Abstract
The translocation t(1;19)(q23;p13) with the resulting chimeric TCF3::PBX1 gene is the third most prevalent recurrent chromosomal translocation in acute lymphoblastic leukemia and accounts for 3–5% of cases. The molecular background of this translocation has been incompletely studied, especially in adult cases. We characterized the chromosomal breakpoints of 49 patients with TCF3::PBX1 and the corresponding reciprocal PBX1::TCF3 breakpoints in 15 cases at the molecular level, thus providing an extensive molecular overview of this translocation in a well-defined study patient population. Breakpoints were found to be remarkably clustered not only in TCF3 but also in PBX1. No association with DNA repeats or putative cryptic recombination signal sequence sites was observed. A simplified detection method for breakpoint identification was developed and the feasibility of patient-specific chromosomal break sites as molecular markers for detecting measurable residual disease (MRD) was explored. A highly sensitive generic real-time PCR for MRD assessment using these breakpoint sequences was established that could serve as a useful alternative to the classical method utilizing rearranged immune gene loci. This study provides the first extensive molecular data set on the chromosomal breakpoints of the t(1;19)/TCF3::PBX1 aberration in adult ALL. Based on the obtained data a generic MRD method was developed that has several theoretical advantages, including an on average higher sensitivity and a greater stability of the molecular marker in the course of disease.
Similar content being viewed by others
Introduction
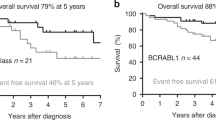
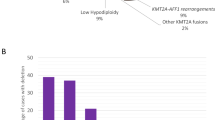
The chromosomal translocation t(1;19)(q23;p13) with the formation of a chimeric TCF3::PBX1 gene (E2A-PBX1 in older nomenclature) is detected in approximately 3–5% of pediatric and adult acute lymphoblastic leukemia (ALL) cases. Despite its relative rarity, the translocation is still the third most frequent recurrent chromosomal translocation in ALL (after t(9;22)/BCR::ABL1 and t(4;11)/KMT2A::AFF1 in adult ALL and after t(12;21)/ETV6::RUNX1 and t(4;11)/KMT2A::AFF1 in pediatric ALL)1. Affected patients exhibit a characteristic B-cell immunophenotype (CD19+/CD10+/CD33−/CD34−/sIg−, and mostly cyIg+), and gene expression analyses have indicated that TCF3::PBX1-positive patients constitute a separate entity among ALL patients2,3,4. The current WHO classification includes TCF3::PBX1-positive ALL as a distinct subgroup of B-lymphoblastic leukemia5. Historically, TCF3::PBX1-positive leukemia has been associated with a poor prognosis, but this has been overcome by modern therapy regimens and TCF3::PBX1 currently defines a group of ALL patients with a good clinical outcome in childhood ALL16,6,7,8,9, although these patients appear to have an increased risk for CNS involvement at diagnosis10. The prognostic impact of TCF3::PBX1 in ALL in older patients (age > 15 years) is less well defined, and relatively few molecular-based and controversial data have been published in this area11,12,13,14,15. TCF3::PBX1-positive ALL has been found to express the receptor tyrosine kinase ROR1, which may serve as a therapeutic target in the future16,17. Some promising therapeutic in vitro effects have been observed with the SRC inhibitor dasatinib18 and the phosphatidylinositide 3-kinase delta (p110δ) inhibitor idelalisib19. However, no established targeted therapy currently exists for TCF3::PBX1-positive patients, and the assessment of measurable residual disease (MRD) remains the most important tool in therapy stratification and prognostication.
Data on the molecular details of the t(1;19)(q23;p13) translocation in adult ALL are scant. The following work analyzed 49 TCF3::PBX1-positive, clinically well-defined adult cases, identified the chromosomal break sites, and characterized the molecular background of this translocation, thus providing a detailed and extensive molecular overview of this translocation. A method for the easy identification of the breakpoint sites is presented, and the potential utilization of these chromosomal breakpoints for detecting measurable residual disease is demonstrated.
