Abstract
Aging is correlated with several complex diseases, including type 2 diabetes, neurodegeneration diseases, and cancer. Identifying the nature of this correlation and treatment of age-related diseases has been a major subject of both modern and traditional medicine. Traditional Persian Medicine (TPM) embodies many prescriptions for the treatment of ARDs. Given that autophagy plays a critical role in antiaging processes, the present study aimed to examine whether the documented effect of plants used in TPM might be relevant to the induction of autophagy? To this end, the TPM-based medicinal herbs used in the treatment of the ARDs were identified from modern and traditional references. The known phytochemicals of these plants were then examined against literature for evidence of having autophagy inducing effects. As a result, several plants were identified to have multiple active ingredients, which indeed regulate the autophagy or its upstream pathways. In addition, gene set enrichment analysis of the identified targets confirmed the collective contribution of the identified targets in autophagy regulating processes. Also, the protein–protein interaction (PPI) network of the targets was reconstructed. Network centrality analysis of the PPI network identified mTOR as the key network hub. Given the well-documented role of mTOR in inhibiting autophagy, our results hence support the hypothesis that the antiaging mechanism of TPM-based medicines might involve autophagy induction. Chemoinformatics study of the phytochemicals using docking and molecular dynamics simulation identified, among other compounds, the cyclo-trijuglone of Juglans regia L. as a potential ATP-competitive inhibitor of mTOR. Our results hence, provide a basis for the study of TPM-based prescriptions using modern tools in the quest for developing synergistic therapies for ARDs.
Similar content being viewed by others
Introduction
Aging is a progressive impairment that ultimately increases one’s vulnerability. Evidence suggests that inducing autophagy—an evolutionarily conserved pathway responsible for recycling the degraded proteins and organelles in the cell aimed at maintaining homeostasis— could prevent the occurrence, delay the progression and decrease the severity of several age-related diseases, such as neurodegenerative diseases, type 2 diabetes, cardiovascular diseases, and many others1,2,3. Autophagy, has a crucial role in reducing the age related pathological changes including accumulation of protein aggregates, inflammaging, cellular senescence and oncogenesis4.
Also, in most cases, longevity regulating pathways modulate autophagy. These include forkhead box O1 (FoxO), sirtuin 1 (SIRT1), AMP-activated protein kinase (AMPK), and mechanistic target of rapamycin (mTOR)5,6,7. In addition, autophagy inhibition, could circumvent the longevity-inducing effects of anti-aging agents such as rapamycin. So it seems that autophagy acts as the core process in longevity assurance mechanisms8.
Considering the complexity of human physiology, adopting a holistic approach capable of integrating different levels of physiological data is warranted. The currently available omics data, together with advances in systems biology tools and algorithms, enable such integrative approaches to treat human multifactorial health issues such as aging9,10.
Recent studies have discovered that combining anti-aging drugs have synergistic anti-aging benefits motivating further systems-oriented studies towards optimal aging treatment design11,12.
Given that autophagy is induced through different pathways, it can be assumed that the anti-aging effects of medicinal plants might be the result of various compounds present in the aforementioned plants, which can simultaneously induce autophagy via different pathways13,14,15.
Li first hypothesized the concept of the relationship between Traditional Chinese Medicine (TCM) syndromes, herbal formula, and molecular networks in 199916. Since then, substantial advances in concepts and methodologies have been made to modernize TCM and turn it from an experience-based medicine into evidence-based medicine17,18,19,20,21. The methodologies of TCM systems pharmacology are mainly based on network construction to uncover the associations between network components and network analysis to identify critical components and functional modules in the network22. Several distinguished studies have been made and validated the proposed concepts and methodologies in this field19,23,24. For instance, Zhang et al. investigated the ancient concept in Traditional Chinese Medicine, ZHENG, have a molecular basis with regard to the neuro-endocrine-immune systems25.
Traditional Persian Medicine (TPM) represents a holistic temperament-dependent approach to human healthcare. According to TPM, illnesses are the result of an imbalance in temperament due to physiological, psychological, or external stimuli. However, apart from the validity of TPM's underlying assumption, it seems that the possible mechanism of herbal medicines is due to the interaction between pharmaceutical compounds with human proteins in a multi-compound/multi-target manner. This prompted us to conduct a systems pharmacological study to investigate the possible effects of herbal medicine in the treatment of ARDs26,27,28.
For this purpose, we searched the medicinal plants that are used to treat three or more ARDs, including cancer, cardiovascular diseases, neurodegeneration diseases, type 2 diabetes, and aging. Then, the phytochemicals of the selected medicinal plants were investigated in order to find their potential autophagy inducing action. Next, genes related to the mechanism of action of the natural products were used for enrichment analysis and network analysis. In addition, we proposed a novel natural product with the ability to inhibit mTOR, the major inhibitor of autophagy, using cheminformatics approaches.
Method
Identification of TPM plants relevant to ARDs therapies
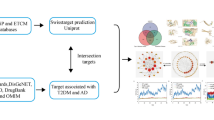
In the first step of our study, the medicinal plants applied in the TPM-based treatment of ARDs, including cancer, cardiovascular diseases, neurodegeneration disorders, type 2 diabetes, and aging, were comprehensively reviewed29,30,31,32,33,34 to identify the relevant herbal medicines (Fig. 1). Noteworthy, the selected diseases are among the leading causes of death in the last years35.
Building on our previous work29,30,31,32and literature review)33,34, herbal medicines in TPM for ARDs were identified. A number of significant Persian medical and pharmaceutical manuscripts, such as Liber Continents by Rhazes36, The Canon of Medicine by Avicenna37, and Tohfat ol Moemenin by Mohammad Tunakabuni38 were used to retrieve geriatric remedies in these studies.
Identification of TPM-related phytochemicals and their targets
After obtaining an extended catalog of TPM-related herbs, each of the herbs was submitted to Dr. Duke’s database (http://phytochem.nal.usda.gov) to identify the relevant phytochemicals. After removing the duplicate compounds, the name of each phytochemical was searched in the literature resources (PubMed and google scholar) together with the “autophagy” keyword to obtain the relevant academic articles. The obtained papers were then reviewed to retrieve the targets of the phytochemicals in question. Also, phytochemicals were classified based on the data retrieved from FooDB (www.foodb.ca). The obtained data on the TPM-based phytochemicals were then visualized.
Gene set enrichment analysis
In order to obtain information regarding the targeted genes, enrichment analysis was done to compare the targets of phytochemicals with annotated gene sets. Consequently, genes related to the medicinal plants’ phytochemicals mechanism of action were enriched, using Enrichr39,40 method at http://amp.pharm.mssm.edu/Enrichr/. Gene ontology (GO)41,42 Biological Process (BP), GO Molecular Function, GO Cellular Component (CC) functional, and Kyoto encyclopedia of genes and genomes (KEGG) pathway43,44 enrichment analysis for the selected genes were performed via the above-mentioned tool.
Protein–protein interactions and network reconstruction
In order to find the essential proteins and pathways in the medicinal plants’ targets, the protein–protein interaction network of the targets was extracted from STRING45 at https://string-db.org/. Then, with the help of Gephi46, the network was visualized. Also, different topological properties (eccentricity, closeness, radiality, betweenness, degree, stress, centroid, eigenvector, bridging) of the network were analyzed by Cytoscape47 plug-in, CentiScaPe48.
The results suggest that mTOR protein has a crucial role in the network. Hence, we used chemoinformatics approaches to find small molecules that can induce autophagy through mTOR inhibition.
