Abstract
The heterogeneity of MRI is one of the major reasons for decreased performance of a radiomics model on external validation, limiting the model’s generalizability and clinical application. We aimed to establish a generalizable radiomics model to predict meningioma grade on external validation through leveraging Cycle-Consistent Adversarial Networks (CycleGAN). In this retrospective study, 257 patients with meningioma were included in the institutional training set. Radiomic features (n = 214) were extracted from T2-weighted (T2) and contrast-enhanced T1 (T1C) images. After radiomics feature selection, extreme gradient boosting classifiers were developed. The models were validated in the external validation set consisting of 61 patients with meningiomas. To reduce the gap in generalization associated with the inter-institutional heterogeneity of MRI, the smaller image set style of the external validation was translated into the larger image set style of the institutional training set using CycleGAN. On external validation before CycleGAN application, the performance of the combined T2 and T1C models showed an area under the curve (AUC), accuracy, and F1 score of 0.77 (95% confidence interval 0.63–0.91), 70.7%, and 0.54, respectively. After applying CycleGAN, the performance of the combined T2 and T1C models increased, with an AUC, accuracy, and F1 score of 0.83 (95% confidence interval 0.70–0.97), 73.2%, and 0.59, respectively. Quantitative metrics (by Fréchet Inception Distance) showed that CycleGAN can decrease inter-institutional image heterogeneity while preserving predictive information. In conclusion, leveraging CycleGAN may be helpful to increase the generalizability of a radiomics model in differentiating meningioma grade on external validation.
Similar content being viewed by others
Introduction
Meningiomas are the most common primary intracranial neoplasms in adults, accounting for approximately one-third of all intracranial tumors1. The majority of meningiomas (80%) are classified as low-grade (World Health Organization [WHO] grade 1; benign) and have an indolent clinical course2. On the other hand, high-grade (WHO grade 2 or 3; atypical or anaplastic) tumors have an aggressive biological behavior, a tendency to recur, and a poor prognosis2. The standard management typically involves surgical resection, and adjuvant radiation therapy is often recommended for high-grade meningiomas3. Therefore, developing a noninvasive generalizable model based on MRI to predict meningioma grade may assist clinical decision making by providing information on treatment planning, including surgical resection strategy4, and care of incidentally detected meningiomas in asymptomatic patients3.
MRI is the key imaging modality for diagnosis and characterization of meningioma and treatment decision5. Several studies applying radiomics, which translates radiological images into high-dimensional mineable imaging data6, have shown promising results in predicting meningioma grade7,8,9,10,11,12. However, majority of them did not perform external validation7. Those studies that performed external validation showed drastically decreased performance in external validation10,11,12, which limits the real-world application of radiomics models. Given that the objective of a prediction model is to predict outcomes in future patients, not to classify previously described characteristics, model generalizability on external validation is critical for model implementation13.
The inter-institutional heterogeneity of MRI protocol is a major reason for decreased performance of a radiomics model in the external validation stage13. Although consensus recommendations for standardized imaging protocol are established in brain tumors such as glioma or brain metastases14,15, consensus imaging protocol for meningiomas is currently lacking, which leads to substantial inter-institutional heterogeneity.
Recently, an approach based on the unpaired image-to-image translation using Cycle-Consistent Adversarial Networks (CycleGAN), a style transfer technique, has been suggested as a promising strategy to overcome poor model performance when dealing with external images16. CycleGAN can transfer the style of the image, while preserving the semantic information within the data16. The approaches using CycleGAN show superior visual similarities between image domains both quantitatively and qualitatively compared with other normalization methods and eliminate manual preparation of the representative reference image because they learn the whole image distribution17,18. We hypothesized that this approach can be applied to convert heterogeneous MRIs and lead to improved performance of a radiomics model to predict meningioma grade on external validation17,18. Thus, the objective of this study was to establish a generalizable radiomics model to predict meningioma grade on external validation through leveraging CycleGAN.