Results
Rationale for use and development of a long range-inverse PCR method
One type of chimeric RNA transcript was predominantly found in TCF3::PBX1-positive patients, showing a fusion of TCF3 exon 16 (reference sequence NG_029953.2) and PBX1 exon 3 (reference sequence NG_028246.2)26. Other transcripts have been described, but they seem to be very rare27. Chromosomal breaks can thus be assumed to occur in the intron 3′ of TCF3 exon 16 (“intron 16”) and in the intron 5′ of PBX1 exon 3 (“intron 2”). The TCF3 reference sequence includes a 3289 bp intron 16 (ncl 1615822–1619110, NC_000019.10, GRCh38.p13 primary assembly). This intron is present in all 41 TCF3 variants listed in the NCBI gene database (updated on 1-Aug-2020). The location of the breakpoint site on chromosome 1 is less clear (8 PBX1 transcript variants with either a 229,182 bp intron 2 (ncl 164563312–164792493) or a 166,397 bp intron 2 (ncl 164626097–164792493). Since the breakpoint region on chromosome 19 appeared to be relatively localized, a long range-inverse PCR (LRI PCR) approach was chosen for the analysis. Commercially available restriction enzymes were screened for those with restriction sites flanking the putative breakpoint region on chromosome 19. Three enzymes were suitable because they had palindromic cutting sites without degenerate nucleotides, produced sticky ends and were frequent cutters: SphI, BamHI and TaqI (Fig. 1A). BamHI had one cutting site 148 bp 5′ of the TCF3 intron end; thus, breakpoints near the intron end could not be detected using this enzyme. The three enzymes provided dense coverage of PBX1 intron 2 with restriction sites (Table S1 and Figure S1).
Various PCR primers and PCR conditions were tested for the development of the inverse PCR method. The efficacy of inverse PCR could be optimized because the three enzymes produced a detectable “control” PCR product when testing normal DNA. The final PCR primer locations are depicted in Fig. 1A.
Analysis of patient samples and chromosomal breakage data
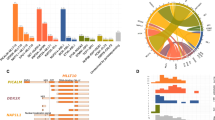
Patient samples were first analyzed with SphI, followed by BamHI and TaqI. Twenty-six TCF3::PBX1 breakpoints were identified with SphI, 12 with BamHI and 11 with TaqI. PCR examples are shown in Fig. 1B. Fifty four TCF3::PBX1-RT-PCR-positive samples were analyzed and the genomic TCF3::PBX1 fusion site was identified in 49 cases. The fact that five samples remained uncharacterized on the genomic level may reflect a limitation of the method. Insufficient DNA quality may also have played a role since some samples were more than 15 years old. Of these 49 TCF3::PBX1 sites, the reciprocal PBX1::TCF3 breakpoint was identified in 15 cases. One sample (5741) harbored an inversion with two breaks in PBX1. The break sites displayed a remarkable nonrandom pattern (Fig. 2, Table 1). Fifty-three (83%; 41 TCF3::PBX1, 12 PBX1::TCF3) were located in a narrow 40 bp region in TCF3 intron 16, while 11 (17%; 8 TCF3::PBX1, 3 PBX1::TCF3) occurred outside this region. In PBX1 intron 2 the clustering was less dense but still apparent. Two major break clusters could be delineated in PBX1: one from ~ 220 to 229 kb (near the intron end), and a second from ~ 120 to ~ 155 kb. These two clusters comprised 34 kb (14.8% of the intron) and 88% of all break events (Fig. 2, Table 1). Twenty-four breaks (37%; 16 TCF3::PBX1, 8 PBX1::TCF3) were located in the first cluster, 33 (51%; 27 TCF3::PBX1, 6 PBX1::TCF3) in the second cluster, and 8 (12%; 7 TCF3::PBX1, 1 PBX1::TCF3) outside both clusters.
The break sites were analyzed bioinformatically for patterns that could explain the observed distribution. There were no apparent DNA microhomologies at the break sites in PBX1 (Table S2). TCF3 intron 16 included 6 DNA repeats comprising 1699 bp (51.6% of the intron), and PBX1 intron 2 included 342 DNA repeats with 51,699 bp (22.6%, Tables S4, S5). The break cluster in TCF3 intron 16 was located at the 5′ end of a repetitive DNA element (MER20). Eight of the 10 break sites in TCF3 outside the cluster were located in or immediately 3′ of other DNA repetitive elements (L2A, MER20, L1MS, Alujb, AluY; Fig. 2A, Tables 1, S6). For the breaks in PBX1, there was no apparent association with repetitive DNA elements. Eight of the 55 breaks in PBX1 (14.5%) occurred inside repetitive elements (# 3120, 3951, 5741, 5974, 6840, 7236, ML4316, ML11220; Tables 1, S4). Bioinformatical analysis revealed 611 potential cryptic recombination signal sequences (cRSSs) in PBX1 intron 2, covering 20.1 kb (8.8%) of the intron. Five of the 65 breaks in PBX1 (7.7%) occurred in or in the vicinity (± 30 bp) of cRSSs (# 3120, 4641, 7281, ML11220, ML11543; Tables 1, S7).
