Abstract
Cocaine epigenetically regulates gene expression via changes in histone post-translational modifications (HPTMs). We previously found that the immediate early gene Nr4a1 is epigenetically activated by cocaine in mouse brain reward regions. However, few studies have examined multiple HPTMs at a single gene. Bivalent gene promoters are simultaneously enriched in both activating (H3K4me3 (K4)) and repressive (H3K27me3 (K27)) HPTMs. As such, bivalent genes are lowly expressed but poised for activity-dependent gene regulation. In this study, we identified K4&K27 bivalency at Nr4a1 following investigator-administered cocaine in male and female mice. We applied sequential chromatin immunoprecipitation and qPCR to define Nr4a1 bivalency and expression in striatum (STR), prefrontal cortex (PFC), and hippocampus (HPC). We used Pearson’s correlation to quantify relationships within each brain region across treatment conditions for each sex. In female STR, cocaine increased Nr4a1 mRNA while maintaining Nr4a1 K4&K27 bivalency. In male STR, cocaine enriched repressive H3K27me3 and K4&K27 bivalency at Nr4a1 and maintained Nr4a1 mRNA. Furthermore, cocaine epigenetically regulated a putative NR4A1 target, Cartpt, in male PFC. This study defined the epigenetic regulation of Nr4a1 in reward brain regions in male and female mice following cocaine, and, thus, shed light on the biological relevance of sex to cocaine use disorder.
Similar content being viewed by others
Introduction
Despite decades of research, cocaine use disorder remains a global health problem for which there is currently no FDA approved treatment. Despite clear epidemiological evidence that cocaine abuse afflicts men and women differently1, the underlying neurobiology of how cocaine impacts each sex is not fully understood. Human sex differences in behavior include higher initiation of cocaine use by men than women, a greater percent of men addicted to cocaine than women2, and a faster shift from casual drug use to addiction in women than men3. Additionally, women enter treatment at an earlier time point than men, have a higher relapse rate3, both of which may reflect women being more vulnerable to cue-induced relapse than men4. Importantly, rodent models of cocaine abuse recapitulate sexually dimorphic behavior, such that female rodents are more responsive to drug-conditioned stimuli5,6 and acquire cocaine-self administration at a faster rate than male rodents7,8. Recent research has indicated there are sexually dimorphic molecular effects of cocaine that might mediate behavioral phenotypes. For example, it was recently shown that cocaine broadly regulates distinct sets of proteins in males relative to females following cocaine self-administration in rodents9. Given the prominent sexually dimorphic cocaine-associated behavioral phenotypes and the distinct neuroadaptations for males and females10, sex must be considered a biological variable within drug research.
On a molecular level, cocaine regulates gene expression in part through changes in histone post-translational modifications (HPTMs). Such changes can be transient or persist across long periods of abstinence11,12,13,14,15,16,17,18. HPTMs tune gene expression in large part by altering chromatin accessibility and recruiting transcriptional machinery13. Acute and chronic cocaine impact HPTMs in multiple brain reward regions17,19,20, such as the striatum (STR)18,21, hippocampus (HPC)22,23,24, and prefrontal cortex (PFC)25,26. However, the complexities of chromatin remodeling following cocaine has yet to be explored in both sexes.
The histone code has historically proposed that engagement of individual HPTMs at a chosen locus can induce changes in protein recruitment and impact gene expression. However, the code also posits that HPTMs can be interdependent and/or work in tandem to impact gene expression27,28. Thus, this model suggests that chromatin accessibility and, correspondingly, gene expression are largely determined not only by the concentration of single HPTMs at a given gene promoter, but also by the combination of co-localized HPTMs27.
While many studies demonstrate genome-wide regulation of individual HPTMs following cocaine exposure13,29,30,31, the literature lacks investigation of how cocaine impacts co-localization of HPTMs. Bivalent gene promoters are comprised of co-localized activating (H3K4me3 (K4)) and repressive (H3K27me3 (K27)) HPTMs32 at a single promoter. H3K4me3 and H3K27me3 may be localized to discrete nucleosomes within the same promoter, or co-exist on the same nucleosome or H3 tail33. The presence of both modifications results in a poised state of gene expression, which can be ‘resolved’ to an active or repressive state by the removal of either H3K27me3 or H3K4me3 in response to transcriptional cues32. Bivalency may also protect against spurious and irreversible silencing of promoters by DNA hypermethylation, maintaining the epigenetic flexibility of stimuli-responsive genes34. While limited studies have investigated bivalency in the brain35,36,37, none to date have specifically examined bivalency with regard to sex or cocaine exposure.
We, and others, recently discovered that cocaine exposure activates expression of nuclear receptor subfamily 4 group A member 1 (Nr4a1) in the nucleus accumbens (NAc), a part of the STR12,38,39. Studies show that activation of Nr4a1 in the NAc attenuates cocaine self-administration38, cocaine conditioned place preference38, cocaine sensitization40, and amphetamine-induced locomotion41. Nr4a1 functions as a transcription factor at target genes involved in drug reward, such as Cocaine- and amphetamine-regulated transcript (Cartpt), Tyrosine Hydroxylase (Th), solute carrier family 6 member 3 (DAT production), and Brain-derived neurotrophic factor (Bdnf)38,42,43, as well as plasticity in reward-related brain regions40,44. However, the precise function of HPTMs in cocaine-induced Nr4a1 expression has yet to be elucidated. Additionally, there is conspicuous lack of data on the regulation of Nr4a1 where sex is a defined variable. Accordingly, we investigated K4&K27 bivalency in the cocaine regulation of Nr4a1 expression in both male and female mice.
In this study, we utilized single sample sequencing (S3EQ)45 and sequential chromatin immunoprecipitation (ChIP)35,46 to independently investigate the transcriptomic and epigenomic profiles of Nr4a1 from three different brain regions in males and females. We also utilized Pearson’s correlation matrices to capture the individual variability in biological changes induced by cocaine exposure in males and females.
Results
K4&K27 bivalency at Nr4a1 promoter in brain reward regions was present in both male and female mice
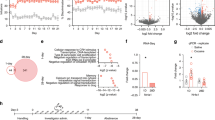
For epigenetic profiling of three reward-associated brain regions following cocaine treatment, we applied sequential ChIP to examine K4&K27 bivalency at Nr4a1 (Fig. 1). ChIP-qPCR and ChIP-Seq protocols that quantify HPTM deposition (the addition of the modification) at monovalent promoters are unable to quantify combinatorial HPTM deposition at single promoter. This weakness stems from the fact that individual HPTM profiling experiments cannot distinguish between bivalent promoter deposition of two HPTMs in a single population of cells or monovalent promoter deposition of two HPTMs from a mixed population of cells33. The sequential ChIP protocol is required to define true bivalency of H3K4me3 and H3K27me3 deposition at the same locus in a single population of cells (and nucleosomes)33. To measure bivalency, we split each single sample and performed H3K4me3 ChIP followed by H3K27me3 ChIP. From the same sample, we also performed H3K27me3 ChIP followed by H3K4me3 ChIP. In this way, sequential ChIP was assayed with either H3K4me3 or H3K27me3 as the first IP, and the other HPTM as the second IP. This approach allowed us to interpret differences in cocaine-regulated K4&K27 bivalency due to differences in rates of deposition or removal between the two HPTMs47. In each case, enrichment based on antibody order was quantified by qCHIP. As we found no differences in results of qCHIP by order of sequential ChIP, we concluded that both approaches demonstrated Nr4a1 K4&K27 promoter bivalency.
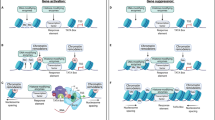
Methods used to investigate regional transcriptional and epigenomic profiles following cocaine exposure. (A) Timeline for investigator-administered cocaine (20 mg/kg; i.p) paradigm. (B) Coronal brain slice illustrations showing the brain regions taken for single sample sequencing (Top to bottom: prefrontal cortex (PFC), striatum (STR), hippocampus (HPC)). (C) Schematic showing the sequential ChIP protocol that includes two immunoprecipitations.
