing the gene expression profiles of primary tumors with CRC liver metastases. KEGG enrichment analysis on screened DEGs revealed that “complement and coagulation cascades” is the most significant enriched pathway related to the primary DEGs. Recent studies have uncovered mounting evidence that complement and coagulation cascades are involved in angiogenesis, tumor cell proliferation, immune response suppression, and metastasis41,42.
Subsequently, we used two machine learning algorithms, P-SVM and LASSO, to select the features. Features picked by both algorithms were considered for further analysis. The results demonstrated that MMP3, WNT11, WNT5a, and TNFSF11 might have essential roles in CRC metastasis. Among them, WNT11 and WNT5a, WNT family members, regulate cell fate, proliferation, migration, and cell death in various ways and are engaged in the process of carcinogenesis and embryogenesis. Several studies have highlighted the significance of these genes in CRC development and metastasis. For instance, Fujii et al. found that WNT5a upregulation promotes the EMT in HT29 cells43. In another study, it was shown that WNT5a suppression by miR-21b initiates the metastatic process in CRC cells44. It's interesting to note that Ki et al.45 demonstrated that the WNT5a expression level is higher in primary tumors than in normal colon samples, while it has significantly lower expression in liver metastasis tumors. WNT11 is another member of the WNT family. Noncanonical WNT11 signaling promotes proliferation and morphological alternations in the intestinal epithelial cells. Several studies have revealed that the WNT11 expression level elevates in CRC and increases the 5-year mortality rates46,47,48,49. This gene is positively related to cell migration and invasion in CRC cells, increasing the likelihood of metastasis. Gorroño-Etxebarria et al. proved that WNT11 is highly expressed in CRC liver metastasis samples through immunohistochemical staining. Besides, some studies have demonstrated the significance of WNT11 in other cancers. For example, Arisen et al. indicated that upregulated WNT11 promotes EMT in aggressive prostate cancer cells50.
TNFRSF11 (RANK) was first found to have a role in bone dissolution and lymph node formation, majorly via the RANK/RANKL/OPG pathway51. Current studies have proved the critical role of RANK/RANKL/OPG in cell migration and invasion. Additionally, it has been observed that TNFRSF11(RANK) is engaged in the development of several types of cancers, including lung cancer52, prostate cancer53, renal cancer54, breast cancer55, and melanoma56. On the other hand, several investigations have shown that the RANKL/RANK system promotes both primary carcinogenesis and metastasis through osteoclast-independent mechanisms57. Furthermore, Ahern et al. found that TNFSF11 knockdown enhances the anti-metastatic effect of antibodies targeting PD1/PD-L1 and suppresses the growth of the subcutaneous tumors of colon cancer animal models58.
MMP3, also known as stromelysin-1, is a member of matrix metalloproteinase (MMP). MMP3 participates in several cellular biological processes, including cell differentiation and inflammation. This gene contributes to the onset and progression of several disorders. MMP3 is capable of degrading ECM, which facilitates tumor invasion and metastasis. Different studies have revealed MMP3 expression alternation and its role in metastasis in various cancers, such as osteosarcoma59 and ovarian cancer60. Moreover, MMP3 expression was also discovered to be substantially elevated in malignant colorectal tumors compared with normal tissue61.
Interestingly, very few studies have shown that MMP3 is downregulated in metastatic lesions compared with primary tumors of different cancers. Maiti et al. revealed that MMP3 downregulates significantly in metastatic sites vs. primary breast tumors using qPCR (P value = 0.0001). In addition, they discovered a correlation between MMP3 and the prognosis of breast cancer patients62. Another study on this issue was conducted by wang et al. to figure out essential DEGs in CRC metastasis using NGS profiling on primary colorectal tumor samples from CRC patients with and without liver metastases and validated their findings by qPCR and immunostaining. The results of this research indicated that MMP3 significantly downregulates in samples with liver metastasis63. The current study found clear support for these findings. We observed that the MMP3 expression level was significantly lower in high-stage tumor samples compared with low-stage samples. This is in contrary to the findings claiming that the expression level of this gene is not associated with the tumor stages64.
