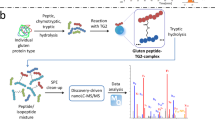
Abstract
Microbial transglutaminase (mTG) is a bacterial survival factor, frequently used as a food additive to glue processed nutrients. As a result, new immunogenic epitopes are generated that might drive autoimmunity. Presently, its contribution to autoimmunity through epitope similarity and cross-reactivity was investigated. Emboss Matcher was used to perform sequence alignment between mTG and various antigens implicated in many autoimmune diseases. Monoclonal and polyclonal antibodies made specifically against mTG were applied to 77 different human tissue antigens using ELISA. Six antigens were detected to share significant homology with mTG immunogenic sequences, representing major targets of common autoimmune conditions. Polyclonal antibody to mTG reacted significantly with 17 out of 77 tissue antigens. This reaction was most pronounced with mitochondrial M2, ANA, and extractable nuclear antigens. The results indicate that sequence similarity and cross-reactivity between mTG and various tissue antigens are possible, supporting the relationship between mTG and the development of autoimmune disorders 150W.
Similar content being viewed by others
Introduction
Genetic predisposition is pivotal for autoimmune diseases (ADs) development, but environmental factors are necessary for their clinical evolvement1,2,3,4. Pending on their association with various ADs, they include: hygiene and diet5, food processed additives6,7,8,9, trace elements10, enteric microbial peptides11, multiple infectious agents3,12,13,14,15, various vaccines16,17, toxic agents or food products4 and recently the checkpoint inhibitors18,19. In fact, many of those environmental factors were cited as part of the autoimmune/inflammatory syndrome induced by adjuvants (ASIA)1,20,21. Zooming into the gastrointestinal tract (GIT), many of the above-mentioned environmental factors inhabit the enteric lumen and are associated with local or peripheral ADs3,4,5,6,7,8,9,10,11,12,13,22.
Many processes were described to operate in the human GIT driving gut-originated autoimmunity. The most reported one is increased gut permeability resulting in leaky gut syndrome6,23,24,25. Among others are posttranslational modification of naïve proteins26, dysbiosis and its harmful mobilome23,27, horizontal gene transfer28, or many immunogenic nutritional compounds, such as gluten4,6,7,8,9,22,27. All those enteric events irradiate peripherally and might induce systemic autoimmunity29.
Indeed, some of those luminal events are blamed to increase the worldwide incidence of ADs23,30. Among the various mechanisms that drive autoimmunity, molecular mimicry is the most reported11,16,31,32. Actually, SARS-CoV-2-associated autoimmunity is suggested to operate through molecular mimicry with self-epitopes33,34.
Transglutaminases are an extensive natural enzymatic family that catalyze the formation of isopeptide bonds by post-translational modification of proteins. In fact, they are considered nature's biological glues35. In the presence of an acyl donor and an acyl acceptor they cross-link the corresponding protein to form a protease-indigestible high molecular mass protein36,37. The accumulated linked complexes can be deposited in various tissues and organs and are involved in multiple human chronic diseases. Inflammatory, cancerous, metabolic, neurodegenerative and ADs7,22,27,38,39 are some examples.
The present study is focused on the frequently consumed processed food additive, namely, microbial transglutaminase (mTG). It is considered a natural family member of the transaminases. Despite having a very low sequence similarity and having a much lower molecular weight, the sequence similarity is much higher on their active site8. Functionally, it imitates the transglutaminases’ cross-linking activity; all of them can deamidate or transamidate their substrates6,7,8,9,22,27,29. In recent years, the mTG enzyme has been reported to functionally join tissue transglutaminase (tTG), which is the autoantigen of celiac disease (CD)6,8,36,37. In fact, mTG has been suggested as a new environmental factor in CD8,9,40,41,42,43,44,45, other ADs22,27,45 and as even being involved in the induction of neurodegenerative diseases7,22,39. Presently, a new aspect of the potential harmful effects of the mTG enzyme in the induction of ADs are being described. The transcytosis of the mTG to the sub-epithelial compartment46, the immune reactions against the mTG-gliadin complexes in CD patients8,9,22,27,39,40,41,42,43,44,45,46,47,48,49,50 and the most recently described resistance of the mTG to oxidative stress in CD51 prompted us to conduct the present study. The cross-reactivity and sequence similarity between mTG and human epitopes have never been explored. The present hypothesis is that cross-reactive antibodies and sequence similarity between the mTG and human self-epitopes further reinforce the relationship of molecular mimicry between the mTG enzyme and the induction of chronic inflammatory, autoimmune and neurodegenerative diseases. Moreover, epitope sharing between these self-proteins and gluten might suggest the involvement of mTG cross-linked gluten complexes in non-celiac chronic conditions.
Results
Cross-reactive polyclonal antibodies to human proteins
The application of affinity-purified polyclonal antibody against mTG to many human tissue antigens resulted in different degrees of reactivity. 57 antigens resulted in ODs of around 0.16 + 0.12, which was very similar to the ELISA background or negative controls.
The cutoff was established based on the optical densities of these 57 antigens, 0.16 + 0.12 = 0.28. Two antigens, cardiolipin and TG6, showed ODs very close to the cutoff point of 0.28. Cardiolipin had an OD of 0.26 and a p value of 0.1, which was not significant. TG6 had an OD of 0.29 and a p value of 0.0004, which was very significant. The higher the OD is above the cutoff point, the more statistically significant the value was. For instance, the next antigen over the cutoff point, somatotropin, had an OD of 0.45 and a p value of 0.0001.
Mitochondria (M2) and ENA had the highest ODs. Although lesser ODs were detected for ANA, TG2, TG6, heparin, α-myosin, chondroitin sulfate, Lupus RO-60, fibrinogen, tyrosinase, β catenin, thyroid peroxidase, claudin 7, sulfatides, somatotropin, S100B, somatostatin and actin, their p values were still very, very significant (Fig. 1).
Rabbit polyclonal antibody to mTG and its reaction to human tissue components expressed as ELISA ODs. This reaction was performed in duplicate, and variation in the ODs between the duplicate wells was less than 7%. As shown, the reaction of this antibody with mitochondrial M2 antigen and ENA resulted in the highest ODs.
Cross reactive mouse monoclonal antibodies to human proteins
The application of affinity-purified mouse monoclonal antibody against mTG to many human tissue antigens resulted in different degrees of reactivity. 67 antigens resulted in ODs of around 0.15 + 0.09, which was very similar to the ELISA background or negative controls. The cutoff was established based on the optical densities of these 67 antigens, 0.15 + 0.09 = 0.24. TG6 had an OD below the cutoff point of 0.24, with an OD of 0.16 and a p value of 0.05, which was not significant. Mitochondria had an OD of 0.33 and a p value of 0.00007, which was very significant. The higher the OD is above the cutoff point, the more statistically significant the value was. For instance, the next antigen over the cutoff point, TG3, had an OD of 0.38 and a p value of 0.0000003.