Cocaine enriched H3K27me3 and bivalency at the Nr4a1 promoter and did not alter Nr4a1 mRNA levels in male striatum
Following cocaine treatment, there was no change in Nr4a1 mRNA (Fig. 2A) or H3K4me3 deposition at Nr4a1 in the male STR (Fig. 2B). In the same samples, there was increased H3K27me3 at Nr4a1 (Fig. 2C) (t10 = 2.3, p = 0.0443) and an increase in K4&K27 bivalency at the Nr4a1 promoter (Fig. 2D,E) (D: t5.973 = 2.663, p = 0.0376; E: t9 = 2.298, p = 0.0476). In the male HPC following cocaine treatment, there was no change in Nr4a1 mRNA (Fig. 2F), but we observed depletion of H3K4me3 at Nr4a1 (Fig. 2G) (t12 = 2.528, p = 0.0265). In the same samples, there was no change in H3K27me3 deposition or K4&K27 bivalency (Fig. 2H,I,J). In the male PFC following cocaine treatment, there was no change in Nr4a1 mRNA (Fig. 2K), H3K4me3, H3K27me3, or K4&K27 bivalency (Fig. 2L–O).
Cocaine alters the epigenetic profile of Nr4a1 in the striatum and hippocampus of male mice. (A) No change in mRNA levels of Nr4a1 (as examined by primer pairs that cover the junction between exon 4/5 and exon 6/7) were observed in the male mouse STR following 10 days of investigator-administered cocaine (n = 5–6/group). There was no change in (B) H3K4me3 (n = 6/group), but an increase in (C) H3K27me3 (n = 6/group), (D) H3K4me3/H3K27me3 bivalency (n = 6/group), and (E) H3K27me3/H3K4me3 bivalency (n = 5–6/group) at Nr4a1 in the male mouse STR following 10 days of investigator-administered cocaine. (F) There was no change in mRNA levels of Nr4a1 (as measured by independent primer pairs that cover the junction between exons 4/5 and exons 6/7) in the male mouse HPC following 10 days of investigator-administered cocaine (n = 7–8/group). There was a decrease in (G) H3K4me3 at Nr4a1 (n = 7/group), but no change in (H) H3K27me3 (n = 6/group), (I) H3K4me3/H3K27me3 bivalency (n = 7–8/group), or (J) H3K27me3/H3K4me3 bivalency (n = 6–7/group) at Nr4a1 in the male mouse HPC following 10 days of investigator-administered cocaine. (K) There was no change in mRNA levels of Nr4a1 (as measured by independent primer pairs that cover the junction between exons 4/5 and exons 6/7) in the male mouse PFC following 10 days of investigator-administered cocaine (n = 7–8/group). There is no change in (L) H3K4me3 (n = 8/group), (M) H3K27me3 (n = 7–8/group), (N) H3K4me3/H3K27me3 bivalency (n = 7–8/group), or (O) H3K27me3/H3K4me3 bivalency at Nr4a1 in the male mouse PFC following 10 days of investigator-administered cocaine (n = 6–7/group). *p < 0.05. Data are shown as mean ± SEM. STR, striatum; HPC, hippocampus; PFC, prefrontal cortex.
Cocaine increased Cartpt mRNA in PFC of male mice
Given the transient nature of immediate early gene expression, we also measured mRNA and HPTM deposition of a downstream transcriptional target of Nr4a138. We found no changes in Cartpt mRNA, or in mRNA of dopamine receptors, Drd1 and Drd2, or adenosine A2a receptor (A2a), in the male STR following cocaine treatment (Fig. 3A). Likewise, we observed no change in H3K4me3 or H3K27me3 deposition at Cartpt (Fig. 3B,C). Similarly, in the male HPC, cocaine treatment did not alter Cartpt, Drd1, Drd2, or A2a mRNA (Fig. 3D), or H3K4me3 or H3K27me3 deposition at Cartpt (Fig. 3E,F). Alternatively, in the male PFC, cocaine treatment increased Cartpt mRNA (t7.165 = 3.182, p = 0.0150), and we observed a trending decrease of Drd1 mRNA (trending, t7.247 = 2.059, p = 0.0771), Drd2 (trending, t7.318 = 2.110, p = 0.0711), but cocaine did not alter A2a mRNA levels (Fig. 3G). This was accompanied by no change in H3K4me3 or H3K27me3 deposition at Cartpt (Fig. 3H,I). These data show that cocaine induces Cartpt in the male PFC.
Nr4a1 downstream target, Cartpt, is regionally activated after 10 days of investigator-administered cocaine in male mice. (A) There was no change in mRNA levels of Cartpt, Drd1, Drd2, or A2a in the male mouse STR following 10 days of investigator-administered cocaine (n = 5–6/group). There was no change in (B) H3K4me3 (n = 6/group) or (C) H3K27me3 (n = 6/group) at Cartpt in the male mouse STR after 10 days of investigator-administered cocaine. (D) There was no change in mRNA levels of Cartpt, Drd1, Drd2, or A2a in the male mouse HPC following 10 days of investigator-administered cocaine (n = 7–8/group). There was no change in (E) H3K4me3 (n = 7/group) or (F) H3K27me3 (n = 6–8/group) at Cartpt in the male mouse HPC after 10 days of investigator-administered cocaine. (G) There was an increase in mRNA levels of Cartpt, a trending decrease in mRNA levels of Drd1 and Drd2, and no change in mRNA levels of A2a in the male mouse PFC following 10 days of investigator-administered cocaine (n = 5–6/group). There was no change in (H) H3K4me3 (n = 8/group) or (I) H3K27me3 at Cartpt in the male mouse PFC after 10 days of investigator-administered cocaine (n = 5–6/group). *p < 0.05. #p < 0.10. Data are shown as mean ± SEM. STR, striatum; HPC, hippocampus; PFC, prefrontal cortex.
Cocaine increased Nr4a1 mRNA in multiple reward-associated brain regions in female mice
Following cocaine treatment, Nr4a1 mRNA was increased in the female STR (Fig. 4A). This increase was independently identified using two different primer sets, each of which spans the junction between two distinct exons within Nr4a1 (Nr4a1 Exon 4/5: t9 = 2.771, p = 0.0217; Nr4a1 Exon 6/7: t9 = 3.030, p = 0.0142). In the same samples, there was no change in H3K4me3, H3K27me3, or K4&K27 bivalency at Nr4a1 (Fig. 4B–E). In the female HPC, there was no change in Nr4a1 mRNA following cocaine treatment when the Nr4a1 Exon 4/5 primer set was used, but we observed a trending increase in Nr4a1 mRNA when mRNA levels were evaluated with Exon 6/7 primers (trending, t14 = 1.1841, p = 0.0869) (Fig. 4F). In the same samples, there was no change in Nr4a1 H3K4me3, H3K27me3 or K4&K27 bivalency (Fig. 4G–J). In the female PFC following cocaine treatment, Nr4a1 mRNA was increased when measured at Nr4a1 E6/7 but not Nr4a1 E4/5 (Nr4a1 Exon 6/7: t13 = 2.327, p = 0.0368) (Fig. 4K). In the same samples, there was no change in H3K4me3 (Fig. 4L), there was H3K27me3 recruitment (t6.392 = 3.363, p = 0.0138) (Fig. 4M), and no change in K4&K27 bivalency (Fig. 4N,O). In sum, these data showed that cocaine treatment in female mice increased Nr4a1 mRNA expression in the STR along with maintenance of H3K4me3, H3K27me3, and K4&K27 bivalency at the Nr4a1 promoter. These data also showed that cocaine treatment in female mice increased Nr4a1 mRNA expression in the PFC, despite recruitment of H3K27me3.
