Abstract
Antimicrobial resistance has been developing fast and incurring a loss of human life, and there is a need for new antimicrobial agents. Naturally occurring antimicrobial peptides offer the characteristics to counter AMR because the resistance development is low or no resistance. Antimicrobial peptides from Paenibacillus peoriae IBSD35 cell-free supernatant were salted out and purified using chromatography and characterized with liquid chromatography–tandem-mass spectrometry. The extract has shown a high and broad spectrum of antimicrobial activity. Combining the strain IBSD35 genome sequence with its proteomic data enabled the prediction of biosynthetic gene clusters by connecting the peptide from LC–MS/MS data to the gene that encode. Antimicrobial peptide databases offered a platform for the effective search, prediction, and design of AMPs and expanded the studies on their isolation, structure elucidation, biological evaluation, and pathway engineering. The genome-based taxonomy and comparisons have shown that P. peoriae IBSD35 is closely related to Paenibacillus peoriae FSL J3-0120. P. peoriae IBSD35 harbored endophytic trait genes and nonribosomal peptide synthases biosynthetic gene clusters. The comparative genomics revealed evolutionary insights and facilitated the discovery of novel SMs using proteomics from the extract of P. peoriae IBSD35. It will increase the potential to find novel bio-molecules to counter AMR.
Similar content being viewed by others
Introduction
The discovery of penicillin has heralded the modern era of drugs and saved millions of lives. However, it was short-lived and antimicrobial resistance (AMR) has been developing fast and incurring a loss of human life1,2. Given this, there is a continuous need for new antimicrobial agents. Prokaryotes perform a vital function for organisms and ecosystems and provide by-products for human and veterinary medicine, agriculture, and manufacturing. They are well-known to mediate microbe-host and microbe-microbe interactions for their survival competition. Consequently, they produced diverse bio-molecules vital for the structure and proper function of living cells or organisms3. At the same time, they caused infectious diseases and extended the AMR through the environment, reproduction, and gene transfer4.
The first functionalized peptide-producing endophyte was isolated from Kennedia nigriscans5. Afterward, Bacillus amyloliquefaciens sp. was isolated from Ophiopogon japonicas which afforded the discovery of antitumor exo-polysaccharides6. We presumed that the plant-microbes association might have affected each other in their respective biochemical properties7. This interaction might enable microbe existence as an endophyte through absorbing or secreting biomolecules8. Furthermore, the discovery of anticancer taxol and taxane production by Taxomyces andreanae has stimulated research interest in bacterial and fungal endophytes9. The understudy P. peoriae IBSD35 (or strain IBSD35) was isolated from the stem of Millettia pachycarpa in our previous experiment10,11,12,13,14, and it has shown antimicrobial activity from its cell-free supernatant (CFS)12. Therefore, we investigated the source genome’s antimicrobial characteristics, extracellular antimicrobial compounds, and the potential to counter AMR.
Naturally occurring antimicrobial peptides (AMPs) offer the characters to counter AMR. They are present in every phylum and kingdom15 and are known for antibacterial, anticancer, antiviral, antifungal, antiparasite, effectors of innate immunity, and antibiotic adjuvant characteristics16,17. The resistance development of AMPs by gene mutation is less expected as mature peptides are created from a non-descript sequence of pro-peptides and lack a definite recognition site. They have multiple attacking target sites in contrast to common antibiotics that target specific proteins16. AMPs are highly conserved, hydrophobic, and mostly amphiphilic despite their diverse biological properties18.
A comprehensive genomics analysis, advanced proteomics techniques, AMP databases, and structural models offer a platform for effective search, prediction, drug development, and elaboration of AMPs19,20. The mass spectrometry (MS) technique can identify AMP sequence and mass from a mixture sample and subsequently disclose their primary, secondary, and tertiary structure21,22. The liquid chromatography-tandem-mass spectrometry (LC–MS/MS), proteomics, and AMP databases have broadly impacted AMP biology and the development of peptide-based drugs23. Moreover, genome mining has enabled the prediction of a biosynthetic gene cluster (BGC), linking the proteins and their peptides from LC–MS/MS data and connecting these natural products to the genes that encode them24. Proteomics has enabled genomic data to guide the discovery of new molecules to be applied in biomedical research25,26,27.
Therefore, the present study analyzes the characteristics of the P. peoriae IBSD35 draft genome and the proteomics of its extracellular AMPs. The understudy genome and its close strain comparisons illustrate its endophytic and antimicrobial traits. It provided an insight into P. peoriae IBSD35 molecular mechanisms and elucidated its potential as a source of novel AMPs to counter AMR.
Results
Genomic deoxyribonucleic acid (DNA) estimation and annotation
Paenibacillus peoriae IBSD35 inoculated on Luria Bertani (LB) agar plate was visible after 12 h of incubation at 38 °C. It is an endospore-forming and neutralophilic bacterium based on Integrated Microbial Genomes Annotation Pipeline (IMG AP) v.4.16.628,29. The genomic deoxyribonucleic acid (gDNA) assessed with 0.8% agarose was estimated to be more than 50-kilo base pairs (kbp), and the concentration was 188.63 µg µl−1 (Supplementary Fig. S1). P. peoriae IBSD35 has a linear DNA topology with a total length of 5,862,582 bp. Its closest species reference genome is Paenibacillus peoriae KCTC 3763 with a symmetrical identity of 62.7495% and a symmetrical identity of 89.4884% with P. peoriae FSL J3-0120 in National Center for Biotechnology Information (NCBI) neighbor genome report30.
Spectrum of antimicrobial activity of the P. peoriae IBSD35 inoculum
The CFS of P. peoriae IBSD35 inoculum retained antimicrobial activity against Gram-positive Staphylococcus aureus American Type Culture Collection (ATCC) 25923. The crude extract has shown antimicrobial activity against the Gram-negative Escherichia coli ATCC 25922 and fungal pathogen Candida albicans ATCC 10231, indicating it’s broad-spectrum of antimicrobial activity. The variation of antimicrobial activity at different stages of purification and concentration suggested the complex nature of biomolecules (Fig. 1A–C).
Cut-well agar diffusion showing antimicrobial activity against (A) Staphylococcus aureus ATCC 25923, (B) Escherichia coli ATCC 25922, (C) Candida albicans ATCC 10231, (D) Klebsiella pneumoniae ATCC 4352, and (E) Salmonella typhimurium ATCC 14028. a, b, c, d, e, and f indicated the cut-wells loaded at different steps of purification and concentration.
Optimum growth conditions for antimicrobial metabolites production
The P. peoriae IBSD35 inoculum’s optimum optical density (OD) was measured to be 0.65 at λ600 nm observed on the 6th day, coinciding with the exponential growth phase in the LB medium. The optimum growth was at pH 6.8–7. P. peoriae IBSD35 is a mesophile bacterium that grows best in moderate temperatures observed at 38 °C (Fig. 2A). The supernatant antimicrobial activity was determined as an arbitrary unit (AU) per milliliter which is the reciprocal of the highest dilution (2n) that resulted in the inhibition of S. aureus ATCC 25923 indicator lawns31. The AU was measured to be 1600 AU ml−1.
(A) The graphic diagram of the P. peoriae IBSD35 optimum growth curve in LB medium. The purple zone indicates the non-growth region, yellow zone represents the growth region, and the black dot indicates the optimum growth zone. (Left Y-axis is the optical density, Right Y- is the optimum temperature, and X-axis is the pH). (B) The chromatogram of the RP-HPLC. X-axis shows the retention time in min while Y-axis shows the absorbance. (The retention times indicate the peaks for practical purposes. 4.165 represents peak P4, and 4.567 represents peak P5). P4 retention time is 4.165 min. The P5 retention time is 4.567 min. Sharp peaks of P4 and P5 indicate they are sufficiently separated and have good biomolecule concentration. The P4 fractions exhibiting higher antimicrobial activity were pooled together and selected.
Salting out and antimicrobial activity retention test of dialysate
The fraction (F4) eluted out with the 700 milimolar (mM) sodium chloride (NaCl) at the flow rate of 2 ml min−1 has shown antimicrobial activity against S. aureus ATCC 25923. The fraction OD was measured to be 1.147 and 2.803 at wavelength (λ)214 and λ280, respectively (Table S1). The fraction (F4) was collected, lyophilized into powder form, and store at − 20 °C. It dissolved in distilled water. It was dialyzed with 12.4 kilo dalton (kDa) molecular weight cut-off (MWCO) dialysis tube (Sigma Aldrich, Germany) to eliminate the larger protein complexes. The dialysate has retained antimicrobial activity against S. aureus ATCC 25923.
Reverse phase-high performance liquid chromatography (RP-HPLC) and estimation of protein
The protein mixture gradient elution was at an isocratic flow rate of 2.5 ml min−1. The P4 (4.165) retention time was 4.165 min with a purity index of 0.86310, and the P5 (4.567) retention time was 4.567 min with a purity index of 0.95597 (Table S2). The RP-HPLC peaks P4 and P5 are sufficiently separated (Fig. 2B), and their absorption at λ205 and λ214 indicated the presence of amino acids and peptide bonds with other impurities. The absorbance at λ280 confirmed the presence of proteins. The P4 fractions had exhibited higher antimicrobial activity than P5. The purification fold of RP-HPLC was 151.72 starting from the initial crude extract (Table S1). The loss of bioactive metabolites from the harvesting of the cells ranged from 0 to 7.7%. The loss of cells after extraction ranged from 0.6 to 2.3%.
Effect of temperature, detergents, inorganic solvents, pH variation, and degradative enzymes on antimicrobial activity
A 0.2 µm filter (Avixa) sterilized supernatant treated at different temperatures retained the antimicrobial activity against S. aureus ATCC 25923. Autoclaving at 121 °C for 15 min and treatment with Tween 20, Tween 80, and sodium dodecyl sulfate (SDS) have slightly increased the antimicrobial activity (from 10 to 14 mm zone of inhibition diameter). It might be due to the disaggregation and exposure of peptide bonds. The inorganic solvents, pH variation, and degradative enzymes treated at a 1:1 ratio with the filtrate did not affect the antimicrobial activity significantly. The antimicrobial activity of the CFS is almost comparable to the ampicillin at 1 mg ml−1 (Table S3). Moreover, the sample persisted in low-temperature freezer storage (− 20 °C) without any preservative for 1 year.
Mass spectrometry and proteomics analysis of peptide mixtures
The P. peoriae IBSD35 proteins searched with Sequest, Mascot, and PEAKS from the uninterpreted experimental LC–MS/MS database confirmed the prediction of AMPs (Supplementary LC–MS Experiment data)32. The number of sequence protein entries predicted in the P. peoriae IBSD35 Uniprot database is 15 protein groups. Different peptide sequences have detected during the process (Table 1). An uncharacterized protein was predicted from the experimental LC–MS/MS with accession no. A0A2S6NUT6. The Uniprot search has shown that it is 94.87% similar to A0A509FZN2_PAEPO of Paenibacillus polymyxa (A0A509FZN2) (Bit score-296, E-value-3.6 e−89)33. Combining the genome mining and proteomics analysis has shown that the gene (ID 2816869382) is 97% similar to the gene (ID 2556935736) of Paenibacillus polymyxa CR1 (NC_023037) (Bit score-286; 1e−95) and it has 134 amino acid residues.