Compound-target interaction study by molecular docking
First, three 3D X-ray crystal structures of mTOR in complex with its inhibitors49 (4JSX, 4JT5, 4JT6) were retrieved from Protein Data Bank (http://www.rcsb.org). In order to prepare the 3D structure for docking, the co-crystallized ligands and water molecules were removed, and polar hydrogens were added using a bash script in Linux operating system. The ligand molecules were minimized and saved in ".pdbqt" format after adding partial charges, and ".pdbqt" file of ligands was prepared in AutoDock tools 1.5.650. Self-docking simulations were performed with AutoDock Vina (1.1.2)51 within a docking box defined by the following parameters: size_x = 30, size_y = 30, size_z = 30 Å. The central atom of the co-crystallized ligands was selected as the center of the docking box in all self-docking simulations. Among the PDB structures, 4JSX-A (RMSD = 0.51) was chosen based on optimal self-docking criteria of RMSD < 2 as a validation protocol. The obtained grid box coordinates from the self-docking procedure for this PDB code (x = 53.482, center = 0.81, center = -46.68) was then used for the final docking of the small molecules.
The 3D structure of the phytochemicals (mol2 format), which were retrieved from Dr. Duke’s website, is required for further analysis. To this end, the open-source program, KNIME52, was used as a platform to perform automated requests on chemical websites, using their application programming interface. ChemSpider (http://chemspider.com) was used for converting chemical names into their formulas and other parameters of small molecules. To use it for big projects, it has set an API for which the users can download it as WSDL file and use their token in platforms such as KNIME. In KNIME’s space, first, a file reader was put to read the compounds’ names of a text file. Then a “Generic Web Service Client” with ChemSpider's token was used to search for these names. Another “Generic Web Service Client” was dedicated to obtaining the results, and another one was linked to get the following parameters: CSID, inCHI, inCHIkey, and SMILES. Then, Open Babel53 software was used to convert SMILES into 3D structure files. Finally, AutoDock tools 1.5.650 was used to save ".pdbqts" from mol2 structures for the small molecules. All docking simulations were accordingly performed, and the binding energies of the poses were retrieved from output files.
Also, the SMILES codes of the phytochemicals were retrieved from the cactus database (at https://cactus.nci.nih.gov/) and the properties were calculated by RDkit54. In addition, a binary vector for each plant was compiled in which the presence or lack of the compounds were represented by 0 and 1. The resulting matrix was subjected to the hierarchical clustering analysis (HCA) method using Hamming distance.
Molecular Dynamics (MD) Simulation of compound-target interactions
MD simulation is a suitable tool to study biological systems with respect to the molecular point of view. This technique helps protein and ligand molecules to acquire the best orientation relative to each other and establish the best interactions so that the desired complex moves toward a more stable situation.
Gromacs 5.0.7 was used to perform MD simulation on the selected complexes from the molecular docking process55. Most of the applied steps and conditions of MD simulation were similar to previous works, although there were slight changes. AMBER99SB-ILDN force field was applied to generate all necessary files for the protein molecule56. ACPYPE (or AnteChamber PYthon Parser interfacE) and the general amber force field were used to generate the topological descriptions of the ligands57. Each of the selected complexes was placed in the center of a cubic box and then filled with TIP3P model of water molecules. A minimal distance of 12 Å from any edge of the box and the periodic boundary conditions was applied. A Concentration of 150 mM of NaCl (physiological conditions) was added to neutralize the simulation system. Particle mesh Ewald (PME) and LINCS algorithm were used to treat long-range electrostatic interactions and constrain all bonds. The non-bonded cutoff of 10 Å was also used. In order to minimize the system, steepest descent algorithm was applied. After that equilibration under NVT ensemble at 300 K was conducted during 500 ps. NPT equilibration was subsequently applied at 1 bar pressure and 300 K temperature during 3000 ps. The velocity rescaling algorithm was used to control the temperature with a time constant 0.1 ps as the temperature coupling while Parrinello-Rahman barostat algorithm with a time constant of 1 ps was applied to control the pressure. Finally, MD production run was conducted for 50 ns under similar conditions at 1 bar and 300 K. Also, A time step of 2.0 fs was applied in all MD simulations.
Calculation of protein–ligand binding free energies
Free energies calculations were performed using the MM/PBSA method58,59. GMXPBSA 2.1 was presented by Paissoni et al.60. The fundamental theory of this calculation was explained in detail in our previous publications61,62. In this study, the GROMACS trajectories were used to compute the affinity of the chosen ligands toward mTOR proteins by GMXPBSA 2.1. Water molecules were removed, and the calculations were conceded on a total of 200 frames.
Results and discussion
Phytochemicals of TPM plants relevant to ARDs treatment
Following our literature review, a total of 215 plants were identified to be practiced TPM for treatment of the ARDs29,30,31,32,33,34 (the entire list of all plants is represented in Supplementary Table S1 online). Among these, eight herbs that were found to be used in the treatment of at least three ARDs were selected for further inspection. The selected plants include Allium sativum L., Juglans regia L., Santalum album L., Cinnamomum verum J. Presl, Terminalia chebula Retz., Coriandrum sativum L., Cocos nucifera L., and Cichorium intybus L. The list of the phytochemicals of each plant and their properties, the resulting matrix of binary vector for each plant, and HCA analysis of the similarity of plants is provided in Supplementary Table S2_a-h online, Supplementary Table S3 online, and Fig. S1 online respectively. Also, the family, major chemical compositions63, parts that were used and their applications in TPM64 were reviewed in Supplementary Table S4 online. Review of modern academic literature confirmed that these herbs contain many compounds (63 out of 1078) that may exert anti-aging effects by inducting autophagy process, through targeting several signaling pathways65,66,67,68,69,70,71,72,73,74,75,76,77,78,79,80,81,82,83,84,85,86,87,88,89,90,91,92,93,94,95,96,97,98,99,100,101,102,103,104,105,106,107,108,109,110,111,112,113,114,115,116,117,118,119,120,121,122,123,124,125,126,127,128,129,130,131.
For instance, Allium sativum L. is documented in several reports 132,133 as an autophagy-inducing agent. In particular, Yung-Lin Chu et al. showed that the major component of the herb, i.e., allicin, could induce p53-mediated autophagy in Hep G2 cells107.
Besides, Poulus et al. showed that feeding mice with 6% or 9% Juglans regia L. (walnut) diet, can significantly reduce the accumulation of polyubiquitinated protein and activated autophagy134. Furthermore, the highest percentage of fatty acid in Juglans regia L. corresponds to linoleic acid, oleic acid, and palmitic acid. The effects of these phytochemicals in inducing autophagy were documented by different research groups84,96,131,135.
α-Santalol, which is one of the primary components of Santalum album L. (Indian sandalwood) oil, has a wide range of health benefits, including anti-cancer and anti-inflammatory effects136. It was shown that East Indian Sandalwood oil (EISO), which constitutes 45–50% of alpha-santalol, could induce autophagy104.
Also, the anti-aging effects of Cinnamomum verum J. Presl (cinnamon) is confirmed in various researches. Evidence shows that cinnamon could increase the life-span of model organisms137,138,139,140,141,142. Cinnamic aldehyde is the main compound of the bark oil of this medicinal plant. Chung, J. et al. showed that cinnamic aldehyde could induce autophagy in macrophages. Another study demonstrated that cinnamic aldehyde exerts a neuroprotective effect in a Parkinson’s disease model through autophagy inhibition143.
