Abstract
The use of data-based species distribution models (SDMs) has increased significantly in recent years. However, studies of determining the minimum requirements of occurrence sites from ecological monitoring datasets used in species distribution modelling remain insufficient. Therefore, this study proposed a framework to determine the threshold of minimum occurrence sites for SDMs by assessing compliance with Benford’s law. The compliance test verified that the national-scale freshwater fish monitoring dataset was natural and reliable. Results derived from true skill statistics (TSS) determined the minimum number of occurrence sites for reliable species distribution modelling was 20 with a TSS value of 0.793 and an overall accuracy of 0.804. The Benford compliance test has shown to be a useful tool for swift and efficient evaluation of the reliability of species occurrence datasets, or the determination of the threshold of occurrence sites before species distribution modelling. Further studies regarding the evaluation of this method’s transferability to other species and validation using SDM performance are required. Overall, the framework proposed in this study demonstrates that Benford compliance test applied to species monitoring datasets can be used to derive a universal and model-independent minimum occurrence threshold for SDMs.
Similar content being viewed by others
Introduction
Species distribution models (SDMs) generate relationships between abiotic and biotic factors and species occurrence records to predict the probability of species presence1,2,3. Over the last several decades, SDMs have been widely used in various applications14,5, including species conservation6, climate change impact assessment7,8, invasive species management9,10, and paleoecology5.
SDM users generally collate species occurrence data from ecological monitoring datasets that pass quality assurance and quality control procedures conducted during the dataset construction phase11,12. These procedures usually focus on defining rules to ensure the integrity of the dataset13 or detecting and correcting errors within the dataset12,14. In addition, sufficient occurrence sites (or sample sizes) are recommended for SDMs, since the model performance deteriorates when the number of occurrence sites is too low15,16,17,18. Previous studies have attempted to determine the minimum number of occurrence sites (e.g., 5 sites to 200 sites) by evaluating model accuracy (e.g., Pearson’s r, area under the receiver operating characteristic curve, weighted kappa, etc.)3,15,17,18,19. However, these thresholds are specific and model-dependent19,20, and should be verified using independent data21,22. Thus, a more generalized procedure is required to determine the reliability of a dataset and the minimum amount of occurrence data for SDM applications.
Hence, this study aimed to develop a novel methodology using Benford’s law as a universal and model-independent criterion to identify the minimum number of occurrence sites required for SDMs from species occurrence datasets. As a case study, the reliability of a national freshwater fish monitoring dataset, which was collected for 13 consecutive years throughout South Korea, was evaluated (Fig. 1a).
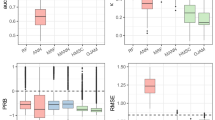
Methodologies used in this study: (a) Monitoring data of freshwater fish (details are available in Table S1) in South Korea were collected (Han, Nakdong, Geum, Yeongsan, and Seomjin river watersheds, represented by their first letter; H, N, G, Y, and S, respectively); (b) Compliance of whole fish dataset (white bars) with Benford’s law (solid line) evaluated by the coefficient of determination (R2); (c) Compliance of single fish dataset with Benford’s law evaluated by the interquartile range (IQR) method; and (d) Determination of the minimum number (threshold) of occurrence sites (blue circle) using true skill statistics (TSS). The maps were generated using ArcGIS Pro (ESRI, ver. 3.1; https://www.esri.com/en-us/arcgis/products/arcgis-pro/overview).
Benford’s law
Benford’s law was discovered by Simon Newcomb23 and justified by Frank Benford24; it refers to a phenomenon in which the distribution of leading (non-zero) digits from a dataset that occurs naturally (or untampered) complies with a logarithm equation (Eq. 1)25,26:
where P is the expected frequency (or probability) of the first digit (li). The expected frequency for each digit is presented in Fig. 1b.
In practice, Benford’s law is frequently applied as a standard when evaluating digit distributions. It is assumed that the dataset will not comply with Benford’s law if the numbers are not natural and influenced by human choice26. Compliance with Benford’s law is determined through goodness-of-fit tests by comparing the frequency of digits that appear in the dataset27. The most popular and widespread use of this law is in fraud detection, including data fabrication and falsification25,28,29. In the field of environmental science, Benford’s law has been applied to secure the reliability or identify anomalies in datasets, which include stream flows27, earthquakes30, tropical cyclones31, ecosystem naturalness32, health/disease report 33, ecotoxicity26, and phytoplankton cells in colonies34 and abundance35.