Cocaine alters the transcriptomic profile of Nr4a1 in the STR of females and the transcriptomic and epigenetic profile of Nr4a1 in the PFC of females. (A) There was an increase in mRNA levels of Nr4a1 (as examined by primer pairs that cover the junction between exons 4/5 and exons 6/7) in the female mouse STR following 10 days of investigator-administered cocaine (n = 4–6/group). There was no change in (B) H3K4me3 (n = 6/group), (C) H3K427me3 (n = 5–6/group), (D) H3K4me3/H3K27me3 bivalency (n = 6/group), or (E) H3K27me3/H3K4me3 bivalency (n = 5–6/group) at Nr4a1 in the female mouse STR after 10 days of investigator-administered cocaine. (F) There was no change in mRNA levels of Nr4a1 as measured by independent primer pairs that cover the junction between exons 4/5, and a trending increase in mRNA levels of Nr4a1 as measured by independent primer pairs that cover the junction between exons 6/7 in the female mouse HPC following 10 days of investigator-administered cocaine (n = 7–8/group). There was no change in (G) H3K4me3 (n = 7–8/group), (H) H3K27me3 (n = 6–8/group), (I) H3K4me3/H3K27me3 (n = 7–8/group), or (J) H3K27me3/H3K4me3 bivalency (n = 6–8/group) at Nr4a1 in the female mouse HPC following 10 days of investigator-administered cocaine. (K) There was an increase in mRNA levels of Nr4a1 (as measured by independent primer pairs that cover the junction between exons 6/7, but not exons 4/5) in the female mouse PFC following 10 days of investigator-administered cocaine (n = 7–8/group). There was no change in (L) H3K4me3 (n = 7/group), but there was an increase in (M) H3K27me3 (n = 7/group) at Nr4a1 following 10 days of investigator-administered cocaine in the female mouse PFC (n = 7/group). (N) There was no change in H3K4me3/H3K27me3 bivalency (n = 7/group) or (O) H3K27me3/H3K4me3 bivalency at Nr4a1 in the female mouse prefrontal cortex following 10 days of investigator-administered cocaine (n = 7/group). *p < 0.05. #p < 0.10. Data are shown as mean ± SEM. STR, striatum; HPC, hippocampus; PFC, prefrontal cortex.
Cocaine did not alter mRNA of drug-related markers, Cartpt mRNA, or Cartpt HPTMs in female mouse brain regions
As we saw independent increases in mRNA levels of Nr4a1 following cocaine in female mice in the PFC and the STR, we expected to find transcriptomic or epigenetic changes at Cartpt, a downstream target of Nr4a138. Surprisingly, we found no transcriptomic or epigenetic changes (H3K4me3, H3K27me3) at Cartpt in female mice following cocaine in the STR (Fig. 5A–C), HPC (Fig. 5D–F), or PFC (Fig. 5G–I), although we observed trending depletion of H3K27me3 levels at Cartpt in the PFC (Fig. 5I) (t11 = 1.953, p = 0.0767). Notably, we also observed no change in mRNA levels of Drd1, Drd2, or A2a in the STR, HPC, or PFC (Fig. 5A,D,G) in female mice following cocaine, but observed a trending decrease in mRNA levels of Drd1 in the PFC (Fig. 5G) (t13 = 1.887, p = 0.0818).
Nr4a1 downstream target, Cartpt, and other drug-related markers are impacted after 10 days of investigator-administered cocaine in the PFC of female mice. (A) There was no change in mRNA levels of Cartpt, Drd1, Drd2, or A2a in the female mouse STR following 10 days of investigator-administered cocaine (n = 6/group). There was no change in (B) H3K4me3 (n = 6/group) or (C) H3K27me3 (n = 6/group) at Cartpt in the female mouse STR after 10 days of investigator-administered cocaine. (D) There was no change in mRNA levels of Cartpt, Drd1, Drd2, or A2a in the female mouse HPC following 10 days of investigator-administered cocaine (n = 8/group). There was no change in (E) H3K4me3 (n = 7–8/group) or (F) H3K27me3 (n = 6–8/group) at Cartpt in the female mouse HPC after 10 days of investigator-administered cocaine. (G) There was a trending decrease in mRNA levels of Drd1 and no change in mRNA levels of Cartpt, Drd2, or A2a in the female mouse PFC following 10 days of investigator-administered cocaine (n = 7–8/group). There was no change in (H) H3K4me3 (n = 7–8/group) but there was a trending decrease in (I) H3K27me3 at Cartpt in the female mouse PFC after 10 days of investigator-administered cocaine (n = 7–8/group). *p < 0.05. #p < 0.10. Data are shown as mean ± SEM. STR, striatum; HPC, hippocampus; PFC, prefrontal cortex.
Cocaine impacted correlational relationships within and between the transcriptomic and epigenetic profiles of Nr4a1 and Cartpt in male and female mice
As we hypothesized that we would see a linear relationship between Nr4a1 and Cartpt following cocaine, we utilized Pearson’s correlation matrices to explore the relationships within our sample population between the transcriptomic and epigenetic profiles of Nr4a1 and Cartpt. Every significant correlation we observed can be found in Supplementary Tables 3–5 which also provides the corresponding figure(s) that show the data within each correlation in Fig. 6 (example: the data used to reveal the positive correlation between Nr4a1 mRNA and Cartpt H3K27me3 in the STR of saline-treated females can been seen in Figs. 4A and 5C.) In the STR we observed positive correlations between (1) Nr4a1 H3K4me3 and Cartpt H3K4me3 and (2) mRNA levels of Nr4a1 measured by the two primers pairs spanning either Exon 4/5 or Exon 6/7 across treatments (saline, cocaine) and sex (male, female) (Fig. 6A,B). In the STR of saline-treated females, we observed a positive correlation between (3) Nr4a1 mRNA and Cartpt H3K27me3 (Fig. 6B). In the STR of males following cocaine, we observed a positive correlation between (4) Cartpt mRNA and Cartpt H3K27me3, (5) Cartpt mRNA and Nr4a1 K4/K27 bivalency, and (6) Cartpt H3K27me3 and Nr4a1 K4/K27 bivalency (Fig. 6A). In the STR of females following cocaine, we observed a positive correlation between (7) Nr4a1 H3K4me3 and H3K4me3 at Cartpt and (8) Nr4a1 H3K27me3 and Cartpt H3K4me3 (Fig. 6B).
Pearson’s Correlation Matrices reveal distinct transcriptomic and epigenetic relationships following cocaine exposure in both sexes. (A) Pearson’s Correlation matrix evaluating transcriptomic and epigenetic relationships of Nr4a1 and Cartpt in the striatum of saline- and cocaine-injected males and (B) females. (C) Pearson’s Correlation matrix evaluating transcriptomic and epigenetic relationships of Nr4a1 and Cartpt in the hippocampus of saline- and cocaine-injected males and (D) females. (E) Pearson’s Correlation matrix evaluating transcriptomic and epigenetic relationships of Nr4a1 and Cartpt in the prefrontal cortex of saline- and cocaine-injected males and (F) females. For all Pearson’s Correlation matrices, red indicates a significant positive correlation (p ≤ 0.05) and blue indicates a significant negative correlation (p ≤ 0.05). A gray box indicates a correlation was not analyzed as it violates the rule of independence for Pearson’s Correlation.