Materials and methods
Patient population
The Yonsei University Institutional Review Board approved this retrospective study and waived the need for obtaining informed patient consent. All methods were performed in accordance with the relevant guidelines and regulations. We identified 297 patients who were pathologically confirmed as having meningioma and underwent baseline conventional MRI between February 2008 and September 2018 in the institutional dataset. Patients with 1) missing MRI sequences or inadequate image quality (n = 17), 2) a previous history of surgery (n = 15), 3) a history of tumor embolization or gamma knife surgery before MRI exam (n = 5), and 4) an error in image processing (n = 2) were excluded. A total of 257 patients (low-grade, 162; high-grade, 95) were enrolled in the institutional cohort.
Identical inclusion and exclusion criteria were applied to identify 62 patients (low-grade, 47; high-grade, 15) from Ewha Mokdong University Hospital between January 2016 and December 2018 for external validation of the model. Patient flowchart is shown in Fig. S1.
Pathological diagnosis
Pathological diagnosis was performed by neuropathologists, according to the WHO criteria19. The criteria for atypical meningioma (WHO grade 2) comprised 4–19 mitoses per 10 high-power fields, the presence of brain invasion, or the presence of at least three of the following features: “sheet-like” growth, hypercellularity, spontaneous necrosis, large and prominent nucleoli, and small cells. The criteria for anaplastic meningioma (WHO grade 3 comprised frank anaplasia (histology resembling carcinoma, sarcoma, or melanoma) or elevated mitoses (> 20 mitoses per 10 high-power fields)19.
MRI protocol
In the institutional training dataset, patients were scanned on 3.0 Tesla MRI units (Achieva or Ingenia; Philips Medical Systems). Imaging protocols included T2-weighted (T2) and contrast-enhanced T1-weighted imaging (T1C). T1C images were acquired after administration of 0.1 mL/kg of gadolinium-based contrast material (Gadovist; Bayer).
In the external validation sets, patients were scanned on 1.5 or 3.0 Tesla MRI units (Avanto; Siemens, or Achieva; Philips Medical Systems), including T2 and T1C images. T1C images were acquired after administration of 0.1 mL/kg of gadolinium-based contrast material (Dotarem; Guerbert, or Gadovist; Bayer). Substantial variation existed between the acquisition parameters for T2 and T1C among the various MRI units between the institutional and external validation sets and reflected the heterogeneity of meningioma imaging data in clinical practice (Supplementary Table 1).
Image preprocessing and radiomics feature extraction
Image resampling to 1-mm isovoxels, low-frequency intensity non-uniformity correction by the N4 bias algorithm, and co-registration of T2 images to T1C images were performed using Advanced Normalization Tools (ANTs)20. After skull stripping by Multi-cONtrast brain STRipping (MONSTR)21, signal intensities were z-score normalized. An affine registration was performed to transform the brain images to the MNI15222.
A neuroradiologist (with 9 years of experience) who was blinded to the clinical information semi-automatically segmented the entire tumor (including cystic or necrotic changes) on the T1C images using 3D Slicer software (v. 4.13.0; www.slicer.org) with edge- and threshold-based algorithms. Another neuroradiologist (with 16 years of experience) re-evaluated and confirmed the segmented lesions.
Radiomic features were calculated with a python-based module (PyRadiomics, version 2.0)23, with a bin size of 32. They included (1) 14 shape features, (2) 18 first-order features, and 3) 75 s-order features (including gray-level co-occurrence matrix, gray-level run-length matrix, gray-level size zone matrix, gray-level dependence matrix, and neighboring gray tone difference matrix) (Supplementary Material S1 and Supplementary Table 2). The features adhered to the standard sets by the Image Biomarker Standardization Initiative 24. A total of 214 radiomic features (107 × 2 sequences) were extracted.
