Abstract
Glioblastoma (GBM) has high metabolic demands, which can lead to acidification of the tumor microenvironment. We hypothesize that a machine learning model built on temporal principal component analysis (PCA) of dynamic susceptibility contrast-enhanced (DSC) perfusion MRI can be used to estimate tumor acidity in GBM, as estimated by pH-sensitive amine chemical exchange saturation transfer echo-planar imaging (CEST-EPI). We analyzed 78 MRI scans in 32 treatment naïve and post-treatment GBM patients. All patients were imaged with DSC-MRI, and pH-weighting that was quantified from CEST-EPI estimation of the magnetization transfer ratio asymmetry (MTRasym) at 3 ppm. Enhancing tumor (ET), non-enhancing core (NC), and peritumoral T2 hyperintensity (namely, edema, ED) were used to extract principal components (PCs) and to build support vector machines regression (SVR) models to predict MTRasym values using PCs. Our predicted map correlated with MTRasym values with Spearman’s r equal to 0.66, 0.47, 0.67, 0.71, in NC, ET, ED, and overall, respectively (p < 0.006). The results of this study demonstrates that PCA analysis of DSC imaging data can provide information about tumor pH in GBM patients, with the strongest association within the peritumoral regions.
Similar content being viewed by others
Introduction
Glioblastoma (GBM) is the most common malignant primary brain tumor in adults, characterized with vasular proliferation, diffuse infiltration in the adjacent brain parenchyma, and resistance to the standard therapies1. The tumor microenvironment plays an important role in abundant infiltration of GBM tumor cells, its resistance to standard therapies, recurrence and therefore, poor patient prognosis2. Due to rapid growth of GBM tumors and actively migrating cell population, hypercellular regions are formed typically surrounding the necrotic foci tissues and have a high metabolic demand3. When the tumor grows, the lack of sufficient circulation compared to the cell population of the tumor results in ischemia and secretion of angiogenic factors, which in turn leads to proliferation of new vessels4.
Neo-angiogenesis forms a tortuous and branched vascular structure with increased blood volume and permeability, and impaired cerebral perfusion with subsequent necrosis3,5. These alterations promote tumor growth, decrease oxygen, increase glycolysis and lactic acid, decrease extracellular pH, facilitate cell invasion6. This augments the probability of mutations, such as vascular endothelial growth factor (VEGF) gene expression triggered by the hypoxia-inducible factor (HIF) family of transcription factors7,8. Even in presence of abundant oxygen, glycolysis is often enhanced in cancers due to elevated concentration of lactic acid, resulting in a substantial decrease in extracellular pH which leads to escalated invasion and aggressiveness of the tumor, and decreased immune response6,9,10.
Various MRI techniques have been used to measure tumor PH. Magnetic resonance spectroscopy (MRS) reveals the amount and spatial distribution of particular biochemical substances involved in metabolic processes in tumoral tissues11. 1H MRS, 13C MRS, and phosphorus MRS (31PMRS) can estimate levels of cellular metabolites and tumoral pH11,12. Chemical exchange saturation transfer (CEST) is a relatively new MR technique that images certain compounds at very low concentrations that are too low to be directly detected by MRS technique13. Tumor blood perfusion may also reflect the underlying tumor pH, as well-perfused tumors have more efficient removal of excess lactic acid, protons, CO2, and other metabolic byproducts. Dynamic susceptibility contrast (DSC) perfusion is a widespread clinical MRI approach for quantification of cerebral perfusion and is often collected for assessment of brain tumors14. As DSC-MRI can measure tissue perfusion and compromised microvasculature in GBMs, it might be able to quantify tumor acidity.
Amine CEST echo-planar imaging (CEST-EPI) is a fast molecular imaging MRI technique to measure tumor pH6. In a recent study, a positive linear correlation between cerebral blood volume (CBV) obtained from DSC perfusion MRI and acidity was demonstrated in areas of T2-hyperintense, non-enhancing tumor in glioma patients15. We hypothesize that principal component analysis (PCA) of DSC-MRI perfusion images in conjunction with machine learning (ML) techniques in patients with GBM may quantify microvascular structure at the voxel level and infer capillary-level hemodynamics that correlates with tumor acidity. PC analysis of DSC-MRI has shown potential in predicting the location of future recurrence16,17, patient’s survival 18, arteriovenous shunting4, and EGFRvIII status19. The aim of this analysis is to use ML methods based on perfusion MRI scans to uncover unique tissue characteristics that correlate with tissue acidity and might provide insights about the tumoral and peritumoral tissue metabolism to guide treatment planning.