It is being known as the “king of medicines” in Tibet144, Terminalia chebula Retz. contains several natural products, such as punicalagin and geraniin, which have shown the ability to induce autophagy92,144. In regard to Coriandrum sativum L. (coriander), Cocos nucifera L. (Coconut), and Cichorium intybus L. (Chicory), the direct evidence on their inducing effect on autophagy is scarce. However, these plants contain several natural products with the known ability to induce autophagy145,146,147,148,149.
The activation/inhibition mechanisms through which the identified phytochemicals can influence autophagy is illustrated in Fig. 2. Also, the precise autophagy inducing effect of each phytochemical is represented in Supplementary Table S5 online.
The schematic molecular mechanism of Autophagy-inducing effects of TPM-related phytochemicals (A) Each phytochemical comes with the number and colored circle, representing its related plant and mechanism of action, respectively. For instance, Allicin, a compound found in Allium sativum L., is represented by 1 and could induce autophagy by activating AMPK(red circle) and inhibiting mTOR(green circle). (B) Herbal medicines could modulate the key regulators of autophagy. The targets of phytochemicals were categorized into 3 groups (green, red, purple). The precise autophagy inducing effect of each phytochemical is also represented in Supplementary Table S5 online.
The targets of phytochemicals of the selected TPM plants
Through an extensive literate survey, we were able to link the phytochemicals of the selected herbs to 20 targets. The identified targets include protein kinase C alpha (PRKCA), autophagy-related 12(ATG12), beclin 1(BECN1), tumor protein p53(TP53), forkhead box O1(FOXO1), sirtuin 1(SIRT1), sirtuin 6(SIRT6), ribosomal protein S6 kinase B1(RPS6KB1), nuclear factor-kappa B subunit 1(NFKB1), phosphatase and tensin homolog(PTEN), signal transducer and activator of transcription 3(STAT3), AKT serine/threonine kinase 1(AKT1), mitogen-activated protein kinase 1(MAPK1), microtubule-associated protein 1 light chain 3 alpha(MAP1LC3A), autophagy-related 5(ATG5), unc-51 like autophagy activating kinase 1(ULK1), autophagy-related 7(ATG7), mitogen-activated protein kinase 8(MAPK8), protein kinase AMP-activated catalytic subunit alpha 1(PRKAA1), and mechanistic target of rapamycin (mTOR).
Gene set enrichment analysis
As can be seen in Fig. 2, genes related to the medicinal plants’ chemical compounds mechanism of action were identified. (AKT1, ATG12, ATG5, ATG7, BECN1, FOXO1, MAP1LC3A, MAPK1, MAPK8, MTOR, NFKB1, PRKAA1, PRKCA, PTEN, RPS6KB1, SIRT1, SIRT6, STAT3, TP53, and ULK1).
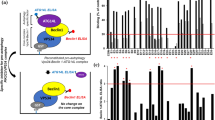
The enrichment analysis is a versatile method to gain insight into the pathways whose activity is influenced by a particular gene group. As seen in Fig. 3, our enrichment analysis indicated that the list of genes targeted by our medicinal plant compounds is most significantly associated with starvation, autophagy, and longevity regulating pathways. This provides a ground for traditional application of these plants in the context of ARDs.
The PPI network of TPM phytochemicals’ targets
We also constructed the protein–protein interaction network for the list of target genes (Fig. 4). Centrality analysis of the PPI network identified the network hub genes. Notably, mTOR was found to be the key hub in the PPI network (Fig. 4).
Network analysis of the medicinal plants’ targets. (a) Network visualization and analysis by Gephi. The size and the color of nodes represent the degree and eigenvector, respectively. (b) Network analysis of the medicinal plants’ targets. mTOR has a crucial role in this network since it has high centralities values for all the computed centralities.
mTOR protein contributes to at least two distinct protein complexes: mTORC1(rapamycin sensitive) and mTORC2 (rapamycin-insensitive)150,151. mTORC1 is considered as a critical modulator of autophagy; most if not all autophagy inducing conditions, such as starvation, result from mTORC1 inhibition14. On the other hand, mTORC2 has a central role in cancer metabolic reprogramming. ATP-competitive mTOR kinase inhibitors (TKIs) like Torin2 _which can inhibit the catalytic domain of mTOR, and consequently, impede the function of both mTORC1 and mTORC2 _ has shown potent antitumor effects in the previous studies152. As such, we further used chemoinformatics tools to explore the potential of ATP-competitive mTOR inhibitors as a target for ARDs.
Compound-target interaction validation via molecular docking
We further examined if the compounds of the TPM-based medicinal plants may indeed strongly inhibit mTOR. In doing so, the binding energy of 572 molecules in complex with mTOR protein were obtained by molecular docking (Supplementary Table S6 online). Amongst these molecules, two were selected as potential ATP-competitive inhibitors.
We considered Torin2, a well-known ATP competitive mTOR inhibitor, as a reference molecule, to evaluate the affinities of other ligands in question. According to the docking results (Supplementary Table S6 online), the binding energy of Torin2, cyclo-trijuglone, and sennoside-A in complex with mTOR were obtained −12.0, −12.6 and −9.6 kcal mol−1, respectively. Therefore, cyclo-trijuglone in complex with mTOR had the most favorable binding energy compared to Torin2, and sennoside A. 2D Structure of these molecules is shown in Supplementary Fig. S2 online. The mode of interaction with mTOR is shown in Supplementary Fig. S3 online. Ile2163, Pro2169, and Leu2185 and more importantly, Trp2239 are identified as crucial residues in mTOR-Torin2 complex49.
According to the docking results, Torin2 established π-π interaction with Trp2239. Furthermore, cation-π interaction and hydrogen bond were seen between Torin2 and Lys2187 and Val2240 residues, respectively. Similar to Torin2, cyclo-trijuglone molecule established π-π and cation-π interactions with Trp2239 and Lys2187, respectively. Moreover, sigma- π interaction was observed between this molecule and Met2345 residue. Hydrogen bonds were established between this molecule and Val2240 and Gly2238 residues. Trp2239 also interacted with sennoside A, through π-π stacking. Hydrogen bonds were also seen between this ligand and Asp2357, Gln2167, Ser2165, Ser2342, Lys2166, Vl2240, and Lys2171 residues. Many other residues also interacted with Torin2, cyclo-trijuglone and sennoside A, through electrostatic and hydrophobic interactions. While all studied ligands interacted with Trp2239, many other residues of this protein interacted with all three ligands.
Molecular dynamics of compound-target interactions for selected phytochemicals
Molecular dynamics simulation can provide a broader landscape of how the compounds in the TPM-based medicinal plants may interact with their potential targets. Contrary to the docking, which assumes a rigid structure for protein, MD simulation allows the proteins residues to freely interact with the ligand and the complex reach to its most stable configuration. Therefore, MD allows for a more rigorous inspection of the extent of interaction between the ligand and the receptor. The selected complexes from molecular docking were used as the initial configuration in MD simulation, and the simulation product was run for 50 s.
