Abstract
Hepatocellular carcinoma (HCC) is the most common lethal malignancy of the liver worldwide. Thus, it is important to dig the key genes for uncovering the molecular mechanisms and to improve diagnostic and therapeutic options for HCC. This study aimed to encompass a set of statistical and machine learning computational approaches for identifying the key candidate genes for HCC. Three microarray datasets were used in this work, which were downloaded from the Gene Expression Omnibus Database. At first, normalization and differentially expressed genes (DEGs) identification were performed using limma for each dataset. Then, support vector machine (SVM) was implemented to determine the differentially expressed discriminative genes (DEDGs) from DEGs of each dataset and select overlapping DEDGs genes among identified three sets of DEDGs. Enrichment analysis was performed on common DEDGs using DAVID. A protein-protein interaction (PPI) network was constructed using STRING and the central hub genes were identified depending on the degree, maximum neighborhood component (MNC), maximal clique centrality (MCC), centralities of closeness, and betweenness criteria using CytoHubba. Simultaneously, significant modules were selected using MCODE scores and identified their associated genes from the PPI networks. Moreover, metadata were created by listing all hub genes from previous studies and identified significant meta-hub genes whose occurrence frequency was greater than 3 among previous studies. Finally, six key candidate genes (TOP2A, CDC20, ASPM, PRC1, NUSAP1, and UBE2C) were determined by intersecting shared genes among central hub genes, hub module genes, and significant meta-hub genes. Two independent test datasets (GSE76427 and TCGA-LIHC) were utilized to validate these key candidate genes using the area under the curve. Moreover, the prognostic potential of these six key candidate genes was also evaluated on the TCGA-LIHC cohort using survival analysis.
Similar content being viewed by others
Introduction
Hepatocellular carcinoma (HCC) is the 3rd leading cause of cancer deaths globally1. Globally, more than of 80% liver cancers are responsible for HCC2 and its prevalence is high in males compared to females3. It usually occurs in people aged 30–50 years3. Different factors such as hepatitis B or hepatitis C4,5, alcohol abuse, smoking, obesity, and type 2 diabetes (T2D) were significantly associated with HCC6. Among them, Hepatitis B is one of the prominent risk factors for the development of HCC, responsible for 50% of cases7. Despite various treatment approaches, namely radiotherapy, chemotherapy, and target therapy have been commonly used to improve the prognosis and recurrence of HCC. Nevertheless, the survival rate of HCC patients is still low8. As a result, the risks of cancer death are still increased due to the lack of early detection and diagnosis of genes and limited treatment facilities. Therefore, it is essential to develop a system for identifying the key or core genes for early detection and better prognosis of HCC.
Recently, bioinformatics analysis has been widely utilized to determine the key prognostic genes or biomarkers as well as their associated molecular pathways for multiple cancers, including HCC8,9,10,11,12,13,14,15,16,17,18,19,20,21,22,23,24,25,26,27,28,29,30,31,32,33,34,35,36,37,38,39,40,41,42,43,44,45,46,47,48,49,50,51,52,53,54,55,56,57,58. Zhou et al.35 identified 15 prognostic biomarkers as well as their associated gene ontology (GO) enrichment and Kyoto Encyclopedia of Genes and Genomes (KEGG) pathway using bioinformatics analysis. Chen et al.39,59,60 also identified 11 potential biomarkers that can play crucial roles in the development and progression of HCC patients. Qiang et al.40 proposed five core genes which were significantly associated with early diagnosis and poor prognosis of HBV-HCC. Wang et al.41 identified 36 hub DEGs and illustrated that 10 candidate genes out of the 36 have significant effect on the tumorigenesis and progression of HCC. Among them, eight candidate genes were inversely related to the survival rate of HCC patients. Dai et al.61 proposed a prognostic model for predicting the prognosis of HCC patients. They identified 17 genes that were potentially associated with the prognosis of HCC patients. These 17 genes were used to make a prognostic model using the Cox hazard regression model and validated its performance using the TCGA and GSE14520 datasets. They showed that six genes were involved in the prognosis of HCC patients. Most researchers simply used hub genes derived from the PPI network to identify the key or core genes. One of the major challenges in studying genetic data was the identification of relevant biomarkers or genes. Recently, machine learning (ML)-based techniques have gained more attraction to address this problem59,60,62,63,64,65,66. Despite the fact that several studies have been carried out for the identification and development of potential candidate genes for HCC8,9,10,11,12,13,14,15,16,17,18,19,20,21,22,23,24,25,26,27,28,29,30,31,32,33,34,35,36,37,38,39,40,41,42,43,44,45,46,47,48,49,50,51,52,53,54,55,56,57,58,67, it remains a challenging issue and still has some scope for more research for the identification of potential genes as well as understanding molecular mechanisms for the development, pathogenies, and progression of HCC.
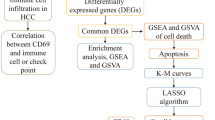
In this work, we used three microarray gene expression (MGE) datasets as training sets to determine the key or core candidate genes for HCC. First, we selected individual DEGs for three datasets. Secondly, support vector machine (SVM) with radial basis function (RBF) was implemented on the identified DEGs from each of the three datasets and calculated the classification accuracy of each DEG. We selected the DEGs from each of the three datasets that provided a classification accuracy of more than 80.0%. At the same time, the overlapping or shared DEGs were identified from three datasets. These overlapping or shared DEGs were called differentially expressed discriminative genes (DEDGs). Thirdly, DAVID was used to perform enrichment analysis on common DEDGs. Fourthly, PPI networks were constructed using STRING and visualized using Cytoscape. Then the hub genes were identified using degree, maximum neighborhood component (MNC), maximal clique centrality (MCC), closeness, and betweenness on the basis of cytoHubba. After that, the central hub genes were determined by overlapping or shared hub genes from the degree, MNC, MCC, centralities of closeness, and betweenness. Molecular Complex Detection (MCODE) was performed for cluster or module analysis and determined the important or significant modules as well as their associated genes. Moreover, the significant meta-hub genes were determined from meta-hub genes, which were extracted from existing studies. The key or core candidate genes were determined among the central hub genes, potential module hub genes, and significant meta-hub genes, which can be easily discriminated against in HCC patients compared to healthy controls. Furthermore, we used another two independent test datasets for the validation as well as to show the discriminative power of the key candidate genes. We also performed a survival analysis of the identified key candidate genes for HCC patients. Therefore, the overall flowchart of our proposed system to determine key candidate genes for HCC is presented in Fig. 1.
Results
Identification of DEGs from each dataset
We implemented limma for identifying DEGs from each of the three GEO datasets (GSE36376, GSE39791, and GSE57957). Using the threshold of \(|log_2 FC|{>1}\), and adj.p-value < 0.01, we identified 699 (up-regulated: 431 vs. down-regulated: 268), 428 (up-regulated: 88 vs. down-regulated: 340 DEGs), and 413 DEGs (up-regulated: 107; down-regulated: 306) DEGs between HCC and healthy controls from GSE36376, GSE39791, and GSE57957 datasets and their volcano plots and heatmap were presented in Fig. 2.
Volcano plot and heatmap of DEGs for each GEO dataset were generated using “ggplot2” version 3.3.6 package110 ( https://cran.r-project.org/package=ggplot2) and “NMF” version 0.24.0 package111 (https://cran.r-project.org/package=NMF) in R . (a) Volcano plot and (b) heatmap of GSE36376 dataset; (c) Volcano plot and (d) heatmap of GSE39791 dataset; (c) Volcano plot and (d) heatmap of GSE57957. Dodger blue represents down-regulated, gray represents no significant genes, and fire brick represents up-regulated DEGs.
Identification of common DEDGs using SVM
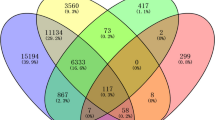
SVM with RBF kernel was applied on the identified DEGs (699 DEGs for GSE36376; 428 DEGs for GSE39791; and 413 DEGs for GSE57957) of each dataset in order to identify the DEDGs of HCC patients. Then, the classification accuracy was computed per gene for DEGs from each dataset. The calculation procedure is clearly discussed in the methodology section. The classification accuracies of all DEGs for individual datasets were ordered in descending order of magnitude, which is presented in Fig. 3. As shown in Fig. 3, we observed that a total of 502 from GSE36376, 169 from GSE39791, and 242 from GSE57957 DEGs were selected as DEDGs because their classification accuracy was more than or equal to 80.0%. Furthermore, 75 common DEDGs were determined among the identified DEDGS from GSE36376, GSE39791, and GSE57957 datasets, which is shown in Fig. 4.