To complement this analysis, all samples were also investigated for intragenic IKZF1 deletions by PCR. These deletions are found in approximately 20% of B-cell precursor ALL cases and are known to be caused by aberrant VDJ recombinase activity. None of the 49 samples showed an intragenic IKZF1 deletion. This does not exclude a possible role of RAG-mediated secondary aberrations in TCF3::PBX1-rearranged ALL as illustrated by the example of ETV6::RUNX1-positive pediatric ALL286,29.
Chromosomal translocations are occasionally associated with DNA secondary structures, such as inverted repeats with hairpin loops30, and thus, the hotspot region of TCF3 was analyzed with RNAfold. The main break site was located in an open loop that was flanked by regions with relatively strong base pair binding (Fig. S8). The analysis of the 15 cases in which reciprocal PBX1::TCF3 were characterized showed mostly no microhomologies at the break sites, with frequent insertion of nontemplate nucleotides, suggesting a nonhomologous end-joining repair (NHEJ) mechanism31. One sample (4297) showed an insertion from the FGF6 gene on chromosome 12, a gene not previously implicated in the pathogenesis of ALL (Fig. S3).
Development and optimization of a real-time qPCR method
The clustering of chromosomal breaks in a narrow region in TCF3 intron 16 suggested a quantitative PCR method with a common forward primer, a common dual-labeled hybridization probe 5′ of the breakpoint cluster region and a patient-specific reverse primer 3′ of the breakpoint. Several forward primers and dual-labeled probes were first tested on patient samples and control DNA to exclude spurious amplifications. Finally, one combination of a common forward primer and a common dual-labelled probe was selected that was tested on 15 randomly chosen patient samples (Table 2, Fig. 3). In all 15 cases, it was possible to design a reverse PCR primer that yielded data with good sensitivity and specificity. The testing of further samples was not possible because of shortage of sample material. This generic real-time qPCR was designed to quantify breakpoints in the TCF3 hotspot region (~ 80%). Breakpoints outside this region (and likewise the PBX1::TCF3 breakpoints) could theoretically also be used as MRD targets, but in these cases, no generic recipe can be given, and individual patient-specific qPCRs would have to be constructed.
Development and optimization of two multiplex long range PCRs
Since breakpoint identification by long range-inverse PCR is an elaborate procedure and since the breakpoints showed clustering in certain regions, efforts were made to simplify the detection procedure. Two multiplex long-range PCRs with a series of PCR oligonucleotides covering the entire breakpoint regions were developed and optimized that allowed the detection of breakpoints in the two breakpoint cluster regions of PBX1. Examples of these multiplex long-range PCRs are shown in Fig. 1C.
Discussion
The translocation t(1;19)(q23;p13) has been described as mostly unbalanced156,32,33,34. This is in accordance with the observation made in this study that in only 14 cases (33%) a reciprocal break site could be characterized.
Clustering of breakpoints
Since the first description of the translocation t(1;19) as a recurrent aberration in ALL in 1984, research has largely focused on cytogenetic aspects of this aberration, and few investigations have been carried out in adult ALL276,346,35. Wiemels et al.36 first systematically investigated the translocation at a molecular level and described 24 cases from various pediatric ALL studies. The median age of the patients was 6.8 years, with only one patient being an adult > 18 years of age. A similar clustering of breakpoints was observed, and the authors speculated that aberrant VDJ recombinase activity might be involved. They identified a reciprocal breakpoint in 5 (21%) cases36.
In this study, no association of t(1;19) chromosomal breaks with repetitive DNA elements was found. While the location of the break cluster in TCF3 intron 16 close to a MER20 element could be coincidental, there was no similar association of the breaks mapping to PBX1 intron 2. Similarly, no direct association with cryptic RSS was observed. None of the 49 patient samples showed an intragenic IKZF1 deletion—an aberration caused by illegitimate VDJ recombination-mediated deletion, and present in approximately 20% of BCR::ABL1-negative B precursor ALLs. Recently, Liu et al. analyzed the TCF3 “fragile zone” and suggested that the initial TCF3 breakage may arise at a CpG site. They found a statistically significant proximity of the activation-induced cytidine deaminase (AID) hotspot motifs WRC and WGCW near the TCF3 breakpoints W = A or T, R = A or G) suggesting AID involvement in the break process 37. This is consistent with the fact that TCF3::PBX1 is predominantly detected in pre-B ALL, which is immunophenotypically the most “mature” entity in B precursor ALL, indicating a relatively late stage of B-cell development.