Firstly, Root Mean Square Deviation (RMSD) was evaluated for backbone atoms of the A-chain of mTOR protein and ligands. As seen in Figs. 5 and 6, the fluctuations of backbone atoms of mTOR protein in complex with sennoside A, cyclo trijoglone, and Torin2 molecules was increased. Since the RMSD parameter does not specify which region of the protein is more stable, the root mean square fluctuation (RMSF) parameter was evaluated for all residues of the A-chain of mTOR protein as well. This parameter identifies the most stable and unstable regions of a protein chain. As seen in Figs. 5 and 6, 1811–1867 residues and their neighbors tolerated significant fluctuation, especially in complex with Torin2. The mentioned residues are exposed to solvent, and they are located near the missing residues in the crystallographic structure; therefore, they generally endure more fluctuations in comparison with intra-residues. However, the missing residues and all residues with high fluctuations are distant from the active site of mTOR protein and cannot exert significant effects on protein–ligand interactions. On the other hand, sennoside A and cyclo-trijuglone have a larger structure than Torin2, so they interact with more residues of mTOR protein. These interactions affect the intramolecular interaction of a protein chain leading to a decrease in the fluctuations of some residues. As shown, the residues with higher fluctuations are seen in complex with Torin2 that has the smallest structure. As seen in the RMSD plots (Figs. 5, 6), sennoside A is more dynamics within the complex compared with Torin2 and cyclo-trijuglone, whereas Torin2 is the most stable within the mTOR pocket. The superimposition of the ligands at the final point of MD simulation over the obtained structure from the docking process confirmed that the ligands reached to their most stable configuration within the mTOR pocket. The obtained structures at the end of simulation time show that all three ligands could maintain their interaction with Trp2239 residue, although some interactions changed over the MD simulation period (Supplementary Fig. S4 online). The ligands modified their interactions with the essential residue of mTOR protein and implanted themselves within the active site of this protein through hydrogen bonds. As seen in Supplementary Fig. S5 online, the number of hydrogen bonds between all three ligands and mTOR protein increased during 50 ns MD simulation. Since sennoside, A molecule has more oxygen atoms than Torin2 and cyclo-trijuglone molecules, the number of hydrogen bonds between the former molecule and mTOR protein is higher than the other two ligands.
RMSD plots of the backbone atoms of the A-chain of mTOR protein and all three ligands. (a): Backbone RMSD plots of the A-chain of mTOR protein in complex with Torin2 (blue), cyclo trijoglone, and sennoside A (green), (b): RMSD plots of Torin2 (blue), cyclo trijoglone, and sennoside A (green) in complex with mTOR protein.
Finally, free energy calculation was performed to compare the affinity of all three ligands relative to mTOR protein, using GMXPBSA 2. The results (Table 1) revealed that the ability of Torin2 and cyclo-trijuglone molecules to inhibit mTOR protein was almost equal. Noteworthy, the binding energy values obtained from the docking study revealed that the affinity of Ciclo-trijoglone to mTOR protein was slightly more than Torin2.
These values showed that polar solvation contributions (ΔGps) were unfavorable components in the complex formation while van der Waals (ΔGvdw), coulombic (ΔGcoul), and non-polar solvation term (ΔGnps) components performed favorable roles in this regard.
Furthermore, the polar contributions, ΔEcoul and ΔGps, revealed that the favorable component, electrostatic interaction, was not able to overcome the larger polar energy barrier. The non-polar contributions include Van der Waals interaction and the non-polar solvation effect. Embedding of hydrophobic fragments of the ligands within the active site of mTOR protein was done through mentioned forces; in this regard, the Van der Waals contribution performed a dominant role. Since the nature of the active site of mTOR protein is hydrophobic, the values of the ΔEcoul is fewer than the ΔEvdw values. Sennoside A has many oxygen atoms, so ΔEcoul value of this molecule was more favorable in comparison with two other molecules; it also had the best ΔGnps and ΔGvdw values, but ΔGps value depredated all of these favorable components.
Study limitations
Since autophagy is triggered via several pathways, many studies merely have uncovered the pathway that is induced/inhibited by the desired compound. Hence, we had to consider the name of the pathways as proteins in PPI-network. For instance, if a study showed that quercetin could trigger autophagy by inhibiting PI3K/AKT/mTOR signaling pathway, we considered PI3K, AKT, and mTOR as this compound’s targets.
In addition, we just searched the “phytochemicals’ names” that were retrieved from Dr. Duke’s website in combination with “autophagy.” Since some phytochemicals have synonyms, it is logical to assume that we missed autophagy-inducing phytochemicals that might have other names. Furthermore, a few phytochemicals were missed in the process of converting their names to 3D structures for further chemoinformatics analysis.
Conclusions
The progressive advances in the field of systems biology and systems pharmacology underscore the ancient medicinal approaches such as Chinese and Persian traditional medicine as a valuable resource to address complex diseases. The present study was among the first to explore the molecular mechanism behind the TPM approach to the treatment of ARDs. According to our study, the plants used in TPM-based prescriptions contain many compounds with the potential to synergistically regulate the autophagy and its upstream pathways, providing a rationale for their documented antiaging effects. The consistency of TPM with chemical/biological data, as exemplified in the present study, provides motivation for future research into the TPM as an inspiring reference for the development of effective alternative medicine. Although further investigations should be performed to confirm the validity of our approach in vitro and in vivo.
Data availability
All data generated and analyzed in this study are included in the main text or Supplementary Information files.
Change history
08 June 2021
A Correction to this paper has been published: https://doi.org/10.1038/s41598-021-91771-6
References
Levine, B., Packer, M. & Codogno, P. Development of autophagy inducers in clinical medicine. J. Clin. Invest. 125, 14–24 (2015).
Rubinsztein, D. C., Codogno, P. & Levine, B. Autophagy modulation as a potential therapeutic target for diverse diseases. Nat. Rev. Drug Discov. 11, 709–730 (2012).
Ding, Q. et al. Natural autophagy regulators in cancer therapy: a review. Phytochem. Rev. 14, 137–154 (2015).
Madeo, F., Tavernarakis, N. & Kroemer, G. Can autophagy promote longevity?. Nat. Cell. Biol. 12, 842–846 (2010).
Nakamura, S. & Yoshimori, T. Autophagy and longevity. Mol. Cells 41, 65–72 (2018).
Madeo, F., Zimmermann, A., Maiuri, M. C. & Kroemer, G. Essential role for autophagy in life span extension. J. Clin. Invest. 125, 85–93 (2015).
Khorraminejad-Shirazi, M. et al. Aging and stem cell therapy: AMPK as an applicable pharmacological target for rejuvenation of aged stem cells and achieving higher efficacy in stem cell therapy. Hematol. Oncol. Stem Cell Ther. 11, 189–194 (2018).
Rubinsztein, D. C., Mariño, G. & Kroemer, G. Autophagy and aging. Cell 146, 682–695 (2011).
Bordbar, A., Monk, J. M., King, Z. A. & Palsson, B. O. Constraint-based models predict metabolic and associated cellular functions. Nat. Rev. Genet. 15, 107–120 (2014).
Song, R., Sarnoski, E. A. & Acar, M. The systems biology of single-cell aging. iScience 7, 154–169 (2018).
Admasu, T. D. et al. Drug synergy slows aging and improves healthspan through IGF and SREBP lipid signaling. Dev. Cell 47, 67-79.e65 (2018).
Blagosklonny, M. V. From rapalogs to anti-aging formula. Oncotarget 8, 35492–35507 (2017).
Foucquier, J. & Guedj, M. Analysis of drug combinations: current methodological landscape. Pharmacol. Res. Perspect. 3, e00149 (2015).
Kim, Y. C. & Guan, K. L. mTOR: a pharmacologic target for autophagy regulation. J. Clin. Invest. 125, 25–32 (2015).
