Abstract
Tooth resorption (TR) in domestic cats is a common and painful disease characterised by the loss of mineralised tissues from the tooth. Due to its progressive nature and unclear aetiology the only treatment currently available is to extract affected teeth. To gain insight into TR pathogenesis, we characterised the transcriptomic changes involved in feline TR by sequencing RNA extracted from 14 teeth (7 with and 7 without signs of resorption) collected from 11 cats. A paired comparison of teeth from the same cat with and without signs of resorption identified 1,732 differentially expressed genes, many of which were characteristic of osteoclast activity and differentiation, in particular matrix metalloproteinase 9 (MMP9). MMP9 expression was confirmed by qPCR and immunocytochemistry of odontoclasts located in TR lesions. A hydroxamate-based MMP9 inhibitor reduced both osteoclast formation and resorption activity while siRNA targeting MMP9 also inhibited osteoclast differentiation although had little effect on resorption activity. Overall, these results suggest that increased MMP9 expression is involved in the progress of TR pathogenesis and that MMP9 may be a potential therapeutic target in feline TR.
Similar content being viewed by others
Introduction
Tooth resorption (TR) of permanent teeth is the pathological loss of dental hard tissues and has been reported in many animals including humans, dogs, horses, domestic cats and wild cats1,2,3,4,5. Affecting multiple teeth, resorption is common in aged domestic cats but is rarer in other animals6,7,8. Destructive and progressive lesions usually initiate on the external surface of the coronal tooth root and extend into the cementum, enamel, dentine and in advanced stages, the pulp7,9,10. TR largely classifies into three Types (Type 1–3) by the Nomenclature Committee of the American Veterinary Dental College (AVDC). Type 1 is characterised by appearance of focal or multifocal lesions of the tooth with normal periodontal ligament and inflammatory changes while Type 2 presents resorption lesions with involvement of periodontal ligament and non-inflammatory replacement resorption11. Type 3 exhibits both features of Type 1 and Type 2. TR is caused by tooth-resorbing odontoclasts, cells functionally similar to bone-resorbing osteoclasts7. Both cell types are derived from circulating blood or bone marrow myeloid lineage progenitors which differentiate into mature multinucleated cells in their respective microenvironments10,12,13,14,15. The main physiological role of odontoclasts is to resorb hard tissues around deciduous teeth allowing tooth eruption to commence during tooth development and to resorb the deciduous teeth for permanent tooth replacement16. In some cats these cells become dysregulated and attack the permanent teeth later in life9,17. Due to recognition of this disease and investigations into the prevalence of TR between the 1990s and 2000s, the aetiology of TR has been widely studied9,18. The central axis of differentiation of osteoclasts and odontoclasts is the pathway comprising colony stimulating factor 1 (CSF-1) and the receptor activator of nuclear factor κ B (RANK), RANK ligand (RANKL), and osteoprotegerin (OPG), a decoy receptor, capturing RANKL12,15. Since the microenvironments of physiological and pathologic TR are different, there has been a focus on identifying co-stimulating factors in pathological conditions. The accumulated evidence suggests that the initiation and progression of TR is complex and multifactorial and is also mediated by local rather than systemic mechanisms. Many possible molecules and risk factors that directly or indirectly impact on the RANK/RANKL/OPG axis pathway have been postulated, including pro-inflammatory cytokines19,20, diet21,22, thyroid hormones23, parathyroid hormone24,25, parathyroid-related peptides26, and vitamin D and its metabolites17,27. However, the precise causes of TR in cats remain unknown.
Transcript identification and the quantification of gene expression have been distinct core activities in molecular biology ever since the discovery of RNA’s role as the key intermediate between the genome and the proteome. To our knowledge, no whole transcriptomic analysis of the feline tooth and dental microenvironment has been reported. We recently reported an optimised protocol for RNA extraction from cat teeth that produced RNA of acceptable quality for generation of cDNA libraries suitable for RNA-seq28. The aims of this study were firstly to perform RNA-seq analysis to compare the feline tooth transcriptome of normal and TR affected teeth. We then identified candidate genes related to feline odontoclastic regulation pathways and targeted them using RNA interference and inhibitors to further evaluate their role in odontoclast differentiation and odontoclast dentine resorption. The transcriptomic changes identified in TR teeth will help us further understand the pathogenesis of TR and help identify genes that can be considered potential therapeutic targets for the treatment of odontoclast dysregulation.
Material and methods
Tooth collection and phenotyping of TR
Teeth were collected from patients with full owner’s consent presented to the Hospital for Small Animals, The Royal (Dick) School of Veterinary Studies, The University of Edinburgh, UK. TR type and staging were confirmed by review of dental radiographs by veterinary dental specialists, S. Thorne and Dr Norman Johnston (MRCVS, RCVS, American & European Specialist in Veterinary Dentistry, DentalVets, Haddington). Further research samples including teeth, maxilla, mandibles and long bones were collected at post mortem from cats euthanized for a wide range of medical or ethical reasons, not related to tooth resorption, and donated to the school for research. Ethical approval for the study was granted by the R(D)SVS veterinary ethical review committee (VERC reference 05.13). All methods were carried out in accordance with the relevant guidelines and regulations.
Tissue processing and RNA extraction
Samples including teeth, mandibles and maxilla were snap frozen in liquid nitrogen and stored at − 80 °C until RNA extraction. Information of samples are described in Table 1. Teeth were extracted from the alveolar sockets using dental equipment or using bone cutters while maintaining cold temperatures by working over dry ice. Tissue processing and RNA extraction protocols were optimised based on a guanidinium thiocyanate-phenol–chloroform extraction method and manufacturer’s instructions for the Qiagen RNeasy Mini Kit. This protocol has been previously documented28.
cDNA library and RNA sequencing
Tested RNA samples were sent to our academic facility (Edinburgh Genomics, The University of Edinburgh, UK) for cDNA library production and RNA sequencing. Briefly, one microgram of RNA from each of the thirteen samples and 0.8 µg of RNA from one low quantity sample (N27C23TR+) were used to generate fourteen cDNA libraries using the Illumina TruSeq stranded mRNA sample preparation kit according to the manufacturer’s instructions (Illumina, San Diego, CA, USA). Paired-end sequencing was performed using the Hiseq 4000 system (Illumina). The low quality reads were filtered by Phred quality score (Q score 30) and 3′ adapter were trimmed with cutadapt (version 1.8.3). All the raw reads have been submitted to the European Nucleotide Archive (ENA) under accession PRJEB24183 (ENA, https://www.ebi.ac.uk/ena). Assessment of quality control of RNA and cDNA library are stated elsewhere28.