Results
In this prospective study, we included 32 patients (19 males, 13 females; age, 64.6 ± 10.11 years old), who were confirmed to have GBM tumors (Table 1). A total of 101 CEST-MRI scans were acquired from the study subjects (12 patients had pre-surgical and 89 had imaging during active treatment with radiation and/or chemotherapy), of which, 78 scans with their corresponding DSC-MRI scans passed our data preparation and pre-processing steps. The SVR machine learning method based on PCA was applied in a cohort of 78 cases.
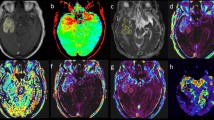
Principal component analysis of perfusion time-series revealed that the tumor subregions, i.e., ET, NC, ED, form characteristic clusters (Fig. 1B), which facilitate specification of the tissues and allow for mapping the heterogeneity within a specific tissue. Figure 2 illustrates structural MRI scans, including T1, T1-Gd, T2, T2-FLAIR images for a male patient (58 years old) with GBM. Furthermore, relative cerebral blood volume (rCBV) map generated from DSC-MRI scans using CaPTk software, PC1–PC3 images derived from PC analysis of the hemodynamic curves, the MTRasym image constructed using our proposed approach, along with the actual MTRasym image quantified from CEST imaging are shown in the whole pathogenic region. As it can be inferred, our constructed MTRasym provides more accurate voxel-wise mapping of the actual MTRasym image, compared to the rCBV map. Among the seven PCs used in building our model, the visual similarity is most striking for the first three PC images since the components progressively capture less variance.
An illustration of the perfusion time-series in tumorous subregions, i.e., ET, NC, and ED (A); and the clustering of each tissue type using PC analysis (B), signifying the potential of the PCs in capturing tissue characteristics. PC1, PC2, and PC3 represent the first, second, and third principal components, respectively. ET Enhancing tumor, NC Necrotic core, ED paeritumoral edema.
Conventional MRI, including T1, T1-Gd, T2, and T2-FLAIR, scans of a 58-year-old male patient included in our study. Map of a proxy to relative cerebral blood volume (ap-rCBV) derived from DSC-MRI scans with CaPTk software. Three principal components (PCs), PC1 to PC3, calculated using PCA of the hemodynamic perfusion curves, along with the MTRasym image constructed using the seven PCs in association with the actual MTRasym image. CaPTk version 1.8.1 (www.med.upenn.edu/cbica/captk/).
The MTRasym image constructed from perfusion PCs using our proposed regression method showed moderate to strong agreement with the MTRasym image, with R of 0.47 (p = 0.006), 0.66 (p = 0.0009), 0.67 (p = 0.00001) in the ET, NC, and ED regions, respectively, and 0.71 (p < 10e-6) in the whole pathogenic region, as a union of ET, NC, and ED areas, averaged over all the patients. Figure 3 demonstrates a strong association of the MTRasym image built from the perfusion PCs and the actual MTRasym image, implying the potential of ML in distinguishing the tumorous regions with specific metabolism characteristics.
This method outperforms the conventional DSC-MRI analysis, as displayed in Fig. 4, where panel (A) presents the signal intensity–time curves of the voxels located in the high (red) and low (blue) MTRasym image. While the voxels have been selected from the regions with different levels of acidic pH, the perfusion curves are not discriminant. Applying PC analysis differentiates the clusters of the same high and low MTRasym, as it can be observed from panel (B) of Fig. 4. The bottom panel (part C) shows three PCs quantified for the higher and lower MTRasym regions. Figure 4C reveals that the first principal component (PC1) is primarily related to the level of the perfusion signal, as evidenced by the large variance throughout the signal time course. The second principal component (PC2) is related to the depth of the perfusion signal drop, in relation to the baseline level, and the third principal component (PC3) relates to the shape of the drop of the perfusion signal, e.g., steepness of the signal drop and recovery. As a comparison, Fig. 5 displays the signal intensity–time curves in the highest (red) and lowest (blue) voxels in PC1, PC2, and PC3 images, suggesting the differentiability of the tissues based on PC analysis. Specifically, PC1 provides a noticeable discrimination between the areas with different hemodynamic properties. The discrimination diminishes in larger PCs as evidenced by this illustration.