Somatotropin and ANA had the highest ODs. Although lesser ODs were detected for TG2, DPP IV, somatotropin, somatostatin, aquaporin, ANA, and ENA, their p values were still very, very significant (Fig. 2).
Molecular similarity between mTG and human immune epitopes
Out of 67,000 epitopes of human tissue antigens, 60 were detected to share significant homology with sub-sections of mTG protein. Out of those, six pairs of similar sequences were detected between human epitopes derived from cross-reactive antigens and mTG sequences that were considered as immunogenic with a strong binding affinity to at least one of the HLA-I and HLA-II alleles. The human epitopes were derived from tissue antigens that are implicated in 10 ADs: rheumatoid arthritis (RA), ankylosing spondylitis (AS), autoimmune atherosclerosis (AIAS), psoriatic arthritis (PA), autoimmune thyroiditis (AIT), Sjogren's syndrome (SS), primary biliary cholangitis (PBC), type 1 diabetes mellitus (T1DM), multiple sclerosis (MS), and autoimmune uveitis (AU).
Table 1 presents sequence similarities driven by six antigens. Four antigens relate to RA, three to T1DM, two to SS, and one relates to AIAS, PBC, MS, PA, AU, AIT, and AS. The similarity of paired sequences is displayed in red, (human on top of mTG), and the isolated amino acid (AA) mismatches are marked in black. The alignment cut-off was kept at a minimum of seven identical AAs, and peptide length > 12 AAs. The resulting human sequences are presented in Table 2 highlighting the antigens’ functionality and their implications in various ADs.
Discussion
The present study aimed to explore several immune mechanisms that operate in the human body, where an external, frequently consumed environmental factor, namely mTG, might drive chronic diseases. The potential role of the microbial enzyme in CD induction8,9,40,41,42,43,44,46,47,48,50,51 and other autoimmune and neurodegenerative diseases was recently extensively described6,7,22,26,27,28,39,46. Various deleterious effects were attributed to this enzymatic food additive and corresponding pathogenic mechanisms were suggested6,7,8,9,22,26,27,28,44,45,46.
Six pairs of similar immunogenic sequences were detected between human endogenous antigens, derived from cross-reactive antibodies, and between mTG immune epitopes (Table 1). All of them showed a strong binding affinity to at least one of the HLA-I and HLA-II alleles and play a crucial role in cellular functions and body homeostasis (Table 2).
Reviewing those six similar pairs of proteins, a functional relationship to the mTG can be suggested:
-
1.
The fibrinogen alpha chain is part of the coagulation system that joins factor XIII to establish an efficient clot. Factor XIII and mTG are integral members of the TG family67, both having the capacity to deamidate or transamidate acyl donors and acceptors molecules. There is no knowledge yet of on circulating mTG, nor its ability to coagulate, however, its intra-enterocytic transport and sub-epithelial deposition was documented46 and its relative resilience to oxidative compounds was recently reported51. Furthermore, restructured meat contains mTG and fibrinogen68, and fibrin gels crosslinked by a mTG are used in the industry, where the mTG reactions are comparable to those of factor XIII and tTG69. The potential pro-coagulant capacity of mTG is still an enigma.
-
2.
Histone H1.2 plays a pivotal role in chromatin and nucleosomes stability and functionality. Interestingly, cross-linking of histone by transglutaminase is well documented. Being a universal protein condenser, transglutaminase can modify core histone and regulate chromatin condensation, thus, impacting gene expression70,71,72,73. The cross-linking might result in free histone deprivation. In fact, epigenetic is a major pathway in ADs initiation and development, including in CD evolvement72,73.
The direct mTG action on histone 1.2 deserves more studies. The question arises whether during the intra-enterocytic transport, can mTG impact the gene expression of the human enterocyte?
-
3.
Dihydrolipoyllysine-residue acetyltransferase component of pyruvate dehydrogenase complex, mitochondrial is an essential enzyme in mitochondrial energy metabolism and preservation. It appears that TG2 is important in mitochondrial functions and dysfunctions74. Upon activation, the enzyme can change the assembly of respiratory chain complexes and modulate the transcription of critical mitochondrial genes. In general, the bacterial enzyme imitates the functions of the human one; however, the impact of mTG on the mitochondrial energetic homeostasis remains to be disclosed.
-
4.
Creatine kinase S-type, mitochondria. Creatine kinases represent a large family of isoenzymes that participate in intracellular energy homeostasis. Mitochondrial creatine kinase is responsible for the transfer of high energy phosphate to the cytosolic carrier, creatine. Creatine kinase S-type is a family member that plays a role in the mitochondrial energy metabolism and production, in organs with large, fluctuating energy demands, such as heart, skeletal muscle and brain75. Indeed, mitochondrial creatine kinase dysfunction was reported in heart, muscle and neurodegenerative conditions76. The observation that creatine reduces transglutaminase-catalyzed protein aggregation77 may connect various neurodegenerative diseases, like Alzheimer's, Parkinson's, and Huntington's diseases to creatine kinase dysfunction, reducing tissue creatine level, resulting in higher cross-linking activity of the local tTG78. Notably, mTG can functionally imitate the posttranslational modification executed by its family member, the tTG. However, the impact of mTG on those tissues is not yet known. Interestingly, abnormal energy metabolism was described in RA and T1DM79,80 (Table 2). In parallel, transglutaminase is implicated in both of the diseases. TG2 participate in synovial inflammation, bone erosion in RA, and in island cell dysfunction in T1DM81,82,83,84. Theoretically, if mTG reaches those target organs, comparable damage might be induced.
-
5.
Dimethyladenosine transferase 2, mitochondrial. By the transfer of methyl groups to specific adenosine residues in mitochondrial tRNAs, this enzyme is essential for the proper folding and function of the tRNAs, which are essential components in protein synthesis. In fact, mitochondrial dysfunction exists in MS85,86 (Table 2) and in RA79,80 (Table 2) and both diseases are affected by tTG, as mentioned above, for RA81,82,83, but also in MS87,88. The place of the tTG functional imitator, namely the mTG, remain to be explored.
-
6.
Cytochrome c1, heme protein, mitochondrial is integral and pivotal for the mitochondrial electron transport chain, responsible for generating ATP through oxidative phosphorylation. It represents a potential clinical marker for mitochondrial and cellular damage89. Mitochondrial failure, accompanied by inadequate energy supply and increased oxidative stress, exists in RA79,80,90,91 (Table 2), in T1DM80,92 (Table 2) and in AU93 (Table 2). Moreover, the mitochondrial Cytochrome c is affected in RA94,95, T1DN96,97 (Table 2) and in AU98 (Table 2). In parallel, the posttranslational modified ability of the tTG to cross-link mitochondrial protein70,71,72,73,74,77,78 and the mTG cross-linking capacity of cytochrome c, using its lysine residue as an acyl acceptor99,100, constitutes a confirmation of the tTG and mTG capacity to regulate mitochondrial proteins, thus contributing to this organelle dysfunction.