Mapping to the feline reference genome and generation of read counts
Alignments to the Felis catus genome (version 6.2) were performed using STAR (version 2.5)29. The read counts from each sample were generated using HTSeq (version 0.6.0.1)30 with mode 'union'. Duplicate reads were found using picard tools (version 1.141). Generation of MDS plots were generated using plotMDS function from edgeR package (version 3.12.1) to visualise the level of similarity of individual cases of a dataset. From the 14 RNA-seq samples, two samples N27C23TR+ and N08C8TR+ were identified as outliers in MDS plots of RNA-seq data therefore they were excluded for final analysis (Fig. S1)28.
Differential expression analysis
‘TR − ve cats’ are defined as teeth collected from a cat without any TR lesion and ‘TR + ve cats’ are teeth collected from a cat with at least one TR lesion. ‘TR tooth’ compares TR status of individual teeth, hence TR − ve teeth were collected from both TR free cats and TR diagnosed cats as long as the tooth was free of TR when assessed on dental radiographs. ‘Paired TR’ compares TR − ve teeth and TR + ve teeth from the same individuals, so a ‘within cat’ comparison.
From the remaining 12 libraries, three comparisons were performed: (1) TR − ve cats (n = 3) and TR + ve cats (n = 5), (2) TR − ve teeth (n = 7) and TR + ve teeth (n = 5), and (3) paired TR, TR − ve (n = 3) and TR + ve (n = 3). Differentially expressed (DE) genes were identified through comparisons between TR − ve and TR + ve groups using the R package edgeR (version 3.12.0)31. The trimmed mean of M-values normalization method was used. To explore sample dispersion between TR − ve and TR + ve groups, distance plot of biological coefficient of variation samples were generated and outliers were excluded for DE gene analysis. Significantly DE genes were identified using the threshold of false discovery rate (FDR)-adjusted p value < 0.05.
Enrichment analyses
We performed two enrichment analyses on the up- and downregulated DE genes: of KEGG (Kyoto Encyclopaedia of Genes and Genomes) metabolic pathways32 and GO (Gene Ontology) terms33. GO term enrichment was assessed using the R/Bioconductor package topGO v2.38.134. KEGG pathway enrichment was performed using the R/Bioconductor package SPIA (Signalling Pathway Impact Analysis) v2.38.035.
Quantitative PCR
Real-time PCR was performed using the Takyon Low ROX SYBR 2X MasterMix kit (Eurogentec) in the Stratagene MX3000P qPCR system (Agilent Technologies). Each reaction contained 300 nM of gene of interest or reference gene primer, 2 × MasterMix, 4 μl of diluted cDNA and nuclease free water, producing a final volume of 20 μl in a 96 well plate format. All samples were performed in duplicate and cycling threshold (Ct) values for each gene were generated. Calculation of relative gene expression analysis was based on delta-delta method36,37. The primers used are described in Table S1.
Generation of feline osteoclasts in vitro
The protocol was modified from a published rodent osteoclast culture protocol38. The long bones were dissected from cats at post mortem and cut off at the proximal and distal ends. Bone marrow was flushed out in α-MEM containing 10% FBS followed by red cell lysis in Lysis Buffer (150 mM NH4Cl, 10 mM NaHCO3, 0.1 mM EDTA) for 5 min, on ice. The bone marrow cells were resuspended in α-MEM containing 10% FBS, 2 mM L glutamine, 100 IU/ml benzyl penicillin, 100 mg/ml streptomycin, and CSF-1 (recombinant mouse CSF-1, 10 ng/ml, R&D Systems) and plated on a T75 flask for 24 h at 37 °C in 5% CO2/95% air. On the following day, non-adherent cells were collected and plated on a dentine disc in each well of a 24 well plate (dentine discs were a generous gift from Professor Timothy R. Arnett, University College London, London, UK) or Corning Osteo Assay Surface multiple well plates (1 × 105 cells/cm2) with CSF-1 (10 ng/ml) at 37 °C in 5% CO2/95% air. On day 4, 90% of the medium was changed and, replaced with a new α MEM containing CSF-1 (10 ng/ml) and RANKL (recombinant mouse RANKL, 3 ng/ml, R&D Systems). The culture media was subsequently changed twice weekly for up to 10–14 days.
Characterisation of cultured osteoclast culture
Cultured osteoclasts on a cell culture plastic or dentine discs were washed twice with phosphate buffered saline (PBS) and fixed in 2.5% glutaraldehyde for 5 min. Tartrate resistant acid phosphatase (TRAP) activity was determined using Acid Phosphatase, Leukocyte (TRAP) Kit (Sigma-Aldrich) according to the manufacturer’s protocol with some modifications including reduced incubation time (30 min) and without haematoxylin counter staining. TRAP positive cells were imaged via light microscopy (Leica DM LB2, Milton Keynes, UK). Cells were considered to be osteoclast-like when TRAP positive and with more than two nuclei when manually counted. Resorption pits in dentine discs were visualised and counted under light microscopy after removal of cells by sonication for 5 min in 0.25 M NH4OH followed by 2 min toluidine blue staining.
Inhibition of MMP9 in osteoclastogenesis
The MMP9 inhibitor containing a hydroxamate (-CONHOH) group was purchased from Calbiochem (Merck Millipore, UK) and was reconstituted in DMSO (1 mg/ml stock solution) and stored at − 20 °C. Primary feline osteoclasts derived from bone marrow were treated with a range of concentrations from 5 nM to 1 µM diluted in α-MEM until termination of osteoclast differentiation. Vehicle controls were treated with DMSO concentrations corresponding to the highest drug concentration used in the experiment. Untreated control cells were also included in each experiment.
Targeting of siRNA of MMP9
The siRNA sequence against the feline MMP9 was designed using an online design tool, i-Score designer (i-score web service program). MMP9 feline siRNA targeting the MMP9 mRNA sequence (CCAGGAGACTTGCGAACTA) was constructed using the Silencer siRNA Construction Kit (Ambion) according to the manufacturer’s protocol. Osteoclast precursors were incubated for two days in α-MEM containing 10% FBS, 2 mM L glutamine, 100 IU/ml benzyl penicillin, 100 mg/ml streptomycin, and CSF-1 (10 ng/ml) at 37 °C/5% CO2. Cells were detached and resuspended for electroporation in 100 μl Mouse Macrophage Nucleofector solution and siRNA. The cell suspension–DNA mixture was transferred to Amaxa electrode cuvettes and electroporated in an Amaxa Nucleofector II (Lonza, UK) using program Y-001. Following electroporation, cells were immediately suspended in medium containing CSF1 (10 ng/ml) and RANKL (3 ng/ml) and seeded into 96-well plates at 1 × 105 cells/well or onto dentine discs. Cells were incubated at 37 °C/5% CO2 until osteoclast formation was observed in control wells. Media were changed every other day.