Real-time qPCR for measurable residual disease detection
Measurable residual disease in ALL is usually assessed by the use of clonally rearranged immunoglobulin (IG) and/or T-cell receptor (TCR) loci for the construction of real-time quantitative PCRs (qPCRs)38. The main advantage of this approach is its universal applicability. Theoretically, it can be applied in any malignant disease of lymphatic origin. However, this method also has some disadvantages. In a significant minority of cases, it is not possible to identify clonal rearrangements, and with the introduction of next generation sequencing techniques it has become apparent that IG/TCR rearrangements are often in fact polyclonal at diagnosis39. IG/TCR-based MRD monitoring is thus often based on only one of several clones, and such an analysis may miss the decisive clone. IG/TCR-based qPCRs frequently show a suboptimal sensitivity (below 10–4), because of the difficulty of constructing a specific PCR against a highly homologous background. In addition, the IG/TCR rearrangements are potentially unstable, and further rearrangements can occur without loss of the malignant cell phenotype, leading to false negative results.
In those cases where chromosomal translocations lead to the expression of a chimeric mRNA transcript, MRD monitoring can also be performed by the relative quantification of this transcript40. However, this approach has been widely discarded in ALL (with the exception of BCR::ABL1), because it only allows a quantification relative to a “housekeeping gene”, assumed to be stably expressed. “Dormant” tumor stem cells with low expression of the oncogene may escape detection by RT-PCR. This is exemplarily illustrated by the observation that in BCR::ABL1-positive ALL, only a limited correlation between BCR::ABL1-mRNA-based and IG/TCR-based MRD levels is found41. Additionally, RNA is relatively unstable and significantly more difficult to handle than DNA.
An alternative approach is targeting the breakpoint sites of chromosomal translocations to detect and monitor MRD by constructing patient-specific qPCR assays. These are stable molecular markers that cannot be lost in the course of disease because they are linked to molecular drivers of the disease. This approach has been exploited in various entities, such as ALL with t(12;21)/ETV6::RUNX1426,43, ALL with 11q23/KMT2A aberrations446,45, ALL or CML with t(9;22)/BCR::ABL1416,46,47 and other hematopoietic malignancies48,49,50. In most of these cases, it could be shown that the break site-specific PCRs were at least as reliable as the IG/TCR-based methods and yielded a superior sensitivity. The main disadvantages of this approach are the technical difficulties posed by the individual characterization of break sites which may in some cases be dispersed over hundreds of kilobases of genomic DNA, precluding this approach for routine clinical studies with the exception of KMT2A-rearranged ALL, where relatively standardized techniques for break site identification are in use51. With the increasing availability of next-generation sequencing techniques and their technical advances (e.g., nanopore sequencing or mate-pair sequencing) and better knowledge of the molecular background these difficulties are likely to be overcome in the future and MRD detection methods based on chromosomal breakpoints will become increasingly important526,53.
Conclusions
The present work characterizes the t(1;19) chromosomal breakpoints of a large number of adult ALL patients from a well-defined study population and is the largest and the first major investigation on this topic in adult ALL. The results provide a representative and relatively unbiased overview of the molecular details of this aberration. Based on the experimental results, a simplified method for the rapid identification of chromosomal breakpoints is proposed and the usefulness of these chromosomal breakpoint data for measurable residual disease detection is demonstrated. While the theoretical advantages of such an MRD approach appear obvious, clinical studies are necessary to validate the TCF3::PBX1 breakpoint fusion as MRD marker in a clinical context. Further testing and comparisons will have to be performed to fully establish TCF3::PBX1 breakpoints as valuable MRD targets.
Methods
Patient samples and ethics statement
Patient samples were collected from residual diagnostic material obtained between 2001 and 2021 in the context of the German Multicenter ALL Therapy Studies (clinicaltrials.gov identifiers: 00199056 and 00198991). Patients gave written informed consent to scientific investigations on study inclusion and the studies were approved by local and central ethics committees, among them an ethics board of the Goethe University, Frankfurt/Main, Germany and the ethics board of the Charité Universitätsmedizin, Berlin, Germany. Our study complied with the principles set forth in the World Medical Association Declaration of Helsinki.
Patient characteristics
Patient clinical details are summarized in Table 1. All patient samples included in this study (31 bone marrow, 17 peripheral blood, one unspecified) had been investigated by flow cytometry and RT-PCR at diagnosis. All patients exhibited a B precursor immunophenotype (CD19+/CD10+/CD33−/CD34−/sIg−). Forty-three (88%) showed a cyIg+ (pre B) and six a cyIg− (common) immunophenotype. All samples were tested negative for BCR::ABL1 and positive for TCF3::PBX1 by RT-PCR. Twenty-seven (55%) of the 49 patients were female and 22 male. The median age was 39.5 years (range 17–77 years).