It can be summarized that tTG is involved in the regulation of the mitochondrial energy productive and regulatory machinery. This ubiquitous enzyme can cross-link histone, control chromatin condensation, determine gene expression70,71, affect mitochondrial functions74, its activity is affected by creatine kinases and by free creatine78,79 and its cross-linking activity can impact key essential mitochondrial molecules responsible for energy equilibrium in several autoimmune81,82,83,84,87,88 and neurodegenerative diseases79. In addition, the tTG is important in degradation of damaged mitochondria, thus playing as a gatekeeper of the mitochondrial functional homeostasis101.
In fact, a lot is still unknown as to whether mTG can replace tTG in all these activities. The fact that bacterial enzyme can cross the enteric epithelial lining46,51, have the capacity to cross-link proteins that contain acyl donors (glutamine) and acceptors (lysine)8,9,22,26,27,37,45,47, mount specific antibodies to its cross-linked complexes7,9,40,41,42,43,45,47,48,49 and be involved in initiation and progression of ADs, is an indication of its disadvantages, being a potential public health concern, and a caveat to public well-being9,22,26,27,45.
The current study brings, for the first time, two new potential pathogenic pathways: (1) relating the mTG enzyme to autoimmune and other chronic human conditions; (2) cross-reactive antibodies and sequence similarity between the environmental enzyme and endogenous human self-antigens. To these two pathogenic mechanisms the epitope sharing between the environmental gluten/gliadin peptides and multiple human antigens should be added. Intriguingly, gluten/gliadin structural segments are prime substrates for mTG de/transamidation6,7,8,9,22,39,40,41,42,43,44,45,46. This posttranslational modification is operating in the processed food industries, in bakeries and more importantly, in the human gut lumen8,9,22,26,27,45. It seems that the mTG-gluten-human self-epitopes axis is interactive and auto-immunogenic. Those three interrelated pathways are the basis for our current novel hypothesis, whereby, two very common environmental domains, plants and microbes, and gluten and mTG, respectively, are joining together to induce autoimmunity and other gluten-dependent inflammatory diseases4,7,9,22,26,27,32,39,41. Interestingly, gluten avoidance was recently reported to alleviate symptoms and disease activity of non-celiac ADs81,82,102,103,104,105,106, although, gluten withdrawal is not devoid of side effects107,108,109. Taken together, both external factors, the mTG and gluten-containing nutrients, can operate as the mythological Trojan horse to drive luminal and extra-intestinal ADs29. Figure 3 presents schematically the cross-reactivity and sequence similarity between mTG-Substrate complexes and gut-antigens that are associated with ADs.
Schematic presentation of cross-reactivity and sequence similarity between mTG-Substrate complexes and gut-antigens that are associated with ADs. (A) Oral consumption of food products that were processed with mTG, such as meat, fish, dairy and bread. (B) mTG-substrate’ complexes, such as mTG-gliadins, reach the gut lumen. (C) Gliadins, and other processed food products, are a substrate for mTG cross-linkage, turning a naïve molecule to immunogenic one. The result is an increase in mtg-induced PTMP that human digestive enzymes cannot break down, thus, inducing gut inflammation and damage to the intestinal epithelium. (D) mTG can potentially damage the lining mucus by breaking its stability and compromising tight junction functional integrity. mTG-Gliadin and other mTG complexes might penetrate into the lamina propria through open junctions or trans-enterocytically. (E) In the lamina propria, mTG-cross-linked complexes induce pro-inflammatory cytokines that drive T cells and B cells activation. (F) CD4 T cells initiate an immune response against mTG-PTMP after APC presentation of epitopes on HLA-II. CD8 T cells can be activated when they are exposed to epitopes presented on HLA-I, and activated by CD4 T cells. Cross-reactivity at the T cell level involves recognition of certain mTG-PTMP epitopes which are similar to self-epitopes. (G) Cross-reactivity at the B cell level when clonal antibodies bind to mTG-PTMP epitopes that are similar to self-epitopes. (H) Autoreactive antibodies, effector B and T cells, and mTG-substrate complexes travel through blood vessels to peripheral organs. They can potentially become autoreactive when they encounter similar self-epitopes, and an autoimmune response will be directed against the host as well.
The list of side effects of the processed food additive, mTG, and its cross-linked complexes is constantly expanding6,7,8,9,22,39,41,42,43,44,45. Multiple mechanisms were offered for those health-targeted detrimental effects. The mTG compromises tight junctional functional integrity, enhancing a leaky gut syndrome9,22,45 and enhances enteric epithelial gliadins uptake and transportation8,9,22,45,105,110. The foreign molecules, mTG and gliadin, are trans-enterocyticaly transported to face and challenge the sub-epithelial immune systems46. The microbial enzyme can compromise the mechanical intestinal protective barriers by introducing resistant isopeptide bonds, thus, perturbating mucin fluidity and stability, resulting in enhanced attachment of pathogenic luminal germs or other harmful factors to the epithelial receptors111. More so, it suppresses mucosal and systemic immune systems. Indeed, Streptococcus suis-originated mTG exerts anti-phagocytic effect, resulting in suppressing a major immune protective barrier112,113,114,115. As a bacterial survival factor, suppressing gut immunity, the mTG is a growth factor for luminal microbiota, dysbiota and pathobionts, as was reported in Lactococcus strain116,117. The problem is accentuated since more sophisticated bioengineered technics produce higher yield and more active mTG for industrial usage118,119,120. The enzyme represents a double-edged sword, a protective bacterial factor in the gut lumen, hence, a human hostile one, compromising human health45. In view of the active horizontal gene transfer in the gut lumen28, a major question arises. Can the harmful mTG be laterally transferred to the physiological microbiome, as is happening for the bacterial resistant genes spread?121,122. On the same line, recently, the trans-membranal region of mTG was suggested to participate in the recognition of host's immune signals and reciprocal bacterial communication, by binding to its corresponding ligand123.
The cross-reactive antibodies warrant some clarification. Polyclonal antibodies contain a heterogenous mixture of antibodies produced by different clones of plasma B cells against different epitopes of a whole antigen, whereas monoclonal antibodies are a homogenous population of antibodies that are produced by a single clone of B cells. Thus, polyclonal antibodies interact with different epitopes on a single antigen, while monoclonal antibodies interact with a particular epitope on the same antigen. This may explain the reactivity of anti-mTG polyclonal antibody with 18 out of 77 autoantigens and the reactivity of anti-mTG monoclonal antibody with only 9 out of 77 human tissue antigens124.