In the HPC of saline-treated male mice we observed a negative correlation between (1) Nr4a1 mRNA levels (Exon 4/5) and Nr4a1 H3K27me3 and (2) Nr4a1 mRNA levels (Exon 4/5) and Cartpt H3K4me3 (Fig. 6C). In the HPC of saline-treated females we observed a positive correlation between (3) Nr4a1 H3K4me3 and Cartpt H3K4me3 and (4) Nr4a1 H3K27me3 and Cartpt H3K27me3 (Fig. 6D). In the HPC of males following cocaine, we observed a positive correlation between (5) Nr4a1 mRNA levels (Exon 4/5 and Exon 6/7), (6) Nr4a1 mRNA levels (Exon 4/5) and Cartpt H3K4me3, and (7) Cartpt mRNA levels and Nr4a1 K27/K4 bivalency (Fig. 6C). In the HPC of females following cocaine, we observed a positive correlation between (8) Nr4a1 mRNA levels (Exon 4/5 and Exon 6/7), (9) Nr4a1 mRNA levels (Exon 4/5) and Cartpt H3K4me3, (10) Nr4a1 H3K27me3 and Cartpt H3K4me3, and (11) Nr4a1 H3K27me3 and Cartpt H3K27me3 (Fig. 6D). We observed a negative correlation in the HPC of females following cocaine between (12) Cartpt mRNA levels and Cartpt H3K4me3 (Fig. 6D).
In the PFC of saline-treated males, we found a positive correlation between (1) mRNA levels of Nr4a1 (Exon 4/5) and H3K4me3/H3K27me3 at Nr4a1 and (2) H3K4me3 at Nr4a1 and H3K4me3 at Cartpt (Fig. 6E). In the PFC of saline-treated females, we found a positive correlation between (3) Nr4a1 mRNA levels (Exon 4/5 and Exon 6/7) as well as (4) Nr4a1 mRNA levels (Exon 6/7) and Nr4a1 K4/K27 bivalency (Fig. 6F). In the PFC of cocaine-injected males, we found multiple positive correlations including: (5) H3K4me3/H3K27me3 bivalency at Nr4a1 and H3K4me3 at Cartpt, (6) H3K4me3 at Nr4a1 and H3K4me3 at Cartpt, (7) Nr4a1 H3K4me3/H3K27me3 bivalency, (8) Nr4a1 mRNA levels (Exon 4/5 and Exon 6/7), and (9) Nr4a1 mRNA levels (Exon 6/7) and Cartpt H3K27me3 (Fig. 6E). In the PFC of females following cocaine, we observed both positive and negative correlations. We found that (10, 11) mRNA levels of Nr4a1 (Exon 4/5 and Exon 6/7) negatively correlate with H3K27me3/H3K4me3 bivalency at Nr4a1. Additionally, (12) mRNA levels of Nr4a1 (Exon 4/5) negatively correlate with mRNA levels of Cartpt (Fig. 6F). We observed a positive correlation between (13) mRNA levels of Cartpt and H3K4me3/H3K27me3 bivalency at Nr4a1 and (14) mRNA levels of Nr4a1 between its primer sets (Exon 4/5 and Exon 6/7) (Fig. 6F).
Discussion
There is emerging literature on the transcriptomic and epigenetic profiles of Nr4a1 and its regulation by cocaine. Despite this progress, no study to date has explored basal and cocaine-activated regulation of Nr4a1 in both males and females in any organism. Additionally, few studies have assessed bivalency in the brain35,36,37, and none, to our knowledge, have investigated cocaine regulation of bivalency. Here, we demonstrate that, following cocaine, Nr4a1 mRNA increases in female mouse STR and PFC, and does not change in male mouse STR, HPC, or PFC. We also find an increase in Nr4a1 K4&K27 promoter bivalency in the male STR following cocaine treatment. Finally, we show that the mRNA level of a putative Nr4a1 target, Cartpt, is activated in the male PFC.
Our results are the first to establish that there is a relationship between bivalency and cocaine in the brain, specifically we show induction of bivalency in the male STR. As this mechanism has not yet been linked to drugs of abuse, it provides a new mechanism of exploration for drug abuse research. Bivalent chromatin regulates DNA accessibility rather elegantly by transitioning euchromatin or heterochromatin to a poised, bivalent state that can quickly react to cellular cues. This flexibility is beneficial for cell differentiation32 and stress responses35, but may underlie detrimental instability of gene expression in the adult brain. Consequently, bivalency is implicated in transcriptional dysregulation observed in Huntington’s Disease48 as well as multiple types of cancer49,50,51,52. We hypothesize that bivalency may similarly underlie transcriptional dysregulation in drug addiction, in both males and females. Our data highlight that the interplay between bivalency and cocaine warrants further investigation.
The current study profiles both ventral and dorsal STR, as well as HPC and PFC, other brain regions associated with reward neuropathology. Both the ventral STR (NAc) and dorsal STR are involved in cocaine reinforcement, and the interconnectivity between these regions mediates cocaine-seeking behavior53. Additionally, pharmacological studies have shown that dopamine transporters in both the NAc and dorsal STR have the same affinity for cocaine binding54, and that the connectivity between dorsal STR and NAc is necessary for the regulation of dopamine to mediate cocaine’s reinforcing properties55. Our lab and others find that cocaine treatment increases Nr4a1 mRNA in the NAc in mixed sex populations38, as well as in males and females when assessed separately39. However, in order to analyze K4&K27 bivalency at the Nr4a1 promoter of a single mouse, it was necessary to isolate chromatin from combined STR and perform sequential ChIP. Both H3K4me3 and H3K27me3 are basally present at Nr4a1, with high enrichment for H3K4me3 in bulk tissue of the nucleus accumbens31 as well as in cell-type specific tissue of the combined striatum56, showing that combined STR and NAc have the same enrichment pattern. Additionally, both H3K4me3 and H3K27me3 are present at baseline at Cartpt, with high co-enrichment for both marks in bulk tissue of the nucleus accumbens31 and in cell-type specific tissue of the striatum56. Using sequential ChIP, we were able to identify the same patterns of enrichment at both modifications at baseline for Nr4a1 and Cartpt.
The strengths of the sequential ChIP approach in deciphering the histone code are limited by the technical requirement of bulk STR starting material, which obscures cell-type specific Nr4a1 regulation by cocaine and sex. The current literature contains complex and conflicting data on the expression pattern of Nr4a1 in direct pathway, Drd1 + , and indirect pathway, Drd2 + and/or A2a + , medium spiny neurons (MSNs)39,40,57,58. Studies suggest that Nr4a1 expression in Drd1 + MSNs, but not A2a + MSNs, functions in cocaine craving and relapse-like behavior. Drd1-specific signaling, such as the phosphoErk (pErk) cascade, increases in the NAc across incubation of cocaine craving59, and may regulate Nr4a1 in the Drd1 + MSN subtype40,57,59,60.
Importantly, the relationships with bivalency (H3K4me3/H3K27me3 vs H3K27me3/H3K4me3) do not always align at the level of a correlation. This is likely inherent to the biological mechanism of bivalency. Induction of H3K4me3/H3K27me3 bivalency is indicative of a transition from an activating to poised state (bivalency results in lower activation), while induction of H3K27me3/H3K4me3 bivalency is indicative of a repressive to poised state (bivalency results in higher activation). Additionally, we found that Nr4a1 mRNA levels did not always correspond with the canonical roles of H3K4me3, H3K27me3, and K4&K27 bivalency, in activating, repressing, and poising gene expression, respectively. For example, in the female PFC following cocaine treatment we observed concomitant increases in Nr4a1 mRNA and H3K27me3 at Nr4a1. We thus applied correlational analyses to examine these relationships more comprehensively. In the male STR, we found a positive correlation between Nr4a1 K4/K27 bivalency and Cartpt mRNA levels. Given that Nr4a1 is a putative activator of Cartpt expression38, this finding went against our expectation that Nr4a1 gene poising would be negatively correlated with Cartpt mRNA levels. Additionally, given the lack of transcriptomic or the epigenomic regulation of Nr4a1 in the male PFC and the vast increase in Cartpt mRNA levels, we hypothesize that Cartpt may additionally be regulated by other distinct transcription factors and/or HPTMs following cocaine. This is supported by our correlational data that shows minimal overlap in the transcriptomic and epigenetic profiles of Nr4a1 and Cartpt in either drug conditions in males or females or in any of the three regions examined. Another reason that some of the epigenetic and mRNA correlations may not have aligned could be due to the fact that our work focused solely on investigating Nr4a1 bivalency at the promoter. Epigenetic investigation of other regulatory regions, such as enhancers, which influence transcription via signaling to promoters61, could shed further light onto epigenetic control of gene regulation. For example, H3K4me3 is predominantly located at promoters, although it has been also been identified as a modifier of enhancers62. Future studies will expand investigation of bivalency beyond promoters to other genomic regulatory regions, provided such regions have been identified and functionally validated.