The other part of our study was identifying vital transcription factors involved in CRC metastasis. We found that EZH2 modulates five interacting DEGs and could be considered an essential TF in this process. EZH2 is an inhibitory transcription factor that plays a role in the histone methylation process65. This protein takes part in the formation of heterochromatin structure, which causes gene silencing66. Various studies have been conducted to indicate EZH2 involvement in different cancers' progression and metastasis. In a study by Zheng et al., they represented that this transcription factor plays a direct role in breast cancer bone metastasis through the TGF-B pathway. They observed that EZH2 knockout in mice prevented bone metastasis67. Also, Chen et al. reported that EZH2 is responsible for poor prognosis in CRC. They proved that EZH2 is upregulated in colorectal tumor tissues by qRT-PCR and western blot analysis. Additionally, they observed that high EZH2 expression was substantially linked with tumor stage, tumor size, histological differentiation, and lymph node metastasis68. Nevertheless, additional studies are required to investigate the role of this TF in CRC metastasis.
In the last part of this study, we investigated the DGIdb database for possible drugs targeting our final genes. In this regard, nine antineoplastic drugs were found. Among them, Bevacizumab, sold under the brand name Avastin is an approved drug for the treatment of metastatic CRC69. Lenalidomide and Plerixafor have also shown potential for metastatic CRC treatment. In a study by Galustian et al., they proved that Lenalidomide can inhibit metastatic CRC in vivo and in vitro70. On the other hand, Plerixafor has completed phase one trials for metastatic CRC71. Other drugs in the drug-DEGs network in this study have also found to be effective in the treatment of different metastatic cancers, such as breast cancer72 and thyroid carcinoma73. These drugs may have the potential to be repurposed as a treatment for metastatic CRC, providing new options for patients and physicians.
The mortality rate of cancer is closely linked to the stage of cancer progression, highlighting the possibility of reducing mortality through early detection and management74. This imperative is underscored by a substantial decrease in the 5-year survival rate, notably in instances of distant metastases, where the survival rate plummets to 10%75,76. The necessity to identify precise genes and pathways is increased aiming for timely diagnosis and personalized therapeutic strategies to effectively confront the intricacies of cancer progression and metastasis77. Importantly, the identification of stage-specific biomarkers in colorectal cancer is of great importance in this context, as it considerably enhances our ability for early detection and targeted interventions, thereby contributing significantly to addressing the challenges associated with colorectal cancer78,79.
The integration of ML algorithms into the analysis of existing datasets holds promising potential for identifying stage-specific biomarkers in colorectal cancer80. this advanced computational approach not only supports the necessity of early detection and intervention but also improves the accuracy and effectiveness of the biomarker selection process81. The application of machine learning, including algorithms like LASSO and P-SVM, introduces a nuanced and targeted methodology, augmenting our capability to discern key biomarkers associated with different stages of colorectal cancer27,82,83. This innovative approach represents an important step toward refining our understanding of cancer progression, establishing a foundation for the development of more effective diagnostic and therapeutic strategies tailored to specific stages of the disease84.
Although further investigations are needed, the present study contributes to a better understanding of CRC metastasis. It is crucial to consider the limitations of this study when interpreting its findings, including low liver metastases sample size due to the scarcity of liver metastasis samples of CRC from participating hospitals. This constraint may have affected the study's ability to detect statistically significant differences and limits the generalizability of the results.
