Abstract
We developed a novel prediction model for recurrence and survival in patients with localized renal cell carcinoma (RCC) after surgery and a novel statistical method of machine learning (ML) to improve accuracy in predicting outcomes using a large Asian nationwide dataset, updated KOrean Renal Cell Carcinoma (KORCC) database that covered data for a total of 10,068 patients who had received surgery for RCC. After data pre-processing, feature selection was performed with an elastic net. Nine variables for recurrence and 13 variables for survival were extracted from 206 variables. Synthetic minority oversampling technique (SMOTE) was used for the training data set to solve the imbalance problem. We applied the most of existing ML algorithms introduced so far to evaluate the performance. We also performed subgroup analysis according to the histologic type. Diagnostic performances of all prediction models achieved high accuracy (range, 0.77–0.94) and F1-score (range, 0.77–0.97) in all tested metrics. In an external validation set, high accuracy and F1-score were well maintained in both recurrence and survival. In subgroup analysis of both clear and non-clear cell type RCC group, we also found a good prediction performance.
Similar content being viewed by others
Introduction
The incidence of renal cell carcinoma (RCC) is increasing worldwide. Approximately 76,000 new cases and almost 14,000 deaths from RCC were reported in the US in 20211. In Korea, we also observed the same trend according to the latest cancer incidence statistics from the Korea Central Cancer Registry2. Among them, clear cell type RCC represents approximately 70% cases in adults3. Estimated 5-year survival rate of localized RCC patients is approximately 90%. However, in about 30% of either recurrence or metastasis cases, the survival rate is drastically reduced4. Thus, it is imperative to predict the high-risk group for recurrence in advance and establish a differentiated surveillance protocol for patients who have undergone a curative surgery.
Over the past decades, several nomograms for recurrence and/or survival of localized RCC have been developed and applied in clinical practice5,6,7,8. Among them, the Kattan nomogram based on pathological T stage, nuclear grade, tumor size, necrosis, vascular invasion, and clinical presentation was the first introduced and widely used model5,6. Subsequently, the Leibovich model was developed by Mayo Clinic to estimate the risk of metastasis or recurrence using tumor stage, regional lymph node status, tumor size, nuclear grade and histologic tumor necrosis7. The most recently developed model known as the GRANT score was based on patient age, nuclear grade, and pathologic T/N stage8. However, these models were developed and validated using a small cohort from a single institution. In addition, they were limited to Western datasets. Moreover, their prediction accuracies were not as high as expected. For most models, their accuracy values were around 0.75,6,7,8.
Thus, we tried to develop a novel prediction model for recurrence and survival in patients with localized RCC after surgery using a large Asian nationwide dataset. We also used a novel statistical method of machine learning (ML) to improve accuracy in predicting outcomes.
Materials and methods
Ethics statement
The Institutional Review Board (IRB) of Seoul National University Bundang Hospital approved this study (approval number: B-2106-688-108). The requirement for obtaining written informed consent from patients was waived by the IRB due to the retrospective nature of this study. Personal identifiers were completely deleted to ensure that data were analyzed anonymously. Our study was conducted according to the ethical standards of the 1964 Declaration of Helsinki and its later amendments.
Data sets
The KOrean Renal Cell Carcinoma (KORCC) database was first established in 2011. It had data from eight academic institutions nationwide9. Recently, data of each institution were updated from March to June 2021. Subsequently, the updated KORCC database covered data of a total of 10,068 patients who had received surgery for RCC with 206 variables, including demographic, perioperative, pathologic, and survival information.
Model development (n = 4,829) and internal validation (n = 2,070) were performed using data from seven centers except data from Seoul National University Bundang Hospital (SNUBH, n = 3,169). External validation was performed using data from the SNUBH to assess the generality of the model performance. SNUBH was suitable for external validation because of its size and diverse patient population.
All study procedures were performed according to the transparent reporting of a multivariable prediction model for individual prognosis or diagnosis (TRIPOD) recommendations10.
All institutions obtained IRB approvals before inputting data into the database. Unified data templates were used for consistent data collection at each institution. Survival data were retrospectively reviewed from medical records or identified from death certificate data.
Data processing and feature selection
Data pre-processing mainly included processing missing values to obtain a reliable set of data. The missing value imputation process was divided into three aspects: patients, predictors, and statistics. At first, we eliminated patients with missing basic information. Subsequently, we performed predictive analytics for variables including total protein, Hb, creatinine. For this method, we used Euclidean distance to determine the similarity between two values and replace the missing one with similar one. Other missing values were corrected using k-nearest neighbor (KNN)11. KNN is non-parametric and instance-based method, and useful for datasets having both qualitative and quantitative attribute values.