These are some strengths of the current study. It combines the human to the mTG epitopes, applying two methods, namely, cross-reactive antibodies and sequence similarity. It describes two members of the transglutaminase's family that shares comparable functions. The environmental mTG has a much broader substrate activity than its endogenous tTG one. So, theoretically, it might cross-react with more human antigens.
As for the study’s limitations, the major one is the lack of proof that the mTG itself or its post-translated modified proteins and cross-linked complexes can circulate systemically to reach peripheral target organs. However, the fact that when active mTG is abandoned in the gut lumen, it reaches the baso-lateral compartment of the enterocytes and its cross-linked complexes are immunogenic, strengthen the present hypothesis. In addition, the presented findings are limited to the curated epitopes that are currently found in the Immune Epitope Database (IEDB, https://www.iedb.org), and to 77 different human tissue antigens that were tested for cross-reactivity. Yet it provides an indication of such antigens that can potentially provoke molecular mimicry.
To further substantiate the present working hypothesis and strengthen the cause-and-effect relationship between the cross-reactive antibodies against mTG in patients with various ADs, those purified antibodies should be checked in the autoimmune affected patient's sera and passively transferred to appropriate animal models. We hope that the present observations will encourage further studies to establish causality between cross-reactivity and sequence similarity and the corresponding ADs.
Methods
Cross-reactive antibodies
To demonstrate cross-reactivity between mTG and various human target tissue antigens, the steps for the ELISA method were extracted from various manuscripts published by Vojdani et al.4,32,125,126,127,128,129,130,131.
In brief, mouse monoclonal antibody to recombinant mTG (MyBioSource, England) and affinity-purified rabbit polyclonal antibodies made against mTG (Zedira GmbH, Germany) were applied to 77 different human tissue antigens using the ELISA method. Different wells of ELISA plates were coated with various tissue antigens representing these categories: coagulation and heart, joints, diabetes-related, skin, epithelial and tight junctions, liver, lung, thyroid, the nervous system, and cellular antigens. The complete list of their source antigens and optimal concentrations is shown in Table S1 in the Supplement section.
Each antigen was first dissolved in 0.01 M of PBS at pH 7.4, then further diluted in carbonate buffer pH 9.5, in optimal amounts ranging from 0.5 to 2 µg per 100 µl, and was added to duplicate wells. Following incubation for 12 H at 25 °C, and an additional 12 H at 4 °C, the plates were washed 5 times, after which 200 µl of blocker containing 1% bovine serum albumin and 1% dried milk was added to each well. After a repeat incubation and washing, mouse monoclonal antibody against mTG, at a dilution of 1:200 and polyclonal antibody at a dilution of 1:400 was added to different sets of ELISA wells, coated with human tissue antigens. Plates were incubated for 1 H at 25 °C, and after another washing, 100 µl of alkaline phosphatase goat anti-mouse IgG at a dilution of 1:600 and goat anti-rabbit, at a dilution of 1:800, were added to different sets of ELISA plates. After another repeat of incubation and washing, 100 µl of substrate was added to each well, and color development was measured at 405 nM.
Sequence similarity
In search of immunoreactive epitopes, all human epitopes that relate to ADs were obtained from IEDB132,133. The IEDB was searched with the following keywords: Epitope: “Linear peptide”, Epitope Source: “Human Organism”, Host: “Human”, Assays included: “T cells”, “B cells”, “HLA-I”, “HLA-II”, Outcome: “Positive Assays”, and Disease: “Autoimmune”. About 67,000 epitopes were extracted from in-vivo experimental studies as antigens implicated with at least one of 61 ADs categories. The complete sequence of mTG protein, Uniprot: P81453, Organism: Streptomyces Mobaraensis, was acquired from the UniProt Knowledgebase (https://www.uniprot.org/)134.
As for sequence alignment, a Pairwise Local Alignment tool, EMBOSS Matcher135,136 was employed to explore sequence similarity between the aggregated human auto-epitopes and the mTG protein. This tool searches for local similarities between any two sequences by implementing an algorithm based on Bill Pearson's Lalign application, version 2.0u4 (Feb. 1996). A cutoff was applied on EMBOSS Matcher’s results to identify those epitopes that could have a higher probability of inducing molecular mimicry. The aligned peptides’ cut-off was kept at a minimum of seven identical AAs, and at peptide length > 12 AAs.
All human epitopes were captured in IEDB from experimental assays published in scientific literature. However, mTG sequences that were extracted and identified by EMBOSS Matcher required additional analysis to assess their immunological potential reactivity. IEDB Immunogenicity Prediction services offer tools to analyze the binding affinity values in terms of half maximal inhibitory concentration (IC50) of peptides binding to HLA-I/II alleles and to assess their potential to elicit an immune response. These tools were utilized to filter out all mTG sequences that were not considered to have immunogenic potential.
As a final selection, the resulting similar sequences were cross-checked with the cross-reactive antigens, and the concluding list includes human epitope sequences that are derived from those antigens. The methodology is presented as a flowchart in Fig. 4.
A graphical representation of the workflow for searching sequence similarity. Data Aggregation: human epitopes that are implicated in ADs were extracted from IEDB and UniProt was searched to retrieve mTG protein sequence. Sequence Alignment: Emboss Matcher was employed; 60 similar sequences were found with a cut-off of at least 7 identical AAs and peptide length > 12 AAs. Data Filtration and Validation: IEDB analysis tools were employed to validate those mTG sequences that are immunogenic, and have HLA-I/II binding affinity. Out of those results, 7 similar sequences were of human antigens that were previously identified to cross-react with mTG protein.
Conclusion
In summary, our findings support the potential contribution of mTG to various autoimmune diseases, which should be the subject of future studies. The presented shared cross-reactive antibodies and sequence similarity between the mTG and human immune epitopes, presents two novel pathological mechanisms that might compromise public health. It is hoped that the current findings will encourage future exploration of the mTG-human enigma.
Data availability
The data and software that supports the findings of this study are openly available in: The Immune Epitope Database (IEDB) at www.iedb.org, reference132,133. UniProt Knowledgebase www.uniprot.org, reference134. Pairwise Local Alignment tool, EMBOSS Matcher, at www.emboss.sourceforge.net, reference135,136, A python script can be found at https://raw.githubusercontent.com/ebi-wp/webservice-clients/master/python/emboss_matcher.py.
Abbreviations
- ADs:
-
Autoimmune diseases
- ASIA:
-
Autoimmune/inflammatory syndrome induced by adjuvants
- GIT:
-
Gastrointestinal tract
- mTG:
-
Microbial transglutaminase
- tTG:
-
Tissue transglutaminase
- CD:
-
Celiac disease
- AAs:
-
Amino acids
- OD:
-
Optical density
References
Mahroum, N. et al. The mosaic of autoimmunity - A taste for more. The 12th international congress of autoimmunity 2021 (AUTO12) virtual. Autoimmun. Rev. 20(11), 102945 (2021).