Radiomics model construction
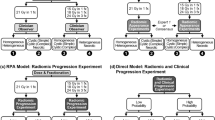
The schematic of radiomics model construction and establishment of an application system based on CycleGAN is shown in Fig. 1a. Radiomic features were MinMax normalized. Because the number of radiomic features was larger than the number of patients, mutual information was applied to select the significant features. The base radiomics classifiers were constructed using extreme gradient boosting with tenfold cross-validation in the training set. Synthetic minority over-sampling technique was applied for oversampling the minority class25. To improve the predictive performance and avoid potential overfitting, Bayesian optimization, which searched the hyperparameter space for optimal hyperparameter combinations, was applied. The area under the curve (AUC), accuracy, sensitivity, specificity, and F1 score (definitions shown in Supplementary Material S2) were obtained. Feature selection and machine learning process were performed using Python 3 with the Scikit-Learn library module (version 0.24.2).
CycleGAN application
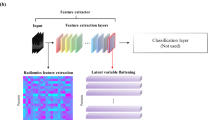
Figure 1b shows the general network architecture of CycleGAN. The generative adversarial network (GAN) has two neural networks, namely, a generator and a discriminator, for distinctive purposes. The CycleGAN uses two sets of GAN for style transfer to train unsupervised image translation models16. Unpaired institutional training and external validation datasets were used to train the discriminators and generators of CycleGAN.
To be delivered into CycleGAN16, the brain MRIs were converted to two-dimensional images in each aspect of the axial, sagittal, and coronal planes. Because the image size was diverse between institutions and individuals, the images were resized to 99 × 117 × 95 pixels after MNI152 template registration and to 116 × 116 pixels before putting them into CycleGAN.
In the first set of GAN, the first generator (G1) in CycleGAN converts the images from the external validation dataset to the domain of the institutional training dataset, while the first discriminator D1 checks if the images computed by G1 are real or fake (generated). Through this process, synthetic images from G1 become better with the feedback of their respective discriminators. In the second set of GAN, the second generator (G2) transfers the synthetic image generated from the first generator (G1) back to the original external validation dataset image, while the second discriminator (D2) checks if the images computed by G2 are real or fake (generated). Through this process, the trained CycleGAN model transferred the style of the external validation images to the training set. The cycle consistency loss, which is the difference between the generated output and the input image, was calculated and used to update the generator models in each training iteration16. L2 loss, which is known to accelerate the training process and generate sharp and realistic images in GAN26,27, was employed to estimate the cycle consistency loss. Inference results were randomly sampled and checked by a neuroradiologist (with 9 years of experience) for plausibility. External validation set images after CycleGAN were subjected to assess the performance of the radiomics model compared with the original external validation dataset. Because original external validation set and external validation set images after CycleGAN were independent from the radiomics modeling in the training process, there is no potential data breach28. Details of the CycleGAN architecture are shown in Supplementary Table 3.
Evaluation of the effect of CycleGAN: Fréchet Inception Distance and t-Distributed Stochastic Neighbor Embedding
The Fréchet Inception Distance (FID) was calculated to measure the similarity between two datasets of images to measure the model quality quantitatively by evaluating the generated data (Supplementary Material S3)29. FID is an extension of the Inception Score30 and compares the distribution of generated images with the distribution of real images that were used to train the generator. FID has been shown to be consistent with human judgments and more robust to noise than inception score29. Three FID scores, namely, “training vs. original external validation,” “original external validation vs. transferred external validation,” and “training vs. transferred external validation” were calculated. To visualize the effect of CycleGAN on the extracted radiomic features, the high-dimensional feature space was projected and visualized into a lower dimensional space by using a two-dimensional t-Distributed Stochastic Neighbor Embedding (t-SNE) manifold31.
Results
Patient characteristics in the institutional training (n = 257) and external validation sets (n = 61) are summarized in Table 1. The proportion of female sex was higher in the high-grade meningiomas in the training set (p < 0.001), but not in the external validation set (p = 0.833). No significant differences were found in other clinical characteristics between the training and external validation sets.