Sun, W., Sanderson, P. E. & Zheng, W. Drug combination therapy increases successful drug repositioning. Drug Discov. Today 21, 1189–1195 (2016).
Li, S. Possible relationship between traditional Chinese medicine ZHENG and molecular networks. in Proceedings of the 1st Academic Annual Meeting of the China Association for Science and Technology (1999).
Li, S. & Lu, L. Computational Systems Biology Approaches for Deciphering Traditional Chinese Medicine. in Basics of Bioinformatics 337–368 (Springer, 2013).
Lai, X. et al. Network pharmacology and traditional medicine. Frontier Pharmacol. 11, 1194 (2020).
Li, S. & Zhang, B. Traditional Chinese medicine network pharmacology: theory, methodology and application. Chin. J. Nat. Med. 11, 110–120 (2013).
Li, S., Zhang, B. & Zhang, N. Network target for screening synergistic drug combinations with application to traditional Chinese medicine. BMC Syst. Biol. 5, S10 (2011).
Wang, Y., Fan, X., Qu, H., Gao, X. & Cheng, Y. Strategies and techniques for multi-component drug design from medicinal herbs and traditional Chinese medicine. Curr. Top. Med. Chem. 12, 1356–1362 (2012).
Liu, Y.-F., Ai, N., Keys, A., Fan, X.-H. & Chen, M.-J. Network pharmacology for traditional Chinese medicine research: methodologies and applications. Chin. Herb. Med. 7, 18–26 (2015).
Li, H. et al. A network pharmacology approach to determine active compounds and action mechanisms of Ge-Gen-Qin-Lian decoction for treatment of type 2 diabetes. Evid. Based Complem. Altern. Med. 2014, 495840 (2014).
Wang, L. et al. A network study of Chinese Medicine Xuesaitong injection to elucidate a complex mode of action with multicompound, multitarget, and multipathway. Evid. Based Complem. Altern. Med. 2013, 652373 (2013).
Li, S. et al. Understanding ZHENG in traditional Chinese medicine in the context of neuro-endocrine-immune network. IET Syst. Biol. 1, 51–60 (2007).
Hosseinkhani, A., Asadi, N., Pasalar, M. & Zarshenas, M. M. Traditional Persian Medicine and management of metabolic dysfunction in polycystic ovary syndrome. J. Tradit. Complem. Med. 8, 17–23 (2018).
Varteresian, T. & Lavretsky, H. Natural products and supplements for geriatric depression and cognitive disorders: an evaluation of the research. Curr. Psychiatry Rep. 16, 456 (2014).
Catana, C. S., Atanasov, A. G. & Berindan-Neagoe, I. Natural products with anti-aging potential: Affected targets and molecular mechanisms. Biotechnol. Adv. 36, 1649–1656 (2018).
Hosseinkhani, A., Sahragard, A., Namdari, A. & Zarshenas, M. M. Botanical sources for Alzheimer’s: a review on reports from traditional Persian Medicine. Am. J. Alzheimers Dis. Other Demen. 32, 429–437 (2017).
Zarshenas, M. M., Jamshidi, S. & Zargaran, A. Cardiovascular aspects of geriatric medicines in traditional Persian medicine; a review of phytochemistry and pharmacology. Phytomedicine 23, 1182–1189 (2016).
Zarshenas, M. M., Khademian, S. & Moein, M. Diabetes and related remedies in medieval Persian medicine. Indian J. Endocrinol. Metab. 18, 142–149 (2014).
Emami, M., Sadeghpour, O. & Zarshenas, M. M. Geriatric management in medieval Persian medicine. J. Midlife Health 4, 210–215 (2013).
Emami, S. A., Sahebkar, A., Tayarani-Najaran, N. & Tayarani-Najaran, Z. Cancer and its treatment in main ancient books of Islamic Iranian traditional medicine (7th to 14th Century AD). Iran Red Crescent Med. J. 14, 747–757 (2012).
Shirbeigi, L., Iranzadasl, M., Mansouri, P., Hejazi, S. & Aliasl, J. Skin aging remedies in traditional Persian medicine. J. Skin Stem Cell 2, e30283 (2015).
Global, regional, and national age-sex-specific mortality for 282 causes of death in 195 countries and territories, 1980–2017: a systematic analysis for the Global Burden of Disease Study 2017. Lancet (London, England) 392, 1736–1788 (2018).
Razes, A. Al Havi [Liber continent] (Ehyaol Toras al-Arabi, Beiruot, 2001).
Avicenna, I. Kitāb al-Qānūn fī al-Tibb (Canon of medicine). (New Delhi, India: Senior Press Superintendent, Jamia Hamdard Printing Press, 1998).
Tunakabuni, D.J.T.R.C.o. Tuhfat al-mu’minin (The Present for the Faithful). (2007).
Kuleshov, M. V. et al. Enrichr: a comprehensive gene set enrichment analysis web server 2016 update. Nucleic Acids Res. 44, W90–W97 (2016).
Chen, E. Y. et al. Enrichr: interactive and collaborative HTML5 gene list enrichment analysis tool. BMC Bioinform. 14, 128 (2013).
Expansion of the Gene Ontology knowledgebase and resources. Nucleic Acids Res. 45, D331-d338 (2017).
Ashburner, M. et al. Gene ontology: tool for the unification of biology. The Gene Ontology Consortium. Nat. Genet. 25, 25–29 (2000).
Kanehisa, M. & Goto, S. KEGG: kyoto encyclopedia of genes and genomes. Nucleic Acids Res. 28, 27–30 (2000).
Kanehisa, M., Sato, Y., Kawashima, M., Furumichi, M. & Tanabe, M. KEGG as a reference resource for gene and protein annotation. Nucleic Acids Res. 44, D457–D462 (2016).
Szklarczyk, D. et al. The STRING database in 2017: quality-controlled protein-protein association networks, made broadly accessible. Nucleic Acids Res. 45, D362-d368 (2017).
Bastian, M., Heymann, S. & Jacomy, M. Gephi: An Open Source Software for Exploring and Manipulating Networks. (2009).
Shannon, P. et al. Cytoscape: a software environment for integrated models of biomolecular interaction networks. Genome Res. 13, 2498–2504 (2003).
Scardoni, G., Petterlini, M. & Laudanna, C. Analyzing biological network parameters with CentiScaPe. Bioinformatics 25, 2857–2859 (2009).
Yang, H. et al. mTOR kinase structure, mechanism and regulation. Nature 497, 217 (2013).
Morris, G. M. et al. AutoDock4 and AutoDockTools4: Automated docking with selective receptor flexibility. J. Comput. Chem. 30, 2785–2791 (2009).
Trott, O. & Olson, A. J. AutoDock Vina: improving the speed and accuracy of docking with a new scoring function, efficient optimization, and multithreading. J. Comput. Chem. 31, 455–461 (2010).
Berthold, M. R. et al. KNIME-the Konstanz information miner: version 2.0 and beyond. AcM SIGKDD Explor. Newsletter 11, 26–31 (2009).
O’Boyle, N. M. et al. Open Babel: An open chemical toolbox. J. Cheminform. 3, 33 (2011).
Landrum, G. RDKit: Open-source cheminformatics. (2006).
Abraham, M., Van Der Spoel, D., Lindahl, E. & Hess, B.J.G.U.M.V. the GROMACS development team. 5 (2014).
Lindorff-Larsen, K. et al. Improved side-chain torsion potentials for the Amber ff99SB protein force field. Proteins 78, 1950–1958 (2010).
da Silva, A. W. S. & Vranken, W. F. J. ACPYPE-Antechamber python parser interface. BMC Res. Notes 5, 367 (2012).