Conclusion
We employed two machine learning algorithms to identify biomarkers associated with CRC metastasis. Through these methods, a total of 11 biomarkers were identified, and four of them were experimentally validated. Also, the SVM model based on these 11 feature genes showed the optimal classification performance in identifying CRC liver metastasis samples. The joint application of these genes could be considered as a diagnostic panel for metastasis assessment, further augmented by the development of an innovative AI predictive model based on these genetic signatures. These findings make a significant contribution to the continuous pursuit of deeper comprehension regarding the intrinsic molecular mechanisms driving CRC metastasis, with potential implications for the advancement of more efficacious diagnostic and therapeutic strategies tailored to this affliction. Additionally, the use of machine learning approaches in this study highlights the potential of this method for identifying biomarkers in complex biological systems.
Data availability
The corresponding author can provide the datasets utilized in this study on a reasonable request. The datasets analyzed during this study are available in the GEO database (https://www.ncbi.nlm.nih.gov/geo/database with GSE41568, GSE41258 and GSE68468 accession numbers) and TCGA database (https://portal.gdc.cancer.gov/).
Abbreviations
- CRC:
-
Colorectal cancer
- DEG:
-
Differentially expressed gene
- mRNA:
-
Messenger RNA
- ML:
-
Machine learning
- DL:
-
Deep learning
- AI:
-
Artificial intelligence
- COAD:
-
Colon adenocarcinoma
- READ:
-
Rectal adenocarcinoma
- GO:
-
Gene ontology
- KEGG:
-
Kyoto Encyclopedia of Genes and Genomes
- GSEA:
-
Gene set enrichment analysis
- LASSO:
-
Least absolute shrinkage and selection operator
- SVM:
-
Support vector machine
- RF:
-
Random Forest
- ROC:
-
Receiver operating characteristic
- AUC:
-
Area under ROC curve
References
Morgan, E. et al. Global burden of colorectal cancer in 2020 and 2040: Incidence and mortality estimates from GLOBOCAN. Gut 72(2), 338–344 (2023).
Zeng, X., Ward, S. E., Zhou, J. & Cheng, A. S. L. Liver immune microenvironment and metastasis from colorectal cancer-pathogenesis and therapeutic perspectives. Cancers 13(10), 2418 (2021).
Maspero, M. et al. Liver transplantation for hepatic metastases from colorectal cancer: Current knowledge and open issues. Cancers 15(2), 345 (2023).
Pavel, M.-C. et al. Impact of neoadjuvant chemotherapy on post-hepatectomy regeneration for patients with colorectal cancer liver metastasis—Systematic review and meta-analysis. Eur. J. Surg. Oncol. 49, 533–541 (2023).
Hasan Abdali, M. et al. Investigating the effect of radiosensitizer for ursolic acid and kamolonol acetate on HCT-116 cell line. Bioorg. Med. Chem. 28(1), 115152 (2020).
Zheng, W. et al. Emerging mechanisms and treatment progress on liver metastasis of colorectal cancer. Onco. Targets. Ther. 14, 3013–3036 (2021).
McAuliffe, J. C., Qadan, M. & D’Angelica, M. I. Hepatic resection, hepatic arterial infusion pump therapy, and genetic biomarkers in the management of hepatic metastases from colorectal cancer. J. Gastrointest. Oncol. 6(6), 699 (2015).
Patz, E. F. Integration of biomarkers and imaging. J. Thorac. Oncol. 1(1), 78–80 (2006).
Zhu, H.-q et al. Diagnostic value of an enhanced MRI combined with serum CEA, CA19-9, CA125 and CA72-4 in the liver metastasis of colorectal cancer. World J. Surg. Oncol. 20(1), 401 (2022).
Sheykhhasan, M. et al. FLVCR1-AS1 and FBXL19-AS1: Two putative lncRNA candidates in multiple human cancers. Non-Coding RNA. 9(1), 1 (2022).
Loktionov, A. Biomarkers for detecting colorectal cancer non-invasively: DNA, RNA or proteins?. World J. Gastrointest. Oncol. 12(2), 124–148 (2020).