Results and discussions
Evaluating the reliability of the fish monitoring dataset
In this study, Benford’s law was applied to evaluate the reliability of the entire fish monitoring dataset (Table S1). The frequency of the first digits extracted from the total dataset complied well with Benford’s law, yielding a coefficient of determination (R2) of 0.994 (Fig. 1b). Considering that R2 > 0.85 is generally accepted as high credibility36,37, it confirms that the national freshwater fish monitoring dataset is highly reliable. In general, compliance with Benford’s law indicates that the dataset is authentic and natural26,27,31. Thus, the national freshwater fish monitoring dataset sufficiently represents the occurrence of freshwater fish in South Korea.
Datasets that do not comply with Benford’s law generally result from insufficient data quantity (or incomplete datasets), excessive rounding of data, and data errors27. For instance, Polidori and Hage38 applied Benford’s law to evaluate the accuracy of elevation, slope, and stream order from a digital elevation model and found large errors in elevation. Moreover, Noleto–Filho et al.39 demonstrated that the compliance assessment of a Brazilian fishing dataset with Benford’s law could identify the cause of unreliability. These findings suggest that Benford’s law can be used as a solid criterion for evaluating the reliability of monitoring datasets40.
Determining the threshold of fish occurrence sites
Benford’s law was also applied to determine the minimum number of fish occurrence sites required for species distribution modelling (Table S2). The interquartile range (IQR) method (Fig. 1c) showed that the species with R2 < 0.698 did not comply with Benford’s law in which 8 species failed among the 148 species tested (Fig. 2). Moreover, true skill statistics (TSS) was used to determine the threshold of minimum occurrence sites for complying with Benford’s law (Fig. 1d), since a reliable threshold can be produced by maximizing the sum of sensitivity and specificity (equivalent to maximizing the TSS value) than other methods (e.g., maximizing overall accuracy, maximizing kappa value, using the mean predicted value, etc.)41. The maximum TSS value of 0.793 was derived at 20 occurrence sites, with an overall accuracy of 0.804 (Fig. 2). Since TSS surpassed the criterion of 0.642,43, the threshold was shown to effectively discriminate compliance with Benford’s law for each fish species according to the number of occurrence sites. As suggested in Szabo et al.40, one of the advantages of Benford’s law is the efficient and swift evaluation process, since modelling each species individually with SDMs entails a higher cost. These findings suggest that Benford’s law can be used as a universal tool for determining the minimum number of fish occurrence sites since it is independent of SDMs.
Determination of the minimum fish occurrence site threshold (blue line) using TSS. Compliance (viz., passed or failed) of the single fish dataset with Benford’s law, evaluated by the IQR method, is denoted in white and red circles, respectively. The maximum TSS value (0.793), with an overall accuracy of 0.804, was yielded when the minimum number was set at 20 sites.
As demonstrated in this study, Benford’s law can be utilized to evaluate the reliability of species monitoring datasets. However, the results obtained from the compliance test provide limited ecological information40. For instance, a species monitoring dataset can be determined to be reliable, but information on species richness or biodiversity is not identified. Meanwhile, failing the compliance test implies additional or detailed investigation is required to ensure the reliability of using the dataset. Thus, Benford compliance test can be used as a screening process to evaluate the reliability of species monitoring or larger ecological datasets40, or to determine the threshold of occurrence sites before developing SDMs.
Further studies of applying this approach to SDMs and validating the threshold of species occurrence with model accuracy are required. Concurrently, datasets from various regions and other groups of organisms (e.g., amphibians, avians, invertebrates, etc.) should be tested for transferability of the approach, because the freshwater fish dataset was the only available dataset with detailed monitoring records that were collectable for this study.
Conclusions
This study demonstrated that Benford compliance test based on species occurrence datasets can provide a universal and model-independent criterion for determining the minimum occurrence threshold for species distribution modelling. A national-scale freshwater fish monitoring dataset was verified to comply with Benford’s law, indicating that the fish monitoring dataset was reliable and natural. Through the TSS, 20 was determined as the minimum occurrence threshold for modelling the distribution of freshwater fish from this dataset. Further studies of testing species occurrence datasets of other groups of organisms or regions are required to verify the transferability of this method. Also, future studies should evaluate the performance of this approach by comparing SDM accuracy divided by the threshold of species occurrence.