After pre-processing, we performed feature selection with an elastic net12. Before implementing elastic net model, we defined four default variables that had been considered as the most significant predictors for recurrence and survival: gender, age at surgery, smoking, and BMI13,14. Elastic net is known as a hybrid of ridge regression and lasso regularization. Thus, elastic net can generate reduced models by generating zero-valued coefficients. Similar to the lasso, elastic net simultaneously perform automatic variable selection and continuous shrinkage15. We subsequently performed a feature importance raking method (Supplemental Fig. 1). Finally, we extracted nine variables for recurrence and 13 variables for survival (Fig. 1).
Synthetic minority oversampling technique (SMOTE)
Imbalanced data problem is a situation in which data are biased toward one class in applying ML classification algorithms16. When modeling using imbalanced data, the ML algorithm attempts to improve the performance by predicting a large number of classes, in which most patients are concentrated, resulting in lower predictability of a small number of classes. Thus, imbalanced data problem should be solved using methods such as oversampling or underdamping. In the current study, we used the SMOTE to the training data set to solve the imbalance problem17.
Statistical analysis and ML model development
We evaluated performances of the following representative ML classification algorithms: logistic regression18, kernel support vector machine (SVM)19, decision tree20, random forest21, naïve Bayes (NB)22, Extreme Gradient Boosting algorithm (XGBoost)23, Natural Gradient Boosting (NGBoost)24, LightGbm25, and CatBoost26. We adopted accuracy and F-1 score to evaluate the prediction performance. The F-1 score is made up of both precision and recall metrics. It is designed to work more accurately on imbalanced data27. We also performed subgroup analysis according to histologic type. Non-clear cell type RCC included eight types: papillary, chromophobe, collecting duct, unclassified, multilocular cystic, mixed, Xp11.2 translocation, and clear cell papillary. All statistical analyses were performed using commercially available software (IBM SPSS Statistics ver. 21.0 and Python ver. 3.7.6).
Ethics statement
The Institutional Review Board (IRB) of Seoul National University Bundang Hospital approved this study (approval number: B-2106–688-108).
Informed consent to patients
The waiver of the informed consent requirement was approved by the local ethics committee of Seoul National University Bundang Hospital considering the retrospective study design involving anonymized data.
Results
Patient characteristics
Distribution of data sets before and after SMOTE for recurrence (n = 6,717) and survival (n = 5,730) is described in Table 1. The ratio of training set to test set was 7:3. Overall survival rates at 3, 5, and 10 years were 94.2%, 90.6%, and 71.9%, respectively; and the recurrence-free rates were 85.2%, 78.8% and 45.3%, respectively.
Subsequently, we compared patient characteristics and distribution of each variable for recurrence and survival (Table 2). In a comparative analysis between recurrence and non-recurrence groups, we found several significantly different variables except for four default variables (gender, age at surgery, smoking, and BMI): Eastern Cooperative Oncology Group (ECOG) performance status, symptoms at diagnosis, transfusion, pathologic T/N stages, sarcomatoid differentiation, necrosis, lymphovascular invasion (LVI), and Fuhrman nuclear grade (all p < 0.05). In terms of survival, ECOG performance status, symptoms at diagnosis, transfusion, pathologic T/N stages, sarcomatoid differentiation, necrosis, LVI, histologic type, Fuhrman nuclear grade, and recurrence were significant variables (all p < 0.05).
Prediction model performance and external validation
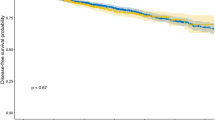
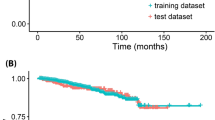
Diagnostic performance of several machine learning algorithms for the prediction of 3-, 5-, and 10-year recurrence and survival are listed in Table 3. All models achieved very high accuracy (range, 0.77–0.94) and F1-score (range, 0.77–0.97) in all tested metrics. Subsequently, external validation with a SNUBH dataset (n = 3,169) was performed using all models (Fig. 2). High accuracy and F1-score were well maintained in external validation in both recurrence and survival (Supplemental Table 1).
Subgroup analysis
In subgroup analysis according to the histologic type (clear vs. non-clear cell type RCC), dataset distribution before and after SMOTE for recurrence and survival is described in Supplemental Table 2. Consequently, we also found very high accuracy (range, 0.64–0.91) and F1-score (range, 0.72–0.94) in all tested metrics (Supplemental Tables 3 and 4).