Kuhn, B. & Kollman, P. A. J. Binding of a diverse set of ligands to avidin and streptavidin: an accurate quantitative prediction of their relative affinities by a combination of molecular mechanics and continuum solvent models. J. Med. Chem. 43, 3786–3791 (2000).
Wang, J., Morin, P., Wang, W. & Kollman, P. A. J. Use of MM-PBSA in reproducing the binding free energies to HIV-1 RT of TIBO derivatives and predicting the binding mode to HIV-1 RT of efavirenz by docking and MM-PBSA. J. Am. Chem. Soc. 123, 5221–5230 (2001).
Paissoni, C., Spiliotopoulos, D., Musco, G. & Spitaleri, A. J. GMXPBSA 21: A GROMACS tool to perform MM/PBSA and computational alanine scanning. Comput. Phys. Commun. 186, 105–107 (2015).
Eslami, M. et al. Deep analysis of N-cadherin/ADH-1 interaction: a computational survey. J. Biomol. Struct. Dyn. 37, 210–228 (2019).
Eslami, M., Hashemianzadeh, S. M., Moghaddam, K. G., Khorsandi-Lagol, A. & Sajadi, S. A. S. Computational evidence to design an appropriate candidate for the treatment of Alzheimer’s disease through replacement of the heptamethylene linker of bis (7) tacrine with S-allylcysteine. RSC Adv. 5, 66840–66851 (2015).
Gruenwald, J., Brendler, T. & Jaenicke, C. PDR for Herbal Medicines (Thomson, Reuters, 2007).
Shirazi, A.J.T., Iran: Tehran University of Medical Sciences. Makhzan Al-adviyah (the Storehouse of medicaments). (2009).
Chen, X. et al. Trehalose, sucrose and raffinose are novel activators of autophagy in human keratinocytes through an mTOR-independent pathway. Sci. Rep. 6, 28423 (2016).
Huang, W. W. et al. Kaempferol induces autophagy through AMPK and AKT signaling molecules and causes G2/M arrest via downregulation of CDK1/cyclin B in SK-HEP-1 human hepatic cancer cells. Int. J. Oncol. 42, 2069–2077 (2013).
Wang, Y., Zhang, W., Lv, Q., Zhang, J. & Zhu, D. The critical role of quercetin in autophagy and apoptosis in HeLa cells. Tumour Biol. 37, 925–929 (2016).
He, Y. et al. Quercetin induces autophagy via FOXO1-dependent pathways and autophagy suppression enhances quercetin-induced apoptosis in PASMCs in hypoxia. Free Radic. Biol. Med. 103, 165–176 (2017).
Granato, M. et al. Quercetin induces apoptosis and autophagy in primary effusion lymphoma cells by inhibiting PI3K/AKT/mTOR and STAT3 signaling pathways. J. Nutr. Biochem. 41, 124–136 (2017).
Kresty, L. A., Weh, K. M., Zeyzus-Johns, B., Perez, L. N. & Howell, A. B. Cranberry proanthocyanidins inhibit esophageal adenocarcinoma in vitro and in vivo through pleiotropic cell death induction and PI3K/AKT/mTOR inactivation. Oncotarget 6, 33438–33455 (2015).
Nie, C. et al. Reduction of apoptosis by proanthocyanidin-induced autophagy in the human gastric cancer cell line MGC-803. Oncol. Rep. 35, 649–658 (2016).
Lijie, S., Zifeng, J. & Rongyu, L. J. Effect of abscisic acid induced autophagy on lung adenocarcinoma cell line spc-A1 in vitro. Acta Universitatis Medicinalis Anhui 4, 009 (2012).
Hu, H. et al. Pentagalloylglucose induces autophagy and caspase-independent programmed deaths in human PC-3 and mouse TRAMP-C2 prostate cancer cells. Mol. Cancer Ther. 8, 2833–2843 (2009).
Shailasree, S., Venkataramana, M., Niranjana, S. R. & Prakash, H. S. Cytotoxic effect of p-Coumaric acid on neuroblastoma, N2a cell via generation of reactive oxygen species leading to dysfunction of mitochondria inducing apoptosis and autophagy. Mol. Neurobiol. 51, 119–130 (2015).
Zhao, C. et al. Daucosterol inhibits cancer cell proliferation by inducing autophagy through reactive oxygen species-dependent manner. Life Sci. 137, 37–43 (2015).
Fu, T. et al. Hyperoside induces both autophagy and apoptosis in non-small cell lung cancer cells in vitro. Acta Pharmacol. Sin. 37, 505–518 (2016).
Wang, S. et al. Overaccumulation of p53-mediated autophagy protects against betulinic acid-induced apoptotic cell death in colorectal cancer cells. Cell Death Dis. 8, e3087 (2017).
Gu, Y., Xiao, L., Ming, Y., Zheng, Z. & Li, W. Corilagin suppresses cholangiocarcinoma progression through Notch signaling pathway in vitro and in vivo. Int. J. Oncol. 48, 1868–1876 (2016).
Kuo, P. L., Hsu, Y. L. & Cho, C. Y. Plumbagin induces G2-M arrest and autophagy by inhibiting the AKT/mammalian target of rapamycin pathway in breast cancer cells. Mol. Cancer Ther. 5, 3209–3221 (2006).
Li, N. et al. D-galactose induces necroptotic cell death in neuroblastoma cell lines. J. Cell Biochem. 112, 3834–3844 (2011).
Banjerdpongchai, R. & Khaw-On, P. Terpinen-4-ol induces autophagic and apoptotic cell death in human leukemic HL-60 cells. Asian Pac. J. Cancer Prev. 14, 7537–7542 (2013).
Kim, H. J., Kim, J., Kang, K. S., Lee, K. T. & Yang, H. O. Neuroprotective effect of chebulagic acid via autophagy induction in SH-SY5Y cells. Biomol. Ther. (Seoul) 22, 275–281 (2014).
De Amicis, F. et al. Bergapten drives autophagy through the up-regulation of PTEN expression in breast cancer cells. Mol. Cancer 14, 130 (2015).
Tan, S. H. et al. Induction of autophagy by palmitic acid via protein kinase C-mediated signaling pathway independent of mTOR (mammalian target of rapamycin). J. Biol. Chem. 287, 14364–14376 (2012).
Bian, Z. et al. Ferulic acid induces mammalian target of rapamycin inactivation in cultured mammalian cells. Biol. Pharm. Bull. 36, 120–124 (2013).
Matboli, M. et al. Caffeic acid attenuates diabetic kidney disease via modulation of autophagy in a high-fat diet/streptozotocin-induced diabetic rat. Sci. Rep. 7, 2263 (2017).
Sung, B., Chung, H. Y. & Kim, N. D. Role of apigenin in cancer prevention via the induction of apoptosis and autophagy. J. Cancer Prev. 21, 216–226 (2016).
Wei, C. C., Chang, C. H. & Liao, V. H. Anti-Parkinsonian effects of beta-amyrin are regulated via LGG-1 involved autophagy pathway in Caenorhabditis elegans. Phytomedicine 36, 118–125 (2017).
Dong, X. et al. Maslinic acid promotes autophagy by disrupting the interaction between Bcl2 and Beclin1 in rat pheochromocytoma PC12 cells. Oncotarget 8, 74527–74538 (2017).