Methods
Data collection
The freshwater fish ecological monitoring data from 2008 to 2020 were collected from the Water Environment Information System (https://water.nier.go.kr; initially accessed on June 05, 2017, and updated on August 03, 2021). Along with the species occurrence results, the attributes regarding the monitoring program were included in the dataset. In addition, all survey stations were assigned to one of the 5 basins (Han, Nakdong, Geum, Seomjin, and Yeongsan River Basins) according to the classification available in WAMIS (https://wamisgo.kr accessed on August 03, 2021.). Details of the collected data and their statistics are listed in Table S1. Among the 159 fish species initially listed, 11 were excluded from the analysis due to non-occurrence. Data archiving and statistical analysis were conducted using MS Excel 2019 (Microsoft Corporation, Redmond, WA, USA).
Evaluating compliance with Benford’s law for species occurrence dataset
Benford’s law was applied by extracting the leading (or first) digit from the collected dataset. Subsequently, the frequency (0–9) of each digit (1–9) was calculated followed by a compliance test. The whole dataset used the entire dataset (W in Table S1), while each fish species used a species-specific dataset, respectively (S in Table S1). The coefficient of determination (R2) was derived by the regression analysis of the first digit frequency from the species monitoring dataset (i.e., whole dataset, datasets of each fish species) and Benford’s law (Fig. 1b). In addition, the occurrence data of freshwater fish in the 5 basins was integrated in the compliance tests assuming that SDM users generally use the largest range of available occurrence data44.
Compliance with Benford’s law was determined using the IQR (interquartile range) method (Fig. 1c). The R2 was classified into pass (R2 ≥ threshold of compliance) and fail (R2 < threshold of compliance) according to the following equations (Fig. 1c; Eqs. 2, 3):
where Q3 is the upper 25% quartile of R2s and Q1 is the lower 25% quartile of R2s acquired from the regression analysis of each species. Meanwhile, q is the coefficient that determines the threshold location, where a common value of 1.5 was applied in this study45,46.
Determining minimum occurrence site threshold
The required minimum occurrence sites for species distribution modelling was determined using true skill statistics (TSS) according to Allouche et al.47. TSS was conducted using a 2 × 2 contingency table (Table S3), where “a” is the number of species that accurately passed the threshold, “b” is the number of species that incorrectly passed the threshold (type I error; false positive), “c” is the number of species that incorrectly did not pass the threshold (type II error; false negative), and “d” is the number of species that correctly did not pass the threshold. The TSS value was calculated by summing sensitivity (Eq. 4) and specificity (Eq. 5) subtracted by 1 (Eq. 6). As presented in Fig. 1d, the minimum occurrence threshold is the point at which the TSS value is initially maximized. In addition, the overall accuracy was calculated using Eq. 7. The indices, excluding the TSS value, range between 0 to 1, where 0 and 1 indicate totally incorrect and correct predictions, respectively. The TSS value ranges from –1 to 1, where –1 and 1 indicate totally incorrect and correct predictions, respectively, and 0 indicates that the prediction is random47. Although there is no specific classification for evaluating the TSS value, a value over 0.6 is generally considered a sufficiently acceptable result42,43.
All calculations were conducted using Microsoft Excel 2019 (Microsoft Corporation, Redmond, WA, USA).
Data and materials availability
All data are available in the main text, supplementary materials, or from accessing the Water Environment Information System (https://water.nier.go.kr/). If website access is difficult, data can be obtained from the corresponding author on request.
References
Araújo, M. B. & Peterson, A. T. Uses and misuse of bioclimatic envelop modeling. Ecology 93, 1527–1539 (2012).
Pletterbauer, F., Graf, W. & Schmutz, S. Effect of biotic dependencies in species distribution models: the future distribution of Thymallus thymallus under consideration of Allogamus auricollis. Ecol. Modell. 327, 95–104 (2016).
van Proosdij, A. S. J., Sosef, M. S. M., Wieringa, J. J. & Raes, N. Minimum required number of specimen records to develop accurate species distribution models. Ecography 39, 542–552 (2016).
Elith, J. & Leathwick, J. R. Species distribution models: ecological explanation and prediction across space and time. Annu. Rev. Ecol. Evol. Syst. 40, 677–697 (2009).
Varela, S., Lobo, J. M. & Hortal, J. Using species distribution models in paleobiogeography: A matter of data, predictors and concepts. Palaeogeogr. Palaeoclimatol. Palaeoecol. 310, 451–463 (2011).
Feijó, A., Ge, D., Wen, Z., Xia, L. & Yang, Q. Identifying hotspots and priority areas for xenarthran research and conservation. Divers. Distrib. 28, 1–13 (2022).
Guisan, A. et al. Predicting species distribution for conservation decisions. Ecol. Lett. 16, 1424–1435 (2013).