Discussion
Using the original KORCC database9, two recent studies have been reported28,29. At first, Byun et al.28 assessed the prognosis of non-metastatic clear cell RCC using a deep learning-based survival predictions model. Harrel’s C-indices of DeepSurv for recurrence and cancer-specific survival were 0.802 and 0.834, respectively. More recently, Kim et al.29 developed ML-based algorithm predicting the probability of recurrence at 5 and 10 years after surgery. The highest area under the receiver operating characteristic curve (AUROC) was obtained from the naïve Bayes (NB) model, with values of 0.836 and 0.784 at 5 and 10 years, respectively.
In the current study, we used the updated KORCC database. It now contains clinical data of more than 10,000 patients. To the best of our knowledge, this is the largest dataset in Asian population with RCC. With this dataset, we could develop much more accurate models with very high accuracy (range, 0.77–0.94) and F1-score (range, 0.77–0.97, Table 3). The accuracy values were relatively high compared to the previous models, including the Kattan nomogram, Leibovich model, the GRANT score, which were around 0.75,6,7,8. Among them, the Kattan nomogram was developed using a cohort of 601 patients with clinically localized RCC, and the overall C-index was 74%5. In a subsequent analysis with the same patient group using an additional prognostic variables including tumor necrosis, vascular invasion, and tumor grade, the C-index was as high as 82%30. Their prediction accuracies were not as high as ours yet.
In addition, we could include short-term (3-year) recurrence and survival data, which would be helpful for developing more sophisticated surveillance strategy. The other strength of current study was that most algorithms introduced so far had been applied18,19,20,21,22,23,24,25,26, showing relatively consistent performance with high accuracy. Finally, we also performed an external validation by using a separate (SNUBH) cohort, and achieved well maintained high accuracy and F1-score in both recurrence and survival (Fig. 2). External validation of prediction models is essential, especially in case of using the multi-institutional dataset, to ensure and correct for differences between institutions.
AUROC has been mostly used as the standard evaluating performance of prediction models5,6,7,8,29. However, AUROC weighs changes in sensitivity and specificity equally without considering clinically meaningful information6. In addition, the lack of ability to compare performance of different ML models is another limitation of AUROC technique31. Thus, we adopted accuracy and F1-score instead of AUROC as evaluation metrics. F1-score, in addition to SMOTE17, is used as better accuracy metrics to solve the imbalanced data problems27.
RCC is not a single disease, but multiple histologically defined cancers with different genetic characteristics, clinical courses, and therapeutic responses32. With regard to metastatic RCC, the International Metastatic Renal Cell Carcinoma Database Consortium and the Memorial Sloan Kettering Cancer Center risk model have been extensively validated and widely used to predict survival outcomes of patients receiving systemic therapy33,34. However, both risk models had been developed without considering histologic subtypes. Thus, the predictive performance was presumed to have been strongly affected by clear cell type (predominant histologic subtype) RCC. Interestingly, in our previous study using the Korean metastatic RCC registry, we found the both risk models reliably predicted progression and survival even in non-clear cell type RCC35. In the current study, after performing subgroup analysis according to the histologic type (clear vs. non-clear cell type RCC), we also found very high accuracy and F1-score in all tested metrics (Supplemental Tables 3 and 4). Taking together, these findings suggest that the prognostic difference between clear and non-clear cell type RCC seems to be offset both in metastatic and non-metastatic RCC. Further effort is needed to develop and validate a sophisticated prediction model for individual subtypes of non-clear cell type RCC.
The current study had several limitations. First, due to the paucity of long-term follow-up cases at 10 years, data imbalance problem could not be avoided. Subsequently, recurrence-free rate at 10-year was reported only to be 45.3%. In the majority of patients, further long-term follow up had not been performed in case of no evidence of disease at five years. However, we adopted both SMOTE and F1-score to solve these imbalanced data problems. The retrospective design of this study was also an inherent limitation. Another limitation was that the developed prediction model only included the Korean population. Validation of the model using data from other countries and races is also needed. In regard of non-clear cell type RCC, the current study cohort is still relatively small due to the rarity of the disease, we could not avoid integrating each subtype and analyzing together. Thus, further studies is still needed to develop and validate a prediction model for each subtypes. In addition, the lack of more accurate classifiers such as cross-validation and bootstrapping is another limitation of current study. Finally, the web-embedded deployment of model should be followed to improve accessibility and transportability.
Conclusions
A novel ML algorithm for predicting recurrence and survival in localized RCC patients after surgery was successfully developed and validated using the updated KORCC database. This prediction model is anticipated to offer a differentiated surveillance protocol. It will be a useful tool for patient counseling.
Data availability
All data enquiries can be directed to the corresponding author.
