Abstract
Clostridium difficile is a spore-forming gram-positive bacterium, recognized as the primary cause of antibiotic-associated nosocomial diarrhoea. Clostridium difficile infection (CDI) has emerged as a major health-associated infection with increased incidence and hospitalization over the years with high mortality rates. Contamination and infection occur after ingestion of vegetative spores, which germinate in the gastro-intestinal tract. The surface layer protein and flagellar proteins are responsible for the bacterial colonization while the spore coat protein, is associated with spore colonization. Both these factors are the main concern of the recurrence of CDI in hospitalized patients. In this study, the CotE, SlpA and FliC proteins are chosen to form a multivalent, multi-epitopic, chimeric vaccine candidate using the immunoinformatics approach. The overall reliability of the candidate vaccine was validated in silico and the molecular dynamics simulation verified the stability of the vaccine designed. Docking studies showed stable vaccine interactions with Toll‐Like Receptors of innate immune cells and MHC receptors. In silico codon optimization of the vaccine and its insertion in the cloning vector indicates a competent expression of the modelled vaccine in E. coli expression system. An in silico immune simulation system evaluated the effectiveness of the candidate vaccine to trigger a protective immune response.
Similar content being viewed by others
Introduction
Clostridium difficile, a spore forming, gram positive bacteria is the causative agent of nosocomial, antibiotic-associated severe diarrhoea1,2,3. These obligate anaerobic bacteria cause Clostridium difficile infection (CDI) that includes symptoms like diarrhoea with or without colitis, abdominal pain, fever with chills and discomfort3,4,5. CDI is mostly acquired and transmitted in hospitals and nursing homes, where the use of efficacious antibiotics and antimicrobials are high6,7. It has a mortality rate ranging from 6 to 17% and a substantial morbidity all over the world8. CDI also shows high recurrence rates (range 15–41%) complicating the infection. This may surge even further with every new episode with larger and more prominent impact, which has been previously discussed by Haubitz et al., and Peery et al.9,10. Likewise, Murphy et al., summarized the mortality data in National Centre for Health Statistics (NCHS) by noting, “Enterocolitis due to Clostridium difficile… has become a growing concern in recent years”11.
CDI contamination occurs after the ingestion of spores, which are shed into the environment by patients with or without disease symptoms. These spores germinate in the gastro-intestinal tract, allowing the colonization of the vegetative cells in the gut and multiplication in the colon12. The colonization is aided by the dysbiosis of internal gut microbiota which mostly results from excessive antibiotic treatments. The spore proteins, other than being agents for disease transmission, play a role in development of the disease. There are number of proteins present on the outermost layer of the spore coat, but CotE protein is of particular significance as it plays an important role in virulence13. This 81 kDa protein is bi-functional, as it carries the N-terminal peroxiredoxin domain as well as the C-terminal chitinase domain13,14 hence it binds to internal mucin glycoproteins enabling adhesion of the spore to the internal GI tract surface. Spore attachment is essential for CDI with the internal colonization being initiated by the CotE spore protein15.
The surface-layer proteins (SLP) are the most abundant cellular proteins in bacteria16. SLPs are capable of adherence to human gastro-intestinal and intestinal epithelial cells in vitro17. Studies on C. difficile SLP shows that they not only play major roles in survival and growth, but also interact with the host and its immune system through TLR4 activation, by inducing the production of cytokines7,18. SlpA protein being the most abundant protein in the S-layer, acts as a major colonization factor19 and recombinant vaccines developed against SlpA showed reduced gut colonization in mice7,20. Thus, this protein can be regarded as a possible candidate for the design of vaccines.
The flagellar proteins, FliC and FliD present in the bacteria play important roles in motility, colonization, biofilm formation and toxin gene expression21,22. It has been observed that the major role of bacterial cell adherence to the gut surface is played by flagellar cap protein (FliC), unlike flagellar subunit protein (FliD) which acts as a weak binder for cell adherence activity23. The attachment of the bacterial cell to the gut enables onset of infection. This marks the triggering of innate immune responses, through the activation of TLR5 which helps in protection against colonization12. FliC protein’s active interaction with TLR5 induces strong immune response in the body, which makes it a preferred choice for vaccine construction2.
For immunization through vaccination against highly morbid CDI, toxin antigens represent the first studied targets and anti-toxin antibodies formed after such vaccinations are concerned with preventing CDI recurrence24. Given that, C. difficile’s virulence factors also include TcdA and TcdB proteins, where efforts were made to develop C. difficile vaccines targeting both TcdA and TcdB. Vaccination with toxoids or recombinant fragments of toxins were tested with varying but minimal success in both animals and humans24. Nonetheless, C. difficile survives in the environment as very stable spore forms, resistant to antibiotics and harsh conditions, and is the root cause of recurrent CDI25. Toxoid vaccines cannot avoid the sporulation or deposition of C. difficile spores into the body, potentially raising the number of asymptomatic carriers of the disease. As a result, vaccine candidates that prevent colonization and adhesion of bacteria to gut epithelia are currently under consideration26. A vaccine strategy that tackles single or multiple factors responsible for colonization, adherence and persistence has a considerable advantage over other developing vaccines. Hence, other proteins like CotE spore protein, SlpA S-layer protein and FliC flagellar cap protein should be considered preferred vaccine candidates over conventional toxoid vaccines for the prevention of C. difficile infections.
Upon injecting into the body, the vaccine candidate is identified by the innate immune system of the host through pattern recognition receptors (PRRs) that bind pathogen-associated molecular patterns (PAMPs). The host cells sense the possible danger when they detect a pathogen through PAMPs embedded in the vaccine and activate dendritic cells, monocytes and neutrophils that patrol the whole body27. The activated immune cells (dendritic cells and monocytes) after sensing ‘danger signals’ elicited by the vaccine, modulate the expression of surface molecules and secrete pro-inflammatory cytokines and chemokines28. Matured dendritic cells formed due to an inflammatory microenvironment, migrate towards lymph nodes where T and B lymphocytes activate29. At around the same time, antigen presenting cells process the antigens into smaller fragments that are then displayed in the grooves of MHC (called Human Leukocyte Antigens, HLA, in humans) molecules on the cell surface. MHC class I molecules bind to antigen peptides produced in the cytosol of infected cells, whereas phagocytized antigens are primarily displayed on MHC class II molecules30,31,32. CD4 + T cells participate in the identification of antigenic peptides, which are displayed by MHC class II molecules, whereas after activation, CD8 + T cells binds to MHC class I- peptide33. Furthermore, the secreted cytokines by the activated CD4 + T cells helps in activation of B cells for proper antibody generation34 (Fig. 1).
Vaccine design which involves either the entire organism or large proteins results in unnecessary antigenic load and chances of allergenic responses35. This difficulty in vaccine development can be overcome by peptide-based vaccines, which involves short immunogenic peptide fragments35. These short peptides are meant to be sufficient in eliciting strong immune responses, while avoiding the induction of allergenic responses. Recent developments in computational biology have opened new opportunities in the design of successful vaccines in silico36,37. Vaccine models designed by this approach have been tested for their efficacy by injecting them into mice models38,39. The results of these experimentations have assured positive immune responses in the animal models, which promises its competency as fast-developing alternative for vaccine designing. In this study, the in silico approach has been used for designing a chimeric vaccine consisting of CotE, SlpA and FliC proteins to prevent Clostridium difficile infection. The designed vaccine candidate induces the cytotoxic T cell and helper T cell activation, along with the ability to induce interferon-γ (IFN-γ), interleukin-2 (IL-2) as well as other pro-inflammatory cytokines (like TNF, IL-18, IL-12, etc.).
Results
T cell epitopes and IFN-γ epitope prediction
A significant role is played by the cytotoxic T lymphocytes (CTL) and helper T lymphocytes (HTL) in generation of a long-lasting adaptive immunity against various microbial infections. Long lasting cellular immunity, generated by CTL epitopes, has the ability of eliminating the pathogen infected cells as well as the circulating antigen in the body40. In contrary to the CTL epitopes, HTL epitopes are necessary for the generation of both humoral and cell mediated immunity41. In fact, HTL epitopes are essential for the development of memory helper T cells which are pivotal to both the activation of cytotoxic T cells and stimulation of B lymphocytes responsible for producing antibodies42. Hence, consideration of CTL and HTL epitopes is preferred in an efficient vaccine candidate design which has the ability to elicit antibodies and cytotoxicity. The CTL epitopes from the selected proteins were predicted using the server NetCTL 1.2 (Table 1) (Supplementary Tables S1, S2 and S3). For the epitope prediction, this server uses an algorithm, which considers a combined score of MHC class I binding, proteasomal C terminal cleavage and TAP transport efficiency43. IEDB consensus method was used to screen the epitopes which are good binders to alleles44. The HTL epitopes were predicted using NetMHC II pan 3.2 server (Table 2) (Supplementary Tables S4, S5 and S6). The selected epitopes were further subjected to filters like antigenicity and immunogenicity where the antigenicity and immunogenicity of the epitopes were checked by VaxiJen v2.0 and IEDB class I immunogenicity servers, respectively. The final epitopes which are included in the vaccine construct were visualized on the modelled 3D structures of the SlpA, CotE and FliC proteins, as shown in Fig. 2.
IFN-γ epitopes also play an important role in inducing innate as well as adaptive immunity as it triggers the release of IFN-γ, a signature cytokine released by the neutrophils during Clostridium difficile infection45,46. Therefore, IFN-γ epitopes were predicted from all the three proteins (Supplementary Table_IFN γ epitope), and the best scoring epitopes from each protein were included in the final vaccine construct.