Immunohistochemistry of feline teeth
Feline tooth samples were collected and fixed by immersion in 10% neutral buffered formalin for 48 h before being processed. All fixed teeth were demineralised in EDTA pH 7.0 for 4–6 weeks at room temperature (RT). Demineralised samples were dehydrated and embedded in paraffin wax following standard procedures. Haematoxylin and Eosin (H & E) staining was performed using an autostainer (Leica Autostainer XL). For MMP9 immunostaining, slides were dewaxed and rehydrated. Antigen retrieval was performed in 0.1 M citrate buffer pH 6.0 for 90 min at 70 °C. Endogenous peroxidase was blocked using 1% hydrogen peroxidase in methanol for 30 min. A further blocking was carried out using a normal goat serum 1:5 in 5% FBS for 30 min. Primary MMP9 antibody (rabbit polyclonal; Abnova, PAB12714) was diluted 1:200 in 5% of FBS in PBS and incubated at 4 °C overnight. Sections were washed and horseradish peroxidase labelled secondary antibodies (Envision Kit, Dako, UK) were added to the sections and incubated for 1 h at RT. Sections were washed 3 times with PBS for 5 min. Labelling was developed using ImmPACT DAB (Vector Labs, UK) at RT for 10 min producing a brown colour in positive sections and was counter-stained with haematoxylin.
Statistical analysis
All experiments were repeated at least twice on two separate occasions gaining similar results (except RNA-seq). In vitro experiments included triple to five technical replicates. Quantitative experiments were analysed using Minitab 15 Statistical Software (Minitab Ltd., UK) and all graphs and diagrams were generated using Microsoft Office 2013 (Microsoft Corporation) or GraphPad prism v5 software. P values of less than 0.05 were deemed statistically significant. The two sample t-test or Mann Whitney U-tests were used to compare differences between two groups.
Results
RNA seq data of TR − ve and TR + ve teeth from the same cat (paired comparison) had the largest number of significantly DE genes
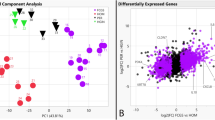
To explore the overall expression profiles between samples, multidimensional scaling (MDS) plotting was used. MDS plot of TR − ve and TR + ve samples in both TR −/+ ve cats and TR −/+ ve teeth comparisons were closely plotted which implied their expression levels were similar (Fig. 1A,B). On the contrary, in the paired comparison, TR − ve samples were widely separated from TR + ve samples indicating possession of more DE genes than the other two comparisons (Fig. 1C). The smear plot allowed for visualisation of the relationship between overall expression level measured in counts per million (CPM) on the x axis and log2 fold-change (FC) on the y axis where DE genes are displayed in red (Fig. 1D–F). The largest number of DE genes was observed in paired TR −/+ ve teeth, with 1,286 up-regulated genes and 466 down-regulated genes identified (Table 2). All expressed genes and significantly differentially expressed genes are listed in Table S2. Additionally, twenty seven differentially expressed genes were common for all three comparisons, including 25 up-regulated genes and 2 down-regulated genes (Table 3).
MDS plot and visualisation of gene expression data by smearplots. MDS plots of (A) TR −/+ cats comparison, (B) TR −/+ teeth comparison, and (C) paired TR −/+ comparison. Smear plots of (D) TR −/+ cats comparison, (E) TR −/+ teeth comparison, and (F) paired TR −/+ comparison. Lined circle: TR + ve samples, dotted circle: TR − ve (control teeth). Horizontal blue lines indicate fold-change of two. Red dot indicates each differentially expressed gene at False Discovery Rate or corrected p value (FDR) of 0.05 or smaller.
Pathway analysis revealed up-regulated genes that are involved in osteoclast differentiation and calcium signalling
For further characterisation of the transcriptomic changes involved in TR, enrichment analyses were performed on the set of 1,732 genes DE between paired TR groups. KEGG downstream pathway analysis identified a total of 28 metabolic pathways within which these genes were involved (Table S3). Of these, the set of upregulated genes were enriched among the genes comprising 25 pathways, with the set of downregulated genes enriched only among 3 pathways. The most biologically relevant pathways were osteoclast differentiation (Fig. 2 and Table S3, KEGG pathway 04380; p = 0.03) and calcium signalling (Fig. 3 and Table S3, KEGG pathway 04020; p = 1.1 × 10–4) which promote the differentiation of osteoclast precursors and induce osteoclastic tooth resorption in teeth. Relative expression of each gene in each pathway was visualised with colour labels in Figs. 2 and 3. Identified pathways and their active status were revealed based on the expression of genes which constitute that pathway (Table S3). For instance, key genes in the osteoclast differentiation pathway were highly expressed in TR + ve teeth (Table S3, total number of genes in osteoclast differentiation pathway = 93, number of up-regulated differentially expressed genes = 15, down-regulated differentially expressed genes = 3). RANK is significantly up-regulated (Table 4, log2 fold change = 1.157, FDR = 0.0013) in TR + ve teeth. RANKL was not significantly up-regulated (Table S2, log2 fold change = 0.810, FDR = 0.253) but OPG was down-regulated (Table 4, log2 fold change = −1.326, FDR = 7.0 × 10–8), which led to an overall increase in the RANKL/OPG ratio. Table 4 highlighted differentially expressed genes in osteoclast differentiation and calcium pathway with putative roles. Gene ontology enrichment analysis presented many gene sets involved in muscle physiology including actin binding, myofibril, actin cytoskeleton, muscle contraction, heart contraction, muscle fibre development, skeletal muscle cell differentiation and muscle organ morphogenesis (Table S4).
Osteoclast differentiation pathway (map04380) obtained from KEGG database32 with official permission and guidance from Kanehisha Laboratories (permission ref 200,290). Expressed genes are coloured with red indicating relative up-regulation and with green indicating down-regulation in TR + ve teeth.
Calcium signalling pathway (map04020) obtained from KEGG database32 with official permission and guidance from Kanehisha Laboratories (permission ref 200,290). Expressed genes are coloured with red indicating relative up-regulation and with green indicating down-regulation in TR + ve teeth.
RT-qPCR validated the differential expression of the selected genes from RNA-seq result
Twelve candidate genes were functionally associated with osteoclast differentiation and activity (Tables 3 and 4) and were selected for validation by qPCR (Table 5). All genes considered significantly DE by RNA-seq (SPI1, MMP9, OPG, RANKL/OPG, ACP5, P2X2R, P2X4R, P2X6R, PCLB4, CA4 and CA6) were confirmed to be up-regulated in TR + ve teeth by qPCR. However, 3 genes considered DE by RNA-seq (CA4, P2X6R, and PCLB4) could not be confirmed by qPCR (Table 5). RANKL expression was unchanged but the gene expression ratio of RANKL/OPG was higher in TR + ve teeth (Fold change = 3.10, p = 0.036) compared to TR –ve teeth. CA6 was identified as the most highly upregulated gene in TR + ve teeth (Fold change = 5.51), followed by MMP9 (Fold change = 4.52), P2X2R (Fold change = 3.04), ACP5 (Fold change = 2.95), SPI1 (Fold change = 2.28) and P2X4R (Fold change = 2.28). CTSK was highly upregulated in TR + ve teeth by qPCR (p = 0.003) but not by RNA-seq (FDR = 0.059). Overall the gene expression profile of TR identified by RNA-seq was largely consistent with that of the qPCR analysis. Differential expression of MMP9 and P2X4R were re-confirmed by qPCR on a further six paired TR − ve and TR + ve teeth samples although this was not the case for SPI1, RANKL, OPG and RANKL/OPG (Fig. S2).