Herrera–R, G. A., Oberdorff, T., Anderson, E. P., Brosse, S., Carvajal–Vallejos, F. M., Frederico, R. G., Hidalgo, M., Jèzèquel, C., Maldonado, M., Maldonado–Ocampo, J. A., Ortega, H., Radinger, J., Torrente–Vilara, G., Zuanon, J. & Tedesco, P. A., The combined effects of climate change and river fragmentation on the distribution of Andean Amazon fishes. Glob. Change Biol. 26, 5509–5523 (2020).
Kim, Z., Shim, T., Ki, S. J., An, K.-G. & Jung, J. Prediction of three–dimensional shift in the distribution of largemouth bass (Micropterus salmoides) under climate change in South Korea. Ecol. Indic. 137, 108731 (2022).
Gallardo, B. & Aldridge, D. C. Priority setting for invasive species management by the water industry. Water Res. 178, 115771 (2020).
Palacio, F. X. et al. A protocol for reproducible functional diversity analyses. Ecography 11, e06287 (2022).
Wicquart, J., Gudka, M., Obura, D., Logan, M., Staub, F., Souter, D. & Planes, S., A workflow to integrate ecological monitoring data from different sources. Ecol. Inform. 68 (2022).
Zuckerberg, B., Huettmann, F. & Frair, J., Proper data management as a scientific foundation for reliable species distribution modeling, In: Predictive Species and Habitat Modeling in Landscape Ecology (Springer, NY, 2011) chap. 4.
Robertson, M. P., Visser, V. & Hui, C. Biogeo: an R package for assessing and improving data quality of occurrence record datasets. Ecography. 39, 394–401 (2016).
Hanberry, B. B., He, H. S. & Dey, D. C. Sample sizes and model comparison metrics for species distribution models. Ecol. Modell. 227, 29–33 (2012).
Papeş, M. & Gaubert, P. Modelling ecological niches from low numbers of occurrences: assessment of the conservation status of poorly known viverrids (Mammalia, Carnivora) across two continents. Divers. Distrib. 13, 890–902 (2007).
Pearson, R. G., Raxworthy, C. J., Nakamura, M. & Pearson, A. T. Predicting species distributions from small numbers of occurrence records: a test case using cryptic geckos in Madagascar. J. Biogeogr. 34, 102–117 (2007).
Wisz, M. S., Hijmans, R. J., Li, J., Peterson, A. T., Graham, C. H., Guisan, A., Elith, J., Dudík, M., Ferrier, S., Huettmann, F., Leathwick, J. R., Lehmann, A., Lohmann, L., Loiselle, B. A., Manion, G., Moritz, C., Nakamura, M., Nakazawa, Y., Overton, J. McC., Phillips, S. J., Richardson, K. S., Scachetti–Pereira, R., Schapire, R. E., Soberón, J., Williams, S. E. & Zimmermann, N. E., Effects of sample size on the performance of species distribution models. Divers. Distrib. 14, 763–773 (2008).
Hernandez, P. A., Graham, C. H., Master, L. L. & Albert, D. L. The effect of sample size and species characteristics on performance of different species distribution modeling methods. Ecography 29, 773–785 (2006).
Stockwell, D. R. B. & Peterson, A. T. Effects of sample size on accuracy of species distribution models. Ecol. Modell. 148, 1–13 (2002).
Zurell, D. et al. A standard protocol for reporting species distribution models. Ecography 43, 1261–1277 (2020).
Lee-Yaw, J. A., McCune, J. L., Pironon, S. & Sheth, S. N. Species distribution models rarely predict biology of real populations. Ecography 2022, e05877 (2022).
Newcomb, S. Note on the frequency of use of the different digits in natural numbers. Am. J. Math. 4, 39–40 (1881).
Benford, F. The law of anomalous numbers. Proc. Am. Philos. Soc. 78, 551–572 (1938).
Maher, M. & Akers, M. Using Benford’s law to detect fraud in the insurance industry. Int. Bus. Econ. Res. 1, 1–12 (2002).
de Vries, P. & Murk, A. J. Compliance of LC50 and NOEC data with Benford’s law: an indication of reliability?. Ecotoxicol. Environ. Saf. 98, 171–178 (2013).
Nigrini, M. J. & Miller, S. J. Benford’s law applied to hydrology data–results and relevance to other geophysical data. Math. Geol. 39, 469–490 (2007).
Berger, A. & Hill, T. P. An Introduction to Benford’s Law 223–230 (Princeton University Press, USA, 2015).
Sambridge, M., Tkalčić, H. & Arroucau, P. Benford’s law of first digits: from mathematical curiosity to change detector. Asia Pac. Math. Newsl. 1, 1–6 (2011).