(A) Perfusion curves calculated within regions of low and high MTRasym (shown in blue and red colors, respectively), suggesting poor discrimination of the regions solely based on hemodynamic curves. (B) Discrimination of low and high MTRasym regions based on PC analysis; PC1 = principal component 1; PC2 = principal component 2; PC3 = principal component 3. (C) The three principal components for high MTRasym regions, yielding a marked differentiation of these regions based on the PCs.
Discussion
Our study showed that high‐resolution pH‐sensitive imaging in brain tumors can be achieved on clinical 3T MR systems using DSC-MRI with strong correlation to CEST-EPI PH imaging. DSC-MRI can characterize microvascular circulation in GBM patients, and the respective acidity can be extracted using PCA and ML techniques. The proposed techniques are ideally suited for evaluation of malignant gliomas since GBM remains the most angiogenic primary brain tumor and therefore exhibits extensive neovascularization, compromised brain blood barrier, and heterogeneous acidity. DSC-MRI without proper processing cannot discern the pH heterogeneity of the tumoral regions, but as proposed in this study, PCA of the perfusion time-series can be used to differentiate tissue acidity.
GBMs have unique pathophysiological characteristics such as significant invasiveness, fast growth, and rapid seeding. Also, similar to the other cancers, GBMs prefer glycolysis over oxidative phosphorylation even in the presence of ample oxygen (Warburg effect) that results in increased intracellular lactate and elevated acidification. These tumors also cause direct destruction of surrounding tissue, including both neuronal death via glutamate excitotoxicity20 and degradation of the extracellular matrix via metalloproteinases and other proteases21 that are pH-dependent22,23. Since, ion movement is directly coupled to movement of other ions, pH not only serves as a regulator of cell activity, but also an indirect surrogate marker of other cellular functions. PH heterogeneity in the tumor microenvironment is critical for surgical and chemo-radiation planning. Weak base and weak acidic drugs get trapped in either the intracellular or extracellular spaces due to “ion trapping” phenomena24.
Wang et al. investigated association between pH-sensitive CEST-EPI and relative cerebral blood volume (CBV) measurements obtained from DSC-MRI in patients with gliomas15. They reported a strong correlation between acidity and CBV in the T2-hyperintense regions, but not in the areas of enhancing tumor. In agreement with this study, our results demonstrated that the correlation between our constructed MTRasym map and the actual MTRasym, as quantified by CEST-EPI, is highest in the peritumoral T2-hyperintense areas, followed by necrotic tumor core, and contrast-enhancing tumor regions. However, in our study, in different subregions of each tumor, a heterogeneous distribution of high-acidity tissues was observed.
The proposed PCA-derived features integrated with SVR method uses temporal dynamics of DSC-MR imaging. Our results indicate that a more comprehensive analysis of DSC along with machine learning, can unravel useful information related to tumor acidity, which is typically not obtained with traditional CBV or CBF maps. Specifically, higher acidotic tissues demonstrate higher level of DSC signal based on PC1, consistent with higher necrosis and free water mobility. PC2 mainly represents the depth of drop in DSC signal, and deeper drop in the signal shows higher acidity. Integrative information of signal drop and recovery is stored in PC3, where lower signal recovery is associated with higher acidotic tissues.
CEST-EPI has been shown to be effective an a non-invasive biomarker to distinguish between IDH1 mutant and wild type gliomas, and also 1p/19q codeleted from intact IDH1 mutant gliomas25,26. In addition, it has been shown to be of value as an early imaging biomarker for bevacizumab treatment response and failure in recurrent glioblastoma27. Our proposed approach could support measuring tumor acidity with DSC-MRI, as a more widely-accessible imaging method compared with CEST-EPI.
There are limitations to our study, including limited sample size and single institutional data collection; these results need to be validated in a large multi-institutional study.
In conclusion, the results of this study indicate that PCA analysis of DSC-MRI in conjunction with machine learning techniques, can potentially enable better localization of highly acidic regions. In turn, this information may be used for tumor prognostication and treatment response evaluation.
Methods
Patients
Institutional review board (IRB) approval of the University of Pennsylvania was obtained for this prospective study and informed consent was collected from the participants. All methods were carried out in accordance with relevant guidelines and regulations. 32 subjects with intra-axial brain mass suggestive of high-grade glioma, who were referred to the radiology department of hospital of University of Pennsylvania from March 2018 to February 2020 and were subsequently proven by histopathology to be GBM were included in this study.