Enrichment analysis of common DEDGS
Enrichment analysis was conducted on 75 shared or overlapping DEDGs clearly grasp the mechanism and development of HCC. The functional characteristics of DEDGs were explored using GO and KEGG pathway analysis. The GO analysis was partitioned into three groups: biological process (BP), cellular component (CC), and morphological component. Using p-values \((< 0.05)\), we identified the significant GO and KEGG pathways, and chose the top five prominent GO terms and KEGG pathway. The top five GO terms, including BP, CC, and MF, are presented in Table 1.
For BP-based GO terms, the common DEDGs were strongly enriched with retinol metabolic process, cellular response to cadmium ion, retinoid metabolic process cellular response to copper ion, and steroid catabolic process. Moreover, the extracellular region, extracellular exosome, extracellular space, high-density lipoprotein particle, and apical plasma membrane were found to be top CC, which were significantly enriched with common DEDGs. As shown in Table 1, MF group GO terms, including retinol dehydrogenase activity; oxidoreductase activity; androsterone dehydrogenase activity; androstan-3-alpha,17-beta-diol dehydrogenase activity; and steroid dehydrogenase activity, were mainly enriched with common DEDGs.
The study of the KEGG pathway for common DEDGs is displayed in Table 2. As shown in Table 2, the common DEDGs were significantly associated with multiple pathways such as retinol metabolism, metabolic pathways, tryptophan metabolism, steroid hormone biosynthesis, and drug metabolism-cytochrome P450.
PPI network construction and central hub genes identification
STRING was utilized to build a PPI network to show the significant connections between proteins encoded by common DEDGs. Cytoscape was used to show the PPI network, which had 51 nodes and 144 edges (see Fig. 5a). Five hub gene-based identification algorithms, including the degree of connectivity, MNC, MCC, closeness, and betweenness in the Cytoscape plug-in cytoHubba, were implemented to determine the hub genes from PPI networks. Then we chose the top 30 hub genes from each algorithm. We made a Venn diagram among the five algorithms, which is shown in Fig. 5b. As shown in Fig. 5b, eight overlapping central hub genes were identified among these algorithms. These eight central hub genes were NUSAP1, TOP2A, CDC20, PRC1, UBE2C, ASPM, PNPLA7, and MT1E, which were utilized to determine the key or core genes for HCC.
PPI network and Venn diagram for common DEDGs and central hub genes. (a) PPI network of common DEDGs with 51 nodes and 144 edges which was generated by Cytoscape 3.9.1118 (www.cytoscape.org); (b) identification of central hub genes among five methods (Degree, MNC, MCC, Closeness, and Betweenness based HGs). Here, HGs represent the hub genes.
Hub modules and its associated genes identification
Module or cluster analysis was performed using MCODE to determine the prominent modules. Three clusters or modules were generated using MCODE and provided 3–6 MCODE scores. We chose the prominent modules that provided the MCODE scores of \(\ge 5\) and the number of nodes \(\ge 5\). Finally, we chose module 1 as a prominent hub module that contained 6 nodes and 30 edges with the highest MCODE scores of 6 and their PPI networks were displayed in Fig. 6. The correspondence six genes were treated as hub module genes.
PPI network of module 1 with 6 nodes and 30 edges which was generated by Cytoscape 3.9.1118 (www.cytoscape.org).
Identification of significant meta-hub genes from metadata
We reviewed 52 existing studies related to gene identification of HCC patients8,9,10,11,12,13,14,15,16,17,18,19,20,21,22,23,24,25,26,27,28,29,30,31,32,33,34,35,36,37,38,39,40,41,42,43,44,45,46,47,48,49,50,51,52,53,54,55,56,57,58. We listed their hub genes in order to make metadata which were presented in Table 3. To make metadata, we extracted 10 hub genes from Maddah et al.9, 5 hub genes from Yan et al.10, 20 from Zhao et al.11, 7 from Zhao et al.12, 10 from Liu et al.13, 11 from Meng et al.14, 42 from Rosli et al.15, 5 from Zhang et al.8, 5 from Li et al.16, 8 from Li et al.17, 5 from Tian et al.18, 12 from Wan et al.19, 10 from Zhu et al.20, 10 from Wang et al.21, 9 from Zhou et al.22, 10 from Zhang et al.23, 18 from Mou et al.24, 8 from Wu et al.25, 9 from Gui et al.26, 10 from Wang et al.27, 28 from Lu and Zhu28, 6 from Bhatt et al.29, 10 from Zhang et al.30, 13 from Jiang et al.31, 20 from Zhang et al.32, 12 from Wu et al.33, 5 from Nguyen et al.34, 15 from Zhou et al.35, 6 from Yu et al.36, 10 from Kakar et al.37, 10 from Ji et al.38, 11 from Chen et al.39, 10 from Qiang et al.40, 10 from Wang et al.41, 10 from Zhang et al.42, 14 from Kim et al.43, 10 from Zhang et al.44, 14 from Sha et al.45, 10 from Chen et al.46, 4 from He et al.47, 10 from Zhang et al.48, 4 from Hu et al.49, 9 from Zhang et al.50, 15 from Li et al.51, 5 from Cao et al.52, 7 from Yang et al.53, 5 from Wang et al.54, 9 from Jiang et al.55, 16 from Li et al.56, 15 from Xing et al.57, 10 from Zhu W et al.58, and 20 from Dai et al.61. Now, we took the union of extracted hub genes and got 214 hub genes as meta-hub genes. At the same time, we also computed the frequency of each meta-hub gene depending on how many studies got that gene as hub gene and selected 52 significant meta-hub genes because their frequency was more than 3. These selected 52 significant meta-hub genes were utilized for the determination of key genes.
Key candidate genes identification
Eight central hub genes were identified from five methods (degree of connectivity, MNC, MCC, closeness, and betweenness), 6 hub module genes from potential hub modules, and 52 significant meta-hub genes from meta-hub genes. Six overlapping genes were identified using the Venn diagram from these three gene identification methods, which is presented in Fig. 7. These six genes (TOP2A, CDC20, ASPM, PRC1, UBE2C, and NUSAP1) were considered as key genes, which can be easily classified into the subjects as HCC and healthy.
Validation of key candidate genes
Discriminative power analysis using ROC curve
Six key or core genes (TOP2A, CDC20, ASPM, PRC1, UBE2C, and NUSAP1) were validated using AUC, computed from ROC curves. We compared the performance of two independent test datasets (GSE76427 and TCGA-LIHC) with one of our train datasets (GSE57957) in order to show the precision of the selected key candidate genes. The ROC curves of six key genes as well as their heatmap for both training and independent test datasets were illustrated in Fig. 8.
Validation of the six key candidate genes using AUC and heatmap: (a), (b) GSE57957-based training dataset; (c), (d) GSE76427-based independent test dataset; and (e), (f) TCGA-LIHC based independent test dataset. Whereas, ROC curves were generated using pROC version 1.18.0 package121 and heatmap was generated using “NMF” version 0.24.0 package in R111.
The ROC curve of six key candidate genes with their AUC values for the training dataset (GSE57957) was displayed in Fig. 8a: TOP2A (AUC: 0.936, 95% CI 0.871–1.000), CDC20 (AUC: 0.917, 95% CI 0.838–0.996), ASPM (AUC: 0.919, 95% CI 0.851–0.987), PRC1 (AUC: 0.938, 95% CI 0.871–1.000), UBE2C (AUC: 0.803, 95% CI 0.703–0.904), and NUSAP1 (AUC: 0.930, 95% CI 0.895–1.000). As displayed in Fig. 8c, the AUC values of six key or core genes were more than almost 0.780. The AUC values of six key or core genes for the GSE76427 dataset were: TOP2A (AUC: 0.900, 95% CI 0.851–0.949), CDC20 (AUC: 0.887, 95% CI 0.883–0.941), ASPM (AUC: 0.893, 95% CI 0.844–0.942), PRC1 (AUC: 0.931, 95% CI 0.889–0.975), UBE2C (AUC: 0.792, 95% CI 0.723–0.863), and NUSAP1 (AUC: 0.881, 95% CI 0.831–0.933).