Based on our findings and the literature, we also hypothesize that additional HPTMs, such as acetylation, regulate expression of Nr4a1 and Cartpt. Intracranial HPC injection of the histone deacetylase (HDAC) inhibitor, Trichostatin A, increases Nr4a1 mRNA and protein levels63, as does HDAC3 inhibition in the dorsal HPC64. In cell lines, Nr4a1 expression is regulated by the recruitment of the histone acetyltransferase p300 or HDAC165. It is interesting to consider that changes in STR histone acetylation, and methylation to a certain extent, are sensitive to the duration of cocaine treatment (acute versus chronic) and time since the last injection21, which may reflect the rapid turnover of acetylation, especially compared to methylation47. Although acetylation turnover is more rapid than methylation, we hypothesize that repeating the endpoints outlined in this study at a different time point, such as 30 min after the final injection versus 24 h, could prompt different levels of H3K4me3, H3K27me3, and K4&K27 bivalency. Specifically, we expect an initial decrease in H3K27me3 and K4&K27 bivalency (< 30 min following cocaine) followed by an increase in H3K27me3 and K4&K27 bivalency at later timepoints (> 24 h following cocaine).
Nonetheless, our findings encourage future studies to investigate combinatorial HPTMs beyond K4&K27 bivalency. For example, H3K27me3 and H3K27ac bivalency are well described in the literature and represent a tractable starting point for further exploration66,67. We hypothesize that cocaine-induced depletion of H3K27ac acts synergistically with increased K4&K27 bivalency to inhibit the activation of Nr4a1 expression in males. We speculate that the persistence of H3K27me3 is sex-dependent, and thus H3K27me3 is likely bivalent with other HPTMs as well. This hypothesis is based in part on the sexually dimorphic expression of H3K27me3-demethylase, Kdm6a, in rodent brain68,69,70. As Kdm6a is X-linked71, it may mediate sex-specific changes in HPTM expression upon cocaine exposure to drive epigenetic changes at Nr4a1 and other cocaine-responsive loci. It is possible that sexually dimorphic molecular changes underlie sexually dimorphic behaviors. For example, male rodents are less responsive to drug-conditioned stimuli5,6 and acquire cocaine-self administration at a slower rate than female rodents7,8. Future studies will explore additional combinatorial HPTMs with H3K27me3 to probe these hypotheses.
Additionally, new methods are being developed to allow for future studies to conduct sequential ChIP profiling of small amounts of input material which will allow for profiling of more specific brain regions and cell-types. For example, the recently developed multi-CUT&Tag72 allows for mapping HPTM co-localization in the same cell, and single-cell multi-CUT&Tag72 can probe for bivalency in specific cell types. While single-cell CUT&Tag has been carried out in mouse brain to profile individual HPTMs73, these multi-CUT&Tag methods that uncover bivalency have yet to be applied in brain tissue as far as we are aware. Finally, in the current study we test and support the hypothesis that K4&K27 bivalency is increased at Nr4a1 in male mice following cocaine treatment, concomitant with increased H3K27me3 deposition. To test the sufficiency of loss of H3K27me3 in increased Nr4a1 mRNA, we can apply in vivo epigenetic editing to exogenously enrich Nr4a1 H3K27me3 in the presence of cocaine. This approach may also shed light onto the role of H3K27ac, given that dCas9-FOG1 interacts with the NuRD complex to cause HDAC1/2-mediated removal of H3K27ac74.
In closing, despite epidemiological evidence that cocaine abuse afflicts both men and women1, the underlying neurobiology of how cocaine impacts each sex is not fully understood. This is problematic as a growing body of knowledge demonstrates that cocaine causes distinct neuroadaptations for males and females10. The inclusion of both sexes within research investigating reward neurobiology is thus vital and necessary to the goal of treating addiction disorders. Future studies should include both males and females to understand how cocaine causes sex-specific neuroadaptations and for uncovering sexually dimorphic mechanisms that could lead to innovations for therapeutic intervention.
Methods and materials
Animals
Male and female mice on the C57BL/6 J background were used in this study. Mice were housed under a 12-h light–dark cycle at 23 °C with access to food and water ad libitum. Note that in compliance with ethical standards to minimize the use of mice, the mice used in this study were cre-negative offspring of R26-CAG-LSL-Sun1-sfGFP;A2a-cre and LSL-Sun1-sfGFP;Drd1-cre, generated for a separate study. All animal procedures were conducted in accordance with the National Institutes of Health Guidelines as well as the Association for Assessment and Accreditation of Laboratory Animal Care. Ethical and experimental considerations were approved by the Institutional Animal Care and Use Committee of The University of Pennsylvania. All experiments complied with ARRIVE guidelines.
Investigator-administered cocaine
Male and female mice were handled daily for three days prior to cocaine administration. Following handling, mice were given a daily cocaine hydrochloride intraperitoneal injection (20 mg/kg dissolved in 0.9% saline) or 0.9% saline injection for 10 days38. Saline injected mice were used as the control group in all experiments. Male and female mice were injected with cocaine at the same time. Mice were sacrificed one day after the final injection. STR, HPC, and PFC tissue was immediately collected from each animal (Fig. 1A,B) and stored at − 80 °C until processing.
S3EQ nuclear and cytosolic fraction separation
S3EQ45 was conducted as previously published with modified buffer volume. Tissue samples were homogenized in 350 μL cell lysis buffer (10 mM Tris–HCl (pH 8.0), 10 mM NaCl, 3 mM MgCl2 and 0.5% NP-40 in H2O) and spun for five minutes (1500 × g, 4 °C). The nuclei-containing pellet and RNA-containing cytosolic supernatant were separated and subjected to sequential ChIP or RNA extraction, respectively (Fig. 1C).
Single and sequential chromatin immunoprecipitation (ChIP) and ChIP-PCR (qCHIP)
Prior to initiating the protocol, the following solutions were prepared as previously described45,46: BSA Blocking Buffer (0.5% Bovine serum albumin in 1 × PBS), Dilution Buffer (16.7 mM Tris–HCl (pH 8.0), 1.1% Triton-X 100, 0.01% SDS, 167 mM NaCl, 1.2 mM EDTA in H2O), Nuclear Lysis Buffer (50 mM Tris–HCl (pH 8.0), 5 mM EDTA, 1% SDS in H2O), Wash Buffer 1 (20 mM Tris–HCl pH 8.0, 150 mM NaCl, 2 mM EDTA, 1% Triton X-100, 0.1% SDS in H2O), Wash Buffer 2 (20 mM Tris–Cl pH 8.0, 500 mM NaCl, 2 mM EDTA, 1% Triton X-100, 0.1% SDS in H2O), Wash Buffer 3 (250 mM LiCl, 10 mM Tris–HCl pH 8.0, 1% sodium deoxycholate, 1 mM EDTA, 1% IGEPAL CA-630 in H2O), TE Buffer (10 mM Tris–HCl pH 8.0, 1 mM EDTA in H2O), Elution Buffer (0.1 M NaHCO3, 1% SDS in H2O).