MMP9 is highly expressed in odontoclasts present in feline tooth resorption pits
MMP9 was successfully localised in feline dental tissues by immunohistochemistry (Fig. 4). Multinucleated odontoclast-like cells based on H&E staining (Fig. 4D,E) and MMP9 immunolabelled odontoclasts were identifiable in TR + ve sections (Fig. 4E,F) and there was no odontoclasts observed in TR − ve sections (Fig. 4A,B). Quantitative analysis was not carried out due to the limited number of sections and number of active odontoclasts. Since aggregated gingival epithelium also expressed some MMP9 (Fig. 4H), only multinucleated cells within an obvious resorption pit were considered to be odontoclasts. High expression of MMP9 was observed in actively resorbing odontoclasts in TR lesions (Fig. 4F,G). Fibroblasts of the periodontal ligament also expressed MMP9 in both TR − ve (Fig. 4B,C) and TR + ve teeth (Fig. 4F,G). It was also noted that the majority of TR + ve teeth had disrupted periodontal ligament fibres, narrowed or complete loss of periodontal ligament space and replacement with bone like tissues demonstrating a degree of ankylosis.
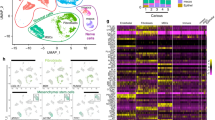
MMP9 is highly expressed in odontoclasts of feline tooth resorption lesions. H&E from TR − ve teeth (A) and TR + ve teeth (D,E). Immunohistochemical labelling for MMP9 protein with haematoxylin counterstaining TR − ve (B,C), TR + ve sections (F,G,H) and isotype control (I). No visible odontoclast is found in TR − ve teeth (B) and circle area of B is magnified in C. Resorbing odontoclasts (D, circle area) are magnified in E. MMP9 expression in TR − ve teeth are observed mainly in periodontal ligament or gingival tissues (B,C). TR + ve teeth contain active odontoclasts with resorption pits (F, dotted circle) with strong MMP9 expression, high magnification of this lesion (G). MMP9 expression was also found in multinucleated like cells but these cells did not reside in a resorption pit (H, arrow). AB alveolar bone, DE dentine, PDL periodontal ligament. Scale bars = 100 µm.
MMP9 expression was confirmed in cultured feline osteoclasts and increased upon osteoclast formation. MMP9 mRNA expression was significantly up-regulated in the early stages of osteoclast differentiation, with a 7.8 ± 0.2 (p < 0.001) fold change on day 3 compared to day 0. MMP9 mRNA expression remained high until day 6 where after it increased further 8.9 ± 0.2 (p = 0.0387) fold change on day 8 (Fig. 5).
MMP9 mRNA expression increased during in vitro feline osteoclast formation over an 8 day culture period. Precursors were treated with M-CSF from day 0, and RANKL was added from day 3. Graphs represent relative expression as fold changes with bars showing standard error of the mean. (n = 3; *p < 0.05, ***p < 0.001 in comparison with the previous time point in culture, ###p < 0.001 in comparison with day 0 by two sample t test).
Treatment with a semi-selective MMP9 inhibitor reduced osteoclast differentiation and resorption activity
In vitro osteoclast cultures were treated with a semi-selective MMP9 inhibitor (hydroxamate-based MMP inhibitor). Inhibitory effects on osteoclast formation and resorption activity were observed in a dose dependent manner (Fig. 6). Osteoclast formation was not significantly reduced at low dose (5 nM, the half maximal inhibitory concentration = 5 nM) compared to vehicle control (p = 0.802), but the inhibitor significantly prevented osteoclast formation at higher doses, whereby 50 ± 6.3% (p < 0.0001) and 48.9 ± 5.8% (p < 0.0001) of the number of osteoclasts formed were observed compare to vehicle at 20 nM and 1000 nM treatments, respectively (Fig. 6A). There was no difference in osteoclast numbers when increasing the concentration from 20 to 1000 nM (p = 0.888). All three concentrations of MMP9 inhibitor resulted in reduction of resorption pits compared to vehicle control (Fig. 6B). This inhibitory effect was dose dependent with 5, 20 and 1000 nM treatments producing 62.2 ± 7.4% (p = 0.002), 55.4 ± 8.8% (p = 0.0003) and 12.2 ± 2.5% (p = 0.001) of the number of resorption pits observed in the vehicle control, respectively.
MMP9 semi-selective inhibitor decreased feline osteoclast formation and resorption activity. Graphs represent percentage of number of osteoclasts (A) or resorption pits (B) with + SEM bars compared to vehicle (DMSO) control. (n = 3; *p < 0.05, **p < 0.01; ***p < 0.001 in comparison with vehicle control, ###p < 0.001 between individual doses comparison by two sample t test).
MMP9 specific siRNAs causes a moderate reduction in mRNA levels and reduced the number of osteoclasts formed
Feline osteoclast precursors were transfected with feline MMP9 siRNA or a scrambled siRNA as control and seeded on dentine discs to investigate the role of MMP9 during osteoclast formation. Following electroporation of the osteoclast precursors with the MMP9 siRNA, a 44.00 ± 0.03% reduction (p = 0.0032) in MMP9 mRNA levels was observed in the transfected cells as assessed by qPCR at 48 h post transfection when compared to negative control transfected precursors (Fig. S3). Electroporation caused some cell death and therefore untransfected cells formed a higher number of osteoclasts and more resorption pits (Fig. 7B,D) but all groups of transfected cells were also able to differentiate into mature osteoclasts. Osteoclast formation was reduced (74.8 ± 7.2%; p < 0.0001) (Fig. 7A,B) in MMP9 siRNA transfected cells in comparison to scrambled siRNA control cells. However, no statistically significant differences in resorption activity was observed between scrambled siRNA and MMP9 siRNA treated wells (p < 0.2298) (Fig. 7).
MMP9 siRNA transfected cells resulted in reduced osteoclast formation in comparison with scrambled control but there was no statistically significant difference in resorption activity. Representative images of TRAP positive osteoclasts on dentin discs from untransfected control, scrambled control and MMP9 siRNA transfected treatment (A). Representative images of toluidine blue stained resorption pits from untransfected control, scrambled control and MMP9 siRNA transfected treatment (C). Graphs represent percentage of number of osteoclasts (B) and resorption pits (D) with + SEM bars. (n = 3; ***p < 0.001 in comparison with scrambled control). Scale bars = 100 µm.