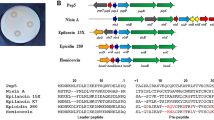
An AMP predicted from the protein lists with accession no. A0A2S6NUT6 is designated peoriaerin IBSD35 based on its source name (Table 2). Its sequence is AAGIQAQAGFGLSDSIQGTGKQKCSFCK. The N-terminal of the sequence considered is alanine (Ala). It has 28 amino acid residues with a molecular weight (MW) of 2802.17 Da (Supplementary Fig. S2). Its sequence backbone has a large portion of helix and residues embedded in the membrane. It suggested peoriaerin IBSD3 might insert the membrane for its antimicrobial activity (Fig. 3). The peptide sequence alignment indicated that it is 38.23% similar to fermentin20. The three-dimensional (3-D) model of peoriaerin IBSD35 was generated using an automated protein structure de novo modeling and deposited at ModelArchive with accession no. 10.5452ma-id7l4 (Fig. 3).
A 3-D model of peoriaerin IBSD35 predicted using de novo modeling. The amino acids composition of peptide is AAGIQAQAGFGLSDSIQGTGKQKCSFCK. Sequence contains high Gly content. It is indicated in black color. Glycine is integral to the formation of alpha-helices in secondary protein structure due to its compact form. The purple color indicated the helix structure property. It included the green color which intersects with the purple color. The green color indicated the residues embedded in membrane. The light orange color indicated the Trp which is the rings in the residues. The red color indicated the Ala. The whole structure represented the backbone and side chain of the AMP except the Gly indicated in black color.
Peoraerin IBSD35 mode of action imaging through scanning electron microscopy
The antimicrobial action had observed after 1 h of incubation. The minimum inhibition concentration (MIC) of peoriaerin IBSD35 was determined to be 0.04 µg µl−1. Scanning electron microscopy (SEM) imaging indicated that the pathogen colonies were slowly disappearing. It correlated with the decreased colony count from more than 400 to 4 in 2 h (Fig. 4). The OD at λ600 nm was 1.409 indicating the cell growth and debris turbidity. There was no evidence of DNA and protein leakage. It showed that the AMP might kill S. aureus ATCC 25923 by penetrating the cell cytoplasm and periplasm.
Mode of action of peoriaerin IBSD35 against S. aureus ATCC 25923 and imaging with scanning electron microscopy. The morphological changes of S. aureus ATCC 25923 when treated with peoriaerin IBSD35 1× MIC at different experimental conditions. (A) S. aureus ATCC 25923 only, (B) S. aureus ATCC 25923 with peoriaerin IBSD35 at 0 h, (C) S. aureus ATCC 25923 with peoriaerin IBSD35 at 1 h, (D) S. aureus ATCC 25923 with peoriaerin IBSD35 at 2 h. The yellow arrows point to S. aureus ATCC 25923 colonies without peoriaerin IBSD35 and at zero hours with peoriaerin IBSD35. The red arrow points to the disappearance of S. aureus ATCC 25923 at 1 and 2 h.
Draft genome sequencing, assembly and annotations of P. peoriae IBSD35
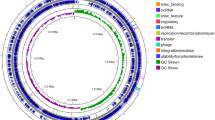
The nucleic acid and protein ratio (A260/A280) was estimated to be 1.9 (Fig. 5A). The draft genome with 250 bp-end reads produced a total number of raw reads (R1 + R2) of 15,497,242 bp34. Total bp are 3,874,320,000 bp with 46.01 guanidine-cytosine (GC) % and a read length of 250. The clean reads produced a total length of 5,862,582 bp with 58.0× fold coverage. The de-novo whole genome assembled with ABySS v.2.035 has produced 66 contigs totaling 5,862,582 bp with N50 of 400,732 and L50 of 6, 42 scaffolds and a GC content of 45.6% (Table S4). The NCBI Prokaryotic Genome Annotation Pipeline (PGAP) v.4.4 has identified 5245 genes, 4983 coding sequences (CDSs), 9, 2, 3 (5S, 16S, 23S) rRNAs (ribosomal ribonucleic acids), 2, 1 (5S, 16S) complete rRNAs, 7, 1, 3 (5S, 16S, 23S) partial rRNAs, 81 tRNAs (transfer ribonucleic acids), and 4 nc RNAs (noncoding RNAs) (Table S4). CRISPR/csd (clustered regularly interspaced short palindromic repeats) is involved in the prokaryotic defense system36. Out of two 16S rRNA bacterial short sequence units (SSU), one has 1560 bp, and the other has 45 bp based on IMG AP v.4.16.6.
(A) The crude gDNA band quality checked on 0.8% agarose gel, where Marker lane is 50 kb DNA ladder and, lane S is DNA whose molecular weight is more than 50 kb. (B) The WEGO analysis of P. peoriae IBSD35 genome. The gene ontology of genome has been classified into cellular component (Red), molecular function (Blue), and biological process (green) categories. The numbers of the genes are indicated in percentage.
Global overview of P. peoriae IBSD35 genome
The maximum number of protein-coding genes (4983) is related to the genetic information process (47%), environment information processing (45%), carbohydrate metabolism protein family (42%), and signaling process (31%) (Supplementary Fig. S3)37,38. Membrane and cell contributed the highest percentage of genes in the cellular component category, while the catalytic activity, binding, and transportation account for the highest number in molecular function. The metabolic and cellular processes have the highest percentage within the biological processes category (Fig. 5B). P. peoriae IBSD35 harbored cytochrome bd ubiquinol oxidase [EC: 7.1.1.7], a component of the aerobic respiratory chain gene (cydA). The ModelSEED predicted its reaction in the cytosol as39
Paenibacillus peoriae IBSD35 genome harbored nitrogenase molybdenum-iron protein beta chain (nifK, nifD) [EC: 1.18.6.1]40,41, phosphotransferase system (PTS), and sugar-specific component. They might be utilizing extra-cellular trehalose, cellobiose, and maltose. It revealed the P. Peoriae IBSD35 auxotrophic lifestyle (www.kegg.jp/kegg/kegg1.html)42,43. The Joint Genome Institute-Integrated Microbial Genomes and Microbiomes System (JGI-IMG/M) analysis revealed a putative horizontally transferred genes count of 10 (0.19%) with the best hits to genes from Proteobacteria and Clostridia. The trans-membrane count is 1392 (26.28%) and might serve as a gateway to transport specific substances across the membrane28,29. The signal peptide is 303 (5.72%) of the total CDS identified. It might prompt the translocation of a cell protein secretion44,45.
Paenibacillus peoriae IBSD35 harbored NisK-NisR biosynthesis sensor histidine kinase [EC: 2.7.13.3], a two-component signal transduction system (TCS), known to affect the biosynthesis of AMP46,47. It also harbored sensor histidine kinase and response regulators, two DesK–DesR of the NarL family, four CheA–CheY of a chemo-taxis family, two Res-ResD, and two DegS-DegU for motility, regulation, and interactions. Flagellar proteins (fliS) in the cellular chaperones might cause microbe motility and signal transduction. A flagellin/flagellar hook-associated protein (fliC) includes glycerol kinase and molecular chaperone (HtpG) that might involve in plant–microbe interaction48. P. peoriae IBSD35 genome also harbored biodegradation capability proteins for xenobiotic compounds49.
Comparison of genomes characteristics
The selected Paenibacillus species have shown varying numbers of DNA scaffolds ranging from 1 to 56. Paenibacillus sp. JDR-2 has the largest genome size (7,184,930 bp), and B. subtilis subtilis AG1839 has the smallest genome size (4,193,640 bp) among the selected genomes (Table 3). The dot plot homology revealed a low level of P. peoriae IBSD35 synteny with Paenibacillus peoriae KCTC 3763 but a higher synteny with Paenibacillus peoriae HS311, P. polymyxa SC2, and P. polymyxa E681 (Supplementary Fig. S4). The predicted numbers of nonribosomal peptide synthase (NRPS) BGCs ranged from 0 to 18 among the eleven genomes indicating their diverse and rich SM natural products (Table 3). P. peoriae IBSD35 harbored 18 NRPS (72% of the total 25 BGCs) and encompassed the highest number of NRPS BGCs among the selected genomes comparisons.
Antibiotics and other secondary metabolites (SMs) gene cluster of P. peoriae IBSD35
The CFS of P. peoriae IBSD35 has shown a high and broad-spectrum antimicrobial activity against S. aureus ATCC 25923, E. coli ATCC 25922, C. albicans ATCC 10231, Klebsiella pneumoniae ATCC 4352, and Salmonella typhimurium ATCC 14028 (Fig. 1A–E). The activity was comparable with ampicillin and nisin at a much lower dose. Antibiotics and Secondary Metabolite Analysis Shell (AntiSMASH) v5 using Integrated Microbial Genomes Atlas of Biosynthetic gene clusters (IMG-ABC) web server tools predicted P. peoriae IBSD35 harbored 25 BGCs50,51. LC–MS sequencing of RP-HPLC purified sample have predicted 15 protein lists (Table 3).
Paenibacillus peoriae IBSD35 harbored resistance genes for beta-lactam, vancomycin, cationic antimicrobial peptide (CAMP), platinum drug, and antifolate drug52,53,54,55. The Rapid Annotation of microbial genomes using Subsystems Technology (RAST) tool predicted that the P. peoriae IBSD35 genome harbored 29 compounds resistant to antibiotics and toxic compounds as compared to the metabolic reconstruction of P. polymyxa E681. They shared 23 compounds, while one is unique in the P. peoriae IBSD35 and nine are unique in P. polymyxa E681.
Paenibacillus peoriae IBSD35 harboring 18 NRPS indicated that nonribosomal peptide compounds are the most abundant SMs of our understudy strain IBSD35 genome (Fig. 6A). The genome encompassed three gene clusters- one Trans-acyltransferase polyketide synthase (Trans AT-PKS) and two clusters of Trans AT-PKS. Two BGCs are associated with Butyrolactone, and one each for Type III-polyketide synthase (T3PKS), bacteriocin, phosphonate, lanthipeptide, NRPS-like, polyketide synthases-like, and lasso peptide. At least 12 BGCs start from a position, and 2 BGCs code rhizomides A, B, and C as predicted by the Antibiotics and Secondary Metabolite Analysis Shell-Minimum Information about a Biosynthetic Gene Cluster (AntiSMASH-MIBiG) comparison (Table 4).