Performance of the classifier for the original external validation and CycleGAN style-transferred external validation images
A total of 27 radiomic features were identified to differentiate meningioma grade (10 features from T2 and 17 features from T1C; 6 shape features, 2 first-order features, and 19 s-order features; details on Supplementary Table 4 and Fig. S2). In the institutional training set, the best performing classifier was achieved in the combined T1C and T2 models, with an AUC, accuracy, sensitivity, specificity, and F1 score of 0.88 (95% confidence interval [CI] 0.77–0.87), 77.7%, 82.8%, and 72.6%, respectively. The T2 and T1C models showed lower performances, with AUCs of 0.84 (95% CI 0.79–0.89) and 0.85 (95% CI 0.82–0.88), respectively.
In the external validation dataset before CycleGAN application, the T2 model showed the highest performance, with an AUC, accuracy, sensitivity, specificity, and F1 score of 0.78 (95% CI 0.64–0.92), 62.7%, 92.3%, 54,4%, and 0.52, respectively. The combined T2 and T1C models showed a similar performance, with an AUC of 0.77 (95% CI 0.63–0.91), whereas the T1C model showed the lowest performance, with an AUC of 0.73 (95% CI 0.58–0.87), respectively.
The style of T2 and T1C images of the original external validation dataset was transformed to match that of the institutional training set images by using CycleGAN (Fig. 2). In the external validation dataset after CycleGAN application, the performance of all three radiomics models increased, although it did not reach statistical significance (Ps > 0.05). The combined T1C and T2 models showed the highest performance, with an AUC, accuracy, sensitivity, specificity, and F1 score of 0.83 (95% CI 0.70–0.97), 73.2%, 84.6%, 69.8%, and 0.59, respectively. T2 and T1C models showed lower performance, with AUCs of 0.80 (95% CI 0.66–0.95) and 0.80 (95% CI 0.67–0.93), respectively. The performance of the radiomics models in the institutional training set and external validation set before and after applying CycleGAN is shown in Table 2. Figure 3 shows the performance of the radiomics models, demonstrating the effect of style transfer by CycleGAN on the classification performance of the radiomics model on external validation. Figure 4 shows representative cases demonstrating the improvement in classifications after CycleGAN. Predictive scores close to 1.0 indicate that the model predicts the meningioma grade with confidence.
Representative cases showing the institutional training dataset and images before and after style transfer of the external validation set. (a) T2 and T1C images of a patient from the institutional training dataset that were leveraged to generate the radiomics model. (b) T2 and T1C images from the original external validation. (c) Style of T2 and T1C images from the external validation set after transformation to match that of the institutional training set images by using CycleGAN. CycleGAN = Cycle-Consistent Adversarial Networks, T1C = postcontrast T1-weighted image, T2 = T2-weighted image.
Radiomics model performance on the (a) AUCs, (b) accuracies, and (b) F1 scores of the radiomics model in the institutional training set and external validation set before and after applying CycleGAN. AUCs = areas under the curve, CycleGAN = Cycle-Consistent Adversarial Networks, T1C = postcontrast T1-weighted image, T2 = T2-weighted image.
Representative cases showing the improvement in classifications after CycleGAN. Predictive scores close to 1.0 indicate that the model predicts the meningioma grade with confidence. (a) A case incorrectly diagnosed as high-grade meningioma before style transfer but correctly diagnosed as low-grade meningioma after style transfer. (b) A case incorrectly diagnosed as low-grade meningioma before style transfer but correctly diagnosed as high-grade meningioma after style transfer. CycleGAN = Cycle-Consistent Adversarial Networks.
Evaluation of the effect of CycleGAN: FID and t-SNE
The FID scores by our proposed model are shown in Fig. S3. The FID score was highest in the “training vs. original external validation,” followed by the “training vs. transferred external validation” and then by the “original external validation vs. transferred external validation.” The FID score decreased to 52.2% after style transfer in the external validation set. However, the “original external validation vs. transferred external validation” showed the lowest FID score, which was 25.6% of the FID score of the “training vs. original external validation”.