He, J. et al. Biomarkers (mRNAs and non-coding RNAs) for the diagnosis and prognosis of colorectal cancer—From the body fluid to tissue level. Front. Oncol. 11, 632834 (2021).
Fang, C. et al. CD133+ CD54+ CD44+ circulating tumor cells as a biomarker of treatment selection and liver metastasis in patients with colorectal cancer. Oncotarget 7(47), 77389 (2016).
Agrawal, R. & Prabakaran, S. Big data in digital healthcare: Lessons learnt and recommendations for general practice. Heredity 124(4), 525–534 (2020).
Zhang, H. et al. Differential diagnosis of hematologic and solid tumors using targeted transcriptome and artificial intelligence. Am. J. Pathol. 193(1), 51–59 (2023).
Janiesch, C., Zschech, P. & Heinrich, K. Machine learning and deep learning. Electron. Mark. 31(3), 685–695 (2021).
Arjmand, B. et al. Machine learning: A new prospect in multi-omics data analysis of cancer. Front. Genet. https://doi.org/10.3389/fgene.2022.824451 (2022).
Samadi, P. et al. An integrative transcriptome analysis reveals potential predictive, prognostic biomarkers and therapeutic targets in colorectal cancer. BMC Cancer 22(1), 1–22 (2022).
Langfelder, P. & Horvath, S. WGCNA: An R package for weighted correlation network analysis. BMC Bioinform. 9, x0026.559 (2008).
Ritchie, M. E. et al. limma powers differential expression analyses for RNA-sequencing and microarray studies. Nucleic Acids Res. 43(7), e47 (2015).
Yu, G., Wang, L.-G., Han, Y. & He, Q.-Y. clusterProfiler: An R package for comparing biological themes among gene clusters. Omics J. Integr. Biol. 16(5), 284–287 (2012).
Walter, W., Sánchez-Cabo, F. & Ricote, M. GOplot: An R package for visually combining expression data with functional analysis. Bioinformatics. 31(17), 2912–2914 (2015).
Taghizadeh, E., Heydarheydari, S., Saberi, A., JafarpoorNesheli, S. & Rezaeijo, S. M. Breast cancer prediction with transcriptome profiling using feature selection and machine learning methods. BMC Bioinform. 23(1), 410 (2022).
Li, J. et al. Feature selection: A data perspective. ACM Comput. Surv. 50(6), 1–45 (2017).
Breiman, L. Random forests. Mach. Learn. 45(1), 5–32 (2001).
Tapak, L., Afshar, S., Afrasiabi, M., Ghasemi, M. K. & Alirezaei, P. Application of genetic algorithm-based support vector machine in identification of gene expression signatures for psoriasis classification: A hybrid model. BioMed Res. Int. 2021, 5520710 (2021).
Becker, N., Werft, W., Toedt, G., Lichter, P. & Benner, A. penalizedSVM: A R-package for feature selection SVM classification. Bioinformatics. 25(13), 1711–1712 (2009).
Wang, Z., Sun, X., Wang, B., Shi, S. & Chen, X. Lasso-Logistic regression model for the identification of serum biomarkers of neurotoxicity induced by strychnos alkaloids. Toxicol. Mech. Methods. 33(1), 65–72 (2023).
Fonti, V. & Belitser, E. Feature selection using lasso. VU Amsterdam Res. Pap. Bus. Anal. 30, 1–25 (2017).
Lee, Y. & Oh, H.-S. A new sparse variable selection via random-effect model. J. Multivar. Anal. 125, 89–99 (2014).
Fan, J. & Li, R. Variable selection via nonconcave penalized likelihood and its oracle properties. J. Am. Stat. Assoc. 96(456), 1348–1360 (2001).
Ma, S. & Huang, J. Penalized feature selection and classification in bioinformatics. Brief. Bioinform. 9(5), 392–403 (2008).
Moradi, S., Tapak, L. & Afshar, S. Identification of novel noninvasive diagnostics biomarkers in the Parkinson’s diseases and improving the disease classification using support vector machine. BioMed Res. Int. 2022, 5009892 (2022).