(A) The Heatmap of the BGCs of the selected genome. The names of the genomes are indicated on the left side. The rows represent the genomes BGCs, and its columns indicate the type of BGC. The BC types are indicated in description on bottom. Cells are colored with hues of green based on the number of copies of the selected protein family (Pfam) in the BC (numbers of BCs are indicated in numbering). Pfams (columns) that occur in all BCs (rows) define the core functions. (B) Venn diagram for the comparison of six closely related strains. (P. terrae HPL-003(NC_016641), P. peoriae HS311 (NZ_CP011512), P. peoriae IBSD35 (NZ_PTJM01000000), Paenibacillus sp. JDR-2 (NC_012914), P. polymyxa E681(NZ_CP048793), and P. polymyxa SC2 (NC_014622). They formed 5615 clusters, 5296 orthologous clusters, and 2125 single-copy gene clusters. They shared 2241 gene clusters. 6-clusters are specific to P. peoriae IBSD35. (C) Tree inferred with FastME 2.1.6.1114 from GBDP distances calculated from genome sequences. P. peoriae IBSD35 has a dDDH of (d4) 76.1, (C.I.) d4 of [73.1–78.9], and a difference G + C% of 0.16 with P. peoriae HS311 indicating their close species relation. The branch lengths are scaled in terms of GBDP distance formula d5. (Leaf labels are annotated by affiliation to species (1) and subspecies (2) clusters, genomic G + C content (3), δ values (4), overall genome sequence length (5), number of proteins (6), and the kind of strain (7). User-provided GenBank accession IDs are shown in parentheses; master record accessions are truncated (Percent G + C—43.5–72.1, No of Proteins—4218–7738, Genome size—4,193,640–8,667,507 bp), delta statistics—0.22–0.373, SSU lengths (in bp)—1331–1546).
Orthologous gene clusters of P. peoriae IBSD35 genome
OrthoVenn analysis has shown the selected Paenibacillus species (P. peoriae HS311, Paenibacillus sp. JDR-2, P. polymyxa E681, P. polymyxa SC2, P. terrae HPL-003, and P. peoriae IBSD35 formed 5615 clusters, 5296 orthologous clusters (at least contains two species), and 2125 single-copy gene clusters (E-value = 1e−2 and inflation value = 1.5) (Fig. 6B). The OrthoVenn diagram has shown that all 6 Paenibacillus species have shared 2241 gene clusters suggesting their conservation in the lineage. Additionally, it has 6-clusters specific to P. peoriae IBSD35 that might be lineage-specific gene clusters. The single-copy genes have suggested they maintained single-copy status through evolutionary time after the divergence of these species.
Central carbon metabolism (CCM) of P. peoriae IBSD35 genome
The metabolite precursors of P. peoriae IBSD35 cell biomass included gluconeogenesis (GG), citrate cycle (TCA), pentose phosphate pathway (PPP) metabolites, and pyruvate metabolism (Table 5)43,56,57,58. The precursors of polyketide-malonyl Coenzyme A (CoA) generated from the pyruvate metabolism- the Acetyl CoA included propinol CoA, and butanol CoA have derived from fatty acid biosynthesis and amino acids metabolisms59. The strain IBSD35 might generate cell energy-adenosine triphosphate (ATP) and reducing agents. The reaction equation of ATP in the cytosol of P. peoriae IBSD35 predicted from the ModelSEED is37,39
Genome-scale phylogeny analysis
Paenibacillus peoriae IBSD35 had a digital DNA-DNA hybridization (dDDH) of Genome-To-Genome Distance Calculator (GGDC) formula 2 (d4) 76.1, confidence intervals (C.I.) d4 of [73.1–78.9], and a difference G + C% of 0.16 with P. peoriae HS311 indicating their close species relation. P. peoriae IBSD35 has a dDDH of d4 86.2, C.I. d4 of (83.6–88.5), and a difference G + C% of 0.00 with P. peoriae FSL J3-0120 which indicated their very close relationship. Additionally, P. polymyxa CR1 and P. peoriae IBSD35 exhibited a close relationship with d4 75.6, C.I. [72.6–78.4] and a difference G + C% of 0.02 (Fig. 6C). The strain IBSD35 exhibited OrthoANI values of 96.37 with P. peoriae FSL H8-0551which is above the species boundary value of average nucleotide identity (ANI) = 95–96%. The result indicated that the G + C% difference computed from the query and subject genome sequences varied by no more than 1% and proved they are within species.
Commensal lifestyle of P. peoriae IBSD35
The genome was predicted to harbor sugar-specific component phenotypes, thus possibly utilizing the host plant trehalose, l-arabinose, sucrose, and maltose. It harbored Tat A (twin-arginine translocation) and Sec (general secretory pathway) protein secretion systems for transportation across the cytoplasmic membrane60,61. Most secretion systems genes related to virulence and pathogenicity, such as types I, II, III, V, and VI secretion systems62, were absent in the P. peoriae IBSD35 genome, which supported its non-pathogenicity characters while handling.
Nitrogen fixation characters of P. peoriae IBSD35
It harbored putative genes encoding Mo-nitrogenase MoFe protein (EC 1.18.6.1), multi-functional system (MFS) transporter, nitrate/nitrite transporter, and ammonium transporter involved in the uptake of nitrate and nitrite (Supplementary Figs. S5, S6). The nitrogen fixation genes include narG, narH, narI, narJ, narW, nirA, nirB, nasA, hcp, gdhA, iscU, nifB, nifD, nifE, nifK, nifN, nifE, nifH, nifV, nifX, nifZ) and other putative nitrogen-fixing proteins plus a hypothetical protein that lacks an assigned function were identified using the Kyoto Encyclopedia of Genes and Genomes (KEGG) pathway database analysis which indicated its nitrogen-fixation nature (Table S5) (www.kegg.jp/kegg/kegg1.html). However, the nitrogen fixation regulator (nifL/A) was not visible in the nitrogen metabolism pathway63. P. peoriae IBSD35 might absorb some of the nitrate sources from the host plant.
Plant-associated endophyte characters of P. peoriae IBSD35
The proteins involved in the flagellum assembly gene ontology (GO): 0044780, flagellum-dependent cell motility (GO: 0071973), and chemotaxis (GO: 0006935) identified from in silico genome analysis of P. peoriae IBSD35 confirmed the bacterium motility traits. It harbored TCS CheA/CheY, methyl-accepting chemoreceptor proteins (mcp, K03406), an adaptor protein CheW (K03408), and flagellar motor switch protein FliM (K02416). They are the core of bacterial chemotaxis signaling genes. They are known to involve in colonization and interaction with host plants29,46,48,49. The analysis also revealed cellular motility proteins for the resources transport and substrate-binding protein of ribose (rbsB, K10439). These traits are required for mobility and thriving in heterogeneous and highly competitive environments. Endoglucanase [EC: 3.2.1.4] enzyme was visible in the understudy genome, and it is necessary for spreading to other intracellular tissues to enter the endophytic life stage (www.kegg.jp/kegg/kegg1.html).
Discussion
Antibiotics have revolutionized the treatment of tropical infections, communicable diseases, food-borne, and other poverty-related infectious diseases. However, the efficacy is short-lived and turned into a resilient clinical problem eliciting innumerable public health crises and causing economic burden1,2,4. Therefore, there is a continuous need for a new antimicrobial agent with high specificity, efficacy, efficiency, and anti-resistant to tackle the AMR challenge2,64. AMPs have prospects because of their antibacterial, anticancer, antiviral, antifungal, anti-aging, and anti-parasite, regulating cell proliferation, extracellular matrix production, cellular immune responses, and adjuvant characteristics15,16,65. Lately, many important SMs have reported from endophytes of medicinal plants from less explored sources which offered unprecedented opportunities to find novel antimicrobial agents5,6. Moreover, the availability of ever-increasing genome sequences, a comprehensive proteomics database, and advanced bioassay techniques offer a platform for the dereplication of old compounds, effective search, prediction, and design of novel AMP to counter AMR19,20.
Paenibacillus peoriae IBSD35 from the stock solution has retained antimicrobial activity against S. aureus ATCC 25923, indicating its inherent antimicrobial potential66. Endospore forming traits may help its endophytic association with the host plant67. Its neutralophilic characters might help neutralize its cytoplasm relative to the plant cell to meet the pH challenges. It might be through direct active uptake or efflux of protons from the cells or tissues29. These indirectly suggested host-plant physiology as well. The optimum growth condition of the strain IBSD35 recorded at 38°, and pH 6.8–7 in LB medium indicated the mesophilic traits. The cultivable characters confirmed P. peoriae IBSD35’s ability to live as a free-living bacterium in laboratory conditions.
An antimicrobial protein mixture precipitated from LB fermentation broth with 70% ammonium sulfate suggests that at least one or more of the putative 25 BGCs of P. peoriae IBSD35 might express antimicrobial metabolites. The 700 mM NaCl eluent from DEAE-C column chromatography has retained activity against S. aureus ATCC 25923. The RP-HPLC peaks P4 and P5, absorption at λ205 and λ214, suggested the presence of amino acids and peptide bonds with other impurities. The absorbance at λ280 confirmed the presence of proteins68,69. Antimicrobial protein total yield was low at 0.05%, which can be enhanced using an advanced purification system or by up-regulating its BGC expression. MIC of peoriaerin IBSD35 at 0.04 µg µl−1 indicating its high specific activity and potency against S. aureus ATCC 25923 compared with ampicillin and nisin at 1 mg ml−1.
The purified AMP has shown antimicrobial activity against S. aureus ATCC 25923, E. coli ATCC 25922, C. albicans ATCC 10231, K. pneumoniae ATCC 4352, and S. typhimurium ATCC 14028 connoted its broad-spectrum antimicrobial character (Tables S6, S7). The sample has retained activity after prolonged storage, and its resistance to degradative enzymes indicated its potential for drug development against AMR. Antimicrobial activity was altering at different stages of purification. It might be due to the disintegration of proteins. The treatment of SDS and high-temperature slightly increased the activity. It might be from the disintegration or aggregation of the proteins and other biomolecule complexes (Table S3). The sample can withstand autoclaving at 121 °C for 15 min presenting a potential antimicrobial agent and food preservatives17,21,70.
The peptide sequences generated using LC–MS/MS coupled to a quadrupole time-of-flight (Q-TOF) had aligned with P. peoriae IBSD35 proteome using proteomics tools21,22,24,33,38. The sequence alignment using the Antimicrobial Peptide Database (APD3) tool indicated that peoriaerin IBSD35 is 38.23% similar to fermentin from Lactobacillus fermentum20,25. Peoriaerin IBSD35 is a trans-membrane helix with an extended strand of 10.71%, beta-turn of 21.43%, and a random coil of 64.00% (Fig. 3, Supplementary Fig. S2). It has a high percentage of helix and residues embedded in the membrane, suggesting at least some part of the sequence may embed in the hydrophobic region of the membrane20. Peoriaerin IBSD35 has a total net charge of + 2 and a theoretical pI of 8.86. These properties indicated an antimicrobial peptide nature. The aliphatic index was 56.07, and the grand average of the hydropathicity index (GRAVY) was − 0.13225. Antimicrobial action was visible after 1 h of incubation. The slower kinetics of killing suggested that peoriaerin IBSD35 might penetrate the cell membrane and transport it inside the cell cytoplasm to cause the effect18. However, the leakage of DNA and proteins was lacking.