The resulting extracted radiomics feature space on t-SNE for the institutional training and external validation sets before and after style transfer is shown in Fig. S4. Before the style transfer, a marked difference was noted between the institutional training and external validation test sets on the values of computed radiomic features. After applying CycleGAN, this influence was markedly decreased. The density plot shows that the distributions of the selected radiomic features of the external validation set became more similar to the institutional training set after applying CycleGAN (Fig. S5).
Discussion
We demonstrated that leveraging CycleGAN is an effective approach to increase the generalizability of radiomics model on external validation. The areas under the curve of the combined T2 and T1C models in the external validation set increased from 0.77 to 0.83 after CycleGAN application. The FID score and t-SNE showed that data distributions between the institutional training and external validation sets in the image and radiomic feature levels became more similar after applying CycleGAN. To the best of our knowledge, this study is the first to investigate CycleGAN strategy for brain tumor imaging to develop a generalizable radiomics model.
CycleGANs are a relatively novel type of conditional generative adversarial networks, which have received considerable attention because of their ability to capture the characteristics of a single image collection and to generate synthetic images in the absence of any paired training examples16,32. Previous generative adversarial network studies on brain imaging have mainly focused on generating missing brain MRI data33,34,35 or creating high-resolution images from low-resolution images36, which requires ground truth sequences. However, no “ground truth” dataset of paired training examples (consisting of internal and external MRI examinations of identical patients at the same period) exists in real-world clinical practice. In our study, we focused on a practical and crucial problem encountered in implementing a machine learning model in medical imaging, which is increasing the generalizability of a radiomics model on external validation. External validation is a crucial process in models with artificial intelligence, because internal validation itself cannot guarantee model generalizability13,37. However, classical preprocessing steps, such as isovoxel resampling, bias field correction, and signal intensity normalization, are insufficient to counter image heterogeneity. We speculate that CycleGAN may be a practical approach to solve the image heterogeneity of an external dataset. A recent study has shown that CycleGAN can reduce the heterogeneity between radiomic features and increase reproducibility in chest radiographs, which is in line with our study38.
A notable finding in our study was that the T2 radiomics model showed relatively less decreased performance in the external validation set before applying CycleGAN, whereas the T1 radiomics model showed a larger decrease in performance in the external validation set before applying CycleGAN. Compared with the T1C protocols with different protocols, T2 protocols are relatively similar between institutions and less prone to failures from image acquisition artifacts39, which may lead to higher performance on external validation than the T1C model. Nonetheless, after CycleGAN application, the combined T2 and T1C models showed the highest performance. This finding suggests that CycleGAN may preserve the biological information from T2 and T1C sequences while effectively removing inter-institutional variation. Our results are in concordance with other studies that demonstrate that single sequence models have limited ability to reflect the underlying pathophysiology of meningiomas9,40.
Our external validation dataset included different scanner vendors, acquisition protocols, image reconstruction algorithms, and field strengths, resulting in large heterogeneity, which reflects the real-world clinical dataset in meningiomas41. Apart from the different MRI vendors with different field strengths, the resolution, sequence, echo time, repetition time, and inversion time have also not reached consensus in meningioma imaging. All of these differences induce heterogeneity of the MRI datasets, which poses as a unique challenge in the generalizability of the artificial intelligence in this area. Collecting heterogeneous labeled data from multiple institutions worldwide is the best solution to overcome this challenge. Nonetheless, even if we tackle this daunting challenge, the generalizability of the resulting artificial intelligence model cannot be fully guaranteed, as the data in another institution are possibly out-of-distribution. In this study, we demonstrate that leveraging an image harmonizing technique based on deep learning is feasible to increase generalizability in radiomics application for grading meningiomas.
The FID score was lowest in the “original external validation vs. transferred external validation,” rather than in the “training vs. transferred external validation” datasets. Considering the equation in FID42, which calculates the difference between the synthetic and real data distributions, the transferred external validation dataset has understandably the most close resemblance to the original external validation dataset. Nonetheless, the FID score from the “original external validation vs. transferred external validation” datasets decreased to 52.2% compared with that from the “training vs. original external validation” datasets. This result demonstrates that the data distributions between the training and external validation sets became more similar after applying CycleGAN.