DNA isolation
DNA was isolated from archived or fresh samples using either the Gentra PureGene method (QIAGEN, Hilden, Germany), the AllPrep DNA/RNA Kit (QIAGEN) or in a few cases the DNA preparation from TRIzol (ThermoFisher Scientific, Darmstadt, Germany) with subsequent DNA purification.
Long range-inverse PCR (LRI PCR)
The LRI PCR methods were developed and optimized for this study. The following restriction enzymes were used: SphI (GCATG|C), BamHI (G|GACC) and TaqI (T|CGA). FastDigest enzymes were used according to the manufacturer’s recommendations (ThermoFisher Scientific, Darmstadt, Germany). The conditions for the long range-inverse PCR were partially adopted from previous work20. Five hundred nanograms of genomic DNA was digested in a 50 µl volume, the reaction mix was inactivated, purified using the MaXtract High Density kit (QIAGEN, Hilden, Germany), ethanol-precipitated and dissolved in a final volume of 30 µl. The entire volume was used in the ligation procedure (50 µl final volume, 5 U T4 ligase, 16 °C overnight). After purification and ethanol precipitation as described above, the ligation mix was dissolved in 30 µl H2O. Five microliters was used in the long-range PCR with the Expand Long Template PCR System kit (Roche, Mannheim, Germany) with buffer 2 and the following cycler program: 95 °C 2 min, 15 cycles (94 °C 30 s, 65 °C 30 s, 68 °C 6 min), 20 cycles (94 °C 30 s, 65 °C 30 s, 68 °C 5 min with 20 s increment/cycle), and 68 °C 10 min, 4 °C. One enzyme-specific reverse (R) PCR primer was combined with a forward (F) primer. The following primer combinations were used: SphI: TCF3-F2/TCF3-R5, BamHI: TCF3-F2/TCF3-R4, TaqI: TCF3-F2/TaqI-R. If a PCR product was visible the PCR was repeated and primer TCF3-F2 replaced by primers TCF3-F7 or TCF3-F6 to try to generate a smaller PCR product for easier sequencing. PCR products of interest were excised from the agarose gel, purified, and analyzed by Sanger sequencing.
Oligonucleotide sequences for the long range-inverse PCR
All oligonucleotides were obtained from tib molbiol (Berlin, Germany). The LRI PCR primer sequences were (5′-3′): TCF3-R4 GAAGGCCTGGGCTACGGAGGGGAACAGCT, TCF3-F2 CTCCCTGACCTGTCTCGGCCTCCCGACT, TCF3-F6 ACCTTGATTCTATCACTCCTAGGCCAGGGCA, TCF3-R5 CACAGGCCTCCATTCATGTCCCTTCCGCA, TaqI-R AGGCCGTGGAGACCCCCGTCGTAGCT. Normal DNA (without t(1;19) translocation) generated “control bands” of the following sizes: TCF3-F2/TCF3-R4 3308 bp, TCF3-F6/TCF3-R4 2131 bp, TCF3-F2/TCF3-R5 5026 bp, TCF3-F6/TCF3-R5 3849 bp, TCF3-F2/TaqI-R 7567 bp, TCF3-F6/TaqI-R 6390 bp.
Sanger sequencing
Apart from the oligonucleotides detailed above several ad hoc designed oligonucleotides were used for Sanger sequencing of individual samples. Technical Sanger sequencing of PCR products was performed by Microsync SeqLab (Göttingen, Germany). Analysis of chromatograms and sequence data assembly was performed at the Charité laboratory in Berlin.
Sequence data
All nucleotide sequence data (91 328 bp) were submitted to the GenBank/ENA/DDBJ database and are available under the accession numbers OK334233-OK334288, ON383218-ON383224, ON809522.
Real-time quantitative PCR
Real-time qPCR was performed on a RotorGene RG-3000 cycler (formerly Corbett Research, now subsidiary of QIAGEN, Hilden, Germany) using the ABgene PCR QPCR Mix (Thermo Fisher Scientific, Darmstadt, Germany). The forward primer TCF3-qF 5′-CAGGCAGACTTTCCAAGTACCTT-3′ was used with the dual-labelled probe TCF3-FAM 5′-6FAM-CTATCACTCCTAGGCCAGGGCATCT-BHQ1-3′ and a patient-specific reverse primer.