The draft genome sequencing and analysis of P. peoriae IBSD35 provided insights into its endophytic characters, antimicrobial traits, and interaction with the host plant10,11,13. The characteristics contributing to strain IBSD35 epiphytic fitness are iron acquisition, capacity to colonize plants, and stress-tolerant. Additional functional genes suggested P. peoriae IBSD35’s close association with the environment through nitrogen fixation14,37,40. However, there are about 980 (18.50%) protein-coding genes without assigning any function, as per IMG AP v.4.16.6 (Table S4), and their relationship to molecular networks of the database is unknown28,29,49. Possibly, they may be endophytic specific, which needs further study to ascertain.
The genes encoding flagellins and flagellar motor proteins are visible in P. peoriae IBSD35. They helped bacterial motility in the plant-associated lifestyle28,46,71. The flagellin proteins (Flg) also facilitated the endophyte to pass through the first line of the plant immune defense system. Moreover, the presence of flagellin (fliC), glycerol kinase [EC: 2.7.1.30], elongation factor (Tu), and molecular chaperone (htpG) are crucial for plant-pathogen interaction indicating P. peoriae IBSD35 close association with the host plant72. Generally, motility enabled bacteria to move and colonize plants and systematically spread within the plant, promoting the endophytic lifestyle. The TCS genes reinforced traits needed for thriving in heterogeneous, highly competitive, and adverse environments44,73. The proteins or SMs might transport from the bacterial cytoplasm through its Tat and Sec secretion systems61,74.
The TCS represented a primary means by which an organism sensed and responded to stress and changing physiological environments45,71. The putative genes encoding small acid-soluble proteins (SASPs) annotated in P. peoriae IBSD35 generally helped the bacterial endospore DNA against heat, UV-radiation, and enzymic degradation45. They are responsible for sporulation septum, engulfment, spore morphogenesis, and germination in harsh environments. The genome ATP-binding cassette (ABC) transporter proteins are other plausible candidates for genes mediating the export of small molecules across the cells44. The presence of siderophores and genes conferring tolerance to oxidative stress in the strain IBSD35 genome supported the proposed importance of oxidative stress tolerance to fitness in symbiotic association. It is consistent with P. peoriae IBSD35 endophytic lifestyle60,61. However, there was no evidence of the biosynthesis of phytotoxins or cellulases.
The KEGG pathway database analysis (www.kegg.jp/kegg/kegg1.html) indicated that carbohydrate and amino acid metabolism accounted for a large proportion of the P. peoriae IBSD35 genome44,45,50, and its CCM network analysis included GG, TCA, PPP metabolites, and pyruvate metabolism57. Most organisms have shared a common core to their metabolic networks58,59. The CCM pathways convert sugars into metabolic precursors and provide the entire cell biomass, including the SMs57. P. peoriae IBSD35 harbored a collection of metabolite and drug efflux systems and bioremediation gene clusters which indicated a need to protect against toxic concentrations of metabolites or metabolic analogs54. It has shown the strain’s capability for bioremediation, survival defense against oxidative stress, and production of antimicrobial compounds and toxins. A repertoire of NRPS (72%) and their expressions might neutralize the competing microbes and antagonistic biomolecules. Gene resistance to β-lactam, cationic AMPs, and antifolate with the antibiotic BGCs in the strain IBSD35 genome supported that the clusters might involve in antibiotic production, which is vital to avoid self-toxicity from its own antibiotics production54,55.
The SMs gene clusters predicted from the P. peoriae IBSD35 genome showed low similarity with other BGCs. It exhibited 100% similarity for fusaricidin and polymyxin BGCs in the MIBiG cluster database (Table 4)50,51,75. A repertoire of NRPS SMs might help microbes in existence and communication, and their elucidation will facilitate the finding of novel bio-molecules52. They might also provide a competitive advantage in the battle for resources and adaptive mechanisms for survival in host plant cells or tissues52,53,55. The different number of BGCs in the analysis of the genomes reflected their habitats, genetics, and functional differences. Thus, a difference in genomes’ SM is a difference in environment and resource competition55. Generally, two closely related strains share most of their genes and will not be unique to the individual species76,77. The in-silico identification of PKS, NRPS, or SM gene clusters revealed considerable differences between the closely related species supporting that P. peoriae IBSD35 is a unique species slightly different from free-living or symbiotic rhizobium bacteria.
The terpenoids and polyketides biosynthetic are the backbones of other SMs78. Pathwhiz has been used to illustrate the type II polyketide biosynthesis pathway of the P. peoriae IBSD35 genome predicting its proteins, enzymes, SMs, and other products79. Dihydrokalafungin, oxytetracycline, tetracycline, nogalavinone, elloramycin, and tetranomycin F are predicted to be derived from simple carbon building blocks of the pathway (Supplementary Fig. S7). The KEGG pathway analysis (www.kegg.jp/kegg/kegg1.html) indicated that terpenoids have derived from 2-C-methyl-d-erythritol 4-phosphate/1-deoxy-d-xylulose 5-phosphate (MEP/DOXP) pathway in P. peoriae IBSD35. The non-mevalonate Pathway of strain IBSD35 has the potential to produce isoprenoids, antibiotics, and anticancer alkaloids80. However, the expressions of SM clusters are typically under environmental and developmental control mediated by complex regulatory cascades that relay signals to the pathway-specific switches53.
The P. peoriae IBSD35 genome harbored the putative genes of the nitrogen cycle. Nitrogen fixation by nitrate to ammonia and other biologically reduced forms might incorporate into amino acids and other vital compounds through nitrogen metabolism40. Nitrogen fixation is energetically costly, and therefore, it is tightly regulated at the transcriptional level by sophisticated regulatory networks that respond to multiple environmental cues40,41,63. However, the nitrogen fixation regulator (nifL–nifA) was not visible, but it was difficult to discern how oxygen levels affect the regulation of nitrogen fixation in our present study (www.kegg.jp/kegg/kegg1.html). However, it is clear that the nitrogen fixation is different from those previously described for other γ-proteobacteria and possibly depends on the host plant for its nitrogen source63. It is one of the unique characteristics of P. peoriae IBSD35 endophytic traits.
The Type (Strain) Genome Server annotation (TYGS) and PGAP annotation predicted that the strain IBSD35 has a close species relation with other P. peoriae strains. Phylogenetically, P. peoriae IBSD35 is closely related to P. peoriae FSL J3-0120. However, the protein Basic Local Alignment Search Tool (BLAST) indicated P. peoriae IBSD35 close similarity with P. polymyxa SC2 (E-value: 0.0, Score: 2273, Ident.: 98.9%). Therefore, the understudy genome is within the species of P. peoriae and related to the P. polymyxa species. The genomic data quality and analysis need further improvement as some of the selected genomes are draft and limited for web server analysis. The variation in genome size suggested that different strains of the same species from different habitats potentially produced their own unique NRPS products. Several gene clusters are common in selected Paenibacillus species suggesting their conservation in the lineage49. Paenibacillus has only been designated a separate species since 199312,13. Some lineage-specific clusters and the predicted unique clusters in our understudy genome could be involved in SMs biosynthesis12.
The unique genes or 2125 single-copy genes from the six Paenibacillus genomes might encode some of the functions in the cell, as they are found only in one organism. These genes might be involved in environmental adaptation, specific competition with other species, or speciation55,76. The synteny dot plot indicated P. peoriae IBSD35 resemblance to its genus. There is also evidence of gene sharing (Mitogen-activated protein kinase (MAPK) pathway from plant, nitrogen fixation gene cluster) and horizontal transfer of genes.
Given the P. peoriae IBSD35 potential applications in agriculture, medicine, industries, and bio-remedy14, its genome sequencing and its extracellular metabolite analysis are crucial to the basic and applied research to find novel antimicrobial agents. Genome mining has enabled the prediction of the putative BGC region in the P. peoriae IBSD35 by linking a protein group from LC–MS/MS data and connecting these natural products to the genes24. The genome information and the proteomics analysis provided here will further enhance the study of environmental information processing and its antimicrobial pathways55,64,65,81,82. It will also facilitate the understanding of evolutionary relationships among Paenibacillus species. Moreover, the less explored ecological niches provide new sources of an unprecedented opportunity to find novel microbes and metabolites to counter AMR to our already exhausted sources of bio-molecules.
Conclusions
Paenibacillus peoriae IBSD35 is an endophytic bacterium. It is a mesophile, endospore forming, and neutralophile bacterium. It has a linear DNA topology with a total length of 5,862,582 bp. It harbored 25 SMs BGCs, out of which 18 BGCs are NRPS. The P. peoriae IBSD35 closest species reference genome is P. peoriae KCTC 3763 with a symmetrical identity of 62.7495% and a symmetrical identity of 89.4884% with P. peoriae FSL J3-0120. The CFS of P. peoriae IBSD35 exhibited a broad spectrum of antimicrobial activity against S. aureus ATCC 25923, E. coli ATCC 25922, C. albicans ATCC 10231, K. pneumoniae ATCC 4352, and S. typhimurium ATCC 14028.
P. peoriae IBSD35 inoculum's optimum growth pH was observed at 6.8–7. It is a mesophile bacterium that grows best at 38 °C. The supernatant antimicrobial activity arbitrary unit was 1600 AU ml−1. The putative extracellular AMPs were purified using salting out and chromatography. A 15 proteins list and AMP sequences have predicted from LC–MS coupled to Q-TOF. An AMP from the protein lists with accession no. A0A2S6NUT6 was selected. It has been given the name peoriaerin IBSD35, based on its source name. The amino acid sequence of peoriaerin IBSD35 is AAGIQAQAGFGLSDSIQGTGKQKCSFCK with a MW of 2802.17 Da. Its sequence backbone has a large portion of helix and residues embedded in the membrane. It exhibited high antimicrobial activity against S. aureus ATCC 25923 at range of 0.04 µg µl−1.
Paenibacillus peoriae IBSD35 exhibited a close phylogenetic relationship with P. peoriae FSL J3-0120 and P. polymyxa CR1. The genome analysis indicated that it harbored nitrogen fixation genes and other putative nitrogen-fixing proteins but without a nitrogen fixation regulator (nifL/A). Its genome has a repertoire of 18 NRPS BGCs. The type II polyketide biosynthesis pathway revealed that terpenoids might be exclusively through (MEP/DOXP) pathway in P. peoriae IBSD35. Combining the genomics, mass spectrometry, and proteomics results obtained from P. peoriae IBSD35 through its extracellular biomolecules will enhance our search for AMPs that are stable, specific, and effective to counter AMR. The potential for applying comparative genomics to get evolutionary insights and discover novel AMPs from less explored regions will increase the probability of finding novel bio-molecules.