Image acquisition
All MRI scans were performed on a Magnetom Tim Trio 3 Tesla scanner (Siemens, Erlangen, Germany) using a 12-channel phased array head coil. Conventional MRI sequences included axial T1-weighted (T1) before and after administration of gadolinium contrast agent (T1-Gd) with matrix size = 192 × 256 × 192, resolution = 0.98 × 0.98 × 1.00 mm3, repetition time (TR in ms)/echo time (TE in ms) = 1760/3.1, T2-weighted (T2) with matrix size = 208 × 256 × 64, resolution = 0.94 × 0.94 × 3.00, TR/TE = 4680/85; T2 fluid-attenuated inversion recovery (T2-FLAIR) with matrix size = 192 × 256 × 60, resolution = 0.94 × 0.94 × 3.00, TR/TE = 9420/141.
DSC-MR imaging was performed by a gradient-echo echo-planar (GE-EPI) imaging sequence during a second 0.1-mmol/kg bolus of Dotarem (Gadoterate Meglumine) with the following parameters: TR/TE = 2000/45 ms, FOV = 22 × 22 cm2, resolution = 1.72 × 1.72 × 3 mm3, 20 sections. A bolus of contrast agent with a dose of 0.1 mmol/kg which was done for DCE (dynamic-contrast-enhanced) imaging served as a preload dose for DSC imaging to reduce the effect of contrast agent leakage on relative cerebral blood volume (rCBV) measurements.
Acquisition of pH‐sensitive information was performed through an amine contrast specific for single-echo CEST-EPI sequence15,28. MR imaging acquisition parameters included the following: FOV = 240–256 × 217–256 mm, matrix size = 128 × 116–128, slice thickness = 4 mm with no inter-slice gap, 25 consecutive slices, excitation pulse flip angle = 90°, TE = 27 ms, bandwidth = 1628 Hz, and generalized auto-calibrating partially parallel acquisition factor = 2–3. Off-resonance saturation was applied using a pulse train of 3 × 100 ms Gaussian pulses with a peak amplitude of 6 microtesla. A total of 29 off-resonance or z-spectral points were sampled at frequency offsets of − 3.5 to − 2.5 ppm, − 0.3 to + 0.3 ppm, and − 2.5 to + 3.5 ppm, all in increments of 0.1 ppm. A reference scan (S0) was obtained with the same acquisition parameters, without the saturation pulses. Total scan time for CEST-EPI was approximately 6 min.
MRI pre-processing
For each patient, all MRI volumes (T1, T2, T2-FLAIR, DSC-MRI and MTRasym) were rigidly co-registered with their corresponding T1-Gd using the Greedy registration method29 (https://github.com/pyushkevich/greedy). Subsequently, all conventional MRI scans (T1, T1-Gd, T2, T2-FLAIR) were smoothened to remove any high frequency intensity variations (i.e., noise)30, corrected for magnetic field inhomogeneities using N4ITK method31 and skull-stripped using FSL BET32 followed by manual revision when needed. For brain tumor segmentation in the images, DeepMedic33, a Deep Learning (DL)-based segmentation algorithm in Cancer Imaging Phenomics Toolkit (CaPTk) v.1.7.834,35 which had been trained on BraTS 2017 training data, was applied to the co-registered conventional MRI scans. Brain tumor segmentation delineated three regions of interest (ROIs), i.e., enhancing tumor (ET), necrosis (NC), and peritumoral edema (ED), in the GBM tumors.
Amine CEST-EPI post-processing
Clinical post-processing of CEST-EPI consisted of affine motion correction (MCFLIRT; FSL, https://fsl.fmrib.ox.ac.uk/fsl/fslwiki/MCFLIRT) and B0 correction via a z-spectra-based K-means clustering and Lorentzian fitting algorithm36. An integral of the width of 0.4 ppm was then obtained around both the − 3.0 and + 3.0 ppm (− 3.2 to − 2.8 and + 2.8 to + 3.2 ppm, respectively) spectral points of the inhomogeneity-corrected data. These data points were combined with the S0 image to calculate the asymmetry in the magnetization transfer ratio (MTRasym) at 3.0 ppm as defined by equation MTRasym (ω) = S(− ω) − S(ω)/S, where ω is the offset frequency of interest (3.0 ppm). All resulting maps were registered to high-resolution post-contrast T1-weighted images for subsequent analyses.