Multiplex long range PCR
The two multiplex long range PCRs comprised the following oligonucleotides (100 nM of each oligonucleotide per reaction mix). For breakpoint cluster 1 (5′-3′): TCF3-F7 AGGAGGGTTTCAGGCAGAGGGCGCA, PBX-long1 CCCGGGGTTGTGCTTCCTCCACCCTT, PBX-long2 TGCGCTCTCTCCCTCCCCCTCATCTCT, PBX-long3 ACGTGGTCCTGCGAGGAGCTCTTAGA, PBX-long4 TGCCCATGCAGCAGGTGACAAGGG, and for breakpoint cluster 2 (5′-3′): TCF3-F7, PBX1-long5 ACGAATCAGGCAGCTGTACAGAAAGCA, PBX1-long6 TCGGCCTCACCTAACTGACTTGCAGGT, PBX1-long7 AGCACCATCCTGAAGTTGCTCGGCT, PBX1-long8 TGCGGGAGGCTGGCAACATTGAGTC, PBX1-long9 ACACAGGTGCTACCTCTGCTCTGCCA, PBX1-long10 TCCAGCTACCTCATGGCTCGCTAGA. PCR conditions were the same as for the LRI PCR.
PCR for IKZF1 deletions
The main four intragenic IKZF1 deletion variants Δ2–7, Δ2–8, Δ4–7, Δ4–8 were investigated by four different PCRs, and the intragenic IKZF1 deletion variant Δ2–3 by one single RT-PCR as outlined previously21.
Bioinformatics and software
Genomic repeats were analyzed with RepeatMasker version 4.0.9, RSSSite and the Tandem repeats finder22,23,24. DNA secondary structures were investigated using RNAfold 2.4.1825. Sanger sequence chromatograms were analyzed with 4Peaks (Nucleobytes, Aalsmeer, The Netherlands) and Nucleotide BLAST (blastn) against the GRCh38.13 reference primary assembly human genome.
Data availability
All nucleotide sequences generated and analyzed during the current study are available in the GenBank/ENA/DDBJ database under the accession numbers OK334233-OK334288, ON383218-ON383224, ON809522.
References
Pui, C. H., Relling, M. V. & Downing, J. R. Acute lymphoblastic leukemia. N. Engl. J. Med. 350, 1535–1548 (2004).
Chiaretti, S. et al. Gene expression profiles of B-lineage adult acute lymphocytic leukemia reveal genetic patterns that identify lineage derivation and distinct mechanisms of transformation. Clin. Cancer Res. 11, 7209–7219 (2005).
Li, J. F. et al. Transcriptional landscape of B cell precursor acute lymphoblastic leukemia based on an international study of 1223 cases. Proc. Natl. Acad. Sci. U S A. 115, E11711–E11720 (2018).
Alexander, T. B. & Mullighan, C. G. Molecular Biology of Childhood Leukemia. Ann. Rev. Cancer Biol. 5, 95–117 (2021).
Alaggio, R. et al. The 5th edition of the world health organization classification of haematolymphoid tumours: lymphoid neoplasms. Leukemia 36, 1720–1748 (2022).
Asai, D. et al. Outcome of TCF3-PBX1 positive pediatric acute lymphoblastic leukemia patients in Japan: a collaborative study of Japan Association of Childhood Leukemia Study (JACLS) and Children’s Cancer and Leukemia Study Group (CCLSG). Cancer Med. 3, 623–631 (2014).
Yen, H. J. et al. Pediatric acute lymphoblastic leukemia with t(1;19)/TCF3-PBX1 in Taiwan. Pediatr. Blood Cancer 64, e26557 (2017).
Lin, A. et al. Excellent outcome of acute lymphoblastic leukaemia with TCF3-PBX1 rearrangement in Hong Kong. Pediatr. Blood Cancer. 65, e27346 (2018).
Jia, M. et al. Clinical features and prognostic impact of TCF3-PBX1 in childhood acute lymphoblastic leukemia: A single-center retrospective study of 837 patients from China. Curr. Probl. Cancer 45(6), 100758 (2021).
Pui, C. H. et al. Treating childhood acute lymphoblastic leukemia without cranial irradiation. N Engl J Med. 360, 2730–2741 (2009).
Foà, R. et al. E2A-PBX1 fusion in adult acute lymphoblastic leukaemia: biological and clinical features. Br J Haematol. 120, 484–487 (2003).
Moorman, A. V. New and emerging prognostic and predictive genetic biomarkers in B-cell precursor acute lymphoblastic leukemia. Haematologica 101, 407–416 (2016).
Burmeister, T. et al. Clinical features and prognostic implications of TCF3-PBX1 and ETV6-RUNX1 in adult acute lymphoblastic leukemia. Haematologica 95, 241–246 (2010).
Yilmaz, M. et al. Translocation t(1;19)(q23;p13) in adult acute lymphoblastic leukemia—A distinct subtype with favorable prognosis. Leuk. Lymphoma. 62, 224–228 (2021).