M280 Sheep anti-Rabbit Dynabeads (Invitrogen, 11204D) were washed three times in BSA blocking buffer. The beads were bound to either H3K4me3 antibody (Rabbit Polyclonal, EMD Millipore, 07–473) or H3K27me3 antibody (Rabbit Polyclonal, EMD Millipore, 07–449) in 2X bead volume of dilution buffer and rotated for six hours (4 °C). Nuclei obtained from S3EQ were resuspended in PBS and fixed with 1% formaldehyde for eight minutes (350 rpm, 22 °C). 100 μL 1 M glycine was added and samples were rocked for five minutes to stop cross-linking (500 rpm, 22 °C). Samples were then centrifuged for five minutes (5500 × g, 4 °C) and the supernatant was discarded. The pellet was resuspended in 200 μL nuclear lysis buffer, transferred to TPX tubes, (Diagenode, C30010010) and incubated on ice for ten minutes. Samples were then sonicated in a Bioruptor® (Diagenode) for two runs (high setting, 30 s on, 30 s off, 15 cycles). Dilution buffer was added to reach 1000 μL. 10% input (relative to each IP) was collected from each sample for normalization later used in qChIP analysis. The remaining diluted chromatin was combined with antibody-bound beads (1.2 μL bead-antibody slurry/1 μL chromatin), dilution buffer, and placed on a rotator (O/N, 4 °C). Samples were then washed with 1 mL of ice-cold Wash Buffer 1, Wash Buffer 2, Wash Buffer 3, and TE Buffer. Samples were rotated for five minutes during each wash (RT). Following washes, samples were rocked with 200 μL of elution buffer for 20 min (500 rpm, RT), centrifuged for three minutes (14,000 × g, RT), and placed on a magnet. The supernatant from each sample was divided in half and transferred to fresh tubes, each containing 100 μL of elution buffer. One tube was collected as the single ChIP sample.
The remaining tube was subjected to a second immunoprecipitation (sequential ChIP samples), in which each sample was subjected to antibody-bound beads that were bound to the reciprocal antibody (i.e., H3K4me3 ChIP samples were bound to H3K27me3 antibody-bound beads) and placed on a rotator (O/N, 4 °C). The sequential ChIP samples were processed according to the single ChIP protocol above. Single and sequential ChIP samples were then incubated for 4 h (65 °C) following the addition of 8 μL 5 M NaCl. Proteinase K (0.002 mg) was added, and incubation continued for two hours (65 °C). Samples were heat inactivated (15 min, 78 °C) and DNA was purified using the QiaAmp DNA Micro kit (Qiagen, 56304). qChIP was conducted as previously published38. Briefly, qChIP was run using Power SYBR™ Green PCR Master Mix (Life Technologies). The primer sequences that span − 100 to + 100bps of target promoter regions used to assess the presence of HPTMs38 are noted in Supplemental Table 1.
RNA Extraction and qPCR
The cytosolic fraction following S3EQ was mixed with 600 μL RLT buffer (Qiagen) and RNA was extracted using the RNAeasy Micro Kit (Qiagen, 74004). RNA concentration was obtained using a Qubit 4 Fluorometer and RNA HS Assay Kit (Invitrogen). cDNA was synthesized using the Bio-Rad iScript™ cDNA Synthesis Kit (Bio-Rad, 1708890). qPCR was conducted using Power SYBR™ Green PCR Master Mix (Life Technologies). The primer sequences used to assess mRNA levels are noted in Supplemental Table 2.
Pearson correlation analysis of qCHIP and qPCR
Pearson correlation analysis was used to assess the relationship between K4&K27 bivalent Nr4a1 promoter, Nr4a1 or Cartpt mRNA levels, and Nr4a1 or Cartpt monovalent H3K4me3 or H3K27me3. Scripts were written and executed in R v4.1.1. First, all values corresponding to relevant endpoints were compiled for each sex (male, female) and brain region (STR, HPC, PFC) combination. For a given combination, the dataset was further separated based on treatment (saline, cocaine). Analyses were conducted with input values (fold change) from qChIP and qPCR data. An endpoint (qChIP or qPCR) was excluded if it lacked at least three observations in both the saline and cocaine datasets. An all-against-all Pearson’s correlation matrix was generated for the saline and cocaine datasets separately using the base R cor (method = ”pearson”) function. A correlation heatmap was generated from the Pearson correlation matrix using the corrplot() function from the corrplot package.
Statistical analyses
Statistical tests used reflect the experimental design75. Specifically, while male and female mice were treated with cocaine in a single cohort, the tissue for each sex and each brain region was processed and analyzed separately due to the technical limitations associated with processing large numbers of sample. We therefore did not apply ANOVA to analyze sex or region differences. G*Power: Statistical Power Analyses Software76 (v3.1) was used to determine sample size. Final sample size was reduced during experimental execution (from n = 10 to n = 8 per group) due to previous empirical experimental endpoints and technical limitations. The Grubbs test was used to identify all outliers after an entire data set was complete and before any other statistical tests were applied (alpha = 0.05). Student’s t-tests were used for qChIP and qPCR analyses as these experiments directly compared one factor (drug exposure) from two groups. Sex was not considered a variable within any analyses due to experimental execution. F-tests of variance were conducted in every analysis and Welch’s correction was utilized when outliers were removed and when variances differed between groups. These data are shown as mean ± SEM. Graphpad Prism V9.3 was used for qChIP and qPCR analyses. qChIP data was analyzed by comparing CT values of each experimental group versus the control group following normalization of each sample’s input using the ∆∆Ct method77,78. Bivalency at Cartpt was not assessed, as not all sequential-immunoprecipitated samples reached minimum signal intensity for qChIP (CT values were ≥ 38). qPCR data was analyzed by comparing CT values of each experimental group versus the control group following normalization of a housekeeping gene (Gapdh) using the ∆∆Ct method77,78. All experiments were carried out one to two times, and data replication was observed in all instances of repeated experiments. Additionally, every qChIP and qPCR dataset had a technical replicate conducted by an independent investigator to ensure the validity of the data. All data shown in final analysis come from the first experiment of replicates, and the data specifically comes from the technical replicate conducted by the first author. Regarding visual data representation, each data point in all graphs represent one animal (Figs. 2, 3, 4, 5). For Pearson correlation analysis, the p-value matrix of all correlations corresponding to each comparison was extracted using the two-sided cor.mtest(method = ”pearson”) function from the corrplot package with a 95% confidence interval. This statistical test for Pearson’s product-moment coefficient was calculated following a t-distribution (\(t=\frac{r\sqrt{\mathrm{n}-2}}{\sqrt{1-{r}^{2}}},\) where n = number of observations and r = calculated Pearson correlation coefficient) with degrees of freedom equal to length (number of complete observations)–2. Heatmap tiles were shaded red or blue only when satisfying a p value ≤ 0.05, where the null hypothesis represents r = 0 (no correlation), using p.mat = p.mat_for_comparison and sig.level = 0.05 within the corrplot() function. Heatmap tiles were shaded gray when analysis was not conducted, as analysis would violate the rule of independence79. All scripts are available at: (https://github.com/HellerLAbeats/Nr4a1_bivalency).
Data availability
Raw data or further methodological information from the current study are available upon reasonable request from the corresponding author.
References
Substance Abuse and Mental Health Services Administration. Key substance use and mental health indicators in the United States: Results from the 2018 National Survey on Drug Use and Health. HHS Publ. No. PEP19–5068, NSDUH Ser. H-54 (2018) doi:https://doi.org/10.1016/j.drugalcdep.2016.10.042.
Kasperski, S. J. et al. College students’ use of cocaine: Results from a longitudinal study. Addict. Behav. 36, 408–411 (2011).
Becker, J. B. & Hu, M. Sex differences in drug abuse. Front. Neuroendocrinol. 29, 36–47 (2008).
Becker, J. B. & Koob, G. F. Sex differences in animal models: Focus on addiction. Pharmacol. Rev. 68, 242–263 (2016).