References
Siegel, R. L., Miller, K. D., Fuchs, H. E. & Jemal, A. Cancer statistics, 2021. CA Cancer J. Clin. 71, 7–33. https://doi.org/10.3322/caac.21654 (2021).
National Cancer Registration Statistics, Ministry of Health and Welfare, Republic of Korea. https://www.cancer.go.kr (2019).
Gansler, T., Fedewa, S., Amin, M. B., Lin, C. C. & Jemal, A. Trends in reporting histological subtyping of renal cell carcinoma: Association with cancer center type. Hum. Pathol. 74, 99–108. https://doi.org/10.1016/j.humpath.2018.01.010 (2018).
Ferlay, J. et al. Estimating the global cancer incidence and mortality in 2018: GLOBOCAN sources and methods. Int. J. Cancer 144, 1941–1953. https://doi.org/10.1002/ijc.31937 (2019).
Kattan, M. W., Reuter, V., Motzer, R. J., Katz, J. & Russo, P. A postoperative prognostic nomogram for renal cell carcinoma. J. Urol. 166, 63–67 (2001).
Sorbellini, M. et al. A postoperative prognostic nomogram predicting recurrence for patients with conventional clear cell renal cell carcinoma. J. Urol. 173, 48–51. https://doi.org/10.1097/01.ju.0000148261.19532.2c (2005).
Leibovich, B. C. et al. Prediction of progression after radical nephrectomy for patients with clear cell renal cell carcinoma: A stratification tool for prospective clinical trials. Cancer 97, 1663–1671. https://doi.org/10.1002/cncr.11234 (2003).
Buti, S. et al. Validation of a new prognostic model to easily predict outcome in renal cell carcinoma: The GRANT score applied to the ASSURE trial population. Ann. Oncol. 28, 2747–2753. https://doi.org/10.1093/annonc/mdx492 (2017).
Byun, S. S. et al. The establishment of KORCC (KOrean Renal Cell Carcinoma) database. Investig. Clin. Urol. 57, 50–57. https://doi.org/10.4111/icu.2016.57.1.50 (2016).
Collins, G. S., Reitsma, J. B., Altman, D. G., Moons, K. G. M. & members of the, T. g. Transparent Reporting of a Multivariable Prediction Model for Individual Prognosis or Diagnosis (TRIPOD): The TRIPOD Statement. Eur Urol 67, 1142-1151, doi:https://doi.org/10.1016/j.eururo.2014.11.025 (2015).
Mucherino, A., Papajorgji, P., Pardalos, P. M. & SpringerLink. k-Nearest Neighbor Classification in Data Mining in Agriculture. Vol 34. (Springer, 2009). https://doi.org/10.1007/978-0-387-88615-2_4
Zou, H. & Hastie, T. Regularization and variable selection via the elastic net (vol B 67, pg 301, 2005). J. R. Stat. Soc. B 67, 768–768. https://doi.org/10.1111/j.1467-9868.2005.00527.x (2005).
Scelo, G. & Larose, T. L. Epidemiology and risk factors for kidney cancer. J. Clin. Oncol. https://doi.org/10.1200/JCO.2018.79.1905 (2018).
Capitanio, U. et al. Epidemiology of renal cell carcinoma. Eur. Urol. 75, 74–84. https://doi.org/10.1016/j.eururo.2018.08.036 (2019).
Chen, G. et al. ProAcePred: Prokaryote lysine acetylation sites prediction based on elastic net feature optimization. Bioinformatics 34, 3999–4006. https://doi.org/10.1093/bioinformatics/bty444 (2018).
Li, D. C., Liu, C. W. & Hu, S. C. A learning method for the class imbalance problem with medical data sets. Comput. Biol. Med. 40, 509–518. https://doi.org/10.1016/j.compbiomed.2010.03.005 (2010).
Alghamdi, M. et al. Predicting diabetes mellitus using SMOTE and ensemble machine learning approach: The Henry Ford ExercIse Testing (FIT) project. PLoS ONE 12, e0179805. https://doi.org/10.1371/journal.pone.0179805 (2017).
Liao, J. G. & Chin, K. V. Logistic regression for disease classification using microarray data: model selection in a large p and small n case. Bioinformatics 23, 1945–1951. https://doi.org/10.1093/bioinformatics/btm287 (2007).
Huang, M. W., Chen, C. W., Lin, W. C., Ke, S. W. & Tsai, C. F. SVM and SVM ensembles in breast cancer prediction. PLoS ONE 12, e0161501. https://doi.org/10.1371/journal.pone.0161501 (2017).
Song, Y. Y. & Lu, Y. Decision tree methods: Applications for classification and prediction. Shanghai Arch. Psychiatry 27, 130–135. https://doi.org/10.11919/j.issn.1002-0829.215044 (2015).