Materials and methods
Plant ethics statements
Millettia pachycarpa plant was used to isolate an endophyte, Paenibacillus peoriae IBSD35 which is used in this study. It is a commonly found wild species in Ukhrul, Manipur, India. Experimental research and field studies on plants including the collection of plant is complied with National guidelines and legislation. Millettia pachycarpa is not included in the list of IUCN Species at Risk of Extinction.
The stem of Millettia pachycarpa plant was collected from Phalee village (25.143524 N & 94.28334 E, 1533.14 M), Ukhrul, Manipur, India. Permissions to collect plant material were obtained from Phalee BMC by payment of fee as per NBA guidelines for collecting plant from area falls within the territorial jurisdiction under sub-section (3) of section 41 of the Act (Biological Diversity Act, 2002).
P. peoriae IBSD35 culture and its morphology and genomic characters
An endophytic bacterium P. peoriae IBSD35 isolated from the stem of M. pachycarpa in our previous experiment was picked in our present context7,8,9,66. It was sub-cultured from 15% phosphate buffer saline (PBS) glycerol stock preserved in the Institute of Bioresources and Sustainable Development-Microbial Resources Division (IBSD-MRC) repository (Accession No. MRC-75001) by thawing for 5 min at room temperature (27 °C)83. A 100 µl of stock culture was pipetted using a micropipette and spread on a 25 ml LB agar plate. It was incubated at 38 °C to check the contamination and colony purity84. A single colony was streaked and sub-cultured repeatedly to maintain its growth curve character, colony purity, viability, and productivity83,85. The cut-well agar diffusion antimicrobial assay against S. aureus ATCC 25923 was performed86. Chemicals were added to the inoculum to avoid bacterial contamination in the fermentation system from instruments, flasks, nutrients, and pipetting.
The strain IBSD35 pure colony was smeared on a glass slide and observed under the phase-contrast microscope (Zeiss Imager. Z2)85. The gDNA was extracted from an overnight grown culture using the cetyl trimethyl ammonium bromide (CTAB) method40. The quality and quantity of the gDNA were determined by A260/A280 ratio using UV-spectrophotometer (Nanodrop, Shimadzu Biotech Bio Spec-nano) and assessed in a 0.8% agarose gel electrophoresis. The genomic and molecular characters of the genome were analyzed by draft genome sequencing using Illumina HiSeq 2500 platform87.
Screening of antimicrobial activity
A 200 ml of P. peoriae IBSD35 inoculums was harvested by centrifugation (Sorvall ST 16R Centrifuge) at 3200×g for 15 min at 4 °C, and the supernatant was collected in an Eppendorf tube88. The spectrum of antimicrobial activity was tested using cut-well agar diffusion bioassay against the representative Gram-positive, Gram-negative, and fungal test pathogens (S. aureus ATCC 25923, E. coli ATCC 25922, C. albicans ATCC 10231) with a slight modification of Clinical and Laboratory Standard Institute (CSLI) methods89,90,91. The test pathogen lawn was prepared by spreading 20 µl of pathogen inoculums (0.5 Macfarland standard) over the Muller Hilton Agar (MHA) plate surface. A diameter of 3–5 mm was punched on the MHA surface with a sterile tip, and the wells were loaded with the test solution (20–100 µl) at desired concentration86. The MHA plates were incubated at 37 °C overnight as per the test pathogens’ best growth condition. The activity zones of inhibition diameter were measured to quantify the antimicrobial activity.
Optimum growth pH and temperature for antimicrobial production
2 l of the production medium (pH 7 ± 2) was inoculated with an overnight grown culture of P. peoriae IBSD35 and incubated in a shaker incubator at 80 rpm (Eppendorf innova R42 Rotary shaker). Its optimum pH, OD, and temperature concerning antimicrobial activity were determined by aseptically aspirating out the inoculum from the fermentation broth66. The production medium was harvested by centrifugation at 3200×g for 15 min at 4 °C. The supernatant antimicrobial activity was tested by serial dilution, and its arbitrary unit was calculated against S. aureus ATCC 2592331. To rule out any false-positive inhibition caused by acid production and catalytic enzyme, 0.6% of calcium carbonate and a unit (10 mg/ml) of 0.01% catalase were added to the production medium84. Toluene was added to avoid contamination from other microbes88.
Precipitation and purification of the antimicrobial protein mixture
The 6-day-old inoculum of 2 l of CFS was harvested from the 6-day-old inoculums by centrifugation at 3200×g for 15 min at 4 °C. Its pH was adjusted to 3 with 2 N hydrochloric acids (HCl) to kill the experimental strain and re-adjusted to the original harvesting state (pH 6.8) after 4 h by adding 2 N sodium hydroxide (NaOH)88. The supernatant pH was re-adjusted to the original harvesting state (pH 6.8) by adding 2 N NaOH. The CFS was filtered through a 0.2 µm filter (Avixa) and precipitated in 70% ammonium sulfate (NH4)2SO4) followed by centrifugation (3200×g, 35 min at 4 °C)92. The precipitated protein from centrifugation was collected. The resulting pellet was re-suspended in 30 ml distilled water and dialyzed using PBS in a dialysis membrane (12.4 kDa cut-off) with intermittent buffer changing36.
The pellet was re-dissolved in 30 ml of distilled water and loaded in diethylaminoethyl cellulose (DEAE-C (Sigma Aldrich). A weakly positive resin, DEAE-cellulose was loaded for anion exchange column chromatography93. Sample was eluted out using 0.1, 0.2, 0.3, 0.4, 0.5 and 0.7 M or 100–700 mM NaCl gradient elution and the OD were recorded at selected wavelength of λ205, λ214, and λ28015,16. The fractions were pooled together and lyophilized into powder form and tested for retention of antimicrobial activity according to the CLSI assay89,90. Fractions showing antimicrobial activity were pooled together for lyophilization and further purified with RP-HPLC94.
RP-HPLC purification of protein mixture
The partially purified sample was again purified using ZORBAX 300 SB-C-18 semi-preparative (9.4 × 250 mm) 5 microns91,94. The column was irrigated with 0.1% TFA in 20% ACN for 45 min at an isocratic flow rate. The sample (150 µl per experiment) was loaded on the HPLC (UFLC Shimazdu) with semi-preparative Agilent ZORBAX 300SB reverse-phase C-18 of 5 µm and 9.4 × 250 mm column and run with 0.1% ion-pairing reagent, trifluoroacetic acid (TFA)68,94. The RP-HPLC mobile phase consisted of (i) HPLC-grade water containing 0.1% TFA and (ii) 20% acetonitrile (ACN) in an isocratic system at a flow rate of 2.5 ml min−193. Eluents were monitored at a wavelength of λ205, λ214, and λ280 using a UV-spectrometer (Eppendorf, Biospectrometer) and collected manually. The fractions which exhibited higher antimicrobial activity were pooled together and collected separately for further analysis. The total protein concentration and activity, specific activity, yield percentage, and purification fold of the extract compared to the initial starting crude extract were calculated using UV-Spectroscopy15,16. The UV-spectrophotometric method for antimicrobial protein concentration was calculated as: concentration (mg/ml) = (1.55 × A280) (0.76 × A260) × dilution factor69.
The effect of external factors on sample antimicrobial activity
The sample was evaluated for sensitivity to degradative enzymes, detergents, organic solvents, temperature, and pH changes78,84. The retention of antimicrobial activity after the treatment was qualitatively determined through cut-well agar diffusion bioassay against the reference pathogen, S. aureus ATCC 25923 as described earlier91. For the pH stability test, the sample dissolved in dist. water was pre-adjusted to pH 2.0–8.9, followed by incubation at 38 °C for 12 h. The sample temperature sensitivity was tested by exposing to temperatures ranging from 20 to 100 °C at the interval of 10 °C for 2–3 h, and additionally at 121 °C for 15 min by autoclaving86,91.
The samples were treated with different degradative enzymes at the concentration of 1 mg ml−1 for 2–3 h at room temperature to test the stability against enzymes84. The organic solvent and test sample were mixed in a 1:1 ratio and incubated at 27 °C to evaporate all the organic solvent, and the samples were assessed for retention of antimicrobial activity. The samples were treated with detergents in a 1:1 ratio and checked for retention of antimicrobial activity86,90. Negative control consists of PBS (pH 7) and the untreated sample solution was used as a positive control.
Mass spectrometric analysis of peptide mixture
The experiment was performed using EASY-nLC 1000 system (Thermo Fisher Scientific) coupled to a Q-Exactive mass spectrometer (Thermo Fisher Scientific) equipped with a nano-electrospray ion source23,24. 1.0 µg of the RP-HPLC purified peptide mixture was resolved using a 15 cm PicoFrit column (360 µm outer diameter, 75 µm inner diameter, 10 µm tip) filled with 1.9 µm of C18-resin (Dr. Maeisch, Germany). The peptides were loaded with buffer A and eluted with a 0–40% gradient of buffer B (95% ACN, 0.1% formic acid) at a flow rate of 300 nl/min for 90 min24. The source heater was set at 300 °C to turn the sample into a vapor for analysis. The UPLC-separated samples were then analyzed by electrospray ionization at 4.5 kV and the peptides were dissociated with collision induced dissociation (CID) for MS/MS spectra95. MS data was acquired using a data-dependent top10 method dynamically choosing the most abundant precursor ions from the survey scan.
One sample was processed and 1 RAW file generated was analyzed with a Proteome Discoverer against P. peoriae IBSD35 Uniprot reference proteome database33. For the Sequest search, the precursor and fragment mass tolerances were set at 10 ppm and 0.5 Da, respectively. Trypsin was used to generate peptides, carbamidomethyl on cysteine as fixed modification, and oxidation of methionine, and N-terminal acetylation was considered as variable modifications for database search. Both peptide spectrum match (PSM) and protein false discovery rate (FDR) were set to 0.01 FDR.
Physiochemical properties of the protein and AMPs were analyzed using the ExPASy-ProtParam tool (http://web.expasy.org/protparam) and APD3 Antimicrobial Peptide Calculator23,25. The secondary structure was analyzed using SOPMA96. The 3-D conformation structure of the peptide was predicted using PEP-FOLD 3 de novo modeling and deposited in ModelArchive26,27. The model was built based on the target-template alignment using ProMod397.