Samasca, G. et al. Polyautoimmunity - The missing ingredient. Autoimmun. Rev. 17, 840–841 (2018).
Vojdani, A., Vojdani, E., Rosenberg, A. Z. & Shoenfeld, Y. The role of exposomes in the pathophysiology of autoimmune diseases II: Pathogens. Pathophysiol. Off. J. Int. Soc. Pathophysiol. 29, 243–280 (2022).
Vojdani, A. & Vojdani, E. The role of exposomes in the pathophysiology of autoimmune diseases I: Toxic chemicals and food. Pathophysiol. 28, 513–543 (2021).
Lanone, S. et al. Bilirubin decreases NOS2 expression via inhibition of NAD(P)H oxidase: Implications for protection against endotoxic shock in rats. FASEB J. 19, 1890–1892 (2005).
Lerner, A. & Matthias, T. Changes in intestinal tight junction permeability associated with industrial food additives explain the rising incidence of autoimmune disease. Autoimmun. Rev. 14, 479–489 (2015).
Lerner, A. & Benzvi, C. ‘Let food be thy medicine’: Gluten and potential role in neurodegeneration. Cells 10(4), 756 (2021).
Lerner, A. & Matthias, T. Possible association between celiac disease and bacterial transglutaminase in food processing: A hypothesis. Nutr. Rev. 73, 544–552 (2015).
Lerner, A. & Matthias, T. Processed food additive microbial transglutaminase and its cross-linked gliadin complexes are potential public health concerns in celiac disease. Int. J. Mol. Sci. 21(3), 1127 (2020).
Lerner, A. Aluminum as an adjuvant in Crohn’s disease induction. Lupus 21, 231–238 (2012).
Garabatos, N. & Santamaria, P. Gut microbial antigenic mimicry in autoimmunity. Front. Immunol. 13, 873607 (2022).
Lerner, A., Arleevskaya, M., Schmiedl, A. & Matthias, T. Microbes and viruses are bugging the gut in celiac disease. Are they friends or foes?. Front. Microbiol. 8, 1392 (2017).
Hassan, A. & Blanchard, N. Microbial (co)infections: Powerful immune influencers. PLoS Pathog. 18(2), e1010212 (2022).
Smatti, M. K. et al. Viruses and autoimmunity: A review on the potential interaction and molecular mechanisms. Viruses 11(8), 762 (2019).
Sener, A. G. & Afsar, I. Infection and autoimmune disease. Rheumatol. Int. 32, 3331–3338 (2012).
Segal, Y. & Shoenfeld, Y. Vaccine-induced autoimmunity: The role of molecular mimicry and immune crossreaction. Cell. Mol. Immunol. 15, 586–594 (2018).
Jara, L. J., Vera-Lastra, O., Mahroum, N., Pineda, C. & Shoenfeld, Y. Autoimmune post-COVID vaccine syndromes: Does the spectrum of autoimmune/inflammatory syndrome expand?. Clin. Rheumatol. 41, 1603–1609 (2022).
Lerner, A. & Benzvi, C. Checkpoint inhibitors and induction of celiac disease-like condition. Biomedicines 10(3), 609 (2022).
Lerner, A. & Benzvi, C. When genetic polymorphism meets an immune checkpoint inhibitor in celiac disease. Int. J. Celiac Dis. 10, 11–16 (2022).
Borba, V. et al. Classical examples of the concept of the ASIA syndrome. Biomolecules 10, 1–17 (2020).
Facciolà, A., Visalli, G., Laganà, A. & Di Pietro, A. An overview of vaccine adjuvants: Current evidence and future perspectives. Vaccines 10(5), 819 (2022).
Lerner, A. & Benzvi, C. Microbial transglutaminase is a very frequently used food additive and is a potential inducer of autoimmune/neurodegenerative diseases. Toxics 9(10), 233 (2021).
Akdis, C. A. Does the epithelial barrier hypothesis explain the increase in allergy, autoimmunity and other chronic conditions?. Nat. Rev. Immunol. 21, 739–751 (2021).
Fasano, A. All disease begins in the (leaky) gut: Role of zonulin-mediated gut permeability in the pathogenesis of some chronic inflammatory diseases. F1000Research 9, (2020).
Martel, J. et al. Gut barrier disruption and chronic disease. Trends Endocrinol. Metab. 33, 247–265 (2022).
Lerner, A., Aminov, R. & Matthias, T. Dysbiosis may trigger autoimmune diseases via inappropriate post-translational modification of host proteins. Front. Microbiol. 7, 84 (2016).
Lerner, A., Aminov, R. & Matthias, T. Transglutaminases in dysbiosis as potential environmental drivers of autoimmunity. Front. Microbiol. 8, 66 (2017).
Lerner, A., Matthias, T. & Aminov, R. Potential effects of horizontal gene exchange in the human gut. Front. Immunol. 8, 1630 (2017).
Lerner, A. & Matthias, T. GUT-the Trojan horse in remote organs’ autoimmunity. J. Clin. Cell. Immunol. 7, 1–10 (2016).
Lerner, A., Jeremias, P. & Matthias, T. The world incidence and prevalence of autoimmune diseases is increasing. Int. J. Celiac Dis. 3, 151–155 (2015).
Rojas, M. et al. Molecular mimicry and autoimmunity. J. Autoimmun. 95, 100–123 (2018).
Vojdani, A., Gushgari, L. R. & Vojdani, E. Interaction between food antigens and the immune system: Association with autoimmune disorders. Autoimmun. Rev. 19, 102459 (2020).
Kanduc, D. & Shoenfeld, Y. Molecular mimicry between SARS-CoV-2 spike glycoprotein and mammalian proteomes: Implications for the vaccine. Immunol. Res. 68, 310–313 (2020).
Dotan, A., Mahroum, N., Bogdanos, D. P. & Shoenfeld, Y. COVID-19 as an infectome paradigm of autoimmunity. J. Allergy Clin. Immunol. 149, 63–64 (2022).
Griffin, M., Casadio, R. & Bergamini, C. M. Transglutaminases: Nature’s biological glues. Biochem. J. 368, 377–396 (2002).
Reif, S. & Lerner, A. Tissue transglutaminase - The key player in celiac disease: A review. Autoimmun. Rev. 3, 40–45 (2004).
Lerner, A., Neidhöfer, S. & Matthias, T. Transglutaminase 2 and anti transglutaminase 2 autoantibodies in celiac disease and beyond: TG2 double-edged sword: Gut and extraintestinal involvement. Immunome Res. 11, 1–4 (2015).
Liu, C., Kellems, R. E. & Xia, Y. Inflammation, autoimmunity, and hypertension: The essential role of tissue transglutaminase. Am. J. Hypertens. 30, 756–764 (2017).