Chan, J. C. W. & Paelinckx, D. Evaluation of Random Forest and Adaboost tree-based ensemble classification and spectral band selection for ecotope mapping using airborne hyperspectral imagery. Remote Sens. Environ. 112, 2999–3011. https://doi.org/10.1016/j.rse.2008.02.011 (2008).
Subbalakshmi, G., Ramesh, K. & Chinna, R. M. Decision support in heart disease prediction system using Naive Bayes. Indian J. Comput. Sci. Eng. 2, 170–176 (2011).
Chen, T. Q. & Guestrin, C. XGBoost: A Scalable Tree Boosting System. Kdd'16: Proceedings of the 22nd Acm Sigkdd International Conference on Knowledge Discovery and Data Mining, 785–794, https://doi.org/10.1145/2939672.2939785 (2016).
Duan, T. et al. NGBoost: Natural gradient boosting for probabilistic prediction. Pr. Mach. Learn. Res. 119, 71 (2020).
Ke, G. L. et al. LightGBM: A highly efficient gradient boosting decision tree. Adv. Neur. In. 30, 89 (2017).
Dorogush, A.V., Ershov, V., & Gulin, A. CatBoost: gradient boosting with categorical features support. ArXiv, abs/1810.11363 (2018).
Takahashi, K., Yamamoto, K., Kuchiba, A. & Koyama, T. Confidence interval for micro-averaged F-1 and macro-averaged F-1 scores. Appl. Intell. 52, 4961–4972. https://doi.org/10.1007/s10489-021-02635-5 (2022).
Byun, S. S. et al. Deep learning based prediction of prognosis in nonmetastatic clear cell renal cell carcinoma. Sci. Rep. Uk 11, 21. https://doi.org/10.1038/s41598-020-80262-9 (2021).
Kim, H., Lee, S. J., Park, S. J., Choi, I. Y. & Hong, S. H. Machine learning approach to predict the probability of recurrence of renal cell carcinoma after surgery: Prediction model development study. Jmir Med. Inf. 9, 35 (2021).
Halligan, S., Altman, D. G. & Mallett, S. Disadvantages of using the area under the receiver operating characteristic curve to assess imaging tests: A discussion and proposal for an alternative approach. Eur. Radiol. 25, 932–939. https://doi.org/10.1007/s00330-014-3487-0 (2015).
Carrington, A. M. et al. A new concordant partial AUC and partial c statistic for imbalanced data in the evaluation of machine learning algorithms. BMC Med. Inform. Decis. 20, 1014 (2020).
Ricketts, C. J. et al. The cancer genome atlas comprehensive molecular characterization of renal cell carcinoma. Cell Rep. 23, 313. https://doi.org/10.1016/j.celrep.2018.03.075 (2018).
Heng, D. Y. C. et al. External validation and comparison with other models of the International Metastatic Renal-Cell Carcinoma Database Consortium prognostic model: A population-based study. Lancet Oncol. 14, 141–148. https://doi.org/10.1016/S1470-2045(12)70559-4 (2013).
Motzer, R. J. et al. Prognostic factors for survival in previously treated patients with metastatic renal cell carcinoma. J. Clin. Oncol. 22, 454–463. https://doi.org/10.1200/Jco.2004.06.132 (2004).
Kim, J. K. et al. Application of the international metastatic renal cell carcinoma database consortium and memorial sloan kettering cancer center risk models in patients with metastatic non-clear cell renal cell carcinoma: A multi-institutional retrospective study using the korean metastatic renal cell carcinoma registry. Cancer Res. Treat. 51, 758–768. https://doi.org/10.4143/crt.2018.421 (2019).
Author information
Authors and Affiliations
Contributions
Conception: J.K.K, S.S.B., H.O. Data collection: S.L., S.K.H., C.K., C.W.J., S.H.K., S.H.H., Y.J.K., J.C., E.C.H., T.G.K. Data analyzation: J.K.K., Y.J.J., J.L., J.K., H.O. Draft manuscript: J.K.K., C.W.J., S.H.K., Y.J.K., J.C., T.G.K. Reviewed manuscript: S.K.H., C.K., S.H.H., E.C.H., S.S.B. Revised manuscript: J.K.K, H.O. All the authors showed consent for publication of this study.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Kim, J.K., Lee, S., Hong, S.K. et al. Machine learning based prediction for oncologic outcomes of renal cell carcinoma after surgery using Korean Renal Cell Carcinoma (KORCC) database. Sci Rep 13, 5778 (2023). https://doi.org/10.1038/s41598-023-30826-2
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-023-30826-2
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.