Peoraerin IBSD35 mode of action imaging through scanning electron microscopy
The AMP mode of action against the test pathogens was checked by the absorbance at λ260 and λ280 with UV-spectrometry. A direct visualization of peptide-induced S. aureus cellular damage and morphological changes were evaluated by SEM98. The samples were prepared by incubating S. aureus ATTC 25923 with the test sample as described earlier89,90. S. aureus were treated with antimicrobial agent at 1× MIC for 2–3 h and incubated at 30 °C ± 0.2. The inoculants were harvested by centrifugation as described earlier87. The pellet was washed with 0.1 M PBS (pH 7.2) and fixed in 2.5% glutaraldehyde and 2% paraformaldehyde made in PBS97. The fixed sample was further processed for coating and imaging99. The test pathogen without any treatment served as the experimental control.
Genome sequencing and assembly
The gDNA integrity was confirmed using a 0.8% agarose gel electrophoresis97. The draft genome sequencing was performed through Illumina HiSeq 2500 system34. The cleaned reads were subjected to the Kmer genie to predict the optimal k value and assembly size100. De novo assembly was performed using ABySS v.2.042. The predicted genes were matched with bacterial proteins from the Uniprot database using the BLASTX program101. Genes that could establish homology relationships (E-value ≤ 10−5 and similarity score ≥ 40%) were retained in the annotation pipeline for further annotation while the others remain unannotated28,29,30.
Genome annotation and bioinformatics analysis
JGI-IMG Annotation Pipeline v.4.16.628,29 and the NCBI PGAP were used for gene annotation30. The gene functional prediction was performed by BLAST against the databases101.
Comparative genomic analysis
The close strains of P. peoriae IBSD35 (NZ_PTJM01000000)7,66 were selected from the NCBI database (P. peoriae KCTC 3763102, P. polymyxa E681(NZ_CP048793)75, P. polymyxa SC2 (NC_014622)103,104, P. vortex V453 (NZ_ADHJ01000000)105, P. terrae HPL-003(NC_016641)106, Paenibacillus sp. JDR-2 (NC_012914)107, P. polymyxa CR1(NC_023037)108, B. subtilis subtilis AG1839 (NZ_CP008698)109, S. coelicolor A3(2) (AL645882)110, and P. peoriae HS311 (NZ_CP011512)). Bacillus subtilis subtilis AG1839 is selected because Paenibacillus represents a new genus from Bacillus12,13. S. coelicolor A3(2) is the model organism for SM BGC studies. P. peoriae HS311 represents the completely sequenced genomes of Paenibacillus and serves as the reference for the natural variation within the genus. P. peoriae KCTC 3763 represents the closest species reference genome with a symmetrical identity of 62.7495%.
OrthoVenn was used for Venn diagram drawing which is based on the protein sequences clusters (E-value = 1e−2 and inflation value = 1.5)76. Web Gene Ontology Annotation Plot (WEGO) was used to compare the provided gene datasets and plot the distribution of GO annotation into a histogram38. SMs biosynthesis gene clusters were analyzed with AntiSMASH V.5 and JGI-IMG/ABC, and the type II polyketide biosynthesis pathway was drawn using KEGG and PathWhiz tool43,50,51,79. All comparative analyses were performed using IMG/M and RAST tools29,37,39.
Genome-based taxonomy and phylogenetic tree
Genome-based taxonomy was determined using TYGS which infers genome-scale phylogenies and estimates for species and subspecies boundaries and automatically determined closest type genome sequences111,112. The pair-wise digital DNA:DNA hybridization (dDDH) values between user genomes and the selected types-strain genomes were performed77. The dDDH values are provided along with their confidence intervals (C.I.) and difference of G + C% using the Genome BLAST Distance Phylogeny (GBDP) formula d577,78. The average nucleotide identity with the closely related species was determined using the OAT tool113. The phylogenetic tree was inferred with FastME 2.1.6.1 from GBDP distances calculated from genome sequences114. The AMP model was built based on the target-template alignment using ProMod397.
Data availability
The datasets generated or analyzed during the current study are given in the table form with the name, accession number, and the database or web link.
Name | Accession number | Database/web link |
|---|---|---|
Paenibacillus peoriae IBSD35 | BioProject: PRJNA224116 | NCBI |
Unknown protein from LC–MS experiment | A0A2S6NUT6 | Uniprot |
Peoriaerin IBSD35 | ma-id7l4 (DOI 10.5452ma-id7l4) | ModelArchive |
Nitrogen metabolism (map00910) | Ref: 210413 | |
Biosynthesis of type II polyketide backbone | PW122264 |
References
Fischbach, M. A. & Walsh, C. T. Antibiotics for emerging pathogens. Science 325, 1089–1093. https://doi.org/10.1126/science.1176667 (2009).
World Health Organization. Antimicrobial Resistance: Global Report on Surveillance (World Health Organization, 2014).
Gougoulias, C., Clark, J. M. & Shaw, L. J. The role of soil microbes in the global carbon cycle: Tracking the below-ground microbial processing of plant-derived carbon for manipulating carbon dynamics in agricultural systems. J. Sci. Food Agric. 94, 2362–2371. https://doi.org/10.1002/jsfa.6577 (2014).
European Centre for Disease Prevention and Control. Surveillance of Antimicrobial Resistance in Europe 2018 (ECDC, 2019).
Castillo, U. F. et al. Munumbicins, wide spectrum antibiotics produced by Streptomyces munumbi, endophytic on Kennedia nigriscans. Microbiology 148, 2675–2685. https://doi.org/10.1099/00221287-148-9-2675 (2002).
Chen, Y. T. et al. Antitumor activity of bacterial exopolysaccharides from the endophyte Bacillus amyloliquefaciens sp. isolated from Ophiopogon japonicus. Oncol. Lett. 5, 1787–1792. https://doi.org/10.3892/ol.2013.1284 (2013).
Rosenblueth, M. & Martínez-Romero, E. Bacterial endophytes and their interactions with hosts. Mol. Plant Microbe Interact. 19, 827–837. https://doi.org/10.1094/MPMI-19-0827 (2006).
Reinhold-Hurek, B. & Hurek, T. Living inside plants: Bacterial endophytes. Curr. Opin. Plant Biol. 14, 435–443. https://doi.org/10.1016/j.pbi.2011.04.004 (2011).
Stierle, A. & Strobel, G. A. Taxol and taxane production by Taxomyces andreanae, an endophytic fungus of Pacific yew. Science 260, 214–216. https://doi.org/10.1126/science.8097061 (1993).
Heyndrickx, M. et al. A polyphasic reassessment of the genus Paenibacillus, reclassification of Bacillus lautus (Nakamura 1984) as Paenibacillus lautus comb. nov. and of Bacillus peoriae (Montefusco et al. 1993) as Paenibacillus peoriae comb. nov., and emended descriptions of P. lautus and of P. peoriae. Int. J. Syst. Bacteriol. 46, 988–1003. https://doi.org/10.1099/00207713-46-4-988 (1996).
Grady, E. N., MacDonald, J., Liu, L., Richman, A. & Yuan, Z. C. Current knowledge and perspectives of Paenibacillus: A review. Microb. Cell Fact. 15, 203. https://doi.org/10.1186/s12934-016-0603-7 (2016).
Ngashangva, N., Indira, S. D. & Kalita, M. C. Screening of endophytes from traditionally used medicinal plants of Manipur for their antimicrobial activity: An impact towards future drug discovery. Int. J. Sci. Res. Biol. Sci. 6, 39–47. https://doi.org/10.26438/ijsrbs/v6i5.3947 (2019).
Singhal, A. K., Sharma, R. P., Baruah, J. N., Govindan, S. V. & Herz, W. Rotenoids from roots of Millettia pachycarpa. Phytochemistry 21, 949–951. https://doi.org/10.1016/0031-9422(82)80103-9 (1982).
Srivastava, R. C. Traditional knowledge of Nyishi (Dafla) tribe of Arunachal Pradesh. Indian J. Trad. Knowl. 9, 26–37 (2010).
Hancock, R. E. W. & Chapple, D. S. Peptides antibiotics. Antimicrob. Agents Chemother. 43, 1317–1323. https://doi.org/10.1128/AAC.43.6.1317 (1999).
Habets, M. G. J. L. & Brockhurst, M. A. Therapeutic antimicrobial peptides may compromise natural immunity. Evol. Biol. R. Socy. 8, 416–418. https://doi.org/10.1098/rsbl.2011.1203 (2012).
Baindara, P. et al. Characterization of the antimicrobial peptide penisin, a class Ia novel lantibiotic from Paenibacillus sp. strain A3. Antimicrob. Agents Chemother. 60, 580–591. https://doi.org/10.1128/AAC.01813-15 (2015).
Epand, R. M. & Vogel, H. J. The diversity of antimicrobial peptides and their mechanisms of action. Biochim. Biophys. Acta 1462, 11–28. https://doi.org/10.1016/s0005-2736(99)00198-4 (1999).
Wang, G., Liand, X. & Wang, Z. APD3: The antimicrobial peptide database as a tool for research and education. Nucleic Acids Res. 44, D1087–D1093. https://doi.org/10.1093/nar/gkv1278 (2016).
Pavlova, A. S. et al. Identification of antimicrobial peptides from novel Lactobacillus fermentum strain. Protein J. 39, 73–84. https://doi.org/10.1007/s10930-019-09879-8 (2020).
Perkins, D. N., Pappin, D. J., Creasy, D. M. & Cottrell, J. S. Probability-based protein identification by searching sequence databases using mass spectrometry data. Electrophoresis 20, 3551–3567 (1999).
Ma, B. et al. PEAKS: powerful software for MS/MS peptide de novo sequencing. Rapid Commun. Mass Spectrom. 20, 2337–2342. https://doi.org/10.1002/rcm.1196 (2003).
Seidler, J., Zinn, N., Boehm, M. E. & Lehmann, W. D. De novo sequencing of peptides by MS/MS. Proteomics 10, 634–649. https://doi.org/10.1002/pmic.200900459 (2010).
Cimermancic, P. et al. Insights into secondary metabolism from a global analysis of prokaryotic biosynthetic gene clusters. Cell 158, 412–421. https://doi.org/10.1016/j.cell.2014.06.034 (2014).
Gasteiger, E. et al. Protein identification and analysis tools on the ExPASy Server. In The Proteomics Protocols Handbook (ed. Walker, J. M.) (Humana Press, 2005).
Lamiable, A. et al. PEP-FOLD3: Faster de novo structure prediction for linear peptides in solution and in complex. Nucleic Acids Res. 44, W449–W454. https://doi.org/10.1093/nar/gkw329 (2016).
Schwede, T. et al. Outcome of a workshop on applications of protein models in biomedical research. Structure 17, 151–159. https://doi.org/10.1016/j.str.2008.12.014 (2009).
Mukherjee, S. et al. Genomes OnLine Database (GOLD) vol 8: Overview and updates. Nucleic Acids Res. 49, D723–D733. https://doi.org/10.1093/nar/gkaa983 (2021).