Lerner, A. & Matthias, T. Don’t forget the exogenous microbial transglutaminases: It is immunogenic and potentially pathogenic. AIMS Biophys. 3, 546–552 (2016).
Matthias, T., Jeremias, P., Neidhöfer, S. & Lerner, A. The industrial food additive, microbial transglutaminase, mimics tissue transglutaminase and is immunogenic in celiac disease patients. Autoimmun. Rev. 15, 1111–1119 (2016).
Lerner, A. & Matthias, T. Microbial transglutaminase should be considered as an environmental inducer of celiac disease. World J. Clin. Cases 7, 3912–3914 (2019).
Agardh, D. et al. Antibodies against neo-epitope of microbial and human transglutaminase complexes as biomarkers of childhood celiac disease. Clin. Exp. Immunol. 199, 294–302 (2020).
Matthias, T. & Lerner, A. Microbial transglutaminase is immunogenic and potentially pathogenic in pediatric celiac disease. Front. Pediatr. 6, 389 (2018).
Lerner, A. & Matthias, T. Microbial transglutaminase: A new potential player in celiac disease. Clin. Immunol. 199, 37–43 (2019).
Lerner, A. & Matthias, T. Microbial transglutaminase is beneficial to food industries but a caveat to public health. Med One 4, (2019).
Stricker, S., De Laffolie, J., Rudloff, S., Komorowski, L. & Zimmer, K. P. Intracellular localization of microbial transglutaminase and its influence on the transport of gliadin in enterocytes. J. Pediatr. Gastroenterol. Nutr. 68, E43–E50 (2019).
Lerner, A., Ramesh, A. & Matthias, T. Serologic diagnosis of celiac disease: new biomarkers. Gastroenterol. Clin. North Am. 48, 307–317 (2019).
Lerner, A., Jeremias, P., Neidhofer, S. & Matthias, T. Comparison of the reliability of 17 celiac disease associated bio-markers to reflect intestinal damage. J. Clin. Cell. Immunol. 8(486), 2 (2017).
Lerner, A. More novel diagnostic antibodies for celiac disease. Expert Rev. Gastroenterol. Hepatol. 10, 767–768 (2016).
Lerner, A. & Matthias, T. Food industrial microbial transglutaminase in celiac disease: Treat or trick. Int. J. Celiac Dis. 3, 1–6 (2015).
Stricker, S., Rudloff, S., De Laffolie, J. & Zimmer, K. P. Tissue transglutaminase but not microbial transglutaminase is inhibited by exogenous oxidative substances in celiac disease. Int. J. Mol. Sci. 23(4), 2248 (2022).
Lio, W. M. et al. Sex as a determinant of responses to a coronary artery disease self-antigen identified by immune-peptidomics. Front. Immunol. 11, 694 (2020).
Khatri, S. et al. Citrullinated peptide epitope targets therapeutic nanoparticles to human neutrophils. Bioconjug. Chem. 30, 2584–2593 (2019).
Joshua, V. et al. Antibody responses to de novo identified citrullinated fibrinogen peptides in rheumatoid arthritis and visualization of the corresponding B cells. Arthritis Res. Ther. 18(1), 1–9 (2016).
Goules, J. D., Goules, A. V. & Tzioufas, A. G. Fine specificity of anti-citrullinated peptide antibodies discloses a heterogeneous antibody population in rheumatoid arthritis. Clin. Exp. Immunol. 174, 10–17 (2013).
Van Der Woude, D. et al. Epitope spreading of the anti-citrullinated protein antibody response occurs before disease onset and is associated with the disease course of early arthritis. Ann. Rheum. Dis. 69, 1554–1561 (2010).
Pérez, M. L. et al. Synthesis of overlapping fibrin citrullinated peptides and their use for diagnosing rheumatoid arthritis. Chem. Biol. Drug Des. 68, 194–200 (2006).
Dwivedi, N. et al. Deimination of linker histones links neutrophil extracellular trap release with autoantibodies in systemic autoimmunity. FASEB J. 28, 2840–2851 (2014).
Muixí, L., Alvarez, I. & Jaraquemada, D. Peptides presented in vivo by HLA-DR in thyroid autoimmunity. Adv. Immunol. 99, 165–209 (2008).
Nuti, F. et al. Antibodies to post-translationally modified mitochondrial peptide PDC-E2(167–184) in type 1 diabetes. Arch. Biochem. Biophys. 659, 66–74 (2018).
Pacini, G. et al. Role of lipoylation of the immunodominant epitope of pyruvate dehydrogenase complex: Toward a peptide-based diagnostic assay for primary biliary cirrhosis. J. Med. Chem. 58, 6619–6629 (2015).
Briand, J.-P. et al. Multiple autoepitope presentation for specific detection of antibodies in primary biliary cirrhosis. Hepatology 16, 1395–1403 (1992).
Sudhir, P. R., Lin, TDu. & Zhang, Q. HLA allele-specific quantitative profiling of type 1 diabetic B lymphocyte immunopeptidome. J. Proteome Res. 21, 250–264 (2022).
Lo, K. C. et al. Comprehensive profiling of the rheumatoid arthritis antibody repertoire. Arthritis. Rheumatol. (Hoboken N. J.) 72, 242–250 (2020).
Wang, J. et al. HLA-DR15 molecules jointly shape an autoreactive T cell repertoire in multiple sclerosis. Cell 183, 1264-1281.e20 (2020).
Venema, W. J. et al. Supplemental Figures for: ERAP2 increases the abundance of a peptide submotif highly selective for the Birdshot Uveitis-associated HLA-A29.
Duarte, L., Matte, C. R., Bizarro, C. V. & Ayub, M. A. Z. Transglutaminases: part I—origins, sources, and biotechnological characteristics. World J. Microbiol. Biotechnol. 36, 1–18 (2020).
Jira, W. & Schwägele, F. A sensitive HPLC-MS/MS method for the simultaneous detection of microbial transglutaminase, and bovine and porcine fibrinogen/thrombin in restructured meat. Food Chem. 237, 841–848 (2017).
Sun, Y., Giraudier, O. & Garde, V. L. Rheological characterization and dissolution kinetics of fibrin gels crosslinked by a microbial transglutaminase. Biopolymers 77, 257–263 (2005).
Kumar, M. D., Singh, A. K., Shama, H. & Deshwal, G. K. Histone cross-linking by transglutaminase. Biochem. Biophys. Res. Commun. 293, 1453–1457 (2002).
Kim, J. H., Choy, H. E., Nam, K. H. & Park, S. C. Transglutaminase-mediated crosslinking of specific core histone subunits and cellular senescence. Ann. N. Y. Acad. Sci. 928, 65–70 (2001).
Gnodi, E., Meneveri, R. & Barisani, D. Celiac disease: From genetics to epigenetics. World J. Gastroenterol. 28, 449–463 (2022).