Multi-epitope vaccine construction, structure modelling and validation
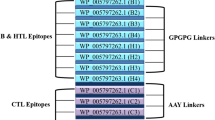
The chimeric vaccine was constructed according to few criteria, namely: (1) it should be antigenic and non-allergenic, (2) it should have overlapping HTL and CTL epitopes (Supplementary Table S7), (3) it must be immunogenic and must be having high affinity to HLA alleles. Following these requirements, a linear vaccine construct was designed using 12 CTL epitopes (4 CTL epitopes from each protein), 9 HTL epitopes (3 HTL epitopes from each protein) and 3 IFN-γ epitopes (1 IFN-γ epitope from each protein). These predicted epitopes were joined with each other using GPGPG linkers. These linkers help in the prevention of potential junctional epitopes and also facilitate immune processing47,48. An adjuvant, Cholera Toxin B (CTB) was attached to the N-terminal of the vaccine using the EAAAK linker (Fig. 3A), as it has been used for the construction of many fusion proteins and acts as a rigid spacer between the protein domains49,50,51. The addition of CTB adjuvants in vaccines improves the formation of a protective immunity as well as prevents chances of autoimmune reactions52. Considering the preliminary criteria for vaccine construction, 6 constructs were made depending upon the position of the epitopes from the three Clostridium difficile proteins (Supplementary Material SM1). The 3D structures of all the constructs were checked for their efficiency based on Z-scores, ERRAT and Ramachandran plot analysis (Supplementary Table S8) and the one with the best scores was chosen as our final vaccine construct. The final vaccine construct has a molecular weight of 51,649.46 Da, consisting of 512 amino acids. A tertiary structure of the final linear vaccine was generated using trRosetta web-server (Fig. 3B)53. The Ramachandran plot of the model structure was generated using RAMPAGE54, which showed that 96.5% residues lie in favoured region, 2.7% lie in allowed and 0.8% residues lie in outlier region (Fig. 3D). This analysis verifies the overall quality of the vaccine model. Also, the generated Z-score of the 3D model was found to be − 8.92, as predicted by ProSA webserver, which indicates that the protein falls in the plot which consists of Z-scores of the already determined structures solved by NMR and X-ray crystallographic experiment (Fig. 3C)55. For further validation, ERRAT analysis was performed which resulted in ERRAT score of 53.3865, representing the percentage of residues falling under 95% rejection limit (Fig. 3E)56. An ERRAT score more than 50 denotes a good quality model; hence a score of 53.3865 further confirms the structural validation of the protein57.
(A) Schematic representation of linear vaccine construct showing CTL epitopes in green colour, HTL epitopes in blue colour and IFN-γ in orange colour. Pale green colour represents GPGPG linkers, which have been used to link epitopes together. CTB adjuvant is represented in yellow colour, joined to the linear vaccine using deep blue EAAAK linker. (B) Tertiary structure of the vaccine. Limon, blue and hot pink colour represents the sheets, loops and helices, respectively. (C) Protein structure validation by ProSA with a Z-score of − 8.92. (D) Ramachandran plot analyses showing 96.5%, 2.7% and 0.8% in favoured, allowed and outlier regions, respectively as predicted by RAMPAGE (E) Protein structure validation by ERRAT score of 53.3865, where X axis denotes residues and Y axis denotes error values.
Antigenicity, allergenicity, immunogenicity and physicochemical properties of the vaccine construct
The capacity to induce humoral and cellular immune responses is defined by immunogenicity while the ability to recognize a specific antigen, which is further accompanied by the generation of an immune response is termed as antigenicity. Hence, in order to recognize an antigen and elicit an immune response with the development of both humoral and cell-mediated immunity, the vaccine should be antigenic as well as immunogenic. The antigenicity of the vaccine was confirmed using VaxiJen v2.0 with a score of 0.9103, where scores of sequences > 0.4 (the predefined cut off or threshold value) are considered to be antigenic58. The immunogenicity of the vaccine was 2.30148, as predicted by the IEDB class I immunogenicity tool (according to IEDB a higher score indicates greater chances in generating an immune response). To assure that no allergenic reactions are triggered by the vaccine, allergenicity of the vaccine was checked. The designed vaccine was predicted to be a non-allergen by AllerTOP webserver. The physicochemical properties of the vaccine were predicted by ExPASy ProtParam tool in order to determine its safety and efficacy (Supplementary Material SM2). The predicted aliphatic index of the vaccine is 76.15, which denotes the vaccine to be stable at high temperatures59. Generally, for a protein, higher the aliphatic index, higher is the protein’s thermostability60. The vaccine has an instability index of 22.16, which means it is quite stable since; in general, an instability index below 40 indicates protein stability60. The grand average hydropathicity (GRAVY) score is − 0.162, which indicates the hydrophilic nature of the vaccine: a lower GRAVY score is the indication of better solubility of the protein in an aqueous environment61. An estimated half-life of the vaccine is 30 h in mammalian reticulocytes, > 20 h in yeast and > 10 h in Escherichia coli by ExPASy ProtParam tool60. There are no transmembrane helices or signal peptides detected in the vaccine structure, which signifies no expression difficulties of the protein and prevention of protein localization, respectively (Supplementary Figs. S1 and S2).
B cell epitope, IL-2 inducing, pro-inflammatory cytokine inducing and IFN-γ inducing HTL epitope predictions
B cells are liable for the humoral immune response62. Helper T cells help in the activation and differentiation of B cells to specific antibody-secreting plasma B cells. Therefore, the presence of B cell epitopes is important in vaccine designing. In this study, the ElliPro server was used to predict the presence of linear/continuous and conformational/discontinuous B cell epitopes in the vaccine construct, under default parameters (Tables 3, 4). The predicted B cell epitopes in the vaccine construct were also visualized using PyMOL (Supplementary Fig. S3).
The selected HTL epitopes were subjected to prediction for IL-2 inducing epitopes using IL2Pred server in order to ensure their effectiveness in inducing interleukin-2 [https://webs.iiitd.edu.in/raghava/il2pred/stat.php]. The results indicated that 5 of our selected HTL epitopes are IL-2 inducing epitopes (Supplementary Table S9), thereby assuring a strong the immune response.
Additionally, the CD4+ epitopes included in our vaccine were checked for pro-inflammatory cytokine inducing epitopes by the server ProInflam63. The server indicates that the epitopes which are pro-inflammatory cytokine inducers can induce cytokines like TNF, IL-18, IL-12, IL-2363. 5 of the 9 HTL epitopes were predicted as inducers of pro-inflammatory cytokines, which has the capacity to induce strong immune response in the body (Supplementary Table S10). The HTL epitopes were also checked for their ability to induce IFN-γ by subjecting them to the Predict Algorithm of IFNepitope server46. Results show that 2 epitopes with positive scores are effective IFN-γ inducers (Supplementary Table S11).
Population coverage
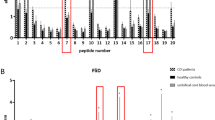
Development of a successful vaccine demands the assessment of HLA allele distribution around the world population64. Based on the difference between regions and ethnicities, the distribution and expression of HLA alleles may vary across the world65. Hence, it is necessary to evaluate if the designed vaccine will be effective against the world population. The selected epitopes in the study have world population coverage of 98.55% (Table 5). In addition, the epitopes showed 99.54%, 99.1%, 94.15%, 95.03%, 95.68% coverage in Europe, United States, China, South Asia and Oceania, respectively (Table 5) (Supplementary Fig. S4). The results are suggestive of the fact that the designed vaccine candidate has the effectiveness to tackle CDI infection globally.
Molecular docking studies
In order to study the interaction between the vaccine construct with the Toll-Like Receptors and MHC molecules, molecular docking studies using HADDOCK server was performed66. TLRs play a major role in adaptive immunity, forming the first line of defence against infections. On interaction with a wide range of microbes and viruses, these TLRs detect the pathogen-associated molecular patterns (PAMPs) on the microbes, which trigger the activation of innate immunity along with orchestration of humoral immune response67. TLR4 is known to recognize bacterial surface and coat proteins while TLR5 recognises flagellin proteins, which leads to an active immune response in body68,69. Hence, it is important for the designed vaccine to interact with target immune receptors like TLRs. The interaction of MHC I and MHC II molecules with the vaccine helps in activating the CTLs and HTLs, which are required for immune response generation70. Thus, the docking analyses of vaccine-MHC molecules is also important for this study.
Docking of the epitope with HLA alleles
The predicted CTL and HTL epitopes were docked with the MHC class I and MHC class II receptor molecules, respectively. This was done for analysing the binding patters of each epitope with their respective receptors (Fig. 4). A detailed overview showing all the binding interactions between the CTL epitopes with MHC I and HTL epitopes with MHC II have been shown in Supplementary Figs. S5 and S6.
Docking of individual epitopes with MHC class I and class II receptors. ‘c’ represents the CTL epitopes docked with MHC class I receptor (grey colour) and ‘h’ represents the HTL epitopes docked with MHC class II receptor (blue colour). All the epitopes are depicted in red colour spheres. ‘F’ denotes epitopes derived from FliC protein, ‘C’ denotes epitopes from CotE protein and ‘S’ denotes the epitopes from SlpA protein. The arrangement of the epitopes are based on their position in the vaccine construct.