Discussion
Current understanding of the aetiologies of feline TR is incomplete. Dysregulation of odontoclasts are responsible for TR but the mechanisms underlying their dysregulation are largely unknown. The central aims of this study were to investigate the transcriptomic profile of TR affected cats, identify candidate genes related to feline odontoclast dysregulation and assess their potential as therapeutic targets.
First, we performed a RNA-seq to study the whole dental transcriptome of feline teeth and identify differentially expressed genes correlated with progression of TR. No previous RNA-seq analysis of feline TR has been reported, but there are several dental transcriptomic studies using RNA-seq on rodent developing teeth and microarray studies on human teeth that suggested that different cells and tissues expressed unique transcriptomic profiles (e.g. between odontoblasts and pre-secretory ameloblasts or deciduous and permanent periodontal ligament and pulp)55,56,57.
In this study, we hypothesised that the transcriptome of TR affected teeth is locally different from TR free teeth and compared transcriptomic changes between TR − ve and TR + ve samples. There were only a few changes in gene expression that were common between all three comparisons of TR −/+ ve teeth, TR −/+ ve cat and paired TR − ve and TR + ve teeth and this may be due to large individual variations in the samples available. There were only limited genes identified in common between TR − ve and TR + ve teeth. The majority of remaining genes were uncharacterised or were not reported to be involved in tooth or osteoclast biology. Of these, carbonic anhydrase 4 (CA4) has been reported to be expressed in the plasma membrane of osteoclasts58 and CA6 is known to be abundantly present in salivary glands and saliva and involved in maintenance of pH homeostasis in oral cavity59,60. An oral microenvironment high in carbonic anhydrase may favour osteoclast differentiation58. Some homeobox genes (e.g. ARX, HOXA4, LBX1, TLX1) has been reported in tooth development61. One of the downregulated genes was SOSTDC1 which has been reported to be largely related to tooth development as an inhibitor of Wnt signalling to control tooth number and morphogenesis62,63. The role of these genes in TR + ve teeth are unclear but may suggest that permanent TR + ve teeth may undergo some form of remodelling process61,64.
The paired TR comparison gave the most interesting results; a large number of transcripts from the TR + ve teeth differed from TR − ve teeth (n = 1732) which implies that these transcriptomic changes occur locally within the same dentition. The fact that there was transcriptomic similarity between teeth of the same phenotype and that the different phenotypes could clearly be separated reveals that the TR affected teeth undergo similar cellular and molecular changes. The transcriptome corresponding to the phenotype may suggest that transcriptomic changes are at the core of TR pathogenesis, and this knowledge has the potential to increase our understanding of the aetiology of TR.
Previous feline TR studies suggested that the local expression of inflammatory cytokines (e.g. IL1B, IL6, TNF and IFNG) and RANKL may be involved in feline TR, however expression of both pro-inflammatory and inhibitory-inflammatory cytokines increased in TR teeth than control by qPCR which makes it difficult to conclude and the local level of actual cytokines (protein) has however never been measured in TR19,20. Therefore overall influence of inflammation cannot be conclusively determined. In human root resorption, P2X7R was considered a stimulator of osteoclast differentiation65.
In this study, both RNA-seq data and qPCR result could not identify any significant changes in the expression of inflammatory cytokine genes between TR –ve and TR + ve teeth (e.g. IL1B, IL6, TNF, IFNG), P2X7R or VDR, Fig. S4). It is possible that inflammation only appears at a particular stage of TR development so that the overall oral environment possesses a similar extent of inflammation within this study population. It has been reported that the prevalence of TR increased with age in random mixed breed cats although TR occurred regardless of age in pure breed cats66,67. In our study population, the age range was broad (from 4 to 16 years), however the mean age (7.2 years) was similar to those in the previous studies (6.2 and 6.5 years)66,68. To avoid bias due to age, young cats were excluded (< 3 years) in our study and there was no statistically difference of the mean age between TR − ve (7.6 ± 3.2) and TR + ve cats (6.7 ± 1.9) (p = 0.68).
Another possible explanation for the lack of any changes in the expression levels of inflammatory cytokines could be due to a bias of TR type in this study. In this study, type 2 was the dominant type of TR (Table 1). It has been proposed that this type might be involved in non-inflammatory or idiopathic TR although other studies have not confirmed this observation11,69,70. The finding that there is no change in RANKL mRNA levels was consistent with previous feline TR studies19,20. It has been reported that RANKL is highly expressed in the periodontal ligament of actively resorbing primary human teeth71,72, whereas the level of RANKL mRNA is very low or not expressed in human adult teeth73. In cats, RANKL mRNA levels in adult teeth and bones are similar19. Since odontoclast activity is tightly controlled by the RANKL/OPG ratio, the level of OPG and RANKL mRNA expression should be considered together. In healthy feline permanent teeth, the level of OPG mRNA is significantly higher than femoral or alveolar bone, providing evidence that the role of OPG may be a key factor, rather than RANKL, in modulating odontoclast function in feline TR19. This is supported by RNA-seq and qPCR data in the comparison of TR -/ + teeth from the same cat; the level of OPG mRNA level was low in the TR + ve teeth thus an increased RANKL/OPG ratio was observed in TR + ve teeth. However, the observation was not verified when repeated in a larger number of samples, hence the RANKL/OPG involvement in feline TR needs to be further investigated.
Based on the large number of DE genes, there were several interesting metabolic pathways identified. Specifically, DE genes were found which are reported to be involved in osteoclast differentiation (SPI1, C-FMS, RANK, RANKL, TREM2, DAP12, Nox1 and Nox2)39,41,48,49,50,51 or highly expressed in functional and mature osteoclast (OCSTAMP, ACP5, MMP9, CALCR and CTSK)42,43,45,46. SPI1, the gene encoding the haematopoietic transcription factor PU.1, is essential for myeloid and B-lymphoid cell development39. This was reported to be the earliest molecule which binds to sites in both the c-fms promoter or intronic regulatory element (FIRE enhancer) for commitment to osteoclast lineage in mice40,74. The increased expression of SPI1 and C-FMS found in TR + ve teeth may suggest that TR + ve teeth possess more odontoclast precursors in their environment which would be likely to form mature odontoclasts and induce tooth resorption. ACP5 and CTSK are recognised osteoclast markers in bone and teeth43,46. Up-regulation of these genes also suggest increased odontoclast number and activity in TR teeth. MMP9 is a matrix metallopeptidase enzyme, gelatinase B and type IV collagenase. Its primary role is to degrade and remodel the extracellular matrix in many tissues including bone and teeth75. Increased MMP9 activity has been implicated in many diseases including osteoclastomas, Paget's disease, bone fracture repair and dental pulp inflammation76,77. It has also been reported that it is highly expressed in early stages of differentiating osteoclasts, mature osteoclasts as well as in odontoclasts in many species including human, mouse, rabbit and cow78,79,80. Recently accumulated evidence suggests that MMP9 have an important role in the progression of caries, specifically in the pulpal and periapical inflammation tissue destruction process. Significant elevation of MMP9 expression was observed in apical periodontitis. MMP9 is synthesized and secreted by various stimulators such as physical agents or cell cytokines (IL1B and TNF-α)77. Interestingly, Zhang et al. (2020) suggested that MMP9 may have anti-inflammatory properties81. They showed that MMP9 knockout mice experimentally induced with apical periodontitis suffered a more severe inflammation than wild type mice. It was also reported that MMP9 induced the reduction of IL1B, TNF-α, RANK, RANKL, TLR2, and TLR4 and increased OPG expression in LPS-stimulated osteoblasts. These findings suggest that the role of MMP9 in bone destruction is complex and diverse and emphasises the role of MMP9 as an inflammatory regulator by osteoblasts81. In our data, MMP9 expression was higher in TR + ve teeth but there was no difference in expression of inflammatory cytokines (IL1B, TNF-α) or RANK. Feline TR starts normally at the external root and neck area rather than within the pulp, and often results in ankylosis between tooth and alveolar bone rather than alveolar bone destruction. Feline TR is a unique phenomenon and might indicate slightly different pathogenesis from typical inflammation. Our study primarily focused on the role of the odontoclast, but it might be necessary to investigate further roles of other cell types comprising of the tooth (odontoblast, periodontal ligament cells). However, expression of MMP9 and its role in bone and tooth resorption has not previously been described in cats. One study reported high expression of MMP9 in feline cancer82.