Similarly, the ROC curves of six key candidate genes with their AUC values for the TCGA-LIHC-independent test dataset were presented in Fig. 8e. As presented in Fig. 8e, it was observed that six key candidate genes were provided the AUC values of more than 0.900 and their individual AUC values were as follows: TOP2A (AUC: 0.961, 95% CI 0.939–0.984), CDC20 (AUC: 0.968, 95% CI 0.949–0.986), ASPM (AUC: 0.960, 95% CI 0.938–0.983), PRC1 (AUC: 0.967, 95% CI 0.948–0.987), UBE2C (AUC: 0.965, 95% CI 0.946–0.985), and NUSAP1 (AUC: 0.919, 95% CI 0.889–0.949). Therefore, these six key genes (TOP2A, CDC20, ASPM, PRC1, UBE2C, and NUSAP1) showed strong discriminative power to classify HCC patients from healthy controls. These validations would be supported our findings and provided them more robust.
Survival analysis
In this work, we adopted survival analysis of six key candidate genes (TOP2A, CDC20, ASPM, PRC1, NUSAP1, and UBE2C) using univariate Cox regression in R and its results are presented in Fig. 9. As shown in Fig. 9, we observed that our identified six key candidate genes for HCC patinets such as TOP2A, CDC20, ASPM, PRC1, NUSAP1, and UBE2C were strongly associated with the survival status of HCC patients (\(\hbox {p}<0.05\)). So, the over-expression levels of TOP2A, CDC20, ASPM, PRC1, NUSAP1, and UBE2C had poor survival periods compared to lower expression levels of that key candidate genes.
Survival analysis of six key candidate genes for HCC: (a) TOP2A; (b) CDC20; (c) ASPM; (d) PRC1; (e) NUSAP1; and (f) UBE2C. The horizontal axis (x-axis) represents the time to event (in days) and the vertical axis (y-axis) represents survival probability. The HCC patients were divided into two groups: high-risk and low-risk and assigned a color. The red line designates the samples with high risk, and the green line represents the samples with low risk. \(\hbox {p} < 0.05\) indicates a statistically significant difference in mortality between groups. The survival plots were generated using the “Survfit” package in R122.
Discussion
In this work, we assessed three datasets, namely GSE36376, GSE39791, and GSE57957, to detect the DEGs for HCC patients. We determined 699, 428, and 413 DEGs using “limma” from the GSE36376, GSE39791, and GSE57957 datasets, which were illustrated in Fig. 2. Moreover, we implemented SVM to determine the DEDGs from individual datasets (see in Fig. 3) and selected overlapping or shared 75 DEDGs among the identified DEDGS from GSE36376, GSE39791, and GSE57957 datasets, which were clearly shown in Fig. 4. At the same time, enrichment analysis was executed on overlapping or shared DEDGs to clear understand their better exploration and molecular mechanism (see in Table 1). We found that the potential BP functional categories were strongly related to the development and progression of HCC patients. Retinol and retinoid metabolic processes have been linked to a variety of liver diseases, including fatty liver disease, which leads to HCC68,69. The rest of the BP categories were also enriched with common DEDGs, which also coincided with existing studies, like cellular response to cadmium ion42,57,70, cellular response to copper ion36,70, and steroid catabolic process42.
The top 5 GO terms were significantly enriched with common DEDGS, which were also consistent with previous results, such as extra cellular region35,37,38,57, extracellular exosome37,38, extracellular space37,38,57, high-density lipoprotein particle57, and apical plasma membrane53. In the case of MFs, common DEDGs were also enriched with top five GO terms. Existing studies supported these enrichment factional categories, including retinol dehydrogenase activity14, and oxidoreductase activity37,38,42. We also analyzed KEGG pathways and chose five pathways that were closely related to our overlapping DEDGs (see in Table 2). Different existing studies supported our findings, such as retinol metabolism35,37,38,40,43,70, metabolic pathways37,38, tryptophan metabolism38,42,70, steroid hormone biosynthesis42,70, and drug metabolism-cytochrome P45035,42,70.
A PPI network was built with shared DEDGs using Cystoscape (see in Fig. 5a and then eight central hub genes (NUSAP1, TOP2A, CDC20, PRC1, UBE2C, ASPM, PNPLA7, and MT1E) were identified from five hub gene selection methods, which were presented in Fig. 5b. The potential modules were identified using MCODE scores and module 1 was identified due to having the highest MCODE scores. We selected six hub module genes from module 1 as well as constructed their PPI network (see in Fig. 6). In addition, we examined 52 papers and took the hub genes from earlier studies8,9,10,11,12,13,14,15,16,17,18,19,20,21,22,23,24,25,26,27,28,29,30,31,32,33,34,35,36,37,38,39,40,41,42,43,44,45,46,47,48,49,50,51,52,53,54,55,56,57,58 in order to make metadata. At the same time, we listed 214 meta-hub genes by taking the union of extracted hub genes, which were presented in Table 3. We selected 52 significant meta-hub genes from the list of meta-hub genes whose frequency was greater than 3. Finally, we identified the six shared genes (TOP2A,CDC20, ASPM, PRC1, UBE2C, and NUSAP1) by intersecting central hub genes, hub module genes, and significant meta-hub genes, extracted from the earlier studies, known as key relevant or candidate genes, which were clearly depicted in Fig. 7. We validated these key relevant or candidate genes using AUC for one training and two independent test datasets (see Fig. 8). We observed that these six key relevant or candidate genes had high discriminative power for the differentiation of HCC patients.
TOP2A is a cell cycle-related gene that encoded a DNA topoisomerase which controls and alters the topologic states of DNA during transcription. TOP2A overexpression has been identified as a core or potential biomarker for ovarian cancers71, glioma72, and lung cancers73. A study showed that TOP2A overexpression in HCC patients was significantly correlated with progression and poor prognosis74,75. In the case of our study, TOP2A was also considered as a key or core gene for the progression and development of HCC. This finding was coincided with previous studies12,14,15,18,20,21,22,23,24,25,27,32,33,34,35,36,39,41,42,43,45,46,48,50,51,54,55,56,57,58,61.
CDC20 is a vital regulator of cell division in humans76,77. Overexpression or high expression of CDC20 has also been linked to lung cancer78, colorectal cancer79, breast cancer80,81, and other cancers. Moreover, CDC20 was strongly correlated with poor prognosis in gastric cancer82, bladder cancer83, and breast cancer84. A study revealed that CDC20 over-expression was significantly associated with HCC85. Another recent study demonstrated that there existed a strong relationship between CDC20 overexpression and the prognosis of HCC86. Our findings also showed that CDC20 was a potential key biomarker that played an crucial or essential role for the development and progression of HCC. Different existing studies also supported our findings10,13,14,17,18,20,24,30,32,37,40,41,44,48,50,55,56,57.
ASPM is a protein that have a major influence in the development of HCC. ASPM is located on chromosome 1 and band 1q31 and consists of 28 exons and 3477 amino-acid proteins87. Lots of studies have identified ASPM as a hub gene or key biomarker for multiple cancers88,89,90. Zhang et al.90 reported that ASPM can be a promising therapeutic target for liver. Moreover, ASPM overexpression was strongly correlated with bladder cancer and consiered as promising predictor91. Our findings also illustrated that ASPM was a novel key biomarker for HCC, which was supported by the existing studies9,22,35,38,39,41,42,43,45,46,47,48,58.
PRC1 is an essential protein that is the regulator of cytokinesis92. The higher expression level of PRC1 was found among HCC patients than healthy controls. The overexpression of PRC1 was associated with a poor prognosis for HCC patients93. Our work also indicated that PRC1 was a promising or key biomarker for the development of HCC, which coincided with previous studies15,22,25,33,35,39,42,43,45,46,56,57,61.