Docking of the vaccine with TLR4
The HADDOCK server necessitates the ambiguous interaction restrains (AIRs), which highlights the active and passive residues of the receptor and the ligand required for docking. The interaction in between molecules and available solvent involves the active residues, while the surface neighbour active residues denote the passive residues. Here, the vaccine candidate is docked with the TLR4 heterodimer to study the interaction between vaccine and TLR4. Among the many predicted clusters, the best reliable cluster with the lowest HADDOCK score was subjected to refinement by the HADDOCK Refinement Server. The statistics of the refined structure is detailed in Table 6 and Supplementary Fig. S7. During refinement, 20 structures were clustered into one cluster, resulting in 100% water refined model. After refinement, the model showed a HADDOCK score of − 204.2 ± 2.0 a.u., which suggests good binding affinity between the receptor (TLR4) and ligand (vaccine) as a negative HADDOCK score suggests better docking. Less water-exposed protein surface and close binding proximity is indicated by the Buried Surface Area (BSA) of 5042.9 ± 160.6 Å271. A low RMSD score of 0.3 ± 0.2 Å of the docked structure indicates a good quality model. A detailed overview of the docked complex along with interacting amino acid residues are shown in Supplementary Fig. S8. The docked complex along with the interacting residues is shown in Fig. 5 and detailed interaction is given in Supplementary Material SM3.
(A) Figure acquired after molecular docking which shows docked TLR4/MD2-vaccine complex. The vaccine construct is shown in green while the TLR4 dimer is shown in hotpink, and the blue colour represents the MD2 co-receptor. (B) Residue interaction between the docked tetramer TLR4 / MD2 (chain A) and the vaccine (chain B). (C) Few hydrogen bonds shown within vaccine-TLR4 complex are focused.
Docking of the vaccine with TLR5
The HADDOCK server necessitates the ambiguous interaction restrains (AIRs), which highlights the active and passive residues of the receptor and the ligand. The interaction in between molecules and available solvent involves the active residues, while the surface neighbour active residues denote the passive residues. Here, the vaccine candidate is docked with TLR5 to study the interaction between vaccine and TLR5 molecule. Among the many predicted clusters, the best reliable cluster with the lowest HADDOCK score was subjected to refinement by the HADDOCK Refinement Server. The statistics of the refined structure is detailed in Table 7 and Supplementary Fig. S9. During refinement, 20 structures were clustered into one cluster, resulting in 100% water refined model. After refinement, the model showed a HADDOCK score of − 259.8 ± 3.1 a.u., which suggests good binding affinity between the receptor (TLR5) and ligand (vaccine) as a negative HADDOCK score suggests better docking. Less water-exposed protein surface and close binding proximity is indicated by the Buried Surface Area (BSA) of 5358.9 ± 45.0 Å271. A low RMSD score of 0.3 ± 0.2 Å of the docked structure indicates a good quality model. A detailed overview of the docked complex along with interacting amino acid residues are shown in Supplementary Fig. S10. The docked complex along with the interacting residues is shown in Fig. 6 and detailed interaction is given in Supplementary Material SM4.
(A) Figure acquired after molecular docking which shows docked TLR5-vaccine complex. The vaccine construct is shown in teal while the TLR5 dimer is shown in red. (B) Residue interaction between the docked tetramer TLR5 (chain A) and the vaccine (chain B). (C) Few hydrogen bonds shown within vaccine-TLR5 complex are focused.
Docking of the vaccine with MHC I and MHC II receptors
The HADDOCK server requires the ambiguous interaction restrains (AIRs), which highlights the active and passive residues of the receptor and the ligand. The interaction in between molecules and available solvent involves the active residues, while the surface neighbour active residues denote the passive residues. Here, the vaccine candidate is docked with the MHC I and MHC II molecules separately to study the interaction between vaccine and MHC molecules. Among the many predicted clusters, the best reliable clusters for both the complexes having lowest HADDOCK scores were subjected to refinement by the HADDOCK Refinement Server. The statistics of the refined structure is detailed in Table 8 and Supplementary Figs. S11–S12. During refinement, 20 structures for each complex were clustered into one cluster, resulting in 100% water refined model. After refinement, the vaccine-MHC I and vaccine-MHC II docked models showed the HADDOCK scores of − 208.6 ± 8.4 a.u. and − 252.4 ± 4.1 a.u., respectively, which suggests good binding affinity between the receptors (MHC I and MHC II) and ligand (vaccine) as negative HADDOCK score suggests better docking. Less water-exposed protein surface and close binding proximity is indicated by the Buried Surface Area (BSA) of 3956.3 ± 75.1 Å2 for vaccine-MHC I complex and 5169.4 ± 45.9 Å2 for vaccine-MHC II complex71. Low RMSD score of 0.3 ± 0.2 Å (vaccine-MHC I complex) and 0.3 ± 0.2 Å (vaccine-MHC II complex) of the docked structures indicates them to be good quality models. A detailed overview of the docked complexes along with interacting amino acid residues are shown in Supplementary Figs. S13 and S14. The docked complexes along with their interacting residues are shown in Fig. 7 (vaccine-MHC I complex) and Fig. 8 (vaccine-MHC II complex). The detailed interactions of vaccine-MHC I and vaccine-MHC II complexes are given in Supplementary Materials SM5 and SM6, respectively.
(A) Figure acquired after molecular docking which shows docked MHC class I receptor-vaccine complex. The vaccine construct is shown in green while the MHC class I receptor is shown in blue. (B) Residue interaction between the docked MHC class I receptor (chain A) and the vaccine (chain B). (C) Few hydrogen bonds shown within vaccine-MHC class I receptor complex are focused.
(A) Figure acquired after molecular docking which shows docked MHC class II receptor-vaccine complex. The vaccine construct is shown in deep blue while the MHC class II receptor is shown in purple. (B) Residue interaction between the docked MHC class II receptor (chain A) and the vaccine (chain B). (C) Few hydrogen bonds shown within vaccine-MHC class II receptor complex are focused.
Binding affinity analysis
The binding affinity of a complex or the Gibbs free energy (∆G) in terms of thermodynamics, is an important quantity to determine if an interaction will actually occur or not in the cell at specific conditions72,73. Hence, the binding affinity of the docked structures were analysed using the PRODIGY web server72, which provided the ∆G values of the vaccine-TLR4, vaccine-TLR5, vaccine-MHC I and vaccine-MHC II complexes as − 17.2, − 18.2, − 16.7 and − 17.2, respectively. It can be concluded that all the interactions are energetically feasible, which is evident from the negative values of Gibbs free energy. The corresponding ∆G values along with the dissociation constant (Kd) values of the docked complexes are provided in the Table 9.
Energy minimization and molecular dynamics simulation (MDS) of vaccine-TLR complexes
The evaluated potential energy of the vaccine-TLR4 complex was − 9.83e + 06 kJ/mol and the plot for the same indicates a steady convergence of potential energy during this step (Supplementary Fig. S15). The consistent low energy of − 9.83e + 06 kJ/mol of the energy minimized structure supported it to a reasonable starting structure for carrying out further steps of MDS. An equilibration phase for 100 ps was done for studying the effect of temperature, pressure and other thermodynamic properties. The NVT ensemble was used for conducting the first equilibration phase to check if the system was stable at the desired temperature of 300 K. Results for this step indicated that the temperature of the system quickly reached 300 K which was maintained during the equilibration phase with very minimal fluctuations indicating the system to be stable at 300 K (Supplementary Fig. S15). In addition, the NPT ensemble evaluated the stability of the system with respect to the desired pressure of 1 bar. In this case the results indicated that the pressure of 1 bar was also maintained during equilibration and negligible fluctuations observed in the plot confirmed its stability for other trajectory analyses. The average density of the system was found to be 1017.3 kg/m3 with a total drift of 1.6 kg/m3. Similar results were observed for vaccine-TLR5 complex at same conditions of temperature and pressure (Supplementary Fig. S15). The computed average potential energy for this system was − 1.12e + 07 kJ/mol, with a density of 1009 kg/m3 having a total drift of 1.5 kg/m3. After a simulation period of 18 ns, a trajectory analysis was also performed to check the flexibility and stability of the vaccine-TLR complexes. The plots for the radius of gyration demonstrated the compactness of the protein along its axes (Supplementary Fig. S15). Very mild fluctuations in the plot of the RMSD backbone indicate that the vaccine-TLR complexes formed strong interactions among themselves hence, attaining greater stability of the complex over time. The RMSF plots for the vaccine-TLR complexes identified regions of high flexibility as indicated by the high peaks observed in the RMSF plot (Fig. 9). Though there were few fluctuating residues, the plots are also suggestive of the fact that those fluctuations were minimal and the vaccine-TLR complexes were mostly stable over time (Fig. 9).
Codon optimization and in silico cloning of vaccine
The Java Codon Adaptation Tool (JCat) has been used to optimize codon usage of the vaccine construct in E. coli (strain K12) for optimal protein expression. The optimized nucleotide sequence has Codon Adaptation Index (CAI) of 1.0 and the average GC content of the adapted sequence is 55.46875, showing the vaccine candidate's potential for good expression in E. coli host74. The ideal GC content percentage range is between 30 and 7074. Finally, the recombinant plasmid sequence was designed using SnapGene tool by inserting the adapted codon sequences into the pET28a (+) vector (Fig. 10).
Immune simulation
The C-IMMSIM server was used for generating an in silico immune response in order to assess the immunogenicity of the multi-epitope vaccine75. The vaccine administration protocol consisted in three injections at day 1, 30 and 60. A bacterial infection was started at day 240 in order to check the efficacy of the vaccination. As expected, the elicited secondary and tertiary responses were significantly higher when compared to the primary response. The decline in antigenic concentration with normal high levels of immunoglobulin activity (i.e., IgM, IgG1 and IgG2) is shown in panel A of Fig. 11. Additionally, a possible class switching and memory development was suggested due to the presence of multiple long lasting B cell isotypes (Fig. 11B). The pre-activation of TCs and a relatively high response in TH (helper) and TC (cytotoxic) cell populations was observed during vaccination (panels C and E of Fig. 11) (also Supplementary Fig. S16). During exposure, macrophages showed higher activities while the NK and dendritic cell activity was recorded to be consistent (Supplementary Fig. S16). The high levels of cytokines like IFN- γ and IL-2 supports the generation of an effective immune response (panel D of Fig. 11). The bacterial challenge following the vaccination shows the efficacy of the vaccine as the bacterial surge is virtually absent; mainly due to the protective action of high concentration of specific antibodies (panel A of Fig. 11). A control simulation was also performed consisting of an injection of replicating bacteria at time zero in absence of vaccination. The results indicate that without the preventive effect of the vaccine, there is an uncontrollable growth of the bacteria; even though the immune system faces the challenge by responding to the pathogen, it fails to control its growth due to the extreme replication speed (Supplementary Fig. S17).