Perry, A. S., Baird, A. M. & Gray, S. G. Epigenetic methodologies for the study of celiac disease. Methods Mol. Biol. 1326, 131–158 (2015).
Lai, T. S., Lin, C. J., Wu, Y. T. & Wu, C. J. Tissue transglutaminase (TG2) and mitochondrial function and dysfunction. Front. Biosci. (Landmark Ed.) 22, 1114–1137 (2017).
Schlattner, U., Kay, L. & Tokarska-Schlattner, M. Mitochondrial proteolipid complexes of creatine kinase. Subcell. Biochem. 87, 365–408 (2018).
Schlattner, U., Tokarska-Schlattner, M. & Wallimann, T. Mitochondrial creatine kinase in human health and disease. Biochim. Biophys. Acta 1762, 164–180 (2006).
Burguera, E. F. & Love, B. J. Reduced transglutaminase-catalyzed protein aggregation is observed in the presence of creatine using sedimentation velocity. Anal. Biochem. 350, 113–119 (2006).
Bürklen, T. S. et al. The creatine kinase/creatine connection to Alzheimer’s disease: CK-inactivation, APP-CK complexes and focal creatine deposits. J. Biomed. Biotechnol. 2006, (2006).
Ma, C., Wang, J., Hong, F. & Yang, S. Mitochondrial dysfunction in rheumatoid arthritis. Biomolecules 12(9), 1216 (2022).
Halmos, T. & Suba, I. Diseases caused by mitochondrial dysfunction. Orv. Hetil. 163, 1383–1393 (2022).
Lerner, A. & Matthias, T. Rheumatoid arthritis-celiac disease relationship: joints get that gut feeling. Autoimmun. Rev. 14, 1038–1047 (2015).
Aaron, L., Patricia, W., Ajay, R., Francois, L. & Torsten, M. The gut feeling of the joints: Celiac disease and rheumatoid arthritis are related. Int. J. Celiac Dis. 7(1), 21–25 (2019).
Lauzier, A., Charbonneau, M., Paquette, M., Harper, K. & Dubois, C. M. Transglutaminase 2 cross-linking activity is linked to invadopodia formation and cartilage breakdown in arthritis. Arthritis Res. Ther. 14(4), 1–14 (2012).
McLaughlin, R. J. et al. Human islets and dendritic cells generate post-translationally modified islet autoantigens. Clin. Exp. Immunol. 185, 133–140 (2016).
de Barcelos, I. P., Troxell, R. M. & Graves, J. S. Mitochondrial dysfunction and multiple sclerosis. Biology (Basel). 8(2), 37 (2019).
Blagov, A. V., Sukhorukov, V. N., Orekhov, A. N., Sazonova, M. A. & Melnichenko, A. A. Significance of mitochondrial dysfunction in the progression of multiple sclerosis. Int. J. Mol. Sci. 23(21), 12725 (2022).
Pearse, D. D., Hefley, A. B., Morales, A. A. & Ghosh, M. Comparative profiling of TG2 and its effectors in human relapsing remitting and progressive multiple sclerosis. Biomedicines 10(6), 1241 (2022).
Pearse, D. D., Otero, P. A., Diaz, A., Pan, X. & Ghosh, M. Neuronal and endothelial transglutaminase-2 expression during experimental autoimmune encephalomyelitis and multiple sclerosis. Neuroscience 461, 140–154 (2021).
Eleftheriadis, T., Pissas, G., Liakopoulos, V. & Stefanidis, I. Cytochrome c as a potentially clinical useful marker of mitochondrial and cellular damage. Front. Immunol. 7, 279 (2016).
Kan, S., Duan, M., Liu, Y., Wang, C. & Xie, J. Role of mitochondria in physiology of chondrocytes and diseases of osteoarthritis and rheumatoid arthritis. Cartilage 13, 1102S-1121S (2021).
Jing, W. et al. Role of reactive oxygen species and mitochondrial damage in rheumatoid arthritis and targeted drugs. Front. Immunol. 14, 1107670 (2023).
Sivitz, W. I. & Yorek, M. A. Mitochondrial dysfunction in diabetes: from molecular mechanisms to functional significance and therapeutic opportunities. Antioxid. Redox Signal. 12, 537–577 (2010).
Saraswathy, S. & Rao, N. A. Posttranslational modification of differentially expressed mitochondrial proteins in the retina during early experimental autoimmune uveitis. Mol. Vis. 17, 1814 (2011).
Becker, Y. L. C., Duvvuri, B., Fortin, P. R., Lood, C. & Boilard, E. The role of mitochondria in rheumatic diseases. Nat. Rev. Rheumatol. 18, 621–640 (2022).
Pullerits, R., Bokarewa, M., Jonsson, I. M., Verdrengh, M. & Tarkowski, A. Extracellular cytochrome c, a mitochondrial apoptosis-related protein, induces arthritis. Rheumatology (Oxford). 44, 32–39 (2005).
Gaignard, P. et al. Mutations in CYC1, encoding cytochrome c1 subunit of respiratory chain complex III, cause insulin-responsive hyperglycemia. Am. J. Hum. Genet. 93, 384–389 (2013).
Jung, S. R. et al. Reduced cytochrome C is an essential regulator of sustained insulin secretion by pancreatic islets. J. Biol. Chem. 286, 17422–17434 (2011).
Wu, G.-S. & Rao, N. A. Cytochrome c is the Major Nitrated Protein in Mitochondria in the Early Phase of Experimental Autoimmune Uveoretinitis. Invest. Ophthalmol. Vis. Sci. 47, 809–809 (2006).
Zhou, J. Q., He, T. & Wang, J. W. PEGylation of cytochrome c at the level of lysine residues mediated by a microbial transglutaminase. Biotechnol. Lett. 38, 1121–1129 (2016).
Santos, J. H. P. M. et al. Lysine-PEGylated cytochrome C with enhanced shelf-life stability. Biosensors 12(2), 94 (2022).
Rossin, F. et al. Transglutaminase 2 ablation leads to mitophagy impairment associated with a metabolic shift towards aerobic glycolysis. Cell Death Differ. 22, 408–418 (2015).
Lerner, A., De Carvalho, J. F., Kotrova, A. & Shoenfeld, Y. Gluten-free diet can ameliorate the symptoms of non-celiac autoimmune diseases. Nutr. Rev. 80, 525–543 (2022).
Lerner, A., Ramesh, A. & Matthias, T. Are non-celiac autoimmune diseases responsive to gluten-free diet?. Int. J. Celiac Dis. 5, 164–167 (2017).
Lerner, A., Shoenfeld, Y. & Matthias, T. Adverse effects of gluten ingestion and advantages of gluten withdrawal in nonceliac autoimmune disease. Nutr. Rev. 75, 1046–1058 (2017).