Similarly, we proposed UBE2C as a key or core predictor for development of HCC, which was supported by various existing studies10,18,33,36,41,44,58. Xiong et al.94 suggested UBE2C as a potential biomarker or gene for HCC. High expression of UBE2C was also found in HCC than healthy subjects95. UBE2C is not play a crucial role HCC but also in variety of cancers: lung cancer, gastric cancer96,97.
NUSAP1 is a protein associated with the nucleolar-spindle that have a vital role in spindle microtubule organization98. overexpression of NUSAP1 was found in a variety of malignancies, including HCC58,99, colon cancer100,101, prostate cancer102,103, and cervical carcinoma104. Moreover, overexpression of NUSAP1 was strongly linked with poor prognosis of prostate cancer103 and colon cancer101. Another study revealed that NUSAP1 is related to HCC105. Roy et al.105 illustrated that NUSAP1 expression might rise in HCC samples with low expression levels of miRNA 193a-5p, and that this overexpression was strongly associated with a shorter patient survival time. Our findings also illustrated that NUSAP1 was one of the key candidate genes that the highest expression levels were found in HCC subjects compared to healthy subjects. These findings were consistent with existing studies15,22,46,56,58,61.
Moreover, two independent test datasets were also used to validate these six key candidate genes using AUC. A survival analysis was also performed of these six candidate genes for HCC patients. In both cases, our identified six key candidate genes (TOP2A, CDC20, ASPM, PRC1, UBE2C, and NUSAP1) showed significant association with the development and progression of HCC. This finding will provide evidence and new insight to physicians and readers in determining the diagnosis of HCC as well as the correlated pathway of HCC.
Materials and methods
Data acquisition and preprocessing
In this work, three publicly available microarray gene expression datasets with GEO accession: GSE3637666, GSE39791106, and GSE57957107 with GPL10558 [Illumina HumanHT-12 V4.0 expression bead chip] were used to determine the key candidate genes. Another two independent test datasets were used to validate key candidate genes. One independent dataset was taken from the GEO database with accession number: GSE76427 with GPL10558 platform102 and another independent test dataset was taken from the Cancer Genome Atlas (TCGA) database. Microarray gene expression datasets were downloaded from the GEO database (www.ncbi.nlm.nih.gov/geo/) and TCGA-liver hepatocellular carcinoma (TCGA-LIHC) dataset was downloaded from the TCGA database (https://portal.gdc.cancer.gov/). The datasets underwent a log2 transformation and quintile normalization. Although these datasets were taken from the publicly available GEO repository, being human data, all methods were performed in accordance with the relevant guidelines and regulations. Table 4 presents a summary of the utilized datasets.
Identification of DEGs from each dataset
To identify the DEGs between HCC and healthy controls, each of the selected datasets was analyzed using the “limma” package108 in R-software with version 4.1.2. We computed the \(|log_2 FC|\) and adj. p-value of each gene from the selected dataset. “Bioconductor annotation”109 package was used to convert microarray data probes into gene symbols. If multiple probes were matched with a gene symbol, take the gene with their associated expression values that provided the lowest or minimum adjusted p-value. The DEGs between HCC and healthy controls were identified with a cutoff of point: \(|log_2FC|>1\) and \(adj. p-value<0.01\) (false discovery rate). The volcano plot of DEGs was generated using the “ggplot2 version 3.3.6” package in R110. Moreover, a heat map of the expression of DEGs was generated with the “NMF” version 0.24.0 package in R111.
SVM-based identification of DEDGs from DEGs for each dataset
The main purpose of SVM is to identify a hyperplane in a high dimensional space112,113 that can easily discriminate HCC patients from healthy control patients using the following discriminate function:
where, b is the bias term.
In this study, we have used radial basis kernel, which is defined as follows:
We set the different values of cost (C) and gamma \((\gamma )\) and tuned these values using a grid search method and select the optimal value of C and \((\gamma )\) to improve classification accuracy. In this current study, we adopted SVM as a gene selection method, and its identification procedure is described as follows:
-
Step 1
Select one gene from a list of identified DEGs.
-
Step 2
Trained SVM-based model with five-fold cross-validation (CV) protocols.
-
Step 3
Calculate the classification accuracy for this selected gene.
-
Step 4
Repeat Step 2 to Step 3 for all identified DEGs.
-
Step 5
Sort the classification accuracy of all DEGs in descending order of magnitude.
-
Step 6
Choose the genes that will produce a classification accuracy of more than 80.0.
Identification of common DEDGs
After selecting differentially expressed discrimination genes (DEDGs) using SVM, we identified the shared or overlapping or common DEDGs among three datasets using the following formula:
where, r is the number of utilizing GEO dataset (here, r = 3).
Enrichment analysis of common DEDGs
To better understand the mechanism and progression of HCC patients, we obtained enrichment analysis, including GO and KEEG analysis114,115 on DEDGs using DAVID version 6.8 tools116 (david.ncifcrf.gov). A p-value < 0.05 was considered for significant.
PPI network analysis and central hub gene identification
The STRING version 11.5 software (www.string-db.org) was utilized to obtain the potential interactions among common DEDGs117. A protein-protein interaction (PPI) with a confidence score of \(> 0.70\) and a maximum number of interactors of 0 was preserved and loaded into Cystoscape version 3.9.1118 to build a PPI network. The degree of connectivity, maximum neighborhood component (MNC), maximal clique centrality (MCC), centralities of closeness, and betweenness were computed using cytoHubba119. Then, we sorted the values of degree of connectivity, MNC, MCC, centralities of closeness, and betweenness in descending order of magnitude and chose the top 30 DEDGs, known as hub genes. The central hub genes were selected by overlapping hub genes, which were computed from the degree of connectivity, MNC, MCC, centralities of closeness, and betweenness. Mathematically, it is defined as follows:
where, hg is the number of hub gene identification methods (Here, hg=5).
Hub modules and its associated genes identification
MCODE was used to determine the most closely connected modules from the PPI network120. We analyzed the modules with the following cutoff points: degree =2, cluster finding =haircut, nodes score =0.2, K-score =2, and max depth =100, respectively. We determined the potential modules that provided the MCODE with scores of \(\ge 6\) and the number of nodes of \(\ge 6\). Then, the hub module genes were identified using the following formula:
where, \(h_m\) is the number of significant modules.
Significant meta-hub genes identification from metadata
We reviewed some existing studies related to HCC-based gene identification. To make metadata, we listed their identified hub genes for HCC, called “meta-hub genes,” which can be written as follows:
where, m is the number of studies obtained from obtaining hub genes (here, m = 52).
We also counted the frequency of each meta-hub gene depending on how many studies identified that gene as a hub gene. Finally, we identified significant meta-hub genes from meta-hub genes whose frequency was greater than or equal to 3, which can be written as follows:
where, \(g_i \in \text {meta-hub gene}\) and n is the number of meta-hub genes whose frequency is \(\ge 3\)
Key candidate genes identification
To identify the key candidate genes, we selected the central hub genes from the PPI network, hub module genes from significant modules, and significant meta-hub genes from existing studies. Therefore, we identified the key candidate genes for HCC using the following formula:
where, k is the number of significant gene identification methods (Here, k = 3). In this work, central hub genes, hub module genes, and significant gene selection methods will be considered “Important Gene Identification Methods”.
Validation of key candidate genes
Discriminative power analysis using ROC curve
In this work, we used two independent test datasets in order to validate the key candidate genes. One independent test dataset (GSE76427) was taken from the GEO database, and another independent dataset was taken from the TCGA database. The description of these independent test datasets is more clearly explained in Table 4. We validated the selected key candidate genes using the area under the curve (AUC), computed from the receiver operating characteristic curve (ROC). In ROC analysis, first, we selected one gene and class label, and then we adopted logistic regression with the leave-one-out CV protocol. We computed AUC values using the “pROC” R-package121. Moreover, we also compared the performances of independent test datasets with one of our training datasets (GSE57957) in order to show the precision of the selected key candidate genes.