This study has several limitations. First, it was conducted with a relatively small amount of data, particularly in the external validation set. As this is a technical feasibility study, a larger multi-institutional validation set is warranted to demonstrate significant performance improvement with CycleGAN. Second, we used two-dimensional CycleGAN rather than three-dimensional CycleGAN because of relative paucity of data. This may lead to slice-to-slice inconsistencies, which may adversely affect the performance. However, despite these limitations, as shortfall in generalization to real-world datasets with heterogeneous imaging data is the major barrier for the adoption of artificial intelligence in medical imaging, the strength of our study is that we demonstrated that CycleGAN is a feasible approach to tackle this challenging issue.
In conclusion, CycleGAN is potentially helpful in increasing the generalizability of a radiomics model in differentiating meningioma grade on external validation.
Abbreviations
- AUC:
-
Area under the curve
- CycleGAN:
-
Cycle-Consistent Adversarial Networks
- FID:
-
Fréchet Inception Distance
- GAN:
-
Generative adversarial network
- T1C:
-
Postcontrast T1-weighted image
- T2:
-
T2-weighted image
- t-SNE:
-
T-Distributed Stochastic Neighbor Embedding
- WHO:
-
World Health Organization
- XGBoost:
-
Extreme gradient boosting
References
Ostrom, Q. T. et al. CBTRUS statistical report: Primary brain and other central nervous system tumors diagnosed in the United States in 2009–2013. Neuro Oncol. 18, v1–v75. https://doi.org/10.1093/neuonc/now207 (2016).
Kshettry, V. R. et al. Descriptive epidemiology of World Health Organization grades II and III intracranial meningiomas in the United States. Neuro Oncol. 17, 1166–1173. https://doi.org/10.1093/neuonc/nov069 (2015).
Goldbrunner, R. et al. EANO guidelines for the diagnosis and treatment of meningiomas. Lancet Oncol 17, e383-391. https://doi.org/10.1016/s1470-2045(16)30321-7 (2016).
Modha, A. & Gutin, P. H. Diagnosis and treatment of atypical and anaplastic meningiomas: A review. Neurosurgery 57, 538–550. https://doi.org/10.1227/01.neu.0000170980.47582.a5 (2005).
Nowosielski, M. et al. Diagnostic challenges in meningioma. Neuro Oncol. 19, 1588–1598. https://doi.org/10.1093/neuonc/nox101 (2017).
Gillies, R. J., Kinahan, P. E. & Hricak, H. Radiomics: Images are more than pictures, they are data. Radiology 278, 563–577. https://doi.org/10.1148/radiol.2015151169 (2016).
Won, S. Y. et al. Quality assessment of meningioma radiomics studies: Bridging the gap between exploratory research and clinical applications. Eur J Radiol 138, 109673. https://doi.org/10.1016/j.ejrad.2021.109673 (2021).
Coroller, T. P. et al. Radiographic prediction of meningioma grade by semantic and radiomic features. PLoS ONE 12, e0187908. https://doi.org/10.1371/journal.pone.0187908 (2017).
Park, Y. W. et al. Radiomics and machine learning may accurately predict the grade and histological subtype in meningiomas using conventional and diffusion tensor imaging. Eur. Radiol. 29, 4068–4076. https://doi.org/10.1007/s00330-018-5830-3 (2019).
Ke, C. et al. Differentiation between benign and nonbenign meningiomas by using texture analysis from multiparametric MRI. J. Magn. Reson. Imaging 51, 1810–1820. https://doi.org/10.1002/jmri.26976 (2020).
Morin, O. et al. Integrated models incorporating radiologic and radiomic features predict meningioma grade, local failure, and overall survival. Neurooncol. Adv. 1, vdz011. https://doi.org/10.1093/noajnl/vdz011 (2019).
Zhu, Y. et al. A deep learning radiomics model for preoperative grading in meningioma. Eur. J. Radiol. 116, 128–134. https://doi.org/10.1016/j.ejrad.2019.04.022 (2019).