Another pathway identified, the calcium signalling pathway, controls multiple cellular functions in many cells via release of calcium ions from internal stores and its entry from the extracellular fluid53. In particular, calcium signals in osteoclasts are responsible for diverse cellular functions including differentiation, bone resorption and gene transcription52. Genes in calcium signalling might induce Ca2+ influx thus contributing to osteoclast differentiation (P2X2R, P2X4R, P2X6R, PLCB4)51,52,53. These genes are known to encode for nonselective cation channels permeable to Ca2+ at the cellular level and thereby contribute to osteoclast differentiation. Originally, purinergic receptors were thought to play a role in ATP production which would in turn induce the formation and activation of osteoclasts51,83. Purinergic receptors e.g. P2X7 or P2Y have been implicated as potent local inhibitors of matrix mineralisation83. Expression of P2X2 receptor has been reported in both osteoblasts and osteoclasts but its precise role in bone is not clear51. Up-regulation of P2X4 receptor has been attributed to inflammation via macrophage invasion84. Although expression of the P2X6 receptor has not been reported in osteoclasts, its expression was reported to modulate the differentiation and migration of human mesenchymal stem cells85. The roles of these genes (and their encoded proteins) in the dental microenvironment is largely unknown, so further studies are required to investigate their possible involvement in tooth resorption.
In addition to osteoclast biology, gene ontology enrichment analysis showed abundant genes involved in muscle physiology. Genes involved in structural myofibril components including ACTA1, ACTN2, ATP2A1, CSRP3, MYH11, MYOZ2, PDLIM3, SYNC, TCAP, TNNI1, TNNT3 and TPM2 were all up-regulated, however, the exact role of these genes in bone and tooth structure/function is unclear. These genes have also been identified to play a role in the development and differentiation of osteocytes, mechanosensing cells that coordinate the remodelling process mediated by osteoblasts and osteoclasts in developing bone and tooth86. This manifestation is unexpected as adult teeth normally possess a limited ability to remodel. However, as TR + ve teeth showed a predominantly disrupted periodontal ligament, new bone formation (ankylosis) and external TR lesions in the root rather than internally, the noted changes in gene expression may reflect repair and some limited remodelling in the roots of the TR + ve teeth, possibly resulting in the level of ankylosis noted in the histological sections. Of the above DE genes, we verified RNA-seq results with qPCR for the genes SPI1, OPG, MMP9, ACP5, CTSK, P2X2R, P2X4R, P2X6R and PCLB4 which all might be involved in osteoclast differentiation and activity, based on previous reports43,46,51,74,75,87,88,89.
Finally, an in vitro feline osteoclast model was established to test the potential role of candidate genes. Since our data showed MMP9 was one of the most up-regulated genes in TR + ve teeth, it was chosen as a target for further investigation. Inhibition of MMP9 by a semi-selective synthetic inhibitor caused both a reduction of osteoclast formation and resorption activity. Synthetic MMP inhibitors have been developed for the treatment of several serious pathologies including periodontitis, although clinical trials gave disappointing results and only a few inhibitors (e.g. Periostat™) has been approved by the FDA for the treatment of periodontal disease90. The synthetic inhibitor possesses a hydroxamic acid analogue which binds to the enzymatic sites of MMPs with high-affinity. However, members of MMPs family share a basic structure which comprises an auto-inhibitory pro-domain rendering them enzymatic latency, the catalytic domain, and the C-terminal hemopexin-like domain, for the recognition of MMP substrates91. Although the inhibition of osteoclastic activity by the MMP9 inhibitor is concentration dependent, it is not clear if the effect of the MMP9 inhibitor is specific to MMP9 or not. Therefore, MMP9 inhibitor experiments were confirmed using the gene specific siRNA approach. When inhibition of MMP9 mRNA was successfully performed by feline MMP9 siRNA, osteoclast formation was inhibited, but not resorption activity. The reduction of resorption activity by synthetic inhibitors but not by the more specific siRNA approach may be a result of off-target inhibition of other members of the MMP superfamily. These results suggest that the role for MMP9 in TR pathogenesis requires further investigation before any firm conclusions can be reached. Strong MMP9 expression both in osteoclasts and odontoclasts has been reported in bovine, rabbit and mice skeletal tissue78,92, and was seen here in the histological tooth sections. Therefore MMP9 may be a potential therapeutic target in feline tooth resorption. The underlying mechanism of inhibition of osteoclast formation by MMP9 needs to be further investigated. Other candidate genes verified in this study could also be potential therapeutic targets in feline TR, and further studies, potentially looking at co-targeting of multiple targets should be performed.
References
Staszyk, C., Bienert, A., Kreutzer, R., Wohlsein, P. & Simhofer, H. Equine odontoclastic tooth resorption and hypercementosis. Vet. J. 178, 372–379 (2008).
Fernandes, M., de Ataide, I. & Wagle, R. Tooth resorption part I—pathogenesis and case series of internal resorption. J. Conserv. Dent. 16, 4–8 (2013).
Peralta, S., Verstraete, F. J. M. & Kass, P. H. Radiographic evaluation of the types of tooth resorption in dogs. Am. J. Vet. Res. 71, 784–793 (2010).
Roux, P., Berger, M., Stich, H. & Schawalder, P. Oral examination and radiographic evaluation of the dentition in wild cats from Namibia. J. Vet. Dent. 26, 16–22 (2009).