Survival analysis
In this work, we used TCGA-LIHC dataset for survival analysis in order to show prognostic status of key candidate genes. We classified HCC patients into high-risk and low-risk groups on the basis of median expression level of each key candidate gene. We performed survival analysis of our identified key candidate genes using the “Survfit” package in R language122. A p-value < 0.05 was considered statistically significant (“Supplementary information”).
Data availability
The datasets generated and/or analyzed during the current study are available in the Gene Expression Omnibus (GEO) repository with accession numbers: GSE36376, GSE39791, GSE57957, and GSE76427 with GPL10558 platforms. One can easily download these datasets from the link: www.ncbi.nlm.nih.gov/geo/.
References
Parkin, D. M., Bray, F., Ferlay, J. & Pisani, P. Global cancer statistics, 2002. CA Cancer J. Clin. 55, 74–108. https://doi.org/10.3322/canjclin.55.2.74 (2005).
Yang, J. D. et al. A global view of hepatocellular carcinoma: Trends, risk, prevention and management. Nat. Rev. Gastroenterol. Hepatol. 16, 589–604. https://doi.org/10.1038/s41575-019-0186-y (2019).
Kumar, V., Abbas, A. K., Fausto, N. & Aster, J. C. Robbins and Cotran Pathologic Basis of Disease 9th edn. (Elsevier Health Sciences, 2015).
Huang, T. et al. The role of hepatitis c virus in the dynamic protein interaction networks of hepatocellular cirrhosis and carcinoma. Int. J. Comput. Biol. Drug Design 4, 5–18. https://doi.org/10.1504/IJCBDD.2011.038654 (2011).
Yuan, W. et al. Comparative analysis of viral protein interaction networks in hepatitis b virus and hepatitis c virus infected hcc. Biochimica et Biophysica Acta (BBA)-Proteins and Proteomics 1844, 271–279. https://doi.org/10.1016/j.bbapap.2013.06.002 (2014).
Llovet, J. M. et al. Hepatocellular carcinoma. Nat. Rev. Dis. Primers 7, 6–34. https://doi.org/10.1038/s41572-020-00240-3 (2021).
Akinyemiju, T. et al. The burden of primary liver cancer and underlying etiologies from 1990 to 2015 at the global, regional, and national level: results from the global burden of disease study 2015. JAMA Oncol. 3, 1683–1691. https://doi.org/10.1001/jamaoncol.2017.3055 (2017).
Zhang, C. et al. The identification of key genes and pathways in hepatocellular carcinoma by bioinformatics analysis of high-throughput data. Med. Oncol. 34, 1–13. https://doi.org/10.1007/s12032-017-0963-9 (2017).
Maddah, R. et al. Identification of critical genes and pathways associated with hepatocellular carcinoma and type 2 diabetes mellitus using integrated bioinformatics analysis. Inform. Med. Unlocked 30, 100956–100963. https://doi.org/10.1016/j.imu.2022.100956 (2022).
Yan, G. & Liu, Z. Identification of differentially expressed genes in hepatocellular carcinoma by integrated bioinformatic analysis. bioRxiv.https://doi.org/10.1101/570846 (2019).
Qian, Z., Yan, Z. & Zhengkui, L. Mining of gene modules and identification of key genes in hepatocellular carcinoma based on gene co-expression network analysis. in Proceedings of the 2020 12th International Conference on Bioinformatics and Biomedical Technology, 18–24. https://doi.org/10.1145/3405758.3405762 (2020).
Zhao, Y. & Xie, Y. Study on differential expression genes in hcc based on geo database. in Proceedings of the 2021 International Conference on Bioinformatics and Intelligent Computing, 63–69. https://doi.org/10.1145/3448748.3448759 (2021).
Liu, J. et al. Identification of multiple hub genes and pathways in hepatocellular carcinoma: A bioinformatics analysis. BioMed Res. Int. 2021, 1–11. https://doi.org/10.1155/2021/8849415 (2021).
Meng, Z. et al. Identification of potential hub genes associated with the pathogenesis and prognosis of hepatocellular carcinoma via integrated bioinformatics analysis. J. Int. Med. Res. 48, 1–23. https://doi.org/10.1177/0300060520910019 (2020).
Rosli, A. F. C., Razak, S. R. A. & Zulkifle, N. Bioinformatics analysis of differentially expressed genes in liver cancer for identification of key genes and pathways. Malaysian J. Med. Health Sci. 15, 18–24 (2019).
Li, Y. et al. Integrated bioinformatics analysis reveals key candidate genes and pathways associated with clinical outcome in hepatocellular carcinoma. Front. Genet. 11, 814–819. https://doi.org/10.3389/fgene.2020.00814 (2020).
Li, Z., Lin, Y., Cheng, B., Zhang, Q. & Cai, Y. Identification and analysis of potential key genes associated with hepatocellular carcinoma based on integrated bioinformatics methods. Front. Genet. 12, 571231–571245. https://doi.org/10.3389/fgene.2021.571231 (2021).
Tian, D., Yu, Y., Zhang, L., Sun, J. & Jiang, W. A five-gene-based prognostic signature for hepatocellular carcinoma. Front. Med. 8, 1–24. https://doi.org/10.3389/fmed.2021.681388 (2021).
Wan, Z., Zhang, X., Luo, Y. & Zhao, B. Identification of hepatocellular carcinoma-related potential genes and pathways through bioinformatic-based analyses. Genet. Testing Mole. Biomarkers 23, 766–777. https://doi.org/10.1089/gtmb.2019.0063 (2019).
Zhu, Q., Sun, Y., Zhou, Q., He, Q. & Qian, H. Identification of key genes and pathways by bioinformatics analysis with tcga rna sequencing data in hepatocellular carcinoma. Mol. Clin. Oncol. 9, 597–606. https://doi.org/10.3892/mco.2018.1728 (2018).
Wang, J., Tian, Y., Chen, H., Li, H. & Zheng, S. Key signaling pathways, genes and transcription factors associated with hepatocellular carcinoma. Mol. Med. Rep. 17, 8153–8160. https://doi.org/10.3892/mmr.2018.8871 (2018).
Zhou, L., Du, Y., Kong, L., Zhang, X. & Chen, Q. Identification of molecular target genes and key pathways in hepatocellular carcinoma by bioinformatics analysis. OncoTargets Therapy 11, 1861. https://doi.org/10.2147/OTT.S156737 (2018).
Zhang, P. et al. Bioinformatics analysis of candidate genes and pathways related to hepatocellular carcinoma in china: A study based on public databases. Pathol. Oncol. Res. 27, 588532–588546. https://doi.org/10.3389/pore.2021.588532 (2021).
Mou, T. et al. Identification and interaction analysis of key genes and micrornas in hepatocellular carcinoma by bioinformatics analysis. World J. Surg. Oncol. 15, 1–9. https://doi.org/10.1186/s12957-017-1127-2 (2017).
Wu, M. et al. Analysis of potential key genes in very early hepatocellular carcinoma. World J. Surg. Oncol. 17, 1–8. https://doi.org/10.1186/s12957-019-1616-6 (2019).
Gui, T., Dong, X., Li, R., Li, Y. & Wang, Z. Identification of hepatocellular carcinoma-related genes with a machine learning and network analysis. J. Comput. Biol. 22, 63–71. https://doi.org/10.1089/cmb.2014.0122 (2015).
Wang, J. et al. Identification and validation of key genes in hepatocellular carcinoma by bioinformatics analysis. Biomed Res. Int. 2021, 6662114–6662127. https://doi.org/10.1155/2021/6662114 (2021).
Lu, H. & Zhu, Q. Identification of key biological processes, pathways, networks, and genes with potential prognostic values in hepatocellular carcinoma using a bioinformatics approach. Cancer Biother. Radiopharm. 36, 837–849. https://doi.org/10.1089/cbr.2019.3327 (2021).
Bhatt, S. et al. Deciphering key genes and mirnas associated with hepatocellular carcinoma via network-based approach. IEEE/ACM Trans. Comput. Biol. Bioinform. 36, 837–849. https://doi.org/10.1109/TCBB.2020.3016781 (2020).