Hu, M. et al. Construction of a 5-feature gene model by support vector machine for classifying osteoporosis samples. Bioengineered. 12(1), 6821–6830 (2021).
Meyer, D. et al. Package ‘e1071’. R J. (2019).
Zhou, G. et al. NetworkAnalyst 3.0: A visual analytics platform for comprehensive gene expression profiling and meta-analysis. Nucleic Acids Res. 47(W1), W234–W241 (2019).
Livak, K. J. & Schmittgen, T. D. Analysis of relative gene expression data using real-time quantitative PCR and the 2−ΔΔCT method. Methods. 25(4), 402–408 (2001).
Carbon, S. et al. AmiGO: Online access to ontology and annotation data. Bioinformatics. 25(2), 288–289 (2009).
Ronaldson-Bouchard, K. et al. Engineering complexity in human tissue models of cancer. Adv. Drug Deliv. Rev. 184, 114181 (2022).
Ganesh, S., Venkatakrishnan, K. & Tan, B. Early detection and prediction of cancer metastasis—Unravelling metastasis initiating cell as a dynamic marker using self-functionalized nanosensors. Sens. Actuators B Chem. 361, 131655 (2022).
Lu, C. et al. Construction of a novel mRNA-miRNA-lncRNA network and identification of potential regulatory axis associated with prognosis in colorectal cancer liver metastases. Aging. 13(11), 14968–14988 (2021).
Zhang, T. et al. Identifying the key genes and microRNAs in colorectal cancer liver metastasis by bioinformatics analysis and in vitro experiments. Oncol. Rep. 41(1), 279–291 (2019).
Fujii, K. et al. Pro-metastatic intracellular signaling of the elaidic trans fatty acid. Int. J. Oncol. 50(1), 85–92 (2017).
Fan, D. et al. MicroRNA 26b promotes colorectal cancer metastasis by downregulating phosphatase and tensin homolog and wingless-type MMTV integration site family member 5A. Cancer Sci. 109(2), 354–362 (2018).
Ki, D. H. et al. Whole genome analysis for liver metastasis gene signatures in colorectal cancer. Int. J. Cancer. 121(9), 2005–2012 (2007).
Gorroño-Etxebarria, I. et al. Wnt-11 as a potential prognostic biomarker and therapeutic target in colorectal cancer. Cancers. 11(7), 908 (2019).
Ouko, L., Ziegler, T. R., Gu, L. H., Eisenberg, L. M. & Yang, V. W. Wnt11 signaling promotes proliferation, transformation, and migration of IEC6 intestinal epithelial cells. J. Biol. Chem. 279(25), 26707–26715 (2004).
He, D. et al. Long noncoding RNA ABHD11-AS1 promote cells proliferation and invasion of colorectal cancer via regulating the miR-1254-WNT11 pathway. J. Cell. Physiol. 234(7), 12070–12079 (2019).
Ji, Y., Lv, J., Sun, D. & Huang, Y. Therapeutic strategies targeting Wnt/β-catenin signaling for colorectal cancer. Int. J. Mol. Med. 49(1), 1–17 (2022).
Arisan, E. D. et al. Upregulated Wnt-11 and miR-21 expression trigger epithelial mesenchymal transition in aggressive prostate cancer cells. Biology. 9(3), 52 (2020).
Fan, Y. et al. Association of genetic polymorphisms in TNFRSF11 with the progression of genetic susceptibility to gastric cancer. J. Oncol. 2020, 4103264 (2020).
Ahern, E. et al. Pharmacodynamics of pre-operative PD1 checkpoint blockade and receptor activator of NFkB ligand (RANKL) inhibition in non-small cell lung cancer (NSCLC): Study protocol for a multicentre, open-label, phase 1B/2, translational trial (POPCORN). Trials. 20(1), 1–9 (2019).