Temporal principal component analysis
Principal component analysis (PCA) is a dimensionality reduction method17 which was used in this study to distill the DSC-MRI time series down to a few components that capture the temporal dynamics of blood perfusion. All hemodynamic perfusion curves were aligned and normalized for the baseline and maximum drop across the patients37. We randomly selected voxels in each tumor subregion, i.e., ET, NC, and ED, and generated their signal intensity–time curves (Fig. 1A). PCA was subsequently applied to capture the variance of the time series in all the ROIs and all subjects. Because of the relative consistency in the perfusion patterns of the various ROIs, seven principal components were sufficient to capture more than 99% of the variance in the perfusion signal for all tumor subregions and all patients.
Generation of MTRasym images based on PCs using machine learning
We built several regression models for tumor subregions using support vector machine regression (SVR) aiming to predict the MTRasym values from the seven PCs on a voxel-by-voxel basis to create a PC-derived MTRasym image, referred to as constructed MTRasym image. Leave-one-subject-out cross-validation of these predictive models was performed to ensure that the model and the associate estimates of accuracy would likely generalize to new patients. We trained the SVR models separately in ET, NC, and ED regions using Gaussian kernel functions with an automatic kernel scale and sequential minimal optimization (SMO) configuration. Performance of the SVR method was evaluated using Spearman’s correlation. All machine learning and statistical analyses was performed in MATLAB 9.4.0.949201 (R2018a) Update 6.
References
Ostrom, Q. T. et al. CBTRUS statistical report: Primary brain and other central nervous system tumors diagnosed in the United States in 2011–2015. Neuro Oncol. 20, iv1–iv86. https://doi.org/10.1093/neuonc/noy131 (2018).
Cooper, L. A. et al. The tumor microenvironment strongly impacts master transcriptional regulators and gene expression class of glioblastoma. Am. J. Pathol. 180, 2108–2119. https://doi.org/10.1016/j.ajpath.2012.01.040 (2012).
Brat, D. J. et al. Pseudopalisades in glioblastoma are hypoxic, express extracellular matrix proteases, and are formed by an actively migrating cell population. Can. Res. 64, 920–927. https://doi.org/10.1158/0008-5472.can-03-2073 (2004).
Nabavizadeh, S. A. et al. Arterial spin labeling and dynamic susceptibility contrast-enhanced MR imaging for evaluation of arteriovenous shunting and tumor hypoxia in glioblastoma. Sci. Rep. 9, 8747. https://doi.org/10.1038/s41598-019-45312-x (2019).
Brat, D. J. & Van Meir, E. G. Vaso-occlusive and prothrombotic mechanisms associated with tumor hypoxia, necrosis, and accelerated growth in glioblastoma. Lab. Investig. J. Tech. Methods Pathol. 84, 397–405. https://doi.org/10.1038/labinvest.3700070 (2004).
Harris, R. J. et al. Simultaneous pH-sensitive and oxygen-sensitive MRI of human gliomas at 3 T using multi-echo amine proton chemical exchange saturation transfer spin-and-gradient echo echo-planar imaging (CEST-SAGE-EPI). Magn. Reson. Med. 80, 1962–1978. https://doi.org/10.1002/mrm.27204 (2018).
Fathi Kazerooni, A., Bakas, S., Saligheh Rad, H. & Davatzikos, C. Imaging signatures of glioblastoma molecular characteristics: A radiogenomics review. J. Magn. Reson. Imaging JMRI 52, 54–69. https://doi.org/10.1002/jmri.26907 (2020).
Maxwell, P. H., Pugh, C. W. & Ratcliffe, P. J. Activation of the HIF pathway in cancer. Curr. Opin. Genet. Dev. 11, 293–299. https://doi.org/10.1016/s0959-437x(00)00193-3 (2001).
Gillies, R. J. & Gatenby, R. A. Metabolism and its sequelae in cancer evolution and therapy. Cancer J. (Sudbury Mass.) 21, 88–96. https://doi.org/10.1097/ppo.0000000000000102 (2015).
Estrella, V. et al. Acidity generated by the tumor microenvironment drives local invasion. Can. Res. 73, 1524–1535. https://doi.org/10.1158/0008-5472.can-12-2796 (2013).
Cichocka, M., Kozub, J. & Urbanik, A. PH measurements of the brain using phosphorus magnetic resonance spectroscopy ((31)PMRS) in healthy men—Comparison of two analysis methods. Pol. J. Radiol. 80, 509–514. https://doi.org/10.12659/pjr.895178 (2015).