Discussion
During the last two decades, the morbidity and mortality rate of Clostridium difficile infection has vastly increased in both community and hospital settings76. Colonization is aided by dysbiosis of internal gut microbiota, which results mostly from elevated antibiotic therapy. Multiple proteins like CotE, SlpA and FliC are responsible for colonization, adherence and persistence of Clostridium difficile19,20,23. Hence, these proteins are selected for development of a multi-epitope vaccine candidate by using immunoinformatic approaches that is both time saving and cost effective73,77,78,79.
In this study, the designed multi-epitope vaccine not only elicits cell-mediated immunity, but also triggers humoral immune responses. Moreover, including only short immunogenic peptide sequences avoids allergic responses and prevents antigenic load80,81. The major advantages of the multi-epitope vaccine over conventional vaccines are as follows: (i) TCRs can identify several MHC Class I and Class II epitopes from diverse T cell subsets; (ii) the overlapping CTL, HTL and B cell epitopes have the capability to simultaneously activate humoral and cellular immune responses; (iii) linking a vaccine adjuvant guarantees long-lasting immune responses with improved immunogenicity; (iv) complications of the in vitro antigen expression and the complexity of pathogen culture can also be prevented82,83,84,85. The design of multi-epitope vaccines is a new field that has already gained significance, and the vaccines developed by this method have not only demonstrated protective immunity in vivo, but have also entered in phase I clinical trials86,87,88.
In this present study, an adjuvant, Cholera Toxin B (CTB) has been attached to N-terminal end of the vaccine using suitable linkers. Studies shows use of non-toxic CTB as a strong bacterial adjuvant in vaccines has successfully triggered an enhanced immune response by activation of CD4 + T cell responses89,90,91. Among various filters used for epitope screening, the CTL and HTL epitopes were checked for antigenicity, immunogenicity and promiscuity. In addition, overlapping CTL and HTL epitopes were also considered for final vaccine construction. The study confirms the antigenicity, immunogenicity and non-allergenicity of the final vaccine construct as predicted by VaxiJen v2.0, IEDB class I immunogenicity and AllerTOP servers, respectively92. Using the ExPASy ProtParam tool, the other physicochemical properties of the vaccine were analysed. The construct's molecular weight was 51,649.46 Da and the instability index was measured at 22.16, which classifies the vaccine as a stable protein. The vaccine's GRAVY index was − 0.162 (lower the GRAVY value, higher the solubility), which represents the polar identity of the vaccine and its efficient interaction with water, indicating high solubility61. The aliphatic index of 76.15 showed the protein as thermostable59. Half-life of 30 h in mammalian reticulocytes, > 20 h in yeast and > 10 h in Escherichia coli refers to the time taken by the protein to achieve 50% of its initial concentration after its synthesis in the cell. Ramachandran plot analysis of the vaccine using RAMPAGE showed 96.5%, 2.7% and 0.8% in favoured, allowed and outlier regions, respectively. An ERRAT score of 53.3865 further validated the overall quality of the vaccine. The ProSA web server predicted a Z-score of − 8.92, which indicates that the protein falls in a plot consisting of Z-scores of previously determined structures, which were solved using NMR and X-ray crystallography.
Our vaccine contains both CD8 + and CD4 + overlapping T cell epitopes which ensures stronger immune response in the body. Additionally, all the HTL epitopes were subjected to the prediction of IFN-γ, IL-2 and pro-inflammatory cytokine inducing epitopes. When the 9 HTL epitopes were screened for IL2Pred prediction, 5 epitopes were predicted to be strong inducers of interleukin-2 (Supplementary Table S9). Similarly, 5 epitopes were predicted to be inducing pro-inflammatory cytokines (including TNF) when subjected to the ProInflam web server (Supplementary Table S10). While checking for IFN-γ inducing HTL epitopes, the 15mer peptide sequences were subjected to the Predict Algorithm of the IFNepitope server, from which two epitopes were predicted as IFN-γ inducing (Supplementary Table S11). From these we can infer that few epitopes among our selected ones are responsible for inducing multiple immune factors. Thus, these epitopes might contribute for a robust immune response in the body.
Studies have shown that Toll-Like Receptor 4 (TLR4) recognizes spore protein CotE and surface layer protein SlpA whereas flagellar protein FliC is recognized by Toll-Like Receptor 5 (TLR5)2,93,94,95,96. TLR4 is expressed in multiple types of immune cells, such as immature dendritic cells, monocytes, granulocytes and macrophages97. In C. difficile-associated infection by SlpA, a significance of TLR4 is producing the immune response, which is required to clear the bacterium94. In addition, direct interaction of TLR4 with CTB is also responsible for CTB mediated activation of TLR498. The ELISA-based assays have already shown that by binding directly to it, CTB can induce NF-κB activation in TLR4 receptor cells. Flagellin, on the other hand, may stimulate innate immunity by CCL20 and IL-8 production through the interaction of flagellar proteins with TLR599,100. Also, a production of the inflammatory cytokine interleukin-22 (IL-22) is observed during TLR5 stimulation in body96,101. Molecular docking studies have assessed the vaccine's pattern of interaction with TLR4 and TLR5 (Figs. 5 and 6). The docking analysis of TLR4 with vaccine construct revealed that during this interaction there were 2 salt bridges and 19 hydrogen bonds formed. Similarly, docking analysis of TLR5 with the designed vaccine showed 3 salt bridges and 18 hydrogen bonds formed during the molecular interaction. Likewise, docking studies of the vaccine with MHC I and MHC II was also performed to check the binding of the vaccine with MHC molecules (Figs. 7 and 8). The docking analysis of MHC I with vaccine construct revealed that in this interaction there were 1 salt bridge and 17 hydrogen bonds formed; and 1 salt bridge and 13 hydrogen bonds were formed in the docked structure of MHC II and vaccine. Molecular Dynamics Simulation (MDS) of the vaccine-TLR complexes was performed for verifying the stability and flexibility of the complexes in various experimental conditions like temperature and pressure. A trajectory run of 18 ns showed stable interactions between the vaccine and the TLRs during the simulation run with limited fluctuations (Fig. 9). The RMSF graphs generated showed regions with high peaks which hint towards high flexibility of the vaccine-TLR complexes (Fig. 9). The immune simulation studies conducted on the designed vaccine construct confirmed its ability to clear the antigen on secondary exposure by eliciting specific immune responses (Fig. 11).
A similar array of in silico analysis was performed by Bazhan and his colleagues where they have designed a T-cell multi epitope vaccine against Ebola virus. The T cell epitopes were predicted using the same server, IEDB—Immune Epitope Database; and the vaccine candidate constructed using the suitable epitopes were found to be immunogenic when expressed in mice38. Similarly, Forountan and his co-workers have designed their experiments, where they have assessed the allergenicity and other physicochemical properties of their vaccine candidate against Toxoplasma gondii by a series of immunoinformatics tools, which are similar to the ones we have used, and their wet lab validation suggested that the multi-epitope vaccine was able to trigger strong humoral and cellular responses in mice39. The antigenicity, allergenicity and physicochemical scores obtained by our vaccine candidate were comparable, and in fact few values like antigenicity, aliphatic index, instability index and Ramachandran plot have better scores than the values reported by Forountan and his team in their published work. These studies strengthen the fact that the vaccine candidates designed in silico using computational tools can be a successful strategy for designing an efficient vaccine candidate against diseases. Similar immunoinformatics concepts have been applied in designing multi-epitope vaccines against SARS-CoV-273, Dengue102, Nipah Virus103, Malaria104, Hendra virus105, and many more. Additionally, development of vaccine candidates against cancer antigens has also been seen35,106. The CTL, HTL, and IFN-γ epitopes found in the vaccine are capable of activating the regulation of the respective immune cells of the host, which may induce the triggering of other immune cells through complex signalling. The vaccine candidate proposed in the study is intended to be delivered via injection following an intramuscular route. Intramuscular administration (I.M) is preferred for the delivery of the vaccine candidate since it is easy to perform and well tolerated, with a low risk for adverse reactions at the site of injection and is the most commonly used route for licensed vaccines107. Recent studies also show that intramuscular immunization is a promising strategy for induction of strong systemic CTL response108. In addition, CDC recommends that vaccines containing an adjuvant should be injected into a muscle rather than any other routes since they can cause local irritation, induration, skin discolouration, inflammation, and granuloma formation109,110.
Materials and methods
Sequence retrieval and PDB structure retrieval
The FASTA sequences of SlpA, FliC and CotE proteins were retrieved from NCBI database having the accession numbers CAJ69681.1, AJP09935.1 and CAJ68298.1, respectively. The PDB structures of TLR4, TLR5, MHC I and MHC II receptors were retrieved from Protein Data Bank with accession ID 3FXI, 3J0A, 1I1Y and 1KG0, respectively.