Wang, Y., Chen, B., Longtine, M. S. & Nelson, D. M. Punicalagin promotes autophagy to protect primary human syncytiotrophoblasts from apoptosis. Reproduction 151, 97–104 (2016).
Cao, K. et al. Punicalagin, an active component in pomegranate, ameliorates cardiac mitochondrial impairment in obese rats via AMPK activation. Sci. Rep. 5, 14014 (2015).
Lee, Y. et al. Terminalia Chebula provides protection against dual modes of necroptotic and apoptotic cell death upon death receptor ligation. Sci. Rep. 6, 25094 (2016).
Guo, X. et al. Corosolic acid protects hepatocytes against ethanol-induced damage by modulating mitogen-activated protein kinases and activating autophagy. Eur. J. Pharmacol. 791, 578–588 (2016).
Potze, L., Mullauer, F. B., Colak, S., Kessler, J. H. & Medema, J. P. Betulinic acid-induced mitochondria-dependent cell death is counterbalanced by an autophagic salvage response. Cell Death Dis. 5, e1169 (2014).
Pietrocola, F. et al. Pro-autophagic polyphenols reduce the acetylation of cytoplasmic proteins. Cell Cycle 11, 3851–3860 (2012).
Jiang, L. et al. Oleic acid induces apoptosis and autophagy in the treatment of Tongue Squamous cell carcinomas. Sci. Rep. 7, 11277 (2017).
Yu, B. et al. The synergic effect of tetramethylpyrazine phosphate and borneol for protecting against ischemia injury in cortex and hippocampus regions by modulating apoptosis and autophagy. J. Mol. Neurosci. 63, 70–83 (2017).
Kim, S. H. et al. Geraniol induces cooperative interaction of apoptosis and autophagy to elicit cell death in PC-3 prostate cancer cells. Int. J. Oncol. 40, 1683–1690 (2012).
Ariyoshi-Kishino, K. et al. Tumor-specific cytotoxicity and type of cell death induced by benzaldehyde. Anticancer Res. 30, 5069–5076 (2010).
Nam, H. & Kim, M. M. Scopoletin has a potential activity for anti-aging via autophagy in human lung fibroblasts. Phytomedicine 22, 362–368 (2015).
Pelissier-Rota, M. A., Pelosi, L., Meresse, P. & Jacquier-Sarlin, M. R. Nicotine-induced cellular stresses and autophagy in human cancer colon cells: a supportive effect on cell homeostasis via up-regulation of Cox-2 and PGE2 production. Int. J. Biochem. Cell Biol. 65, 239–256 (2015).
Doan, K. V. et al. Gallic acid regulates body weight and glucose homeostasis through AMPK activation. Endocrinology 156, 157–168 (2015).
Hsieh, L. C. et al. Induction of alpha-phellandrene on autophagy in human liver tumor cells. Am. J. Chin. Med. 43, 121–136 (2015).
Dickinson, S. E. et al. A novel chemopreventive mechanism for a traditional medicine: East Indian sandalwood oil induces autophagy and cell death in proliferating keratinocytes. Arch. Biochem. Biophys. 558, 143–152 (2014).
Liu, J. et al. β-Elemene-induced autophagy protects human gastric cancer cells from undergoing apoptosis. BMC Cancer 11, 183 (2011).
Chung, J. et al. Trans-cinnamic aldehyde inhibits Aggregatibacter actinomycetemcomitans-induced inflammation in THP-1-derived macrophages via autophagy activation. J. Periodontol. 89, 1262–1271 (2018).
Chu, Y. L., Ho, C. T., Chung, J. G., Rajasekaran, R. & Sheen, L. Y. Allicin induces p53-mediated autophagy in Hep G2 human liver cancer cells. J. Agric. Food Chem. 60, 8363–8371 (2012).
Muhammad, S. et al. The morphology, extractions, chemical constituents and uses of Terminalia chebula: a review. J. Med. Plant Res. 6, 4772–4775 (2012).
Hong, J. F. et al. Anticancer activity of taraxerol acetate in human glioblastoma cells and a mouse xenograft model via induction of autophagy and apoptotic cell death, cell cycle arrest and inhibition of cell migration. Mol. Med. Rep. 13, 4541–4548 (2016).
Yue, L. J. et al. S-allyl-cysteine sulfoxide (alliin) alleviates myocardial infarction by modulating cardiomyocyte necroptosis and autophagy. Int. J. Mol. Med. 44, 1943–1951 (2019).
Yue, Z. et al. Diallyl disulfide induces apoptosis and autophagy in human osteosarcoma MG-63 cells through the PI3K/Akt/mTOR pathway. Molecules 24, 2665 (2019).
Niso-Santano, M. et al. Unsaturated fatty acids induce non-canonical autophagy. EMBO J. 34, 1025–1041 (2015).
Sanches, L. J. et al. Cytotoxicity of citral against melanoma cells: the involvement of oxidative stress generation and cell growth protein reduction. Tumour Biol 39, 1010428317695914 (2017).
Wang, Z.P., et al. Corosolic acid ameliorates cardiac hypertrophy via regulating autophagy. Biosci Rep 39 (2019).
Wu, S. et al. Cyanidin-3-o-glucoside inhibits UVA-induced human dermal fibroblast injury by upregulating autophagy. Photodermatol. Photoimmunol. Photomed. 35, 360–368 (2019).
Choi, J.-A. et al. Enhancement of the antimycobacterial activity of macrophages by ajoene. Innate Immun. 24, 79–88 (2018).
Wen, M. et al. Eicosapentaenoic acid-enriched phosphatidylcholine mitigated Aβ1-42-induced neurotoxicity via autophagy-inflammasome pathway. J. Agric. Food Chem 67, 13767–13774 (2019).
Rockenfeller, P. et al. Phosphatidylethanolamine positively regulates autophagy and longevity. Cell Death Differ. 22, 499–508 (2015).
Lv, H. et al. Unraveling the potential role of glutathione in multiple forms of cell death in cancer therapy. Oxid. Med. Cell Longev. 2019, 3150145 (2019).
Wang, P. et al. Juglone induces apoptosis and autophagy via modulation of mitogen-activated protein kinase pathways in human hepatocellular carcinoma cells. Food Chem. Toxicol. 116, 40–50 (2018).
Boga, J. A. et al. Therapeutic potential of melatonin related to its role as an autophagy regulator: a review. J. Pineal Res. 66, e12534 (2019).
Yin, L., Sun, Z., Ren, Q., Su, X. & Zhang, D. Methyl eugenol induces potent anticancer effects in RB355 human retinoblastoma cells by inducing autophagy, cell cycle arrest and inhibition of PI3K/mTOR/Akt signalling pathway. J. BUON 23, 1174–1178 (2018).
Cao, J. et al. Myricetin induces protective autophagy by inhibiting the phosphorylation of mTOR in HepG2 cells. Anat. Rec. (Hoboken) 301, 786–795 (2018).
Kim, S. W. et al. Niacin alleviates TRAIL-mediated colon cancer cell death via autophagy flux activation. Oncotarget 7, 4356–4368 (2016).
Shi, Y. et al. Oleanolic acid induced autophagic cell death in hepatocellular carcinoma cells via PI3K/Akt/mTOR and ROS-dependent pathway. Korean J. Physiol. Pharmacol. 20, 237–243 (2016).
Sivangala Thandi, R. et al. Ornithine-A urea cycle metabolite enhances autophagy and controls Mycobacterium tuberculosis infection. Nat. Commun. 11, 3535 (2020).