Lerner, A., Ramesh, A. & Matthias, T. Going gluten free in non-celiac autoimmune diseases: The missing ingredient. Expert Rev. Clin. Immunol. 14, 873–875 (2018).
Lerner, A. & Benzvi, C. Should rheumatoid arthritis patients go on a gluten-free diet? Int. J. Nutr. Sci. 8(1), 1070 (2023).
Lerner, A., O’Bryan, T. & Matthias, T. Navigating the Gluten-Free Boom: The Dark Side of Gluten Free Diet. Front. Pediatr. 7, 414 (2019).
Lerner, A. & Matthias, T. Gluten-free diet tough alley in torrid time. Int. J. Celiac Dis. 5, 50–55 (2017).
Lerner, A. & Matthias, T. The Yin and Yang of dietary gluten transgressions in real-life scenarios of celiac patients. BMC Med. 18(1), 1–3 (2020).
Lebreton, C. et al. Interactions among secretory immunoglobulin A, CD71, and transglutaminase-2 affect permeability of intestinal epithelial cells to gliadin peptides. Gastroenterology 143(3), 698–707 (2012).
Recktenwald, C. V. & Hansson, G. C. The reduction-insensitive bonds of the MUC2 mucin are isopeptide bonds. J. Biol. Chem. 291, 13580–13590 (2016).
Yu, J. et al. Functional and structural characterization of the antiphagocytic properties of a novel transglutaminase from streptococcus suis. J. Biol. Chem. 290, 19081–19092 (2015).
Xia, X. et al. How Streptococcus suis serotype 2 attempts to avoid attack by host immune defenses. J. Microbiol. Immunol. Infect. 52, 516–525 (2019).
Pian, Y. et al. Proteomics identification of novel fibrinogen-binding proteins of Streptococcus suis contributing to antiphagocytosis. Front. Cell. Infect. Microbiol. 5, 19 (2015).
Xu, B. et al. hsdS, belonging to the type I Restriction-modification system, contributes to the streptococcus suis serotype 2 survival ability in phagocytes. Front. Microbiol. 8, 1524 (2017).
Fu, R. Y., Chen, J. & Li, Y. Heterologous leaky production of transglutaminase in Lactococcus lactis significantly enhances the growth performance of the host. Appl. Environ. Microbiol. 71, 8911–8919 (2005).
Fu, R. Y., Chen, J. & Li, Y. Influence of expression of transglutaminase on the growth of Lactococcus lactis. Wei Sheng Wu Xue Bao 45, 510–515 (2005).
Zhang, N. et al. Intein-mediated intracellular production of active microbial transglutaminase in Corynebacterium glutamicum. Enzyme Microb. Technol. 142, 109680 (2020).
Rickert, M. et al. Production of soluble and active microbial transglutaminase in Escherichia coli for site-specific antibody drug conjugation. Protein Sci. https://doi.org/10.1002/PRO.2833 (2016).
Duarte, L., Matte, C. R., Bizarro, C. V. & Ayub, M. A. Z. Review transglutaminases: Part II-industrial applications in food, biotechnology, textiles and leather products. World J. Microbiol. Biotechnol. 36, 1–20 (2019).
Lerner, A., Shoenfeld, Y. & Matthias, T. Probiotics: If it does not help it does not do any harm. really?. Microorganisms 7(4), 104 (2019).
Lerner, A., Soprun, L. & Benzvi, C. Antimicrobial resistance along the food chain: Contaminated and industrially processed nutrients. J. Food Nutr. Heal. 3, 1–11 (2022).
Nezametdinova, V. Z., Yunes, R. A., Dukhinova, M. S., Alekseeva, M. G. & Danilenko, V. N. The role of the PFNA Operon of bifidobacteria in the recognition of host’s immune signals: Prospects for the use of the FN3 protein in the treatment of COVID-19. Int. J. Mol. Sci. 22(17), 9219 (2021).
Heil, A. et al. Microbial transglutaminase has a lower deamidation preference than human tissue transglutaminase on a celiac disease relevant wheat gliadin T-cell epitope. J. Cereal Sci. 70, 47–56 (2016).
Vojdani, A. Molecular mimicry as a mechanism for food immune reactivities and autoimmunity. Altern. Ther. Health Med. 21, 34–45 (2015).
Vojdani, A., Afar, D. & Vojdani, E. Reaction of lectin-specific antibody with human tissue: Possible contributions to autoimmunity. J. Immunol. Res. 2020, (2020).
Vojdani, A. Lectins, agglutinins, and their roles in autoimmune reactivities. Altern. Ther. Health Med. 21, 46–51 (2015).
Vojdani, A. & Kharrazian, D. Potential antigenic cross-reactivity between SARS-CoV-2 and human tissue with a possible link to an increase in autoimmune diseases. Clin. Immunol. 217, 108480 (2020).
Vojdani, A., Vojdani, E. & Kharrazian, D. Reaction of human monoclonal antibodies to SARS-CoV-2 proteins with tissue antigens: Implications for autoimmune diseases. Front. Immunol. 11, 3679 (2021).
Vojdani, A., Monro, J., Lanzisera, F. & Sadeghi, H. Serological cross-reactivity between viruses and their contribution to autoimmunity. Autoimmun. Rev. 20(7), 102840 (2021).
Vojdani, A., Lerner, A. & Vojdani, E. Cross-reactivity and sequence homology between Al-Pha-synuclein and food products: A step further for Parkinson’s disease synucleinopathy. Cells 10, 1111 (2021).
Vita, R. et al. The immune epitope database (IEDB): 2018 update. Nucl. Acids Res. 47, D339–D343 (2019).
Fleri, W. et al. The immune epitope database and analysis resource in epitope discovery and synthetic vaccine design. Front. Immunol. 8, 278 (2017).
Bateman, A. et al. UniProt: The universal protein knowledgebase in 2021. Nucl. Acids Res. 49, D480–D489 (2021).
Madeira, F. et al. Search and sequence analysis tools services from EMBL-EBI in 2022. Nucl. Acids Res. 50(W1), W276–W279 (2022).
Rice, P., Longden, L. & Bleasby, A. EMBOSS: The European molecular biology open software suite. Trends Genet. 16, 276–277 (2000).
Author information
Authors and Affiliations
Contributions
A.L.- screened the literature, designed and wrote the manuscript, C.B.- screened the literature, wrote, edited, and revised the manuscript, designed figures 3 and 4 with BioRender.com permission. A.V.- designed and wrote the manuscript, performed the ALISA essays, and analyzed the results. All authors reviewed the manuscript. The three authors agreed to the published version of the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Lerner, A., Benzvi, C. & Vojdani, A. Cross-reactivity and sequence similarity between microbial transglutaminase and human tissue antigens. Sci Rep 13, 17526 (2023). https://doi.org/10.1038/s41598-023-44452-5
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-023-44452-5
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.