Christoph, F. et al. RANKL/RANK/OPG cytokine receptor system: mRNA expression pattern in BPH, primary and metastatic prostate cancer disease. World J. Urol. 36(2), 187–192 (2018).
Bernardi, S. et al. TRAIL, OPG, and TWEAK in kidney disease: Biomarkers or therapeutic targets?. Clin. Sci. 133(10), 1145–1166 (2019).
Wu, X. et al. RANKL/RANK system-based mechanism for breast cancer bone metastasis and related therapeutic strategies. Front. Cell Dev. Biol. 8, 76 (2020).
Ferguson, J. et al. Osteoblasts contribute to a protective niche that supports melanoma cell proliferation and survival. Pigment Cell Melanoma Res. 33(1), 74–85 (2020).
Okamoto, K. Role of RANKL in cancer development and metastasis. J. Bone Miner. Metab. 39(1), 71–81 (2021).
Ahern, E. et al. RANKL blockade improves efficacy of PD1-PD-L1 blockade or dual PD1-PD-L1 and CTLA4 blockade in mouse models of cancer. Oncoimmunology. 7(6), e1431088 (2018).
Huang, J.-F., Du, W.-X. & Chen, J.-J. Elevated expression of matrix metalloproteinase-3 in human osteosarcoma and its association with tumor metastasis. J. BUON. 21(1), 235–243 (2016).
Zheng, J., Zhou, Y., Li, X. & Hu, J. MiR-574-3p exerts as a tumor suppressor in ovarian cancer through inhibiting MMP3 expression. Eur. Rev. Med. Pharmacol. Sci. 23(16), 6839–6848 (2019).
Yu, J. et al. Comprehensive analysis of the expression and prognosis for MMPs in human colorectal cancer. Front. Oncol. https://doi.org/10.3389/fonc.2021.771099 (2021).
Maiti, A. et al. Altered expression of secreted mediator genes that mediate aggressive breast cancer metastasis to distant organs. Cancers. 13(11), 2641 (2021).
Wang, S. et al. Transcriptome analysis in primary colorectal cancer tissues from patients with and without liver metastases using next-generation sequencing. Cancer Med. 6(8), 1976–1987 (2017).
Busuioc, C. et al. Analysis of differentially expressed genes, MMP3 and TESC, and their potential value in molecular pathways in colon adenocarcinoma: A bioinformatics approach. BioMedInformatics. 2(3), 474–491 (2022).
Viré, E. et al. The polycomb group protein EZH2 directly controls DNA methylation. Nature. 439(7078), 871–874 (2006).
Wu, S., Yin, Y. & Wang, X. The epigenetic regulation of the germinal center response. Biochim. Biophys. Acta Gene Regul. Mech. 1865(6), 194828 (2022).
Zhang, L. et al. EZH2 engages TGFβ signaling to promote breast cancer bone metastasis via integrin β1-FAK activation. Nat. Commun. 13(1), 2543 (2022).
Chen, Z. et al. Expression of EZH2 is associated with poor outcome in colorectal cancer. Oncol. Lett. 15(3), 2953–2961 (2018).
Bevacizumab. (2006).
Liu, W. M. et al. Inhibition of metastatic potential in colorectal carcinoma in vivo and in vitro using immunomodulatory drugs (IMiDs). Br. J. Cancer. 101(5), 803–812 (2009).
Martin, M. et al. At the Bedside: Profiling and treating patients with CXCR4-expressing cancers. J. Leukoc. Biol. 109(5), 953–967 (2020).
Mehta, R. S. et al. Overall survival with fulvestrant plus anastrozole in metastatic breast cancer. N. Engl. J. Med. 380(13), 1226–1234 (2019).
Leboulleux, S. et al. Vandetanib in locally advanced or metastatic differentiated thyroid cancer: A randomised, double-blind, phase 2 trial. Lancet Oncol. 13(9), 897–905 (2012).