Najac, C. & Ronen, S. M. MR molecular imaging of brain cancer metabolism using hyperpolarized 13C magnetic resonance spectroscopy. Top. Magn. Reson. Imaging TMRI 25, 187–196. https://doi.org/10.1097/rmr.0000000000000104 (2016).
Wu, B. et al. An overview of CEST MRI for non-MR physicists. EJNMMI Phys. 3, 19. https://doi.org/10.1186/s40658-016-0155-2 (2016).
Boxerman, J. L. et al. Consensus recommendations for a dynamic susceptibility contrast MRI protocol for use in high-grade gliomas. Neuro Oncol. 22, 1262–1275. https://doi.org/10.1093/neuonc/noaa141 (2020).
Wang, Y. L. et al. Association between tumor acidity and hypervascularity in human gliomas using pH-weighted amine chemical exchange saturation transfer echo-planar imaging and dynamic susceptibility contrast perfusion MRI at 3T. AJNR Am. J. Neuroradiol. 40, 979–986. https://doi.org/10.3174/ajnr.A6063 (2019).
Akbari, H. et al. Imaging surrogates of infiltration obtained via multiparametric imaging pattern analysis predict subsequent location of recurrence of glioblastoma. Neurosurgery 78, 572–580. https://doi.org/10.1227/neu.0000000000001202 (2016).
Akbari, H. et al. Pattern analysis of dynamic susceptibility contrast-enhanced MR imaging demonstrates peritumoral tissue heterogeneity. Radiology 273, 502–510. https://doi.org/10.1148/radiol.14132458 (2014).
Macyszyn, L. et al. Imaging patterns predict patient survival and molecular subtype in glioblastoma via machine learning techniques. Neuro Oncol. 18, 417–425. https://doi.org/10.1093/neuonc/nov127 (2016).
Bakas, S. et al. In vivo detection of EGFRvIII in glioblastoma via perfusion magnetic resonance imaging signature consistent with deep peritumoral infiltration: The φ-index. Clin. Cancer Res. Off. J. Am. Assoc. Cancer Res. 23, 4724–4734. https://doi.org/10.1158/1078-0432.ccr-16-1871 (2017).
Ye, Z. C. & Sontheimer, H. Glioma cells release excitotoxic concentrations of glutamate. Can. Res. 59, 4383–4391 (1999).
Nakada, M., Okada, Y. & Yamashita, J. The role of matrix metalloproteinases in glioma invasion. Front. Biosci. 8, e261-269. https://doi.org/10.2741/1016 (2003).
Fasciglione, G. F., Marini, S., D’Alessio, S., Politi, V. & Coletta, M. pH- and temperature-dependence of functional modulation in metalloproteinases. A comparison between neutrophil collagenase and gelatinases A and B. Biophys. J. 79, 2138–2149. https://doi.org/10.1016/s0006-3495(00)76461-7 (2000).
Gioia, M. et al. pH dependence of the enzymatic processing of collagen I by MMP-1 (fibroblast collagenase), MMP-2 (gelatinase A), and MMP-14 ectodomain. J. Biol. Inorg. Chem. JBIC Publ. Soc. Biol. Inorg. Chem. 15, 1219–1232. https://doi.org/10.1007/s00775-010-0680-8 (2010).
Raghunand, N. & Gillies, R. J. pH and drug resistance in tumors. Drug Resist. Updates Rev. Comment. Antimicrob. Anticancer Chemother. 3, 39–47. https://doi.org/10.1054/drup.2000.0119 (2000).
Yao, J. et al. Metabolic characterization of human IDH mutant and wild type gliomas using simultaneous pH- and oxygen-sensitive molecular MRI. Neuro Oncol. 21, 1184–1196. https://doi.org/10.1093/neuonc/noz078 (2019).
Yao, J. et al. Human IDH mutant 1p/19q co-deleted gliomas have low tumor acidity as evidenced by molecular MRI and PET: A retrospective study. Sci. Rep. 10, 11922. https://doi.org/10.1038/s41598-020-68733-5 (2020).
Yao, J. et al. pH-weighted amine chemical exchange saturation transfer echoplanar imaging (CEST-EPI) as a potential early biomarker for bevacizumab failure in recurrent glioblastoma. J. Neurooncol. 142, 587–595. https://doi.org/10.1007/s11060-019-03132-z (2019).