Zakharova, E., Wade, D. & Izenwasser, S. Sensitivity to cocaine conditioned reward depends on sex and age. Pharmacol. Biochem. Behav. 92, 131–134 (2009).
Russo, S. J. et al. Sex differences in the conditioned rewarding effects of cocaine. Brain Res. 970, 214–220 (2003).
Lynch, W. J. & Carroll, M. E. Sex differences in the acquisition of intravenously self-administered cocaine and heroin in rats. Psychopharmacology 144, 77–82 (1999).
Hu, M., Crombag, H. S., Robinson, T. E. & Becker, J. B. Biological basis of sex differences in the propensity to self-administer cocaine. Neuropsychopharmacology 29, 81–85 (2004).
López, A. J. et al. Cocaine self-administration induces sex-dependent protein expression in the nucleus accumbens. Commun. Biol. 4, 883 (2021).
Carroll, M. E. & Lynch, W. J. How to study sex differences in addiction using animal models. Addict. Biol. 21, 1007–1029 (2016).
Walker, D. M. & Nestler, E. J. Neuroepigenetics and addiction. Handb. Clin. Neurol. https://doi.org/10.1016/B978-0-444-64076-5.00048-X (2018).
Walker, D. M. et al. Cocaine self-administration alters transcriptome-wide responses in the brain’s reward circuitry. Biol. Psychiatry 84, 867–880 (2018).
Pierce, R. C. et al. Environmental, genetic and epigenetic contributions to cocaine addiction. Neuropsychopharmacology 43, 1471–1480 (2018).
De Sa Nogueira, D., Merienne, K. & Befort, K. Neuroepigenetics and addictive behaviors: Where do we stand?. Neurosci. Biobehav. Rev. 106, 58–72 (2019).
Robison, A. J. & Nestler, E. J. Transcriptional and epigenetic mechanisms of addiction. Nat. Rev. Neurosci. 12, 623–637 (2011).
Nestler, E. J. & Lüscher, C. The molecular basis of drug addiction: Linking epigenetic to synaptic and circuit mechanisms. Neuron 102, 48–59 (2019).
Walker, D. M., Cates, H. M., Heller, E. A. & Nestler, E. J. Regulation of chromatin states by drugs of abuse. Curr. Opin. Neurobiol. 30, 112–121 (2015).
Kumar, A. et al. Chromatin remodeling is a key mechanism underlying cocaine-induced plasticity in striatum. Neuron 48, 303–314 (2005).
Russo, S. J. & Nestler, E. J. The brain reward circuitry in mood disorders. Nat. Rev. Neurosci. https://doi.org/10.1038/nrn3381 (2013).
Koob, G. F. & Volkow, N. D. Neurobiology of addiction: A neurocircuitry analysis. The Lancet Psychiatry 3, 760–773 (2016).
Jordi, E. et al. Differential effects of cocaine on histone posttranslational modifications in identified populations of striatal neurons. Proc. Natl. Acad. Sci. 110, 9511–9516 (2013).
Zhou, Z., Yuan, Q., Mash, D. C. & Goldman, D. Substance-specific and shared transcription and epigenetic changes in the human hippocampus chronically exposed to cocaine and alcohol. Proc. Natl. Acad. Sci. 108, 6626–6631 (2011).
Gajewski, P. A. et al. Epigenetic regulation of hippocampal fosb expression controls behavioral responses to cocaine. J. Neurosci. 39, 8305–8314 (2019).
De Sa Nogueira, D. et al. Hippocampal cannabinoid 1 receptors are modulated following cocaine self-administration in male rats. Mol. Neurobiol. 59, 1896–1911 (2022).
Sadri-Vakili, G. et al. Cocaine-induced chromatin remodeling increases brain-derived neurotrophic factor transcription in the Rat Medial prefrontal cortex, which alters the reinforcing efficacy of cocaine. J. Neurosci. https://doi.org/10.1523/jneurosci.2328-10.2010 (2010).
Zhang, Y. X. et al. The histone demethylase KDM6B in the medial prefrontal cortex epigenetically regulates cocaine reward memory. Neuropharmacology 141, 113–125 (2018).
Jenuwein, T. & Allis, C. D. Translating the histone code. Science 293, 1074–1080 (2001).
Strahl, B. D. & Allis, C. D. The language of covalent histone modifications. Nature 403, 41–45 (2000).
Sadri-Vakili, G. Cocaine triggers epigenetic alterations in the corticostriatal circuit. Brain Res. 1628, 50–59 (2015).
Werner, C. T., Altshuler, R. D., Shaham, Y. & Li, X. Epigenetic mechanisms in drug relapse. Biol. Psychiatry 89, 331–338 (2021).
Feng, J. et al. Chronic cocaine-regulated epigenomic changes in mouse nucleus accumbens. Genome Biol. 15, R65 (2014).
Bernstein, B. E. et al. A bivalent chromatin structure marks key developmental genes in embryonic stem cells. Cell 125, 315–326 (2006).
Voigt, P., Tee, W.-W. & Reinberg, D. A double take on bivalent promoters. Genes Dev. 27, 1318–1338 (2013).
Kumar, D., Cinghu, S., Oldfield, A. J., Yang, P. & Jothi, R. Decoding the function of bivalent chromatin in development and cancer. Genome Res. 31, 2170–2184 (2021).
Nasca, C. et al. Role of the astroglial glutamate exchanger xCT in ventral hippocampus in resilience to stress. Neuron 96, 402–413 (2017).
Yan, L. et al. Epigenomic landscape of human fetal brain, heart, and liver. J. Biol. Chem. 291, 4386–4398 (2016).
Weiner, A. et al. Co-ChIP enables genome-wide mapping of histone mark co-occurrence at single-molecule resolution. Nat. Biotechnol. 34, 953–961 (2016).
Carpenter, M. D. et al. Nr4a1 suppresses cocaine-induced behavior via epigenetic regulation of homeostatic target genes. Nat. Commun. 11, 504 (2020).
Savell, K. E. et al. A dopamine-induced gene expression signature regulates neuronal function and cocaine response. Sci. Adv. 6, eaba4221 (2020).
Cirnaru, M.-D. et al. Nuclear receptor nr4a1 regulates striatal striosome development and dopamine D 1 receptor signaling. Eneuro 6, ENEURO.0305-19.2019 (2019).
Bourhis, E. et al. The transcription factors Nur77 and retinoid X receptors participate in amphetamine-induced locomotor activities. Psychopharmacology 202, 635–648 (2009).
Volpicelli, F. et al. Bdnf gene is a downstream target of Nurr1 transcription factor in rat midbrain neurons in vitro. J. Neurochem. 102, 441–453 (2007).
Kadkhodaei, B. et al. Nurr1 is required for maintenance of maturing and adult midbrain dopamine neurons. J. Neurosci. 29, 15923–15932 (2009).
Bridi, M. S., Hawk, J. D., Chatterjee, S., Safe, S. & Abel, T. Pharmacological activators of the NR4A nuclear receptors enhance LTP in a CREB/CBP-dependent manner. Neuropsychopharmacology 42, 1243–1253 (2017).
Xu, S. J. & Heller, E. A. Single sample sequencing (S3EQ) of epigenome and transcriptome in nucleus accumbens. J. Neurosci. Methods 308, 62–73 (2018).
Peng, G.-H. & Chen, S. Double chromatin immunoprecipitation: Analysis of target co-occupancy of retinal transcription factors. In Methods in Molecular Biology (eds Weber, B. H. F. & Langmann, T.) 311–328 (Humana Press, 2012). https://doi.org/10.1007/978-1-62703-080-9_22.
Barth, T. K. & Imhof, A. Fast signals and slow marks: The dynamics of histone modifications. Trends Biochem. Sci. 35, 618–626 (2010).