Ribera, J. et al. Prognostic heterogeneity of adult B-cell precursor acute lymphoblastic leukaemia patients with t(1;19)(q23;p13)/TCF3-PBX1 treated with measurable residual disease-oriented protocols. Br. J. Haematol. 196, 670–675 (2022).
Karvonen, H., Niininen, W., Murumägi, A. & Ungureanu, D. Targeting ROR1 identifies new treatment strategies in hematological cancers. Biochem. Soc. Trans. 45, 457–464 (2017).
Zhao, Y. et al. Tyrosine kinase ROR1 as a target for anti-cancer therapies. Front. Oncol. 11, 680834 (2021).
Bicocca, V. T. et al. Crosstalk between ROR1 and the Pre-B cell receptor promotes survival of t(1;19) acute lymphoblastic leukemia. Cancer Cell 22, 656–667 (2012).
Eldfors, S. et al. Idelalisib sensitivity and mechanisms of disease progression in relapsed TCF3-PBX1 acute lymphoblastic leukemia. Leukemia 31, 51–57 (2017).
Burmeister, T. et al. Fine structure of translocation breakpoints within the major breakpoint region in BCR-ABL1-positive leukemias. DNA Repair 10, 1131–1137 (2011).
Kobitzsch, B. et al. Loss-of-function but not dominant-negative intragenic IKZF1 deletions are associated with an adverse prognosis in adult BCR-ABL-negative acute lymphoblastic leukemia. Haematologica 102, 1739–1747 (2017).
Tempel, S. Using and understanding repeatmasker. Methods Mol. Biol. 859, 29–51 (2012).
Merelli, I. et al. RSSsite: a reference database and prediction tool for the identification of cryptic recombination signal sequences in human and murine genomes. Nucleic Acids Res. 38, W262–W267 (2010).
Benson, G. Tandem repeats finder: A program to analyze DNA sequences. Nucleic Acids Res. 27, 573–580 (1999).
Lorenz, R. et al. ViennaRNA Package 2.0. Algorithms Mol. Biol. 6, 26 (2011).
van Dongen, J. J. et al. Standardized RT-PCR analysis of fusion gene transcripts from chromosome aberrations in acute leukemia for detection of minimal residual disease. Report of the BIOMED-1 Concerted Action: Investigation of minimal residual disease in acute leukemia. Leukemia 13, 1901–1928 (1999).
Paulsson, K. et al. Characterisation of genomic translocation breakpoints and identification of an alternative TCF3/PBX1 fusion transcript in t(1;19)(q23;p13)-positive acute lymphoblastic leukaemias. Br. J. Haematol. 138, 196–201 (2007).
Papaemmanuil, E. et al. RAG-mediated recombination is the predominant driver of oncogenic rearrangement in ETV6-RUNX1 acute lymphoblastic leukemia. Nat. Genet. 46, 116–125 (2014).
Brady, S. W. et al. The genomic landscape of pediatric acute lymphoblastic leukemia. Nat. Genet. 54, 1376–1389 (2022).
Thys, R. G., Lehman, C. E., Pierce, L. C. & Wang, Y. H. DNA secondary structure at chromosomal fragile sites in human disease. Curr. Genom. 16, 60–70 (2015).
Aplan, P. D. Causes of oncogenic chromosomal translocation. Trends Genet. 22, 46–55 (2006).
Andersen, M. K. et al. Paediatric B-cell precursor acute lymphoblastic leukaemia with t(1;19)(q23;p13): Clinical and cytogenetic characteristics of 47 cases from the Nordic countries treated according to NOPHO protocols. Br. J. Haematol. 155, 235–243 (2011).
Mullighan, C. G. Molecular genetics of B-precursor acute lymphoblastic leukemia. J Clin Invest. 122, 3407–3415 (2012).
Paulsson, K., Horvat, A., Fioretos, T., Mitelman, F. & Johansson, B. Formation of der(19)t(1;19)(q23;p13) in acute lymphoblastic leukemia. Genes Chromosomes Cancer 42, 144–148 (2005).
Carroll, A. J. et al. Pre-B cell leukemia associated with chromosome translocation 1;19. Blood 63, 721–724 (1984).
Wiemels, J. L. et al. Site-specific translocation and evidence of postnatal origin of the t(1;19) E2A-PBX1 fusion in childhood acute lymphoblastic leukemia. Proc. Natl. Acad. Sci. U S A. 99, 15101–15106 (2002).