Gorrel, C. Tooth resorption in cats: pathophysiology and treatment options. J. Feline Med. Surg. 17, 37–43 (2015).
Sogur, E., Sogur, H. D., Baksi Akdeniz, B. G. & Sen, B. H. Idiopathic root resorption of the entire permanent dentition: systematic review and report of a case. Dent. Traumatol. 24, 490–495 (2008).
Okuda, A. & Harvey, C. E. Etiopathogenesis of feline dental resorptive lesions. Vet. Clin. N. Am. Small Anim. Pract. 22, 1385–1404 (1992).
Gorrel, C. & Larsson, A. Feline odontoclastic resorptive lesions: unveiling the early lesion. J. Small Anim. Pract. 43, 482–488 (2002).
Reiter, A. M. Feline ‘odontolysis’ in the 1920s: the forgotten histopathological study of feline odontoclastic resorptive lesions (FORL). J. Vet. Dent. 15, 35–41 (1998).
DeLaurier, A., Boyde, A., Horton, M. A. & Price, J. S. A scanning electron microscopy study of idiopathic external tooth resorption in the cat. J. Periodontol. 76, 1106–1112 (2005).
DuPont, G. A. & DeBowes, L. J. Comparison of periodontitis and root replacement in cat teeth with resorptive lesions. J. Vet. Dent. 19, 71–75 (2002).
Arnett, T. R. Chapter 8. Osteoclast biology. In Osteoporosis (4th Edition) 149–160 (ed. Cauley, R. M. F. W. D. L. A.) (Academic Press, New York, 2013). https://doi.org/10.1016/B978-0-12-415853-5.00008-X.
Sahara, N., Toyoki, A., Ashizawa, Y., Deguchi, T. & Suzuki, K. Cytodifferentiation of the odontoclast prior to the shedding of human deciduous teeth: an ultrastructural and cytochemical study. Anat. Rec. 244, 33–49 (1996).
Luchin, A. et al. The microphthalmia transcription factor regulates expression of the tartrate-resistant acid phosphatase gene during terminal differentiation of osteoclasts. J. Bone Miner. Res. 15, 451–460 (2000).
Boyle, W. J., Simonet, W. S. & Lacey, D. L. Osteoclast differentiation and activation. Nature 423, 337–342 (2003).
Sasaki, T. Differentiation and functions of osteoclasts and odontoclasts in mineralized tissue resorption. Microsc. Res. Tech. 61, 483–495 (2003).
Reiter, A. M., Lewis, J. R. & Okuda, A. Update on the etiology of tooth resorption in domestic cats. Vet. Clin. N. Am. Small Anim. Pract. 35, 913–942 (2005).
Lund, E. M. et al. Prevalence and risk factors for odontoclastic resorptive lesions in cats. J. Am. Vet. Med. Assoc. 212, 392–395 (1998).
DeLaurier, A., Allen, S., deFlandre, C., Horton, M. A. & Price, J. S. Cytokine expression in feline osteoclastic resorptive lesions. J. Comp. Pathol. 127, 169–177 (2002).
Booij-Vrieling, H. E., Tryfonidou, M. A., Riemers, F. M., Penning, L. C. & Hazewinkel, H. A. Inflammatory cytokines and the nuclear vitamin D receptor are implicated in the pathophysiology of dental resorptive lesions in cats. Vet. Immunol. Immunopathol. 132, 160–166 (2009).
Donoghue, S., Scarlett, J. M., Williams, C. A. & Saidla, J. Diet as a risk factor for feline external odontoclastic resorption. J. Nutr. 124, 2693S-2694S (1994).
Scarlett, J. M., Saidla, J. & Hess, J. Risk factors for odontoclastic resorptive lesions in cats. J. Am. Anim. Hosp. Assoc. 35, 188–192 (1999).
Kanatani, M. et al. Thyroid hormone stimulates osteoclast differentiation by a mechanism independent of RANKL-RANK interaction. J. Cell. Physiol. 201, 17–25 (2004).
Lombardi, G. et al. The roles of parathyroid hormone in bone remodeling: prospects for novel therapeutics. J. Endocrinol. Investig. 34, 18–22 (2011).
Reiter, A. M. & Mendoza, K. A. Feline odontoclastic resorptive lesions an unsolved enigma in veterinary dentistry. Vet. Clin. N. Am. Small Anim. Pract. 32, 791–837 (2002).
Carter, P. H. & Schipani, E. The roles of parathyroid hormone and calcitonin in bone remodeling: prospects for novel therapeutics. Endocr. Metab. Immune Disord. Drug Targets 6, 59–76 (2006).
Booij-Vrieling, H. E. et al. Increased vitamin D-driven signalling and expression of the vitamin D receptor, MSX2, and RANKL in tooth resorption in cats. Eur. J. Oral Sci. 118, 39–46 (2010).
Lee, S., Trivedi, U., Johnson, C., Farquharson, C. & Bergkvist, G. T. Optimised isolation method for RNA extraction suitable for RNA sequencing from feline teeth collected in a clinical setting and at post mortem. Vet. Res. Commun. 43, 17–27 (2019).
Hubbard, T. et al. The Ensembl genome database project. Nucleic Acids Res. 30, 38–41 (2002).
Anders, S., Pyl, P. T. & Huber, W. HTSeq—a Python framework to work with high-throughput sequencing data. Bioinformatics 31, 166–169 (2015).
Robinson, M. D., McCarthy, D. J. & Smyth, G. K. edgeR: a bioconductor package for differential expression analysis of digital gene expression data. Bioinformatics 26, 139–140 (2010).
Kanehisa, M. & Goto, S. KEGG: kyoto encyclopedia of genes and genomes. Nucleic Acids Res. 28, 27–30 (2000).
Ashburner, M. et al. Gene ontology: tool for the unification of biology. The gene ontology consortium. Nat. Genet. 25, 25–29 (2000).
Alexa A, R. J. topGO: Enrichment Analysis for Gene Ontology. (2019).
Tarca, A. L. et al. A novel signaling pathway impact analysis. Bioinformatics 25, 75–82 (2009).
Pfaffl, M. W. A new mathematical model for relative quantification in real-time RT-PCR. Nucleic Acids Res. 29, e45 (2001).
Rao, X., Huang, X., Zhou, Z. & Lin, X. An improvement of the 2(−ΔΔCT) method for quantitative real-time polymerase chain reaction data analysis. Biostat. Bioinform. Biomath. 3, 71–85 (2013).
Orriss, T. R. & Arnett, I. R. Rodent osteoclast cultures. In Bone Research Protocols. In Bone Research Protocols 816 (eds Helfrich, M. H. & Ralston, S. H.) (Humana Press, Totowa, 2012).
Tondravi, M. M. et al. Osteopetrosis in mice lacking haematopoietic transcription factor PU.1. Nature 386, 81 (1997).
Ross, F. P. M-CSF, c-Fms, and signaling in osteoclasts and their precursors. Ann. N. Y. Acad. Sci. 1068, 110–116 (2006).