Zsindely, N. & Bodai, L. Histone methylation in Huntington’s disease: Are bivalent promoters the critical targets?. Neural Regen. Res. 13, 1191 (2018).
Rodriguez, J. et al. Bivalent domains enforce transcriptional memory of DNA methylated genes in cancer cells. Proc. Natl. Acad. Sci. 105, 19809–19814 (2008).
Hall, A. W. et al. Bivalent chromatin domains in glioblastoma reveal a subtype-specific signature of glioma stem cells. Cancer Res. 78, 2463–2474 (2018).
Curry, E. et al. Genes predisposed to DNA hypermethylation during acquired resistance to chemotherapy are identified in ovarian tumors by bivalent chromatin domains at initial diagnosis. Cancer Res. 78, 1383–1391 (2018).
Bernhart, S. H. et al. Changes of bivalent chromatin coincide with increased expression of developmental genes in cancer. Sci. Rep. 6, 37393 (2016).
Belin, D. & Everitt, B. J. Cocaine seeking habits depend upon dopamine-dependent serial connectivity linking the ventral with the dorsal striatum. Neuron 57, 432–441 (2008).
Boja, J. W. & Kuhar, M. J. [3H]cocaine binding and inhibition of [3H]dopamine uptake is similar in both the rat striatum and nucleus accumbens. Eur. J. Pharmacol. 173, 215–217 (1989).
Veeneman, M. M. J., Damsteegt, R. & Vanderschuren, L. J. M. J. The nucleus accumbens shell and the dorsolateral striatum mediate the reinforcing effects of cocaine through a serial connection. Behav. Pharmacol. 26, 193–199 (2015).
Carpenter, M. D. et al. Cell-type specific profiling of histone post-translational modifications in the adult mouse striatum. bioRxiv 17, 11 (2022).
Davis, M. I. & Puhl, H. L. Nr4a1-eGFP is a marker of striosome-matrix architecture, development and activity in the extended striatum. PLoS ONE 6, e16619 (2011).
Lobo, M. K., Karsten, S. L., Gray, M., Geschwind, D. H. & Yang, X. W. FACS-array profiling of striatal projection neuron subtypes in juvenile and adult mouse brains. Nat. Neurosci. 9, 443–452 (2006).
Edwards, S., Bachtell, R. K., Guzman, D., Whisler, K. N. & Self, D. W. Emergence of context-associated GluR1 and ERK phosphorylation in the nucleus accumbens core during withdrawal from cocaine self-administration. Addict. Biol. 16, 450–457 (2011).
Salinas, A. G., Davis, M. I., Lovinger, D. M. & Mateo, Y. Dopamine dynamics and cocaine sensitivity differ between striosome and matrix compartments of the striatum. Neuropharmacology 108, 275–283 (2016).
Calo, E. & Wysocka, J. Modification of enhancer chromatin: What, how, and why?. Mol. Cell 49, 825–837 (2013).
Pekowska, A. et al. H3K4 tri-methylation provides an epigenetic signature of active enhancers. EMBO J. 30, 4198–4210 (2011).
Hawk, J. D. et al. NR4A nuclear receptors support memory enhancement by histone deacetylase inhibitors. J. Clin. Invest. 122, 3593–3602 (2012).
Kwapis, J. L. et al. Epigenetic regulation of the circadian gene Per1 contributes to age-related changes in hippocampal memory. Nat. Commun. 9, 3323 (2018).
Kang, S.-A. et al. Regulation of Nur77 protein turnover through acetylation and deacetylation induced by p300 and HDAC1. Biochem. Pharmacol. 80, 867–873 (2010).
Voigt, P. et al. Asymmetrically modified nucleosomes. Cell 151, 181–193 (2012).
Du, Y. et al. Structural mechanism of bivalent histone H3K4me3K9me3 recognition by the spindlin1/C11orf84 complex in rRNA transcription activation. Nat. Commun. 12, 949 (2021).
Cabrera Zapata, L. E. et al. X-linked histone H3K27 demethylase Kdm6a regulates sexually dimorphic differentiation of hypothalamic neurons. Cell. Mol. Life Sci. 78, 7043–7060 (2021).
Phillips, R. A. et al. An atlas of transcriptionally defined cell populations in the rat ventral tegmental area. Cell Rep. 39, 110616 (2022).
Jaric, I., Rocks, D., Greally, J. M., Suzuki, M. & Kundakovic, M. Chromatin organization in the female mouse brain fluctuates across the oestrous cycle. Nat. Commun. 10, 2851 (2019).
Arnold, A. P. et al. The importance of having two X chromosomes. Philos. Trans. R. Soc. B. Biol. Sci. 371, 20150113 (2016).
Gopalan, S., Wang, Y., Harper, N. W., Garber, M. & Fazzio, T. G. Simultaneous profiling of multiple chromatin proteins in the same cells. Mol. Cell 81, 4736-4746.e5 (2021).
Bartosovic, M., Kabbe, M. & Castelo-Branco, G. Single-cell CUT&Tag profiles histone modifications and transcription factors in complex tissues. Nat. Biotechnol. 39, 825–835 (2021).
O’Geen, H. et al. dCas9-based epigenome editing suggests acquisition of histone methylation is not sufficient for target gene repression. Nucleic Acids Res. 45, 9901–9916 (2017).
Nieuwenhuis, S., Forstmann, B. U. & Wagenmakers, E.-J. Erroneous analyses of interactions in neuroscience: A problem of significance. Nat. Neurosci. 14, 1105–1107 (2011).
Faul, F., Erdfelder, E., Lang, A.-G. & Buchner, A. G*Power 3: A flexible statistical power analysis program for the social, behavioral, and biomedical sciences. Behav. Res. Methods 39, 175–191 (2007).
Livak, K. J. & Schmittgen, T. D. Analysis of relative gene expression data using real-time quantitative PCR and the 2−ΔΔCT method. Methods 25, 402–408 (2001).
Yuan, J. S., Reed, A., Chen, F. & Stewart, C. N. Statistical analysis of real-time PCR data. BMC Bioinformatics 7, 85 (2006).
H. J. Motulsky. GraphPad statistics guide. http://www.graphpad.com/guides/prism/7/statistics/index.htm.
Acknowledgements
Financial support was provided by NIH-NIDA Avenir Director’s Pioneer Award (E.A.H, DP1 DA044250), NIDA Research Project Grant (E.A.H, 1R01DA052465-01A1), Pilot Award from the Epigenetics Institute at the University of Pennsylvania, T32 Predoctoral Training Grant in Addiction (D.K.F. NIDA T32DA028874), Penn Undergraduate Research & Mentoring program (C.H.) and The Endowed Dean’s Research Award at the University of Pennsylvania (C.H.). We thank Dr. Heath Schmidt (University of Pennsylvania) for his continuous support and meticulous review of the data and manuscript. Figure 1 was created with Biorender.com. This paper has been approved for preprint by BioRxiv and can be accessed at: https://www.biorxiv.org/content/10.1101/2022.04.22.489203v1. The paper has been edited since the preprint was available. The authors report no biomedical financial interests or potential conflicts of interest.
Author information
Authors and Affiliations
Contributions
D.K.F. and E.A.H. wrote the manuscript, D.K.F., K.S.K, C.H., E.A.H. designed experiments, D.K.F., C.H., M.T.W. performed injections and tissue collection, D.K.F. and C.H. performed all molecular experiments, D.K.F. and K.S.K analyzed all data, K.S.K performed all computational analyses. All authors read, edited, and approved the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Fischer, D.K., Krick, K.S., Han, C. et al. Cocaine regulation of Nr4a1 chromatin bivalency and mRNA in male and female mice. Sci Rep 12, 15735 (2022). https://doi.org/10.1038/s41598-022-19908-9
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-022-19908-9
This article is cited by
-
Cell-type specific profiling of histone post-translational modifications in the adult mouse striatum
Nature Communications (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.