Park, J. E., Park, S. Y., Kim, H. J. & Kim, H. S. Reproducibility and generalizability in radiomics modeling: Possible strategies in radiologic and statistical perspectives. Korean J. Radiol. 20, 1124–1137. https://doi.org/10.3348/kjr.2018.0070 (2019).
Boxerman, J. L. et al. Consensus recommendations for a dynamic susceptibility contrast MRI protocol for use in high-grade gliomas. Neuro Oncol. 22, 1262–1275. https://doi.org/10.1093/neuonc/noaa141 (2020).
Kaufmann, T. J. et al. Consensus recommendations for a standardized brain tumor imaging protocol for clinical trials in brain metastases. Neuro Oncol 22, 757–772. https://doi.org/10.1093/neuonc/noaa030 (2020).
Zhu, J.-Y., Park, T., Isola, P. & Efros, A. A. In: Proceedings of the IEEE international conference on computer vision. 2223–2232.
Shaban, M. T., Baur, C., Navab, N. & Albarqouni, S. Staingan: Stain style transfer for digital histological images. 2019 IEEE 16th International Symposium on Biomedical Imaging (ISBI 2019), 953–956 (2019).
Shin, S. J. et al. Style transfer strategy for developing a generalizable deep learning application in digital pathology. Comput. Methods Programs Biomed. 198, 105815. https://doi.org/10.1016/j.cmpb.2020.105815 (2021).
Louis, D. N. et al. The 2016 World Health Organization classification of tumors of the central nervous system: A summary. Acta Neuropathol. 131, 803–820. https://doi.org/10.1007/s00401-016-1545-1 (2016).
Avants, B. B., Tustison, N. & Song, G. Advanced normalization tools (ANTS). Insight J. 2, 1–35 (2009).
Roy, S., Butman, J. A. & Pham, D. L. Robust skull stripping using multiple MR image contrasts insensitive to pathology. Neuroimage 146, 132–147. https://doi.org/10.1016/j.neuroimage.2016.11.017 (2017).
Fonov, V. et al. Unbiased average age-appropriate atlases for pediatric studies. Neuroimage 54, 313–327. https://doi.org/10.1016/j.neuroimage.2010.07.033 (2011).
van Griethuysen, J. J. M. et al. Computational radiomics system to decode the radiographic phenotype. Can. Res. 77, e104–e107. https://doi.org/10.1158/0008-5472.Can-17-0339 (2017).
Zwanenburg, A., Leger, S., Vallières, M. & Löck, S. Image biomarker standardisation initiative. arXiv preprint https://arxiv.org/abs/1612.07003 (2016).
Lusa, L. Improved shrunken centroid classifiers for high-dimensional class-imbalanced data. BMC Bioinformatics 14, 1–13 (2013).
Isola, P., Zhu, J.-Y., Zhou, T. & Efros, A. A. in Proceedings of the IEEE conference on computer vision and pattern recognition. 1125–1134.
Wu, H., Zheng, S., Zhang, J. & Huang, K. In: Proceedings of the 27th ACM international conference on multimedia. 2487–2495.
Russell, S. & Norvig, P. Artificial intelligence: A modern approach. (2002).
Heusel, M., Ramsauer, H., Unterthiner, T., Nessler, B. & Hochreiter, S. Gans trained by a two time-scale update rule converge to a local nash equilibrium. arXiv preprint https://arxiv.org/abs/1706.08500 (2017).
Salimans, T. et al. Improved techniques for training gans. arXiv preprint https://arxiv.org/abs/1606.03498 (2016).
Van der Maaten, L. & Hinton, G. Visualizing data using t-SNE. J. Mach. Learn. Res. 9 (2008).
Liu, M.-Y., Breuel, T. & Kautz, J. Unsupervised image-to-image translation networks. arXiv preprint https://arxiv.org/abs/1703.00848 (2017).
Lee, D., Moon, W.-J. & Ye, J. C. Assessing the importance of magnetic resonance contrasts using collaborative generative adversarial networks. Nat. Mach. Intell. 2, 34–42 (2020).