T cell epitope and IFN-γ epitope prediction
NetCTL 1.2 server was used for predicting the 9-mer long CTL epitopes with 54–89% sensitivity and 94–99% specificity43. These epitopes were recognized by commonly occurring HLA class I supertypes in human population, which are: A1, A2, A3, A24, A26, B7, B8, B27, B39, B44, B58 and B62. In addition, Immune Epitope Consensus (IEDB) tool also detected CTL epitopes which were recognized by other HLA class I alleles44. In contrary, 15-mer HTL epitopes, having an affinity to HLA class II alleles were predicted by NetMHCII pan 3.2 server111. Based on the idea of percentile rank given by the server, all the epitopes are classified as strong, intermediate and non-binders to HLA alleles with threshold values of 2, 10 and > 10% respectively. All the predicted epitopes were verified for antigenicity by VaxiJen v2.0 server58, along with immunogenicity check using IEDB class I immunogenicity web server112.
IFN-γ epitopes from the chosen bacterial proteins were predicted using the Scan Algorithm of the IFNepitope server46. This server uses various approaches like as machine learning strategy, motive-based analysis and accuracy hybrid approach for epitope prediction and has a maximum accuracy of 81.39%46.
Multi-epitope vaccine construction, modelling and validation of vaccine structure
The selected CTL, HTL and IFN-γ epitopes screened from the SlpA, CotE and FliC proteins were linked together with GPGPG linkers. Also, EAAAK linker was used to attach Cholera Toxin B (CTB) adjuvant to the vaccine at the N terminal. The designed vaccine linear construct was subjected to tertiary modelling by trRosetta webserver53. The resulted 3D model was validated using ERRAT, Rampage and ProSA webservers54,55,56 and these servers were used for the prediction of ERRAT score, Ramachandran plot and Z-score of the protein 3D model.
Antigenicity, immunogenicity, allergenicity and physicochemical properties of the vaccine model
To predict the antigenicity of the vaccine, VaxiJen v2.0 server was used. Sequences with a score more than 0.4 threshold value are considered antigenic. VaxiJen v2.0 server uses viral and bacterial databases which contain 100 identified antigens and 100 non-antigens. Models are evaluated using these datasets with 70–89% accuracy, using internal leave-one-out cross-validation and external validation. The immunogenicity of the vaccine construct was checked using IEDB class I immunogenicity server, where more the score, more is the immunogenicity. Similarly, the allergenicity of the vaccine was verified using AllerTOP server, which employs auto-cross-covariance (ACC) grouping of protein sequences into uniform equal-length vectors92. To evaluate other physicochemical properties like aliphatic index, theoretical pI, half-life, molecular weight, instability index and GRAVY score of the vaccine, ExPASy ProtParam tool was used60. The vaccine construct was also checked by SignalP4.1 and TMHMM server v2.0 for the presence of any signal peptides and transmembrane helices, respectively113,114.
B cell epitope, IL2 inducing, pro-inflammatory cytokine inducing and IFN-γ inducing HTL epitope predictions
The presence of B cell epitopes in vaccine construct was confirmed using ElliPro webserver from IEDB115. The server predicted continuous/linear and discontinuous/conformational B cell epitopes in the vaccine.
The IL-2 epitopes were predicted by IL2Pred webserver (https://webs.iiitd.edu.in/raghava/il2pred/). The server predicts the IL-2 inducing positive and negative epitopes based on the derived epitopes from IEDB dataset. Similarly, Pro-inflammatory cytokine inducing CD4 + T cell epitopes were predicted by using the ProInflam web server (http://metagenomics.iiserb.ac.in/proinflam/) This server identifies the epitopes which are pro-inflammatory cytokine inducers and can induce cytokines like TNF, IL-18, IL-12, IL-2363. The IFN-γ inducing HTL epitopes, selected for the vaccine were identified using IFNepitope server’s Predict Algorithm46. This server identifies the IFN-γ epitopes on the basis of their respective scores, where a positive score indicates the epitopes to be IFN-γ secreting46.
Population coverage
The population coverage for the epitopes of the designed vaccine was checked for the world population, United States, Europe, China, South Asia and Oceania using IEDB population coverage analysis tool (http://tools.iedb.org/population/)65. All the parameters for the study were kept at default and the coverage was checked against MHC class I and MHC class II alleles.
Docking of the vaccine construct with TLR4, TLR5, MHC I and MHC II and binding affinity
The vaccine was subjected to molecular docking studies for better understanding of its interaction with Toll-Like Receptors and MHC molecules, which aid in better and more stable immune response. For this interaction study, firstly active and passive residues for the vaccine and receptor molecules used in this study are predicted using CPORT116 and HADDOCK 2.4 server then helps in performing docking of the vaccine with TLR4 and TLR5 as well as MHC I and MHC II66. Among the docked clusters, the best cluster was chosen based on lowest HADDOCK score. The chosen model was subjected to refinement by HADDOCK Refinement Server. The final docked structures after refinement were subjected to PRODIGY web-server (https://bianca.science.uu.nl/prodigy/) for prediction of binding energy72. In addition, PDBsum was used to map the interacting residues between the docked chains (i.e., the vaccine, the TLRs and the MHCs)117.
Energy minimization and molecular dynamics simulation
Vaccine-TLR complexes
For performing Molecular Dynamics Simulation (MDS) and energy minimization of the vaccine-TLR complexes, a Linux-based command line program, GROMACS (GROningen MAchine for Chemical Simulations) was used118. In order to mimic the experimental conditions, the complexes were subjected to MDS. Physical conditions like pressure and temperature were imitated in this study using the canonical ensemble NVT and the isobaric and isothermal ensemble NPT. Mimicking the experimental conditions, temperature set for the desired simulation was 300 K. OPLS-AA (Optimized Potential for Liquid Simulation-All Atom) force field constraint for energy minimization and equilibration was used to generate a topology file used for the simulation. In order to obtain the periodic image of the complex structures 2 nm apart, the structures were positioned at a distance of 1 nm from the edge of the cube filled with water molecules. An equilibrated three-point water model, spc216 was used as the solvent to simulate the vaccine with periodic boundary conditions. After neutralizing the system by addition of charged ions, the net charge of the vaccine-TLR constructs were evaluated. The system’s temperature and pressure was also stabilized by performing the NVT and NPT equilibration for 100 pico-seconds (ps) consisting of 50,000 steps. In order to find the Root Mean Square Deviation (RMSD) of backbone and Root Mean Square Fluctuation (RMSF) of side chain, a 18 ns Molecular Dynamics (MD) simulation run was carried out for the energy minimised structure of the complexes. To analyse the compactness of the protein structure, the radius of gyration was also plotted. Xmgrace, software based on Linux, was employed to visualize the simulation graphs generated after the simulation runs119.
Codon optimization and in silico cloning of vaccine
The Java Codon Adaptation Tool (JCat) was used to optimize codon and for reverse translation that generated the vaccine's cDNA sequence that can be used for an efficient expression in E. coli K-12 strain74. Additionally, the SnapGene software (from Insightful Science; available at snapgene.com) was used to insert the optimized multi-epitope vaccine’s DNA sequence into the pET28a (+) vector.
Immune simulation
The C-IMMSIM server (https://kraken.iac.rm.cnr.it/C-IMMSIM/) was used to perform in silico immune simulation in order to characterize the immunogenicity and immune response of the engineered peptide75. C-IMMSIM is an agent-based model that uses position-specific scoring matrices (PSSM) derived from machine learning techniques for predicting immune interactions. For most of the vaccines currently in use, 4 weeks is the minimum recommended time between the first and second dose120. The entire simulation ran for 1400 time steps which are about 15 months (a time step is about 8 h). Three peptide injections were given at day 1, day 30 (i.e., after 1 month) and day 60 (i.e., after 2 months), followed by the bacterial challenge at day 240 (i.e., after 8 months = 6 months after end of vaccination) in order to check the efficacy of the vaccination process.
Conclusion
Increased repetitive occurrence of CDI is observed in patients who receive antibiotic therapies in hospital environment. No proper prevention except antibiotic treatments and toxoid vaccines prescribed after incidence of Clostridium difficile infection is available for patients. Toxoid vaccines act on the secreted toxin A and toxin B after C. difficile colonize in gastro-intestinal tract. However, the concern regarding bacterial attachment, colonization and spore formation remains untouched. The in silico approaches can be used to develop an efficient vaccine using target antigenic proteins in less time and at minimal cost. In this study, immunoinformatic tools have been applied for constructing a multi-epitope chimeric vaccine candidate against Clostridium difficile. The designed vaccine showed both antigenic as well as immunogenic properties with no allergenic responses. Strong immune responses were validated using Immune Simulation studies. Also, detailed molecular docking and molecular dynamics simulation studies have shown the vaccine to be stable.
References
Johnson, S. & Gerding, D. N. Clostridium difficile-associated diarrhea. Clin. Infect. Dis. 26, 1027–1034. https://doi.org/10.1086/520276 (1998) (quiz 1035–1026).
Ghose, C. et al. Immunogenicity and protective efficacy of recombinant Clostridium difficile flagellar protein FliC. Emerg. Microbes Infect. 5, e8. https://doi.org/10.1038/emi.2016.8 (2016).
Ofosu, A. Clostridium difficile infection: A review of current and emerging therapies. Ann. Gastroenterol. 29, 147–154. https://doi.org/10.20524/aog.2016.0006 (2016).
Bartlett, J. G. Clinical practice. Antibiotic-associated diarrhea. N. Engl. J. Med. 346, 334–339. https://doi.org/10.1056/NEJMcp011603 (2002).
Korman, T. M. Diagnosis and management of Clostridium difficile infection. Semin. Respir. Crit. Care Med. 36, 31–43. https://doi.org/10.1055/s-0034-1398741 (2015).