Markowitz, V. M. et al. The integrated microbial genomes (IMG) system in 2007: Data content and analysis tool extensions. Nucleic Acids Res. 36, 846. https://doi.org/10.1093/nar/gkm846 (2008).
Tatusova, T. et al. NCBI prokaryotic genome annotation pipeline. Nucleic Acids Res. https://doi.org/10.1093/nar/gkw569 (2016).
Papagianni, M., Avramidis, N., Filioussis, G., Dasiou, D. & Ambrosiadis, I. Determination of bacteriocin activity with bioassays carried out on solid and liquid substrates: Assessing the factor indicator microorganism. Microb. Cell Fact. 5, 1–14. https://doi.org/10.1186/1475-2859-5-30 (2006).
Junqueira, M. et al. Protein identification pipeline for the homology-driven proteomics. J. Proteom. 71, 346–356. https://doi.org/10.1016/j.jprot.2008.07.003 (2008).
Apweiler, R. et al. UniProt: The universal protein knowledgebase. Nucleic Acids Res. 1, D115–D119. https://doi.org/10.1093/nar/gkh131 (2004).
Korostin, D. et al. Comparative analysis of novel MGISEQ-2000 sequencing platform vs Illumina HiSeq 2500 for whole-genome sequencing. PLoS ONE 15, e0230301. https://doi.org/10.1371/journal.pone.0230301 (2020).
Jackman, S. D. et al. ABySS 2.0: Resource-efficient assembly of large genomes using a bloom filter. Gen. Res. 27, 768. https://doi.org/10.1101/gr.214346.116 (2017).
Makarova, K. S. et al. Evolution and classification of the CRISPR-Cas systems. Nat. Rev. Microbiol. 9, 467–477. https://doi.org/10.1038/nrmicro2577 (2011).
Overbeek, R. et al. The SEED and the Rapid Annotation of microbial genomes using Subsystems Technology (RAST). Nucleic Acids Res. 42, D206–D214. https://doi.org/10.1093/nar/gkt1226 (2014).
Jia, Y. et al. WEGO: A web tool for plotting GO annotations. Nucleic Acids Res. 34, W293–W297. https://doi.org/10.1093/nar/gkl031 (2006).
Aziz, R. K. et al. SEED servers: High-performance access to the SEED genomes, annotations, and metabolic models. PLoS ONE 7, e48053. https://doi.org/10.1371/journal.pone.0048053 (2012).
Wang, L. et al. A minimal nitrogen fixation gene cluster from Paenibacillus sp. WLY78 enables expression of active nitrogenase in Escherichia coli. PLoS Genet. 9, 1–11. https://doi.org/10.1371/journal.pgen.1003865 (2013).
Leigh, J. A. & Dodsworth, J. A. Nitrogen regulation in bacteria and archaea. Annu. Rev. Microbiol. 61, 349–377. https://doi.org/10.1146/annurev.micro.61.080706.093409 (2007).
Capra, E. J. & Laub, M. T. The evolution of two-component signal transduction systems. Annu. Rev. Microbiol. 66, 325–347. https://doi.org/10.1146/annurev-micro-092611-150039 (2012).
Kanehisa, M., Sato, Y. & Morishima, K. BlastKOALA and GhostKOALA: KEGG tools for functional characterization of genome and metagenome sequences. J. Mol. Biol. 428, 726–731. https://doi.org/10.1016/j.jmb.2015.11.006 (2016).
Lewis, V. G., Ween, M. P. & McDevitt, C. A. The role of ATP-binding cassette transporters in bacterial pathogenicity. Protoplasma 249, 919–942. https://doi.org/10.1007/s00709-011-0360-8 (2012).
Hathout, Y., Setlow, B., Cabrera-Martinez, R. M., Fenselau, C. & Setlow, P. Small, acid-soluble proteins as biomarkers in mass spectrometry analysis of Bacillus spores. Appl. Environ. Microbiol. 69, 1100–1107. https://doi.org/10.1128/AEM.69.2.1100-1107.2003 (2003).
Falke, J. J., Bass, R. B., Butler, S. L., Chervitz, S. A. & Danielson, M. A. The two-component signaling pathway of bacterial chemotaxis: A molecular view of signal transduction by receptors, kinases, and adaptation enzymes. Annu. Rev. Cell Dev. Biol. 13, 457–512. https://doi.org/10.1146/annurev.cellbio.13.1.457 (1997).
Lubelski, J., Rink, R., Khusainov, R., Moll, G. & Kuipers, O. Biosynthesis, immunity, regulation, mode of action and engineering of the model lantibiotic nisin. Cell. Mol. Life Sci. 65, 455–476. https://doi.org/10.1007/s00018-007-7171-2 (2008).
Yu, J. & Shapiro, L. Early Caulobacter crescentus genes fliL and fliM are required for flagellar gene expression and normal cell division. J. Bacteriol. 174, 3327–3338. https://doi.org/10.1128/jb.174.10.3327-3338.1992 (1992).
Chen, I. A. et al. IMG/M vol 5.0: An integrated data management and comparative analysis system for microbial genomes and microbiomes. Nucleic Acids Res. 847, D666–D677. https://doi.org/10.1093/nar/gky901 (2019).
Blin, K. AntiSMASH 5.0: Updates to the secondary metabolite genome mining pipeline. Nucleic Acids Res. 47, W81–W87. https://doi.org/10.1093/nar/gkz310 (2019).
Palaniappan, K. IMG-ABC vol 5.0: An update to the IMG/Atlas of biosynthetic gene clusters knowledgebase. Nucleic Acids Res. 48, D422–D430. https://doi.org/10.1093/nar/gkz932 (2020).
Demain, A. L. & Fang, A. The natural functions of secondary metabolites. Adv. Biochem. Eng. Biotechnol. 69, 1–39. https://doi.org/10.1007/3-540-44964-7_1 (2000).
Bachmann, B. O. & Ravel, J. Methods for in silico prediction of microbial polyketide and nonribosomal peptide biosynthetic pathways from DNA sequence data. Methods Enzymol. 458, 181–217. https://doi.org/10.1016/S0076-6879(09)04808-3 (2009).
Blanco, P. et al. Bacterial multidrug efflux pumps: Much more than antibiotic resistance determinants. Microorganisms 4, 14. https://doi.org/10.3390/microorganisms4010014 (2016).
Osbourn, A. Secondary metabolic gene clusters: Evolutionary toolkits for chemical innovation. Trends Genet. 26, 449–457. https://doi.org/10.1016/j.tig.2010.07.001 (2010).
Papagianni, M. Recent advances in engineering the central carbon metabolism of industrially important bacteria. Microb. Cell Fact. 11, 50. https://doi.org/10.1186/1475-2859-11-50 (2012).
Noor, E., Eden, E., Milo, R. & Alon, U. Central carbon metabolism as a minimal biochemical walk between precursors for biomass and energy. Mol. Cell 39, 809–820. https://doi.org/10.1016/j.molcel.2010.08.031 (2010).
Fell, D. A. Evolution of central carbon metabolism. Mol. Cell 39, 663–664. https://doi.org/10.1016/j.molcel.2010.08.034 (2010).
Shi, L. & Tu, B. P. Acetyl-CoA and the regulation of metabolism: Mechanisms and consequences. Curr. Opin. Cell Biol. 33, 125–131. https://doi.org/10.1016/j.ceb.2015.02.003 (2015).
Green, E. R. & Mecsas, J. Bacterial secretion systems: An overview. Microbiol. Spectr. 4, 1. https://doi.org/10.1128/microbiolspec.VMBF-0012-2015 (2016).
Pugsley, A. P. The complete general secretory pathway in gram-negative bacteria. Microbiol. Rev. 57, 50–108. https://doi.org/10.1128/mr.57.1.50-108.1993 (1993).
Kirienko, N. V., Ausubel, F. M. & Ruvkun, G. Mitophagy confers resistance to siderophore-mediated killing by Pseudomonas aeruginosa. Proc. Nat. Acad. Sci. U.S.A. 112, 1821–1826. https://doi.org/10.1073/pnas.1424954112 (2015).
Martinez-Argudo, I., Little, R., Shearer, N., Johnson, P. & Dixon, R. The NifL-NifA system: A multidomain transcriptional regulatory complex that integrates environmental signals. J. Bacterial. 186, 601–610. https://doi.org/10.1128/JB.186.3.601-610.2004 (2004).
Thaker, M. N. et al. Identifying producers of antibacterial compounds by screening for antibiotic resistance. Nat. Biotechnol. 31, 922–927. https://doi.org/10.1038/nbt.2685 (2013).
Jia, N., Ding, M. Z. & Luo, H. Complete genome sequencing and antibiotics biosynthesis pathways analysis of Streptomyces lydicus 103. Sci. Rep. 7, 44786. https://doi.org/10.1038/srep44786 (2017).
Ngashangva, N., Mukherjee, P., Sharma, K. C., Kalita, M. C. & Indira, S. Analysis of antimicrobial peptide metabolome of bacterial endophyte isolated from traditionally used medicinal plant Millettia pachycarpa Benth. Front. Microbiol. 12, 656896. https://doi.org/10.3389/fmicb.2021.65689 (2021).
Ito, C. et al. Isoflavonoids with antiestrogenic activity from Millettia pachycarpa. J. Nat. Prod. 69, 138–141. https://doi.org/10.1021/np050341w (2006).
Simonian, M. H. Spectrophotometric determination of protein concentration. Curr. Protoc. Toxicol. 21, 1–7 (2004).
Scopes, R. K. Protein Purification: Principles and Practice 3rd edn. (Springer, 1994).
Liu, S., Takala, T. M., Wan, X., Reunanen, J. & Saris, P. E. J. Cell-mediated killing of L. monocytogenes by leucocin C producing Escherichia coli. Microbiol. Res. 168, 300–304. https://doi.org/10.1016/j.micres.2012.11.011 (2013).
Ryzhykau, Y. L. et al. Molecular model of a sensor of two-component signaling system. Sci. Rep. 11, 10774. https://doi.org/10.1038/s41598-021-89613-6 (2021).
Muir, R. E. & Gober, J. W. Regulation of late flagellar gene transcription and cell division by flagellum assembly in Caulobacter crescentus. Mol. Microbiol. 41, 117–130. https://doi.org/10.1046/j.1365-2958.2001.02506.x (2001).
Koretke, K. K., Lupas, A. N., Warren, P. V., Rosenberg, M. & Brown, J. R. Evolution of two-component signal transduction. Mol. Biol. Evol. 17, 1956–1970. https://doi.org/10.1093/oxfordjournals.molbev.a026297 (2000).
Tseng, T. T., Tyler, B. M. & Setubal, J. C. Protein secretion systems in bacterial-host associations, and their description in the Gene Ontology. BMC Microbiol. 9(Suppl 1), S2. https://doi.org/10.1186/1471-2180-9-S1-S2 (2009).