Sharma, A. & Hamarneh, G. Missing MRI pulse sequence synthesis using multi-modal generative adversarial network. IEEE Trans. Med. Imaging 39, 1170–1183 (2019).
Conte, G. M. et al. Generative adversarial networks to synthesize missing T1 and FLAIR MRI sequences for use in a multisequence brain tumor segmentation model. Radiology 299, 203786 (2021).
Chen, Y. et al. in 2018 IEEE 15th International Symposium on Biomedical Imaging (ISBI 2018). 739–742 (IEEE).
Sounderajah, V. et al. Developing specific reporting guidelines for diagnostic accuracy studies assessing AI interventions: The STARD-AI Steering Group. Nat Med 26, 807–808. https://doi.org/10.1038/s41591-020-0941-1 (2020).
Marcadent, S. et al. Generative adversarial networks improve the reproducibility and discriminative power of radiomic features. Radiology: Artificial Intelligence 2, e190035 (2020).
Bangalore Yogananda, C. G. et al. A novel fully automated MRI-based deep-learning method for classification of IDH mutation status in brain gliomas. Neuro Oncol. 22, 402–411. https://doi.org/10.1093/neuonc/noz199 (2020).
Hamerla, G. et al. Comparison of machine learning classifiers for differentiation of grade 1 from higher gradings in meningioma: A multicenter radiomics study. Magn. Reson. Imaging 63, 244–249. https://doi.org/10.1016/j.mri.2019.08.011 (2019).
Hagiwara, A., Fujita, S., Ohno, Y. & Aoki, S. Variability and standardization of quantitative imaging: Monoparametric to multiparametric quantification, radiomics, and artificial intelligence. Invest Radiol. 55, 601–616. https://doi.org/10.1097/rli.0000000000000666 (2020).
Borji, A. Pros and cons of gan evaluation measures. Comput. Vis. Image Underst. 179, 41–65 (2019).
Acknowledgements
This research received funding from the Basic Science Research Program through the National Research Foundation of Korea funded by the Ministry of Science, Information and Communication Technologies & Future Planning (2020R1A2C1003886). This research was also supported by the Basic Science Research Program through the National Research Foundation of Korea funded by the Ministry of Education (2020R1I1A1A01071648). This research was supported by a grant of the Korea Health Technology R&D Project through the Korea Health Industry Development Institute, funded by the Ministry of Health & Welfare, Republic of Korea (HI21C1161), and by a Severance Hospital Research fund for Clinical excellence (C-2021-0019). This research was also funded by the Bio Industrial Strategic Technology Development Program (20003883, 20005021) funded by the Ministry of Trade, Industry & Energy (MOTIE, Korea) and a grant from the Korea Health Technology R&D Project through the Korea Health Industry Development Institute (KHIDI), funded by the Ministry of Health &Welfare, Republic of Korea (HR16C0001).
Author information
Authors and Affiliations
Contributions
S.C.Y. and S.S.A. designed and conceptualized the study. Y.W.P. and S.M.L. collected the data. J.E. and H.L. performed machine learning analysis.Y.W.P. and S.J.S. wrote the main manuscript text. S.C.Y. and S.S.A. revised the manuscript. All authors reviewed and approved the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Park, Y.W., Shin, S.J., Eom, J. et al. Cycle-consistent adversarial networks improves generalizability of radiomics model in grading meningiomas on external validation. Sci Rep 12, 7042 (2022). https://doi.org/10.1038/s41598-022-10956-9
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-022-10956-9
This article is cited by
-
Multicenter PET image harmonization using generative adversarial networks
European Journal of Nuclear Medicine and Molecular Imaging (2024)
-
Intelligent noninvasive meningioma grading with a fully automatic segmentation using interpretable multiparametric deep learning
European Radiology (2023)
-
Preoperative surgical risk assessment of meningiomas: a narrative review based on MRI radiomics
Neurosurgical Review (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.