Zhang, Y. et al. A gene module identification algorithm and its applications to identify gene modules and key genes of hepatocellular carcinoma. Sci. Rep. 11, 1–14. https://doi.org/10.1038/s41598-021-84837-y (2021).
Jiang, X. & Hao, Y. Analysis of expression profile data identifies key genes and pathways in hepatocellular carcinoma. Oncol. Lett. 15, 2625–2630. https://doi.org/10.3892/ol.2017.7534 (2018).
Zhang, X. et al. Identification of hub genes associated with hepatocellular carcinoma prognosis by bioinformatics analysis. J. Cancer Therapy 12, 186–207. https://doi.org/10.4236/jct.2021.124019 (2021).
Wu, M., Liu, Z., Zhang, A. & Li, N. Identification of key genes and pathways in hepatocellular carcinoma: A preliminary bioinformatics analysis. Medicine 98, 1–7. https://doi.org/10.1097/MD.0000000000014287 (2019).
Nguyen, T. B., Do, D. N., Nguyen-Thanh, T., Tatipamula, V. B. & Nguyen, H. T. Identification of five hub genes as key prognostic biomarkers in liver cancer via integrated bioinformatics analysis. Biology 10, 957–970. https://doi.org/10.3390/biology10100957 (2021).
Zhou, Z. et al. Screening hub genes as prognostic biomarkers of hepatocellular carcinoma by bioinformatics analysis. Cell Transplant. 28, 76S-86S. https://doi.org/10.1177/0963689719893950 (2019).
Yu, C., Chen, F., Jiang, J., Zhang, H. & Zhou, M. Screening key genes and signaling pathways in colorectal cancer by integrated bioinformatics analysis. Mol. Med. Rep. 20, 1259–1269. https://doi.org/10.3892/mmr.2019.10336 (2019).
Kakar, M. et al. Identification of novel potential biomarkers in hepatocarcinoma cancer; a transcriptome analysis. Preprint (Version 3) available at Research Square (02 March 2021) 1–21. https://doi.org/10.21203/rs.3.rs-154350/v2 (2021).
Ji, Y., Yin, Y. & Zhang, W. Integrated bioinformatic analysis identifies networks and promising biomarkers for hepatitis b virus-related hepatocellular carcinoma. Int. J. Genom. 2020, 1–18. https://doi.org/10.1155/2020/2061024 (2020).
Chen, D. et al. Bioinformatic evidence reveals that cell cycle correlated genes drive the communication between tumor cells and the tumor microenvironment and impact the outcomes of hepatocellular carcinoma. BioMed Res. Int. 2021, 4092635–4092660. https://doi.org/10.1155/2021/4092635 (2021).
Qiang, R. et al. Identification of 5 hub genes related to the early diagnosis, tumour stage, and poor outcomes of hepatitis b virus-related hepatocellular carcinoma by bioinformatics analysis. Comput. Math. Methods Med. 2021, 1–20. https://doi.org/10.1155/2021/9991255 (2021).
Wang, J. et al. Global analysis of gene expression signature and diagnostic/prognostic biomarker identification of hepatocellular carcinoma. Sci. Progress 104, 1–7. https://doi.org/10.1177/00368504211029429 (2021).
Zhang, Y., Tang, Y., Guo, C. & Li, G. Integrative analysis identifies key mrna biomarkers for diagnosis, prognosis, and therapeutic targets of hcv-associated hepatocellular carcinoma. Aging (Albany NY) 13, 12865–12895. https://doi.org/10.18632/aging.202957 (2021).
Kim, S.-H. et al. Identification of key genes and carcinogenic pathways in hepatitis b virus-associated hepatocellular carcinoma through bioinformatics analysis. Ann. Hepato-biliary-pancreatic Surg. 26, 58–68. https://doi.org/10.14701/ahbps.21-108 (2022).
Zhang, G., Kang, Z., Mei, H., Huang, Z. & Li, H. Promising diagnostic and prognostic value of six genes in human hepatocellular carcinoma. Am. J. Transl. Res. 12, 1239–1254 (2020).
Sha, M. et al. Identification of genes predicting unfavorable prognosis in hepatitis b virus-associated hepatocellular carcinoma. Ann. Transl. Med. 9, 975–985. https://doi.org/10.21037/atm-21-2085 (2021).
Chen, H. et al. Identification of hub genes associated with immune infiltration and predict prognosis in hepatocellular carcinoma via bioinformatics approaches. Front. Genet. 11, 575762–575779. https://doi.org/10.3389/fgene.2020.575762 (2021).
He, B. et al. Bioinformatics analysis of key genes and pathways for hepatocellular carcinoma transformed from cirrhosis. Medicine. 96, 6938–6946. https://doi.org/10.1097/MD.0000000000006938 (2017).
Zhang, S., Peng, R., Xin, R., Shen, X. & Zheng, J. Conjoint analysis for hepatic carcinoma with hub genes and multi-slice spiral ct. Medicine 99, e23099–e23110. https://doi.org/10.1097/MD.0000000000023099 (2020).
Hu, W. Q. et al. Identification of biological targets of therapeutic intervention for hepatocellular carcinoma by integrated bioinformatical analysis. Med. Sci. Monitor 24, 3450–3461. https://doi.org/10.12659/MSM.909290 (2018).
Zhang, Q. et al. Prediction and analysis of weighted genes in hepatocellular carcinoma using bioinformatics analysis. Mol. Med. Rep. 19, 2479–2488. https://doi.org/10.3892/mmr.2019.9929 (2019).
Li, N., Li, L. & Chen, Y. The identification of core gene expression signature in hepatocellular carcinoma. Oxidative Med. Cell. Longevity 2018, 1–15. https://doi.org/10.1155/2018/3478305 (2018).
Cao, J., Zhang, R., Zhang, Y. & Wang, Y. Combined screening analysis of aberrantly methylated-differentially expressed genes and pathways in hepatocellular carcinoma. J. Gastrointestinal Oncol. 13, 311–325. https://doi.org/10.21037/jgo-21-866 (2022).
Yang, L., Zeng, L.-F., Hong, G.-Q., Luo, Q. & Lai, X. Construction of a novel clinical stage-related gene signature for predicting outcome and immune response in hepatocellular carcinoma. J. Immunol. Res. 2022, 1–10. https://doi.org/10.1155/2022/6535009 (2022).
Wang, M., Wang, L., Wu, S., Zhou, D. & Wang, X. Identification of key genes and prognostic value analysis in hepatocellular carcinoma by integrated bioinformatics analysis. Int. J. Genom. 2019, 1–22. https://doi.org/10.1155/2019/3518378 (2019).
Jiang, N. et al. Identification of core genes related to progression and prognosis of hepatocellular carcinoma and small-molecule drug predication. Front. Genet. 12, 608017–608036. https://doi.org/10.3389/fgene.2021.608017 (2021).
Li, L., Lei, Q., Zhang, S., Kong, L. & Qin, B. Screening and identification of key biomarkers in hepatocellular carcinoma: Evidence from bioinformatic analysis. Oncol. Rep. 38, 2607–2618. https://doi.org/10.3892/or.2017.5946 (2017).
Xing, T., Yan, T. & Zhou, Q. Identification of key candidate genes and pathways in hepatocellular carcinoma by integrated bioinformatical analysis. Exp. Therap. Med. 15, 4932–4942. https://doi.org/10.3892/etm.2018.6075 (2018).
Zhu, W., Xu, J., Chen, Z. & Jiang, J. Analyzing roles of nusap1 from clinical, molecular mechanism and immune perspectives in hepatocellular carcinoma. Front. Genet. 12, 689159–689181. https://doi.org/10.3389/fgene.2021.689159 (2021).
Jiang, M. et al. Identification of hepatocellular carcinoma related genes with k-th shortest paths in a protein-protein interaction network. Mol. BioSyst. 9, 2720–2728. https://doi.org/10.1039/C3MB70089E (2013).
Huang, T., Wang, J., Cai, Y.-D., Yu, H. & Chou, K.-C. Hepatitis c virus network based classification of hepatocellular cirrhosis and carcinoma. PloS One 7, e34460. https://doi.org/10.1371/journal.pone.0034460 (2012).