Sambridge, M., Tkalčić, H. & Jackson, A. Benford’s law in the natural sciences. Geophys. Res. Lett. 37, L22301 (2010).
Joannes-Boyau, R., Bodin, T., Scheffers, A., Sambridge, M. & May, S. M. Using Benford’s law to investigate Natural Hazard dataset homogeneity. Sci. Rep. 5, 12046 (2015).
Özkan, K. Estimating ecosystem naturalness using Benford’s law and generalized Benford’s law. Turk. J. For. 22, 73–82 (2021).
Lee, K. B., Han, S. & Jeong, Y. COVID–19, flattening the curve, and Benford’s law. Phys. A. 559, 125090 (2020).
Costas, E., López-Rodas, V., Toro, F. J. & Flores-Moya, A. The number of cells in colonies of the cyanobacterium Microcystis aeruginosa satisfies Benford’s law. Aquat. Bot. 89, 341–343 (2008).
Özkundakci, D. & Pingram, M. A. Nature favours “one” as the leading digit in phytoplankton abundance data. Limnologica 78, 125707 (2019).
Moriasi, D. N., Gitau, N. W., Pai, N. & Daggupati, P. Hydrologic and water quality models: performance measures and evaluation criteria. Trans. Am. Soc. Agric. Biol. Eng. 58, 1763–1785 (2015).
Shim, T. et al. Integrating hydraulic and physiologic factors to develop an ecological habitat suitability model. Environ. Modell. Softw. 131, 104760 (2020).
Polidori, L. & El Hage, M. Application de la Loi de Benford AU Contrôle de Qualité Des Modèles Numériques de Terrain. Rev. XYZ. 158, 19–22 (2019).
Noleto-Filho, E. M., Carvalho, A. R., Thomè-Souza, M. J. F. & Angelini, R. Reporting the accuracy of small–scale fishing data by simply applying Benford’s law. Front. Mar. Sci. 9, 947503 (2022).
Szabo, J. K., Forti, L. R. & Callaghan, C. T. Large biodiversity datasets conform to Benford’s law: Implications for assessing sampling heterogeneity. Biol. Conserv. 280, 109982 (2023).
Liu, C., White, M. & Newell, G. Selecting thresholds for the prediction of species occurrence with presence-only data. J. Biogeogr. 40, 778–789 (2013).
Gallien, L., Douzet, R., Pratte, S., Zimmermann, N. E. & Thuiller, W. Invasive species distribution models – how violating the equilibrium assumption can create new insights. Glob. Ecol. Biogeogr. 21, 1126–1136 (2012).
Shrestha, U. B., Sharma, K. P., Devkota, A., Siwakoti, M. & Shrestha, B. B. Potential impact of climate change on the distribution of six invasive alien plants in Nepal. Ecol. Indic. 95, 99–107 (2018).
Huang, J. & Frimpong, E. A. Limited transferability of stream-fish distribution models among river catchments: reasons and implications. Freshw. Biol. 61, 729–744 (2016).
Zhang, X. D. et al. Robust statistical methods for hit selection in RNA interference high-throughput screening experiments. Pharmacogenomics 7, 299–309 (2006).
Respondek, I. & Benoit, D. M. Fast degenerate correlation-corrected vibrational self-consistent field calculations of the vibrational spectrum of 4-mercaptopyridine. J. Chem. Phys. 131, 054109 (2009).
Allouche, O., Tsoar, A. & Kadmon, R. Assessing the accuracy of species distribution models: prevalence, kappa and the true skill statistics (TSS). J. App. Ecol. 43, 1223–1232 (2006).
Acknowledgements
The authors would like to thank the National Institute of Environmental Research for providing details of the freshwater fish monitoring data and the support of a Korea University Grant.
Funding
National Research Foundation of Korea grant NRF–2021R1I1A1A01060115.
Author information
Authors and Affiliations
Contributions
Conceptualization: TS, Methodology: TS, ZK, Investigation: TS, ZK, Visualization: ZK, Funding acquisition: TS, Project administration: TS, Supervision: JJ, Writing – original draft: TS, Writing – review & editing: JJ.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Ethical approval for animal experiments.
No experiments with live animals were conducted in this research.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Shim, T., Kim, Z. & Jung, J. A Benford’s law-based framework to determine the threshold of occurrence sites for species distribution modelling from ecological monitoring databases. Sci Rep 13, 16777 (2023). https://doi.org/10.1038/s41598-023-44010-z
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-023-44010-z
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.