Harris, R. J. et al. Simulation, phantom validation, and clinical evaluation of fast pH-weighted molecular imaging using amine chemical exchange saturation transfer echo planar imaging (CEST-EPI) in glioma at 3 T. NMR Biomed. 29, 1563–1576. https://doi.org/10.1002/nbm.3611 (2016).
Yushkevich, P. A. et al. Fast automatic segmentation of hippocampal subfields and medial temporal lobe subregions in 3 Tesla and 7 Tesla T2-weighted MRI. Alzheimer’s Dement. 7, 126–127 (2016).
Smith, S. M. & Brady, J. M. SUSAN—A new approach to low level image processing. Int. J. Comput. Vis. 23, 45–78 (1997).
Sled, J. G., Zijdenbos, A. P. & Evans, A. C. A nonparametric method for automatic correction of intensity nonuniformity in MRI data. IEEE Trans. Med. Imaging 17, 87–97. https://doi.org/10.1109/42.668698 (1998).
Smith, S. M. Fast robust automated brain extraction. Hum. Brain Mapp. 17, 143–155. https://doi.org/10.1002/hbm.10062 (2002).
Kamnitsas, K. et al. Efficient multi-scale 3D CNN with fully connected CRF for accurate brain lesion segmentation. Med. Image Anal. 36, 61–78. https://doi.org/10.1016/j.media.2016.10.004 (2017).
Davatzikos, C. et al. Cancer imaging phenomics toolkit: quantitative imaging analytics for precision diagnostics and predictive modeling of clinical outcome. J. Med. Imaging (Bellingham, Wash.) 5, 011018. https://doi.org/10.1117/1.jmi.5.1.011018 (2018).
Fathi Kazerooni, A. et al. Cancer imaging phenomics via CaPTk: Multi-institutional prediction of progression-free survival and pattern of recurrence in glioblastoma. JCO Clin. Cancer Inform. 4, 234–244. https://doi.org/10.1200/cci.19.00121 (2020).
Yao, J. et al. Improving B(0) correction for pH-weighted amine proton chemical exchange saturation transfer (CEST) imaging by use of k-means clustering and Lorentzian estimation. Tomography (Ann Arbor, Mich.) 4, 123–137. https://doi.org/10.18383/j.tom.2018.00017 (2018).
Akbari, H. et al. Histopathology-validated machine learning radiographic biomarker for noninvasive discrimination between true progression and pseudo-progression in glioblastoma. Cancer 126, 2625–2636. https://doi.org/10.1002/cncr.32790 (2020).
Acknowledgements
The authors wish to thank Gamaliel Isaac, PhD for his help in implementing and testing CEST-EPI pulse sequence.
Funding
Research reported in the publication was supported by the McCabe foundation award of the University of Pennsylvania, National Institutes of Health (NIH) R01 Grant on “Predicting brain tumor progression via multiparametric image analysis and modeling” (R01-NS042645), National Institutes of Health (NIH) U24 Grant of “Cancer imaging phenomics software suite: application to brain and breast cancer” (U24-CA189523), American Cancer Society (ACS) Research Scholar Grant (RSG-15-003-01-CCE), UCLA Brain SPORE (P50CA211015), DoD CDMRP (CA200290), and NCI (R21CA223757).
Author information
Authors and Affiliations
Contributions
Conception and design: H.A., A.F.K., C.D., A.N. Development of methodology: H.A., A.F.K., C.D., A.N. Acquisition of data: J.W., E.M., H.A., S.G., C.S., C.R., B.E., A.N. Preprocessing of images: H.A., A.F.K., J.W., E.M., H.A., S.G., C.S., J.Y. Analysis and interpretation of data (e.g., statistical analysis, biostatistics, computational analysis): H.A., A.F.K., A.N. Writing, review, and/or revision of the manuscript: H.A., A.F.K., J.W., E.M., H.A., S.G., C.S., C.R., S.B., D.M.O., A.S.D., S.J.B., B.E., C.D., A.N.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Akbari, H., Kazerooni, A.F., Ware, J.B. et al. Quantification of tumor microenvironment acidity in glioblastoma using principal component analysis of dynamic susceptibility contrast enhanced MR imaging. Sci Rep 11, 15011 (2021). https://doi.org/10.1038/s41598-021-94560-3
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-021-94560-3
This article is cited by
-
Functional biomaterials for biomimetic 3D in vitro tumor microenvironment modeling
In vitro models (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.