Bartlett, J. G. Historical perspectives on studies of Clostridium difficile and C. difficile infection. Clin. Infect. Dis. 46(Suppl 1), S4-11. https://doi.org/10.1086/521865 (2008).
Mori, N. & Takahashi, T. Characteristics and immunological roles of surface layer proteins in Clostridium difficile. Ann. Lab. Med. 38, 189–195. https://doi.org/10.3343/alm.2018.38.3.189 (2018).
Hota, S. S. et al. Determining mortality rates attributable to Clostridium difficile infection. Emerg. Infect. Dis. 18, 305–307. https://doi.org/10.3201/eid1802.101611 (2012).
Haubitz, S. et al. Outcome of Clostridioides difficile infections treated in a Swiss tertiary care hospital: an observational study. Swiss. Med. Wkly. 150, w20173. https://doi.org/10.4414/smw.2020.20173 (2020).
Peery, A. F. et al. Burden of gastrointestinal disease in the United States: 2012 update. Gastroenterology 143, 1179–1187. https://doi.org/10.1053/j.gastro.2012.08.002 (2012).
Murphy, S., Xu, J. & Kochanek, K. Deaths: Preliminary data for 2010. National vital statistics reports, vol. 60, No. 4. (Centers for Disease Control and Prevention, 2012).
Pechine, S. et al. Immunological properties of surface proteins of Clostridium difficile. J. Med. Microbiol. 54, 193–196. https://doi.org/10.1099/jmm.0.45800-0 (2005).
Permpoonpattana, P. et al. Surface layers of Clostridium difficile endospores. J. Bacteriol. 193, 6461–6470. https://doi.org/10.1128/JB.05182-11 (2011).
Permpoonpattana, P. et al. Functional characterization of Clostridium difficile spore coat proteins. J. Bacteriol. 195, 1492–1503. https://doi.org/10.1128/JB.02104-12 (2013).
Hong, H. A. et al. The spore coat protein CotE facilitates host colonization by Clostridium difficile. J. Infect. Dis. 216, 1452–1459. https://doi.org/10.1093/infdis/jix488 (2017).
Fagan, R. P. & Fairweather, N. F. Biogenesis and functions of bacterial S-layers. Nat. Rev. Microbiol. 12, 211–222. https://doi.org/10.1038/nrmicro3213 (2014).
Calabi, E., Calabi, F., Phillips, A. D. & Fairweather, N. F. Binding of Clostridium difficile surface layer proteins to gastrointestinal tissues. Infect. Immun. 70, 5770–5778. https://doi.org/10.1128/iai.70.10.5770-5778.2002 (2002).
Bianco, M. et al. Immunomodulatory activities of surface-layer proteins obtained from epidemic and hypervirulent Clostridium difficile strains. J. Med. Microbiol. 60, 1162–1167. https://doi.org/10.1099/jmm.0.029694-0 (2011).
Kirk, J. A., Banerji, O. & Fagan, R. P. Characteristics of the Clostridium difficile cell envelope and its importance in therapeutics. Microb. Biotechnol. 10, 76–90. https://doi.org/10.1111/1751-7915.12372 (2017).
Bruxelle, J. F. et al. Immunogenic properties of the surface layer precursor of Clostridium difficile and vaccination assays in animal models. Anaerobe 37, 78–84. https://doi.org/10.1016/j.anaerobe.2015.10.010 (2016).
Aubry, A. et al. Modulation of toxin production by the flagellar regulon in Clostridium difficile. Infect. Immun. 80, 3521–3532. https://doi.org/10.1128/IAI.00224-12 (2012).
Dapa, T. & Unnikrishnan, M. Biofilm formation by Clostridium difficile. Gut Microbes 4, 397–402. https://doi.org/10.4161/gmic.25862 (2013).
Tasteyre, A., Barc, M. C., Collignon, A., Boureau, H. & Karjalainen, T. Role of FliC and FliD flagellar proteins of Clostridium difficile in adherence and gut colonization. Infect. Immun. 69, 7937–7940. https://doi.org/10.1128/IAI.69.12.7937-7940.2001 (2001).
Bruxelle, J. F., Pechine, S. & Collignon, A. Immunization strategies against Clostridium difficile. Adv. Exp. Med. Biol. 1050, 197–225. https://doi.org/10.1007/978-3-319-72799-8_12 (2018).
Wang, S., Wang, Y., Cai, Y., Kelly, C. P. & Sun, X. Novel chimeric protein vaccines against Clostridium difficile infection. Front. Immunol. 9, 2440. https://doi.org/10.3389/fimmu.2018.02440 (2018).
Henderson, M., Bragg, A., Fahim, G., Shah, M. & Hermes-DeSantis, E. R. A review of the safety and efficacy of vaccines as prophylaxis for Clostridium difficile infections. Vaccines (Basel). https://doi.org/10.3390/vaccines5030025 (2017).
Palm, N. W. & Medzhitov, R. Pattern recognition receptors and control of adaptive immunity. Immunol. Rev. 227, 221–233. https://doi.org/10.1111/j.1600-065X.2008.00731.x (2009).
Iwasaki, A. & Medzhitov, R. Regulation of adaptive immunity by the innate immune system. Science 327, 291–295. https://doi.org/10.1126/science.1183021 (2010).
Sakaguchi, S., Miyara, M., Costantino, C. M. & Hafler, D. A. FOXP3+ regulatory T cells in the human immune system. Nat. Rev. Immunol. 10, 490–500. https://doi.org/10.1038/nri2785 (2010).
Groothuis, T. A., Griekspoor, A. C., Neijssen, J. J., Herberts, C. A. & Neefjes, J. J. MHC class I alleles and their exploration of the antigen-processing machinery. Immunol. Rev. 207, 60–76. https://doi.org/10.1111/j.0105-2896.2005.00305.x (2005).
Shastri, N., Cardinaud, S., Schwab, S. R., Serwold, T. & Kunisawa, J. All the peptides that fit: The beginning, the middle, and the end of the MHC class I antigen-processing pathway. Immunol. Rev. 207, 31–41. https://doi.org/10.1111/j.0105-2896.2005.00321.x (2005).
Jutras, I. & Desjardins, M. Phagocytosis: At the crossroads of innate and adaptive immunity. Annu. Rev. Cell Dev. Biol. 21, 511–527. https://doi.org/10.1146/annurev.cellbio.20.010403.102755 (2005).
Krogsgaard, M. & Davis, M. M. How T cells “see” antigen. Nat. Immunol. 6, 239–245. https://doi.org/10.1038/ni1173 (2005).
Ahlers, J. D. & Belyakov, I. M. Molecular pathways regulating CD4(+) T cell differentiation, anergy and memory with implications for vaccines. Trends Mol. Med. 16, 478–491. https://doi.org/10.1016/j.molmed.2010.07.007 (2010).
Chauhan, V., Rungta, T., Goyal, K. & Singh, M. P. Designing a multi-epitope based vaccine to combat Kaposi Sarcoma utilizing immunoinformatics approach. Sci. Rep. 9, 2517. https://doi.org/10.1038/s41598-019-39299-8 (2019).
Faisal, A. M., Imtiaz, S. H., Zerin, T., Rahman, T. & Shekhar, H. U. Computer aided epitope design as a peptide vaccine component against Lassa virus. Bioinformation 13, 417–429. https://doi.org/10.6026/97320630013417 (2017).
Atapour, A. et al. Designing a fusion protein vaccine against HCV: An in silico approach. Int. J. Pept. Res. Ther. 25, 861–872 (2019).
Bazhan, S. I. et al. In silico designed ebola virus T-cell multi-epitope DNA vaccine constructions are immunogenic in mice. Vaccines 7, 34 (2019).
Foroutan, M., Ghafarifar, F., Sharif, Z. & Dalimi, A. Vaccination with a novel multi-epitope ROP8 DNA vaccine against acute Toxoplasma gondii infection induces strong B and T cell responses in mice. Comp. Immunol. Microbiol. Infect. Dis. 69, 101413 (2020).
Doherty, P. C., Allan, W., Eichelberger, M. & Carding, S. R. Roles of alpha beta and gamma delta T cell subsets in viral immunity. Annu. Rev. Immunol. 10, 123–151. https://doi.org/10.1146/annurev.iy.10.040192.001011 (1992).
Panina-Bordignon, P. et al. Universally immunogenic T cell epitopes: Promiscuous binding to human MHC class II and promiscuous recognition by T cells. Eur. J. Immunol. 19, 2237–2242. https://doi.org/10.1002/eji.1830191209 (1989).
Wilson, C. C. et al. Identification and antigenicity of broadly cross-reactive and conserved human immunodeficiency virus type 1-derived helper T-lymphocyte epitopes. J. Virol. 75, 4195–4207. https://doi.org/10.1128/JVI.75.9.4195-4207.2001 (2001).
Larsen, M. V. et al. Large-scale validation of methods for cytotoxic T-lymphocyte epitope prediction. BMC Bioinform. 8, 424. https://doi.org/10.1186/1471-2105-8-424 (2007).
Moutaftsi, M. et al. A consensus epitope prediction approach identifies the breadth of murine T(CD8+)-cell responses to vaccinia virus. Nat. Biotechnol. 24, 817–819. https://doi.org/10.1038/nbt1215 (2006).
El-Zaatari, M. et al. Tryptophan catabolism restricts IFN-gamma-expressing neutrophils and Clostridium difficile immunopathology. J. Immunol. 193, 807–816. https://doi.org/10.4049/jimmunol.1302913 (2014).
Dhanda, S. K., Vir, P. & Raghava, G. P. Designing of interferon-gamma inducing MHC class-II binders. Biol. Direct 8, 30. https://doi.org/10.1186/1745-6150-8-30 (2013).