Sharma, L., Lone, N. A., Knott, R. M., Hassan, A. & Abdullah, T. Trigonelline prevents high cholesterol and high fat diet induced hepatic lipid accumulation and lipo-toxicity in C57BL/6J mice, via restoration of hepatic autophagy. Food Chem. Toxicol. 121, 283–296 (2018).
Chandra, S., Roy, A., Jana, M. & Pahan, K. Cinnamic acid activates PPARα to stimulate Lysosomal biogenesis and lower Amyloid plaque pathology in an Alzheimer’s disease mouse model. Neurobiol. Dis. 124, 379–395 (2019).
Suzuki, J. I., Miki, S., Ushijima, M. & Kodera, Y. Regulation of immune response by S-1-propenylcysteine through autophagy-mediated protein degradation. Exp. Ther. Med. 19, 1570–1573 (2020).
Xiao, J. et al. Garlic-Derived S-Allylmercaptocysteine Ameliorates nonalcoholic fatty liver disease in a rat model through inhibition of apoptosis and enhancing autophagy. Evid. Based Complem. Altern. Med. 2013, 642920 (2013).
Yang, B. et al. ω-6 Polyunsaturated fatty acids (linoleic acid) activate both autophagy and antioxidation in a synergistic feedback loop via TOR-dependent and TOR-independent signaling pathways. Cell Death Dis. 11, 607 (2020).
Chu, Y. L. et al. Autophagy therapeutic potential of garlic in human cancer therapy. J. Tradit. Complem. Med. 3, 159–162 (2013).
Zhao, R., Xie, E., Yang, X. & Gong, B. Alliin alleviates myocardial ischemia-reperfusion injury by promoting autophagy. Biochem. Biophys. Res. Commun. 512, 236–243 (2019).
Poulose, S. M., Bielinski, D. F. & Shukitt-Hale, B. Walnut diet reduces accumulation of polyubiquitinated proteins and inflammation in the brain of aged rats. J. Nutr. Biochem. 24, 912–919 (2013).
Saxena, R., Joshi, D. D. & Singh, R. Chemical composition and antimicrobial activity of walnut oil, (2009). Int. J. Essent. Oil Ther. 5, 6 (2009).
Bommareddy, A. et al. Medicinal properties of alpha-santalol, a naturally occurring constituent of sandalwood oil: review. Nat. Prod. Res. 33, 527–543 (2019).
Yu, Y. B. et al. Cinnamomum cassia bark in two herbal formulas increases life span in Caenorhabditis elegans via insulin signaling and stress response pathways. PLoS ONE 5, e9339 (2010).
Shen, Y. et al. Cinnamon extract enhances glucose uptake in 3T3-L1 adipocytes and C2C12 myocytes by inducing LKB1-AMP-activated protein kinase signaling. PLoS ONE 9, e87894 (2014).
Mollazadeh, H. & Hosseinzadeh, H. Cinnamon effects on metabolic syndrome: a review based on its mechanisms. Iran. J. Basic Med. Sci. 19, 1258–1270 (2016).
Schriner, S. E. et al. Extension of Drosophila lifespan by cinnamon through a sex-specific dependence on the insulin receptor substrate chico. Exp. Gerontol. 60, 220–230 (2014).
Kazemi, M. & Mokhtariniya, S. Essential oil composition of bark of Cinnamomum zeylanicum. J. Essent Oil Bear Plants 19, 786–789 (2016).
Vazirian, M. et al. Antimicrobial effect of cinnamon (Cinnamomum verum J. Presl) bark essential oil in cream-filled cakes and pastries. Res. J. Pharmacogn. 2, 11–16 (2015).
Bae, W. Y., Choi, J. S. & Jeong, J. W. The neuroprotective effects of cinnamic aldehyde in an MPTP mouse model of Parkinson’s disease. Int. J. Mol. Sci. 19, 786 (2018).
Bag, A., Bhattacharyya, S. K. & Chattopadhyay, R. R. The development of Terminalia chebula Retz. (Combretaceae) in clinical research. Asian Pac. J. Trop. Biomed. 3, 244–252 (2013).
Laribi, B., Kouki, K., M’Hamdi, M. & Bettaieb, T. Coriander (Coriandrum sativum L.) and its bioactive constituents. Fitoterapia 103, 9–26 (2015).
Sahib, N. G. et al. Coriander (Coriandrum sativum L.): a potential source of high-value components for functional foods and nutraceuticals—a review. Phytother. Res. 27, 1439–1456 (2013).
Lima, E. B. et al. Cocos nucifera (L) (Arecaceae): a phytochemical and pharmacological review. Braz. J. Med. Biol. Res. 48, 953–964 (2015).
DebMandal, M. & Mandal, S. Coconut (Cocos nucifera L.: Arecaceae): in health promotion and disease prevention. Asian Pac. J. Trop. Med. 4, 241–247 (2011).
Street, R. A., Sidana, J. & Prinsloo, G. Cichorium intybus: traditional uses, phytochemistry, pharmacology, and toxicology. Evid. Based Complem. Altern. Med. 2013, 579319 (2013).
Laplante, M. & Sabatini, D. M. mTOR signaling in growth control and disease. Cell 149, 274–293 (2012).
Dorvash, M. et al. Dynamic modeling of signal transduction by mTOR complexes in cancer. J. Theor. Biol. 483, 109992 (2019).
Moschetta, M., Reale, A., Marasco, C., Vacca, A. & Carratu, M. R. Therapeutic targeting of the mTOR-signalling pathway in cancer: benefits and limitations. Br. J. Pharmacol. 171, 3801–3813 (2014).
Acknowledgments
This study was conducted by Pouria Mosaddeghi in partial fulfillment of the requirements for the Pharm.D. degree of Shiraz University of Medical Sciences (Thesis #99-01-36-23771). The authors wish to thank Mr. H. Argasi at the Research Consultation Center (RCC) of Shiraz University of Medical Sciences for his invaluable assistance in editing this manuscript.
Author information
Authors and Affiliations
Contributions
P.M., M.K.-S., and M.D. conceived the idea of the research. M.F., M.A., and R.R. comprehensively reviewed articles and retrieved the precise autophagy inducing mechanism of action of the phytochemicals. M.K.-S. prepared Fig. 2 by Adobe Illustrator. F.S. retrieved the SMILES codes of the compounds from the cactus database and calculated the properties by RDkit. Also, he used KNIME to perform automated requests on ChemSpider to convert chemical names into their formulas and other parameters of small molecules. A.S. conducted Compound-target interaction study by molecular docking. M.E. performed Molecular Dynamics (MD) Simulation of compound-target interactions. M.T. performed enrichment and PPI network analysis. M.M.Z. introduced distinguished research in Traditional Persian Medicine that are related to ARDs treatment. In addition, he reviewed the validity of our study from the perspective of the philosophy of science. N.N., M.M. and Y.G. extensively reviewed the manuscript and analyzed the results. All authors wrote the manuscript and reviewed the last version of the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
The original online version of this Article was revised: The original version of this Article contained errors in the Affiliations, where the affiliation 9 was incorrectly given and affiliation 10 was omitted for Meysam Mobasheri.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Mosaddeghi, P., Eslami, M., Farahmandnejad, M. et al. A systems pharmacology approach to identify the autophagy-inducing effects of Traditional Persian medicinal plants. Sci Rep 11, 336 (2021). https://doi.org/10.1038/s41598-020-79472-y
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-020-79472-y
This article is cited by
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.