Choi, S. K. et al. Identification and functional analysis of the fusaricidin biosynthetic gene of Paenibacillus polymyxa E681. Biochem. Biophys. Res. Commun. 365, 89–95. https://doi.org/10.1016/j.bbrc.2007.10.147 (2008).
Wang, Y., Coleman-Derr, D., Chen, G. & Gu, Y. Q. OrthoVenn: A web server for genome wide comparison and annotation of orthologous clusters across multiple species. Nucleic Acids Res. 43, W78–W84. https://doi.org/10.1093/nar/gkv487 (2015).
Goris, J. et al. DNA-DNA hybridization values and their relationship to whole-genome sequence similarities. Int. J. Syst. Evol. Microbiol. 57, 81–91. https://doi.org/10.1099/ijs.0.64483-0 (2007).
Pattanaik, B. & Lindberg, P. Terpenoids and their biosynthesis in cyanobacteria. Life. 5, 269–293. https://doi.org/10.3390/life5010269 (2015).
Ramirez-Gaona, M. et al. A web tool for generating high quality machine-readable biological pathways. J. Vis. Exp. 120, 54869. https://doi.org/10.3791/54869 (2017).
Rohdich, F., Kis, K., Bacher, A. & Eisenreich, W. The non-mevalonate pathway of isoprenoids: Genes, enzymes and intermediates. Curr. Opin. Chem. Biol. 5, 535–540. https://doi.org/10.1016/s1367-5931(00)00240-4 (2001).
Jeong, H., Choi, S. K., Ryu, C. M. & Park, S. H. Chronicle of a soil bacterium: Paenibacillus polymyxa E681 as a tiny guardian of plant and human health. Front. Microbiol. 10, 467. https://doi.org/10.3389/fmicb.2019.00467 (2019).
Kjærbøllinga, I. et al. Linking secondary metabolites to gene clusters through genome sequencing of six diverse Aspergillus species. Proc. Nat. Acad. of Sci. U.S.A. https://doi.org/10.1073/pnas.1715954115 (2018).
Brock, T. D. Milestones in Microbiology 2nd edn, 1546–1940 (ASM Press, 1999).
Green, G., Dicks, L. M. T., Bruggeman, G., Vandamme, E. J. & Chikindas, M. L. Pediocin PD-1, a bactericidal antimicrobial peptide from Pediococcusm damnosus NCFB1832. J. Appl. Microbiol. 83, 127–132. https://doi.org/10.1046/j.1365-2672.1997.00241.x (1997).
Holt, J. G., Krieg, N. R., Sneath, P. H. A., Staley, J. T. & Williams, S. T. Bergey’s Manual of Determinative Bacteriology 9th edn. (Lippincott Williams & Wilkins, 1994).
Balouiri, M., Sadiki, M. & Ibnsouda, S. K. Methods for in vitro evaluating antimicrobial activity. A review. J. Pharm. Anal. 6, 71–79. https://doi.org/10.1016/j.jpha.2015.11.005 (2016).
Wilson, K. Preparation of genomic DNA from bacteria. Curr. Prot. Mol. Biol. https://doi.org/10.1002/0471142727.mb0204s56 (1997).
He, Z. et al. Isolation and identification of a Paenibacillus polymyxa strain that coproduces a novel lantibiotic and polymyxin. Appl. Environ. Microbiol. 73, 168–178. https://doi.org/10.1128/AEM.02023-06 (2007).
Clinical and Laboratory Standards Institute. Method for Dilution Antimicrobial Susceptibility Test for Bacteria that Grow Aerobically 7th edn. (Wayne, 2006).
Clinical and Laboratory Standards Institute. Performance Standards for Antimicrobial Susceptibility Testing (Approved Standard-18th Informational Supplement) (Wayne, 2008).
Magaldi, S. et al. Well diffusion for antifungal susceptibility testing. Int. J. Infect. Dis. 8, 39–45. https://doi.org/10.1016/j.ijid.2003.03.002 (2004).
Wingfield, P. Protein precipitation using ammonium sulfate. In Current Protocols in Protein Science (eds Coligan, J. E. et al.) (Wiley, 2001).
Herraiz, T. Sample preparation and reversed phase-high performance chromatography analysis of food-derived peptides. Anal. Chim. Acta 352, 119–139. https://doi.org/10.1016/S0003-2670(97)00199-2 (1997).
Conlon, J. M. Purification of naturally occurring peptides by reversed-phase HPLC. Nat. Protoc. 2, 191–197. https://doi.org/10.1038/nprot.2006.437 (2006).
Sonia, K. S. & Jata, S. Proteomic analysis revealed ROS-mediated growth inhibition of Aspergillus terreus by shikonin. J. Proteom. 224, 103849. https://doi.org/10.1016/j.jprot.2020.103849 (2020).
Geourjon, C. & Deléage, G. SOPMA: Significant improvements in protein secondary structure prediction by consensus prediction from multiple alignments. Comput. Appl. Biosci. 11, 681–684. https://doi.org/10.1093/bioinformatics/11.6.681 (1995).
Studer, G. et al. ProMod3-A versatile homology modelling toolbox. PLoS Comput. Biol. 17, e1008667. https://doi.org/10.1371/journal.pcbi.1008667 (2021).
Carol, H., Sucheta, K., Marilyn, C. & Jong, P. Preparation of cultured cells for scanning electron microscope. Protocol Exchange 2007, 1–5. https://doi.org/10.1038/nprot.2007.504 (2007).
Hartmann, M. et al. Damage of the bacterial cell envelope by antimicrobial peptides gramicidin S and PGLa as revealed by transmission and scanning electron microscopy. Antimicrob. Agents Chemother. 54, 3132–3142. https://doi.org/10.1128/AAC.00124-10 (2010).
Chor, B., Horn, D., Goldman, N., Levy, Y. & Massingham, T. Genomic DNA k-mer spectra: Models and modalities. Gen. Biol. 10, R108. https://doi.org/10.1186/gb-2009-10-10-r108 (2009).
Altschul, S. F., Gish, W., Miller, W., Myers, E. W. & Lipman, D. J. Basic local alignment search tool. J. Mol. Biol. 215, 403–410. https://doi.org/10.1016/S0022-2836(05)80360-2 (1990).
Jeong, H., Choi, S. K., Park, S. Y., Kim, S. H. & Park, S. H. Draft genome sequence of Paenibacillus peoriae strain KCTC 3763T. J. Bacterial. 194, 1237–1238. https://doi.org/10.1128/JB.06577-11 (2012).
Alvarez, V. M., von der Weid, I., Seldin, L. & Santos, A. L. Influence of growth conditions on the production of extracellular proteolytic enzymes in Paenibacillus peoriae NRRL BD-62 and Paenibacillus polymyxa SC2. Lett. Appl. Microbiol. 43, 625–630. https://doi.org/10.1111/j.1472-765X.2006.02015.x (2006).
Ma, M. et al. Complete genome sequence of Paenibacillus polymyxa SC2, a strain of plant growth-promoting Rhizobacterium with broad-spectrum antimicrobial activity. J. Bacterial. 193, 311–312. https://doi.org/10.1128/JB.01234-10 (2011).
Shin, S. H. et al. Genome sequence of Paenibacillus terrae HPL-003, a xylanase-producing bacterium isolated from soil found in forest residue. J. Bacteriol. 194, 1266. https://doi.org/10.1128/JB.06668-11 (2012).
Chow, V. et al. Complete genome sequence of Paenibacillus sp. strain JDR-2. Stand. Genomic Sci. 6, 1–10. https://doi.org/10.4056/sigs.2374349 (2012).
Weselowski, B., Nathoo, N., Eastman, A. W., MacDonald, J. & Yuan, Z. C. Isolation, identification and characterization of Paenibacillus polymyxa CR1 with potentials for biopesticide, biofertilization, biomass degradation and biofuel production. BMC Microbiol. 16, 244. https://doi.org/10.1186/s12866-016-0860-y (2016).
Bin-hui, J., Jin-liang, L. & Xiao-min, H. Draft genome sequence of the efficient bioflocculant-producing bacterium Paenibacillus sp. Strain A9. Gen. Announc. 1, e00131–e00213. https://doi.org/10.1128/genomeA.00131-13 (2013).
Smith, J. L., Goldberg, J. M. & Grossman, A. D. Complete genome sequences of Bacillus subtilis subsp. subtilis laboratory strains JH642 (AG174) and AG1839. Gen. Announc. 2, e00663. https://doi.org/10.1128/genomeA.00663-14 (2014).
Bentley, S. et al. Complete genome sequence of the model actinomycete Streptomyces coelicolor A3(2). Nature 417, 141–147. https://doi.org/10.1038/417141a (2002).
Meier-Kolthoff, J. P. & Göker, M. TYGS is an automated high-throughput platform for state-of-the-art genome-based taxonomy. Nat. Commun. 10, 2182. https://doi.org/10.1038/s41467-019-10210-3 (2019).
Meier-Kolthoff, J. P., Auch, A. F., Klenk, H.-P. & Göker, M. Genome sequence-based species delimitation with confidence intervals and improved distance functions. BMC Bioinform. 14, 60. https://doi.org/10.1186/1471-2105-14-60 (2013).
Lee, I., Kim, Y. O., Park, S. C. & Chun, J. OrthoANI: An improve algorithm and software for calculating average nucleotide identity. Int. J. Syst. Evol. Microbiol. 66, 1100–1103. https://doi.org/10.1099/ijsem.0.000760 (2015).
Lefort, V., Desper, R. & Gascuel, O. FastME 2.0: A comprehensive, accurate, and fast distance-based phylogeny inference program. Mol. Biol. Evol. 32, 2798–2800. https://doi.org/10.1093/molbev/msv150 (2015).
Acknowledgements
The authors thank the C-CAMP, Bangalore and Valerian Chem. Pvt. Ltd., New Delhi for the LC-MS experiment. They also thank the Jawaharlal Nehru University’s Advanced Instrumentation Research Facility (AIRF) for the Scanning Electron Microscope. The draft genome sequencing was performed at AgriGenome Labs (P) Ltd, Kerala.
Funding
This work was supported by Department of Biotechnology of the Ministry of Science and Technology, Government of India.
Author information
Authors and Affiliations
Contributions
N.N. performed the experiment and wrote the main manuscript and prepared all the figures and tables. C.K.S. performed the RP-HPLC. P.K.M., N.N., S.I., and M.C.K. initiated the ideas and proofread the manuscript. The IBSD manuscript no. is IBSD/MS/2020/01/094.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Ngashangva, N., Mukherjee, P.K., Sharma, C. et al. Integrated genomics and proteomics analysis of Paenibacillus peoriae IBSD35 and insights into its antimicrobial characteristics. Sci Rep 12, 18861 (2022). https://doi.org/10.1038/s41598-022-23613-y
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-022-23613-y
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.