Dai, Q. et al. Six genes involved in prognosis of hepatocellular carcinoma identified by cox hazard regression. BMC Bioinform. 22, 1–12. https://doi.org/10.1186/s12859-021-04095-7 (2021).
Qing, J.-B., Song, W.-Z., Li, C.-Q. & Li, Y.-F. The diagnostic and predictive significance of immune-related genes and immune characteristics in the occurrence and progression of iga nephropathy. J. Immunol. Res. 2022, 1–20. https://doi.org/10.1155/2022/9284204 (2022).
Yu, S.-H. et al. Lasso and bioinformatics analysis in the identification of key genes for prognostic genes of gynecologic cancer. J. Pers. Med. 11, 1177. https://doi.org/10.3390/jpm11111177 (2021).
Basith, S., Hasan, M. M., Lee, G., Wei, L. & Manavalan, B. Integrative machine learning framework for the identification of cell-specific enhancers from the human genome. Brief. Bioinform. 22, bbab252. https://doi.org/10.1093/bib/bbab252 (2021).
Hasan, Al Mehedi, Maniruzzaman, M. & Shin, J. Identification of key candidate genes for iga nephropathy using machine learning and statistics based bioinformatics models. Sci. Rep. 12, 1–14. https://doi.org/10.1038/s41598-022-18273-x (2022).
Lim, H.-Y. et al. Prediction of disease-free survival in hepatocellular carcinoma by gene expression profiling. Ann. Surg. Oncol. 20, 3747–3753. https://doi.org/10.1245/s10434-013-3070-y (2013).
Zeng, L. et al. Differential combinatorial regulatory network analysis related to venous metastasis of hepatocellular carcinoma. BMC Genom. 13, 1–14. https://doi.org/10.1186/1471-2164-13-s8-s14 (2012).
Shirakami, Y., Sakai, H. & Shimizu, M. Retinoid roles in blocking hepatocellular carcinoma. Hepatobiliary Surg. Nutr. 4, 222–228. https://doi.org/10.3978/j.issn.2304-3881.2015.05.01 (2015).
Pettinelli, P. et al. Altered hepatic genes related to retinol metabolism and plasma retinol in patients with non-alcoholic fatty liver disease. PLoS One 13, e0205747–e0205763. https://doi.org/10.1371/journal.pone.0205747 (2018).
Lai, X. et al. A novel gene signature based on cdc20 and fcn3 for prediction of prognosis and immune features in patients with hepatocellular carcinoma. J. Immunol. Res. 2022, 1–22. https://doi.org/10.1155/2022/9117205 (2022).
Gao, Y. et al. Top2a promotes tumorigenesis of high-grade serous ovarian cancer by regulating the tgf-\(\beta\)/smad pathway. J. Cancer 11, 4181–4192. https://doi.org/10.7150/jca.42736 (2020).
Zhou, T., Wang, Y., Qian, D., Liang, Q. & Wang, B. Over-expression of top2a as a prognostic biomarker in patients with glioma. Int. J. Clin. Exp. Pathol. 11, 1228–1237 (2018).
Ma, W. et al. Prognostic significance of top2a in non-small cell lung cancer revealed by bioinformatic analysis. Cancer Cell Int. 19, 1–17. https://doi.org/10.1186/s12935-019-0956-1 (2019).
Cai, H., Shao, B., Zhou, Y. & Chen, Z. High expression of top2a in hepatocellular carcinoma is associated with disease progression and poor prognosis. Oncol. Lett. 20, 1–9. https://doi.org/10.3892/ol.2020.12095 (2020).
Meng, J., Wei, Y., Deng, Q., Li, L. & Li, X. Study on the expression of top2a in hepatocellular carcinoma and its relationship with patient prognosis. Cancer Cell Int. 22, 1–18. https://doi.org/10.1186/s12935-021-02439-0 (2022).
Weinstein, J., Jacobsen, F. W., Hsu-Chen, J., Wu, T. & Baum, L. G. A novel mammalian protein, p55cdc, present in dividing cells is associated with protein kinase activity and has homology to the saccharomyces cerevisiae cell division cycle proteins cdc20 and cdc4. Mol. Cell. Biol. 14, 3350–3363. https://doi.org/10.1128/mcb.14.5.3350-3363.1994 (1994).
Weinstein, J. Cell cycle-regulated expression, phosphorylation, and degradation of p55cdc: A mammalian homolog of cdc20/fizzy/slp1. J. Biol. Chem. 272, 28501–28511. https://doi.org/10.1074/jbc.272.45.28501 (1997).
Kato, T. et al. Overexpression of cdc20 predicts poor prognosis in primary non-small cell lung cancer patients. J. Surg. Oncol. 106, 423–430. https://doi.org/10.1002/jso.23109 (2012).
Wu, W.-J. et al. Cdc20 overexpression predicts a poor prognosis for patients with colorectal cancer. J. Transl. Med. 11, 1–8. https://doi.org/10.1186/1479-5876-11-142 (2013).
Karra, H. et al. Cdc20 and securin overexpression predict short-term breast cancer survival. Br. J. Cancer 110, 2905–2913. https://doi.org/10.1038/bjc.2014.252 (2014).
Tang, J. et al. Overexpression of aspm, cdc20, and ttk confer a poorer prognosis in breast cancer identified by gene co-expression network analysis. Front. Oncol. 9, 310–324. https://doi.org/10.3389/fonc.2019.00310 (2019).
Ding, Z.-Y., Wu, H.-R., Zhang, J.-M., Huang, G.-R. & Ji, D.-D. Expression characteristics of cdc20 in gastric cancer and its correlation with poor prognosis. Int. J. Clin. Exp. Pathol. 7, 722–727 (2014).
Choi, J.-W., Kim, Y., Lee, J.-H. & Kim, Y.-S. High expression of spindle assembly checkpoint proteins cdc20 and mad2 is associated with poor prognosis in urothelial bladder cancer. Virchows Archiv 463, 681–687. https://doi.org/10.1007/s00428-013-1473-6 (2013).
Alfarsi, L. H. et al. Cdc20 expression in oestrogen receptor positive breast cancer predicts poor prognosis and lack of response to endocrine therapy. Breast Cancer Res. Treatment 178, 535–544. https://doi.org/10.1007/s10549-019-05420-8 (2019).
Li, J., Gao, J.-Z., Du, J.-L., Huang, Z.-X. & Wei, L.-X. Increased cdc20 expression is associated with development and progression of hepatocellular carcinoma. Int. J. Oncol. 45, 1547–1555. https://doi.org/10.3892/ijo.2014.2559 (2014).
Zhang, X. et al. Connection between cdc20 expression and hepatocellular carcinoma prognosis. Med. Sci. Monitor 27, e926760–e926765. https://doi.org/10.12659/MSM.926760 (2021).
Bond, J. et al. Aspm is a major determinant of cerebral cortical size. Nat. Genet. 32, 316–320. https://doi.org/10.1038/ng995 (2002).
Pai, V. C. et al. Aspm promotes prostate cancer stemness and progression by augmenting wnt- dvl-3- \(\beta\)-catenin signaling. Oncogene 38, 1340–1353. https://doi.org/10.1038/s41388-018-0497-4 (2019).
Hsu, C.-C. et al. The differential distributions of aspm isoforms and their roles in wnt signaling, cell cycle progression, and pancreatic cancer prognosis. J. Pathol. 249, 498–508. https://doi.org/10.1002/path.5341 (2019).
Zhang, H. et al. Aspm promotes hepatocellular carcinoma progression by activating wnt/\(\beta\)-catenin signaling through antagonizing autophagy-mediated dvl2 degradation. FEBS Open Bio 11, 2784–2799. https://doi.org/10.1002/2211-5463.13278 (2021).
Xu, Z., Zhang, Q., Luh, F., Jin, B. & Liu, X. Overexpression of the aspm gene is associated with aggressiveness and poor outcome in bladder cancer. Oncol. Lett. 17, 1865–1876. https://doi.org/10.3892/ol.2018.9762 (2019).
Jiang, W. et al. Prc1: A human mitotic spindle-associated cdk substrate protein required for cytokinesis. Mol. Cell 2, 877–885. https://doi.org/10.1016/S1097-2765(00)80302-0 (1998).