Saadi, M., Karkhah, A. & Nouri, H. R. Development of a multi-epitope peptide vaccine inducing robust T cell responses against brucellosis using immunoinformatics based approaches. Infect. Genet. Evol. 51, 227–234. https://doi.org/10.1016/j.meegid.2017.04.009 (2017).
Livingston, B. et al. A rational strategy to design multiepitope immunogens based on multiple Th lymphocyte epitopes. J. Immunol. 168, 5499–5506. https://doi.org/10.4049/jimmunol.168.11.5499 (2002).
Amet, N., Lee, H. F. & Shen, W. C. Insertion of the designed helical linker led to increased expression of tf-based fusion proteins. Pharm. Res. 26, 523–528. https://doi.org/10.1007/s11095-008-9767-0 (2009).
Chen, X., Zaro, J. L. & Shen, W. C. Fusion protein linkers: Property, design and functionality. Adv. Drug Deliv. Rev. 65, 1357–1369. https://doi.org/10.1016/j.addr.2012.09.039 (2013).
Ahmad, S. et al. Immuno-informatics analysis of Pakistan-based HCV subtype-3a for chimeric polypeptide vaccine design. Vaccines (Basel). https://doi.org/10.3390/vaccines9030293 (2021).
Stratmann, T. Cholera toxin subunit b as adjuvant—An accelerator in protective immunity and a break in autoimmunity. Vaccines (Basel) 3, 579–596. https://doi.org/10.3390/vaccines3030579 (2015).
Yang, J. et al. Improved protein structure prediction using predicted interresidue orientations. Proc. Natl. Acad. Sci. U. S. A. 117, 1496–1503. https://doi.org/10.1073/pnas.1914677117 (2020).
Lovell, S. C. et al. Structure validation by Calpha geometry: Phi, psi and Cbeta deviation. Proteins 50, 437–450. https://doi.org/10.1002/prot.10286 (2003).
Wiederstein, M. & Sippl, M. J. ProSA-web: Interactive web service for the recognition of errors in three-dimensional structures of proteins. Nucleic Acids Res. 35, W407-410. https://doi.org/10.1093/nar/gkm290 (2007).
Colovos, C. & Yeates, T. O. Verification of protein structures: Patterns of nonbonded atomic interactions. Protein Sci. 2, 1511–1519. https://doi.org/10.1002/pro.5560020916 (1993).
Messaoudi, A., Belguith, H. & Ben Hamida, J. Homology modeling and virtual screening approaches to identify potent inhibitors of VEB-1 beta-lactamase. Theor. Biol. Med. Model. 10, 22. https://doi.org/10.1186/1742-4682-10-22 (2013).
Doytchinova, I. A. & Flower, D. R. VaxiJen: A server for prediction of protective antigens, tumour antigens and subunit vaccines. BMC Bioinform. 8, 4. https://doi.org/10.1186/1471-2105-8-4 (2007).
Ikai, A. Thermostability and aliphatic index of globular proteins. J. Biochem. 88, 1895–1898 (1980).
Walker, J. M. The Proteomics Protocols Handbook. (Springer, 2005).
Kyte, J. & Doolittle, R. F. A simple method for displaying the hydropathic character of a protein. J. Mol. Biol. 157, 105–132. https://doi.org/10.1016/0022-2836(82)90515-0 (1982).
Getzoff, E. D., Tainer, J. A., Lerner, R. A. & Geysen, H. M. The chemistry and mechanism of antibody binding to protein antigens. Adv. Immunol. 43, 1–98. https://doi.org/10.1016/s0065-2776(08)60363-6 (1988).
Gupta, S., Madhu, M. K., Sharma, A. K. & Sharma, V. K. ProInflam: A webserver for the prediction of proinflammatory antigenicity of peptides and proteins. J. Transl. Med. 14, 178. https://doi.org/10.1186/s12967-016-0928-3 (2016).
Nain, Z. et al. Proteome-wide screening for designing a multi-epitope vaccine against emerging pathogen Elizabethkingia anophelis using immunoinformatic approaches. J. Biomol. Struct. Dyn. 38, 4850–4867. https://doi.org/10.1080/07391102.2019.1692072 (2020).
Bui, H. H. et al. Predicting population coverage of T-cell epitope-based diagnostics and vaccines. BMC Bioinform. 7, 153. https://doi.org/10.1186/1471-2105-7-153 (2006).
van Zundert, G. C. P. et al. The HADDOCK2.2 Web Server: User-friendly integrative modeling of biomolecular complexes. J. Mol. Biol. 428, 720–725. https://doi.org/10.1016/j.jmb.2015.09.014 (2016).
Carty, M. & Bowie, A. G. Recent insights into the role of Toll-like receptors in viral infection. Clin. Exp. Immunol. 161, 397–406. https://doi.org/10.1111/j.1365-2249.2010.04196.x (2010).
Lester, S. N. & Li, K. Toll-like receptors in antiviral innate immunity. J. Mol. Biol. 426, 1246–1264. https://doi.org/10.1016/j.jmb.2013.11.024 (2014).
Feuillet, V. et al. Involvement of Toll-like receptor 5 in the recognition of flagellated bacteria. Proc. Natl. Acad. Sci. U. S. A. 103, 12487–12492. https://doi.org/10.1073/pnas.0605200103 (2006).
Kumar, N., Sood, D., Sharma, N. & Chandra, R. Multiepitope subunit vaccine to evoke immune response against acute encephalitis. J. Chem. Inf. Model. 60, 421–433 (2019).
Kaliamurthi, S. et al. Exploring the papillomaviral proteome to identify potential candidates for a chimeric vaccine against cervix papilloma using immunomics and computational structural vaccinology. Viruses https://doi.org/10.3390/v11010063 (2019).
Vangone, A. et al. Large-scale prediction of binding affinity in protein-small ligand complexes: the PRODIGY-LIG web server. Bioinformatics 35, 1585–1587. https://doi.org/10.1093/bioinformatics/bty816 (2019).
Kar, T. et al. A candidate multi-epitope vaccine against SARS-CoV-2. Sci. Rep. 10, 10895. https://doi.org/10.1038/s41598-020-67749-1 (2020).
Grote, A. et al. JCat: A novel tool to adapt codon usage of a target gene to its potential expression host. Nucleic Acids Res. 33, W526-531. https://doi.org/10.1093/nar/gki376 (2005).
Rapin, N., Lund, O., Bernaschi, M. & Castiglione, F. Computational immunology meets bioinformatics: The use of prediction tools for molecular binding in the simulation of the immune system. PLoS One 5, e9862. https://doi.org/10.1371/journal.pone.0009862 (2010).
Goudarzi, M., Seyedjavadi, S. S., Goudarzi, H., Mehdizadeh Aghdam, E. & Nazeri, S. Clostridium difficile infection: Epidemiology, pathogenesis, risk factors, and therapeutic options. Scientifica (Cairo) 2014, 916826. https://doi.org/10.1155/2014/916826 (2014).
Tosta, S. F. O. et al. Multi-epitope based vaccine against yellow fever virus applying immunoinformatics approaches. J. Biomol. Struct. Dyn. 39, 219–235. https://doi.org/10.1080/07391102.2019.1707120 (2021).
Goodman, A. G. et al. A human multi-epitope recombinant vaccinia virus as a universal T cell vaccine candidate against influenza virus. PLoS One 6, e25938. https://doi.org/10.1371/journal.pone.0025938 (2011).
Nosrati, M., Behbahani, M. & Mohabatkar, H. Towards the first multi-epitope recombinant vaccine against Crimean-Congo hemorrhagic fever virus: A computer-aided vaccine design approach. J. Biomed. Inform. 93, 103160. https://doi.org/10.1016/j.jbi.2019.103160 (2019).
Sbai, H., Mehta, A. & DeGroot, A. S. Use of T cell epitopes for vaccine development. Curr. Drug Targets Infect. Disord. 1, 303–313. https://doi.org/10.2174/1568005014605955 (2001).
Sette, A. & Fikes, J. Epitope-based vaccines: An update on epitope identification, vaccine design and delivery. Curr. Opin. Immunol. 15, 461–470. https://doi.org/10.1016/s0952-7915(03)00083-9 (2003).
He, R. et al. Efficient control of chronic LCMV infection by a CD4 T cell epitope-based heterologous prime-boost vaccination in a murine model. Cell Mol. Immunol. 15, 815–826. https://doi.org/10.1038/cmi.2017.3 (2018).
Lu, I. N., Farinelle, S., Sausy, A. & Muller, C. P. Identification of a CD4 T-cell epitope in the hemagglutinin stalk domain of pandemic H1N1 influenza virus and its antigen-driven TCR usage signature in BALB/c mice. Cell Mol. Immunol. 14, 511–520. https://doi.org/10.1038/cmi.2016.20 (2017).
Davies, M. N. & Flower, D. R. Harnessing bioinformatics to discover new vaccines. Drug Discov. Today 12, 389–395. https://doi.org/10.1016/j.drudis.2007.03.010 (2007).
Zhu, S. et al. Hepatitis B virus surface antigen as delivery vector can enhance Chlamydia trachomatis MOMP multi-epitope immune response in mice. Appl. Microbiol. Biotechnol. 98, 4107–4117. https://doi.org/10.1007/s00253-014-5517-x (2014).
Jiang, P. et al. Evaluation of tandem Chlamydia trachomatis MOMP multi-epitopes vaccine in BALB/c mice model. Vaccine 35, 3096–3103. https://doi.org/10.1016/j.vaccine.2017.04.031 (2017).