Liu, D., Loh, Y. E., Hsieh, C. L. & Lieber, M. R. Mechanistic basis for chromosomal translocations at the E2A gene and its broader relevance to human B cell malignancies. Cell Rep. 36, 109387 (2021).
van der Velden, V. H. et al. Analysis of minimal residual disease by Ig/TCR gene rearrangements: guidelines for interpretation of real-time quantitative PCR data. Leukemia 21, 604–611 (2007).
Brüggemann, M. et al. Standardized next-generation sequencing of immunoglobulin and T-cell receptor gene recombinations for MRD marker identification in acute lymphoblastic leukaemia; a EuroClonality-NGS validation study. Leukemia 33, 2241–2253 (2019).
Gabert, J. et al. Standardization and quality control studies of ‘real-time’ quantitative reverse transcriptase polymerase chain reaction of fusion gene transcripts for residual disease detection in leukemia—A Europe Against Cancer program. Leukemia 17, 2318–2357 (2003).
Hovorkova, L. et al. Monitoring of childhood ALL using BCR-ABL1 genomic breakpoints identifies a subgroup with CML-like biology. Blood 129, 2771–2781 (2017).
Metzler, M. et al. Minimal residual disease analysis in children with t(12;21)-positive acute lymphoblastic leukemia: comparison of Ig/TCR rearrangements and the genomic fusion gene. Haematologica 91, 683–686 (2006).
Hoffmann, J. et al. High sensitivity and clonal stability of the genomic fusion as single marker for response monitoring in ETV6-RUNX1-positive acute lymphoblastic leukemia. Pediatr. Blood Cancer 66, e27780 (2019).
Van der Velden, V. H. et al. Prognostic significance of minimal residual disease in infants with acute lymphoblastic leukemia treated within the Interfant-99 protocol. Leukemia 23, 1073–1079 (2009).
Burmeister, T. et al. Monitoring minimal residual disease by quantification of genomic chromosomal breakpoint sequences in acute leukemias with MLL aberrations. Leukemia 20, 451–457 (2006).
Bartley, P. A. et al. Sensitive detection and quantification of minimal residual disease in chronic myeloid leukaemia using nested quantitative PCR for BCR-ABL DNA. Int. J. Lab. Hematol. 32, e222–e228 (2010).
Krumbholz, M. et al. Large amplicon droplet digital PCR for DNA-based monitoring of pediatric chronic myeloid leukaemia. J. Cell. Mol. Med. 23, 4955–4961 (2019).
Zerkalenkova, E. et al. Molecular characteristic of acute leukemias with t(16;21)/FUS-ERG. Ann. Hematol. 97, 977–988 (2018).
Duployez, N. et al. Minimal residual disease monitoring in t(8;21) acute myeloid leukemia based on RUNX1-RUNX1T1 fusion quantification on genomic DNA. Am. J. Hematol. 89, 610–615 (2014).
Krumbholz, M. et al. Characterization and diagnostic application of genomic NPM-ALK fusion sequences in anaplastic large-cell lymphoma. Oncotarget 9, 26543–26555 (2018).
Meyer, C. et al. Diagnostic tool for the identification of MLL rearrangements including unknown partner genes. Proc. Natl. Acad. Sci. U S A. 102, 449–454 (2005).
Cumbo, C. et al. Genomic BCR-ABL1 breakpoint characterization by a multi-strategy approach for “personalized monitoring” of residual disease in chronic myeloid leukemia patients. Oncotarget 9, 10978–10986 (2018).
Rowsey, R. A. et al. Characterization of TCF3 rearrangements in pediatric B-lymphoblastic leukemia/lymphoma by mate-pair sequencing (MPseq) identifies complex genomic rearrangements and a novel TCF3/TEF gene fusion. Blood Cancer J. 9, 81 (2019).
Acknowledgements
The authors are grateful to all participating clinics and physicians of the GMALL study group for their support. This work was supported by a grant from the Alfred und Angelika Gutermuth Stiftung, Frankfurt to TB.
Funding
Open Access funding enabled and organized by Projekt DEAL.
Author information
Authors and Affiliations
Contributions
T.B. was the principal investigator, designed research, analyzed data and wrote the manuscript. D.G. performed technical work (PCR, etc.). N.G. is head of the GMALL study group, D.H. is the former head of the GMALL study group, B.S., M.S., and A.E. are major patient samples contributors, U.K. is head of the Charité Dept. of Hematology, S.S. performed immunophenotyping. All authors critically read and made contributions to the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Burmeister, T., Gröger, D., Gökbuget, N. et al. Molecular characterization of TCF3::PBX1 chromosomal breakpoints in acute lymphoblastic leukemia and their use for measurable residual disease assessment. Sci Rep 13, 15167 (2023). https://doi.org/10.1038/s41598-023-42294-9
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-023-42294-9
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.