Yang, Z. et al. Ccnb2, cdc20, aurka, top2a, melk, ncapg, kif20a, ube2c, prc1, and aspm may be potential therapeutic targets for hepatocellular carcinoma using integrated bioinformatic analysis. Int. J. General Med. 14, 10185–10194. https://doi.org/10.2147/IJGM.S341379 (2021).
Xiong, Y. et al. Ube2c functions as a potential oncogene by enhancing cell proliferation, migration, invasion, and drug resistance in hepatocellular carcinoma cells. Biosci. Rep. 39, 1–8. https://doi.org/10.1042/BSR20182384 (2019).
Ieta, K. et al. Identification of overexpressed genes in hepatocellular carcinoma, with special reference to ubiquitin-conjugating enzyme e2c gene expression. Int. J. Cancer 121, 33–38. https://doi.org/10.1002/ijc.22605 (2007).
Dastsooz, H., Cereda, M., Donna, D. & Oliviero, S. A comprehensive bioinformatics analysis of ube2c in cancers. Int. J. Mol. Sci. 20, 2228–22247. https://doi.org/10.3390/ijms20092228 (2019).
Zhang, H. et al. Overexpression of ube2c correlates with poor prognosis in gastric cancer patients. Eur. Rev. Med. Pharmacol. Sci. 22, 1665–1671. https://doi.org/10.26355/eurrev_201803_14578 (2018).
Petry, S. Mechanisms of mitotic spindle assembly. Ann. Rev. Biochem. 85, 659–683. https://doi.org/10.1146/annurev-biochem-060815-014528 (2016).
Hou, S., Hua, L., Wang, W., Li, M. & Xu, L. Nucleolar spindle associated protein 1 (nusap1) facilitates proliferation of hepatocellular carcinoma cells. Transl. Cancer Res. 8, 2113–2120. https://doi.org/10.21037/tcr.2019.09.28 (2019).
Han, G. et al. Nusap1 gene silencing inhibits cell proliferation, migration and invasion through inhibiting dnmt1 gene expression in human colorectal cancer. Exp. Cell Res. 367, 216–221. https://doi.org/10.1016/j.yexcr.2018.03.039 (2018).
Liu, Z. et al. High nusap1 expression predicts poor prognosis in colon cancer. Pathol.-Res. Practice 214, 968–973. https://doi.org/10.1016/j.prp.2018.05.017 (2018).
Gulzar, Z. G., McKenney, J. K. & Brooks, J. D. Increased expression of nusap in recurrent prostate cancer is mediated by e2f1. Oncogene 32, 70–77. https://doi.org/10.1038/onc.2012.27 (2013).
Gordon, C. A., Gong, X., Ganesh, D. & Brooks, J. D. Nusap1 promotes invasion and metastasis of prostate cancer. Oncotarget 8, 29935–29950. https://doi.org/10.18632/oncotarget.15604 (2017).
Li, H. et al. Nucleolar and spindle associated protein 1 promotes metastasis of cervical carcinoma cells by activating wnt/\(\beta\)-catenin signaling. J. Exp. Clin. Cancer Res. 38, 1–18. https://doi.org/10.1186/s13046-019-1037-y (2019).
Roy, S. et al. microrna 193a–5p regulates levels of nucleolar-and spindle-associated protein 1 to suppress hepatocarcinogenesis. Gastroenterology 155, 1951–1966. https://doi.org/10.1053/j.gastro.2018.08.032 (2018).
Kim, J. H. et al. Genomic predictors for recurrence patterns of hepatocellular carcinoma: Model derivation and validation. PLoS Med. 11, e1001770–e1001786. https://doi.org/10.1371/journal.pmed.1001770 (2014).
Mah, W.-C. et al. Methylation profiles reveal distinct subgroup of hepatocellular carcinoma patients with poor prognosis. PloS One 9, e104158–e104168. https://doi.org/10.1371/journal.pone.0104158 (2014).
Ritchie, M. E. et al. limma powers differential expression analyses for rna-sequencing and microarray studies. Nucleic Acids Res. 43, 1–13. https://doi.org/10.1093/nar/gkv007 (2015).
Carlson, M. R. et al. Genomic annotation resources in r/bioconductor. Stat. Genom. 67, 90. https://doi.org/10.1007/978-1-4939-3578-9_4 (2016).
Wickham, H. et al. ggplot2: Create elegant data visualisations using the grammar of graphics (3.3. 6)[computer software]. https://cran.r-project.org/package=ggplot2. Accessed 25 June 2022 (2022).
Gaujoux, R. & Seoighe, C. Nmf: Algorithms and framework for nonnegative matrix factorization (nmf). R Package Version 0.20 6, http://CRAN.R-project.org/package=NMF (2015).
Hasan, M. A. M., Nasser, M., Pal, B. & Ahmad, S. Support vector machine and random forest modeling for intrusion detection system (ids). J. Intell. Learn. Syst. Appl. 2014, 1. https://doi.org/10.4236/jilsa.2014.61005 (2014).
Jan, S. U., Lee, Y.-D., Shin, J. & Koo, I. Sensor fault classification based on support vector machine and statistical time-domain features. IEEE Access 5, 8682–8690. https://doi.org/10.1109/ACCESS.2017.2705644 (2017).
Kanehisa, M. Toward understanding the origin and evolution of cellular organisms. Protein Sci. 28, 1947–1951. https://doi.org/10.1002/pro.3715 (2019).
Kanehisa, M., Furumichi, M., Sato, Y., Kawashima, M. & Ishiguro-Watanabe, M. Kegg for taxonomy-based analysis of pathways and genomes. Nucleic Acids Res.https://doi.org/10.1093/nar/gkac963 (2022).
Huang, D. W., Sherman, B. T. & Lempicki, R. A. Systematic and integrative analysis of large gene lists using david bioinformatics resources. Nat. Protocols 4, 44–57. https://doi.org/10.1038/nprot.2008.211 (2009).
Szklarczyk, D. et al. The string database in 2017: Quality-controlled protein-protein association networks, made broadly accessible. Nucleic Acids Res. 45, D362–D368. https://doi.org/10.1093/nar/gkw937 (2016).
Shannon, P. et al. Cytoscape: A software environment for integrated models of biomolecular interaction networks. Genome Res. 13, 2498–2504. https://doi.org/10.1101/gr.123930 (2003).
Chin, C.-H. et al. cytohubba: Identifying hub objects and sub-networks from complex interactome. BMC Syst. Biol. 8, 1–7. https://doi.org/10.1186/1752-0509-8-S4-S11 (2014).
Bader, G. D. & Hogue, C. W. An automated method for finding molecular complexes in large protein interaction networks. BMC Bioinform. 4, 1–27. https://doi.org/10.1186/1471-2105-4-2 (2003).
Robin, X. et al. Proc: An open-source package for r and s+ to analyze and compare roc curves. BMC Bioinform. 12, 1–8. https://doi.org/10.1186/1471-2105-12-77 (2011).
Therneau, T. & Lumley, T. R survival package. R Core Team. https://rweb.webapps.cla.umn.edu/R/library/survival/doc/survival.pdf. Accessed 30 June 2022 (2013).
Funding
This work was supported by the Graduate School Research Fund of The University of Aizu, Japan.
Author information
Authors and Affiliations
Contributions
All listed authors participated meaningfully in the study, and they have seen and approved the submission of this manuscript. Conceptualization, M.A.M.H.; Methodology, M.A.M.H., M.M.; Data collection and curation: M.A.M.H., M.M., J.S.; Interpreted and analyzed the data, M.A.M.H., M.M., J.S.; Writing-original draft preparation, M.A.M.H., M.M.; Writing-review and editing, M.A.M.H., M.M., J.S.; Supervision, J.S., M.A.M.H.; Project administration and funding, J.S.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Hasan, M.A.M., Maniruzzaman, M. & Shin, J. Differentially expressed discriminative genes and significant meta-hub genes based key genes identification for hepatocellular carcinoma using statistical machine learning. Sci Rep 13, 3771 (2023). https://doi.org/10.1038/s41598-023-30851-1
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-023-30851-1
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.