Yang, J. D. et al. A global view of hepatocellular carcinoma: Trends, risk, prevention and management. Nat. Rev. Gastroenterol. Hepatol. 16(10), 589–604 (2019).
Shokrollah, N. et al. A systems biology approach to identify novel biomarkers in progression from Crohn’s disease to colorectal cancer. Asian Pac. J. Cancer Prev. APJCP. 24(6), 1993–2001 (2023).
He, J.-H. et al. A nomogram model for predicting distant metastasis of newly diagnosed colorectal cancer based on clinical features. Front. Oncol. https://doi.org/10.3389/fonc.2023.1186298 (2023).
Housini, M. et al. Colorectal cancer: Genetic alterations, novel biomarkers, current therapeutic strategies and clinical trails. Gene. 892, 147857 (2023).
Palaniappan, A., Ramar, K. & Ramalingam, S. Computational identification of novel stage-specific biomarkers in colorectal cancer progression. PLoS ONE. 11(5), e0156665 (2016).
Fadaka, A. O. et al. Stage-specific treatment of colorectal cancer: A microRNA-nanocomposite approach. J. Pharm. Anal. https://doi.org/10.1016/j.jpha.2023.07.008 (2023).
Sufyan, M., Shokat, Z. & Ashfaq, U. A. Artificial intelligence in cancer diagnosis and therapy: Current status and future perspective. Comput. Biol. Med. 165, 107356 (2023).
Dhillon, A., Singh, A. & Bhalla, V. K. A systematic review on biomarker identification for cancer diagnosis and prognosis in multi-omics: From computational needs to machine learning and deep learning. Arch. Comput. Methods Eng. 30(2), 917–949 (2023).
Maurya, N. S., Kushwaha, S., Vetukuri, R. R. & Mani, A. Unlocking the potential of the CA2, CA7, and ITM2C gene signatures for the early detection of colorectal cancer: A comprehensive analysis of RNA-Seq data by utilizing machine learning algorithms. Genes. 14(10), 1836 (2023).
Al-Tashi, Q. et al. Machine learning models for the identification of prognostic and predictive cancer biomarkers: A systematic review. Int. J. Mol. Sci. 24(9), 7781 (2023).
Skrede, O.-J. et al. Deep learning for prediction of colorectal cancer outcome: A discovery and validation study. Lancet. 395(10221), 350–360 (2020).
Acknowledgements
We would like to express our deepest gratitude to ProfessorMasoud Saidijam. Our special thanks also go to the staff of Mortaz Hospital for their assistance in providing the necessary data and resources.
Funding
This work is supported by a grant from Hamadan University of Medical Sciences, Hamadan, Iran (No. 140007276125).
Author information
Authors and Affiliations
Contributions
S.A., L.T. and A.A. conceived and designed the analysis. A.A. collected the data. A.A., S.A. and L.T. contributed to analysis tools. A.A. and L.T. performed the analysis. A.A., S.A., L.T., A.T. and F.N. contributed to the interpretation of the results. A.A. wrote the manuscript in consultation with all authors. A.M., S.A. and L.T. edited and revised the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Ahmadieh-Yazdi, A., Mahdavinezhad, A., Tapak, L. et al. Using machine learning approach for screening metastatic biomarkers in colorectal cancer and predictive modeling with experimental validation. Sci Rep 13, 19426 (2023). https://doi.org/10.1038/s41598-023-46633-8
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-023-46633-8
This article is cited by
-
Glycosylation-related genes mediated prognostic signature contribute to prognostic prediction and treatment options in ovarian cancer: based on bulk and single‑cell RNA sequencing data
BMC Cancer (2024)
-
A pan-cancer analysis indicates long noncoding RNA HAND2-AS1 as a potential prognostic, immunomodulatory and therapeutic biomarker in various cancers including colorectal adenocarcinoma
Cancer Cell International (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.