Slingluff, C. L. Jr. et al. A randomized phase II trial of multiepitope vaccination with melanoma peptides for cytotoxic T cells and helper T cells for patients with metastatic melanoma (E1602). Clin. Cancer Res. 19, 4228–4238. https://doi.org/10.1158/1078-0432.CCR-13-0002 (2013).
Toledo, H. et al. A phase I clinical trial of a multi-epitope polypeptide TAB9 combined with Montanide ISA 720 adjuvant in non-HIV-1 infected human volunteers. Vaccine 19, 4328–4336. https://doi.org/10.1016/s0264-410x(01)00111-6 (2001).
Baldauf, K. J., Royal, J. M., Hamorsky, K. T. & Matoba, N. Cholera toxin B: One subunit with many pharmaceutical applications. Toxins (Basel) 7, 974–996. https://doi.org/10.3390/toxins7030974 (2015).
Antonio-Herrera, L. et al. The nontoxic cholera B subunit is a potent adjuvant for intradermal DC-targeted vaccination. Front. Immunol. 9, 2212. https://doi.org/10.3389/fimmu.2018.02212 (2018).
Bowman, C. C. & Clements, J. D. Differential biological and adjuvant activities of cholera toxin and Escherichia coli heat-labile enterotoxin hybrids. Infect. Immun. 69, 1528–1535. https://doi.org/10.1128/IAI.69.3.1528-1535.2001 (2001).
Dimitrov, I., Bangov, I., Flower, D. R. & Doytchinova, I. AllerTOP vol 2-a server for in silico prediction of allergens. J. Mol. Model 20, 2278. https://doi.org/10.1007/s00894-014-2278-5 (2014).
Riddell, J. R., Wang, X. Y., Minderman, H. & Gollnick, S. O. Peroxiredoxin 1 stimulates secretion of proinflammatory cytokines by binding to TLR4. J. Immunol. 184, 1022–1030. https://doi.org/10.4049/jimmunol.0901945 (2010).
Ryan, A. et al. A role for TLR4 in Clostridium difficile infection and the recognition of surface layer proteins. PLoS Pathog. 7, e1002076. https://doi.org/10.1371/journal.ppat.1002076 (2011).
Pechine, S., Bruxelle, J. F., Janoir, C. & Collignon, A. Targeting Clostridium difficile surface components to develop immunotherapeutic strategies against Clostridium difficile infection. Front. Microbiol. 9, 1009. https://doi.org/10.3389/fmicb.2018.01009 (2018).
Jarchum, I., Liu, M., Lipuma, L. & Pamer, E. G. Toll-like receptor 5 stimulation protects mice from acute Clostridium difficile colitis. Infect. Immun. 79, 1498–1503. https://doi.org/10.1128/IAI.01196-10 (2011).
Vaure, C. & Liu, Y. A comparative review of toll-like receptor 4 expression and functionality in different animal species. Front. Immunol. 5, 316. https://doi.org/10.3389/fimmu.2014.00316 (2014).
Phongsisay, V., Iizasa, E., Hara, H. & Yoshida, H. Evidence for TLR4 and FcRgamma-CARD9 activation by cholera toxin B subunit and its direct bindings to TREM2 and LMIR5 receptors. Mol. Immunol. 66, 463–471. https://doi.org/10.1016/j.molimm.2015.05.008 (2015).
Yoshino, Y. et al. Clostridium difficile flagellin stimulates toll-like receptor 5, and toxin B promotes flagellin-induced chemokine production via TLR5. Life Sci. 92, 211–217. https://doi.org/10.1016/j.lfs.2012.11.017 (2013).
Batah, J. et al. Clostridium difficile flagella predominantly activate TLR5-linked NF-kappaB pathway in epithelial cells. Anaerobe 38, 116–124. https://doi.org/10.1016/j.anaerobe.2016.01.002 (2016).
Van Maele, L. et al. TLR5 signaling stimulates the innate production of IL-17 and IL-22 by CD3(neg)CD127+ immune cells in spleen and mucosa. J. Immunol. 185, 1177–1185. https://doi.org/10.4049/jimmunol.1000115 (2010).
Ali, M. et al. Exploring dengue genome to construct a multi-epitope based subunit vaccine by utilizing immunoinformatics approach to battle against dengue infection. Sci. Rep. 7, 9232. https://doi.org/10.1038/s41598-017-09199-w (2017).
Ojha, R., Pareek, A., Pandey, R. K., Prusty, D. & Prajapati, V. K. Strategic development of a next-generation multi-epitope vaccine to prevent Nipah virus zoonotic infection. ACS Omega 4, 13069–13079. https://doi.org/10.1021/acsomega.9b00944 (2019).
Pandey, R. K., Bhatt, T. K. & Prajapati, V. K. Novel Immunoinformatics approaches to design multi-epitope subunit vaccine for malaria by investigating anopheles salivary protein. Sci. Rep. 8, 1125. https://doi.org/10.1038/s41598-018-19456-1 (2018).
Kamthania, M. et al. Immunoinformatics approach to design T-cell epitope-based vaccine against hendra virus. Int. J. Pept. Res. Ther. 25, 1627–1637 (2019).
Mishra, S. & Sinha, S. Immunoinformatics and modeling perspective of T cell epitope-based cancer immunotherapy: A holistic picture. J. Biomol. Struct. Dyn. 27, 293–306. https://doi.org/10.1080/07391102.2009.10507317 (2009).
Ols, S. et al. Route of vaccine administration alters antigen trafficking but not innate or adaptive immunity. Cell Rep. 30, 3964–3971. https://doi.org/10.1016/j.celrep.2020.02.111 (2020).
Schmidt, S. T. et al. Induction of cytotoxic T-lymphocyte responses upon subcutaneous administration of a subunit vaccine adjuvanted with an emulsion containing the toll-like receptor 3 ligand poly (I: C). Front. Immunol. 9, 898 (2018).
Shaw, F. E., Kohl, K. S., Lee, L. M., Thacker, S. B. & Centers for Disease Control and Prevention (CDC). Public health then and now: celebrating 50 years of MMWR at CDC: introduction. MMWR Suppl. 60, 2–6 (2011).
Zhang, L., Wang, W. & Wang, S. Effect of vaccine administration modality on immunogenicity and efficacy. Expert Rev. Vaccines 14, 1509–1523. https://doi.org/10.1586/14760584.2015.1081067 (2015).
Jensen, K. K. et al. Improved methods for predicting peptide binding affinity to MHC class II molecules. Immunology 154, 394–406. https://doi.org/10.1111/imm.12889 (2018).
Calis, J. J. et al. Properties of MHC class I presented peptides that enhance immunogenicity. PLoS Comput. Biol. https://doi.org/10.1371/journal.pcbi.1003266 (2013).
Nielsen, H. Predicting secretory proteins with SignalP. Methods Mol. Biol. 1611, 59–73. https://doi.org/10.1007/978-1-4939-7015-5_6 (2017).
Krogh, A., Larsson, B., von Heijne, G. & Sonnhammer, E. L. Predicting transmembrane protein topology with a hidden Markov model: Application to complete genomes. J. Mol. Biol. 305, 567–580. https://doi.org/10.1006/jmbi.2000.4315 (2001).
Ponomarenko, J. et al. ElliPro: A new structure-based tool for the prediction of antibody epitopes. BMC Bioinform. 9, 514. https://doi.org/10.1186/1471-2105-9-514 (2008).
de Vries, S. J. & Bonvin, A. M. CPORT: A consensus interface predictor and its performance in prediction-driven docking with HADDOCK. PLoS One 6, e17695. https://doi.org/10.1371/journal.pone.0017695 (2011).
Laskowski, R. A., Jablonska, J., Pravda, L., Varekova, R. S. & Thornton, J. M. PDBsum: Structural summaries of PDB entries. Protein Sci. 27, 129–134. https://doi.org/10.1002/pro.3289 (2018).
Abraham, M. J. et al. GROMACS: High performance molecular simulations through multi-level parallelism from laptops to supercomputers. SoftwareX 1, 19–25 (2015).
Turner, P. XMGRACE, Version 5.1. 19. Center for Coastal and Land-Margin Research, Oregon Graduate Institute of Science and Technology, Beaverton, OR (2005).
Castiglione, F., Mantile, F., De Berardinis, P. & Prisco, A. How the interval between prime and boost injection affects the immune response in a computational model of the immune system. Comput. Math. Methods Med. 2012, 842329. https://doi.org/10.1155/2012/842329 (2012).
Acknowledgements
F.C. wish to thank the Italian Ministry of Education, University and Research, for partial support under the frame of JPI AMR (project MAGIcIAN, N. 0000873) and the European Commission for partial support under the IMI 2 Joint Undertaking (project ERA4TB, Grant agreement N. 853989).
Author information
Authors and Affiliations
Contributions
S.B., designed and performed experiments and helped in the writing, D.D., designed and performed epitope selection prediction experiment, binding affinity analyses and helped in the writing, U.N., performed molecular dynamics simulation experiment and helped in the writing, T.K., performed the docking experiments and helped in the writing, F.C., performed immune simulation experiment and helped in writing. I.S. devised experiments and helped in writing. P.D.B devised and supervised experiments, analyzed data, and helped in writing. A.P.S. devised and supervised experiments, analyzed data, and helped in writing the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Basak, S., Deb, D., Narsaria, U. et al. In silico designing of vaccine candidate against Clostridium difficile. Sci Rep 11, 14215 (2021). https://doi.org/10.1038/s41598-021-93305-6
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-021-93305-6
This article is cited by
-
A Multi-epitope Vaccine Candidate Against Bolivian Hemorrhagic fever Caused by Machupo Virus
Applied Biochemistry and Biotechnology (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.