Wada, T., Nakashima, T., Hiroshi, N. & Penninger, J. M. RANKL-RANK signaling in osteoclastogenesis and bone disease. Trends Mol. Med. 12, 17–25 (2006).
Miyamoto, H. et al. Osteoclast stimulatory transmembrane protein and dendritic cell-specific transmembrane protein cooperatively modulate cell–cell fusion to form osteoclasts and foreign body giant cells. J. Bone Min. Res. 27, 1289–1297 (2012).
Halleen, J., Hentunen, T. A., Hellman, J. & Väänänen, H. K. Tartrate-resistant acid phosphatase from human bone: purification and development of an immunoassay. J. Bone Min. Res. 11, 1444–1452 (1996).
Sundaram, K. et al. RANK ligand signaling modulates the matrix metalloproteinase-9 gene expression during osteoclast differentiation. Exp. Cell. Res. 313, 168–178 (2007).
Granholm, S., Lundberg, P. & Lerner, U. H. Expression of the calcitonin receptor, calcitonin receptor-like receptor, and receptor activity modifying proteins during osteoclast differentiation. J. Cell. Biochem. 104, 920–933 (2008).
Wilson, S. R., Peters, C., Saftig, P. & Bromme, D. Cathepsin K activity-dependent regulation of osteoclast actin ring formation and bone resorption. J. Biol. Chem. 284, 2584–2592 (2009).
Park, J. H., Lee, N. K. & Lee, S. Y. Current understanding of RANK signaling in osteoclast differentiation and maturation. Mol. Cells 40, 706–713 (2017).
Despars, G. et al. DAP12 overexpression induces osteopenia and impaired early hematopoiesis. PLoS ONE 8, e65297 (2013).
Sasaki, H. et al. NADPH oxidase-derived reactive oxygen species are essential for differentiation of a mouse macrophage cell line (RAW264.7) into osteoclasts. J. Med. Investig. 56, 33–41 (2009).
Quinn, J. M. et al. Transforming growth factor beta affects osteoclast differentiation via direct and indirect actions. J. Bone Min. Res. 16, 1787–1794 (2001).
Burnstock, G., Arnett, T. R. & Orriss, I. R. Purinergic signalling in the musculoskeletal system. Purinergic. Signal. 9, 541–572 (2013).
Kajiya, H. Calcium signaling in osteoclast differentiation and bone resorption. Adv. Exp. Med. Biol. 740, 917–932 (2012).
Hwang, S. Y. & Putney, J. W. Jr. Calcium signaling in osteoclasts. Biochim. Biophys. Acta. 1813, 979–983 (2011).
Wang, N., Agrawal, A., Jorgensen, N. R. & Gartland, A. P2X7 receptor regulates osteoclast function and bone loss in a mouse model of osteoporosis. Sci. Rep. 8, 3507 (2018).
Hu, S., Parker, J. & Wright, J. T. Towards unraveling the human tooth transcriptome: the dentome. PLoS ONE 10, e0124801 (2015).
Song, J. S. et al. Comparative gene expression analysis of the human periodontal ligament in deciduous and permanent teeth. PLoS ONE 8, e61231 (2013).
Kim, J.-H. et al. Distinctive genetic activity pattern of the human dental pulp between deciduous and permanent teeth. PLoS ONE 9, e102893 (2014).
Riihonen, R. et al. Membrane-bound carbonic anhydrases in osteoclasts. Bone 40, 1021–1031 (2007).
Reibring, C. G. et al. Expression patterns and subcellular localization of carbonic anhydrases are developmentally regulated during tooth formation. PLoS ONE 9, e96007 (2014).
Leinonen, J., Kivela, J., Parkkila, S., Parkkila, A. K. & Rajaniemi, H. Salivary carbonic anhydrase isoenzyme VI is located in the human enamel pellicle. Caries Res. 33, 185–190 (1999).
Raju, K. et al. Characterization and developmental expression of Tlx-1, the murine homolog of HOX11. Mech. Dev. 44, 51–64 (1993).
Ahn, Y., Sanderson, B. W., Klein, O. D. & Krumlauf, R. Inhibition of Wnt signaling by Wise (Sostdc1) and negative feedback from Shh controls tooth number and patterning. Development 137, 3221–3231 (2010).
Amri, N. et al. Distorted patterns of dentinogenesis and eruption in Msx2 null mutants: involvement of sost/sclerostin. Am. J. Pathol. https://doi.org/10.1016/j.ajpath.2016.06.013 (2016).
Ramanathan, A., Srijaya, T. C., Sukumaran, P., Zain, R. B. & Abu Kasim, N. H. Homeobox genes and tooth development: understanding the biological pathways and applications in regenerative dental science. Arch. Oral Biol. 85, 23–39 (2018).
Hartsfield, J. K. Jr. Pathways in external apical root resorption associated with orthodontia. Orthod. Craniofac. Res. 12, 236–242 (2009).
Girard, N., Servet, E., Biourge, V. & Hennet, P. Feline tooth resorption in a colony of 109 cats. J. Vet. Dent. 25, 166–174 (2008).
Verhaert, L. & Wetter, C. Survey of oral diseases in cats in Flanders. Vlaams Diergeneeskd. Tijdschr. 73, 331–340 (2004).
Mestrinho, L. A., Runhau, J., Braganca, M. & Niza, M. M. Risk assessment of feline tooth resorption: a Portuguese clinical case control study. J. Vet. Dent. 30, 78–83 (2013).
Arzi, B. et al. Presence and quantification of mast cells in the gingiva of cats with tooth resorption, periodontitis and chronic stomatitis. Arch. Oral Biol. 55, 148–154 (2010).
DeLaurier, A., Boyde, A., Jackson, B., Horton, M. A. & Price, J. S. Identifying early osteoclastic resorptive lesions in feline teeth: a model for understanding the origin of multiple idiopathic root resorption. J. Periodontal. Res. 44, 248–257 (2009).
Fukushima, H., Kajiya, H., Takada, K., Okamoto, F. & Okabe, K. Expression and role of RANKL in periodontal ligament cells during physiological root-resorption in human deciduous teeth. Eur. J. Oral Sci. 111, 346–352 (2003).
Hasegawa, T. et al. Human periodontal ligament cells derived from deciduous teeth induce osteoclastogenesis in vitro. Tissue Cell 34, 44–51 (2002).
Kamat, M., Puranik, R., Vanaki, S. & Kamat, S. An insight into the regulatory mechanisms of cells involved in resorption of dental hard tissues. J. Oral Maxillofac. Pathol. 17, 228–233 (2013).
Krysinska, H. et al. A two-step, PU.1-dependent mechanism for developmentally regulated chromatin remodeling and transcription of the c-fms gene. Mol. Cell. Biol. 27, 878–887 (2007).
Christensen, J. & Shastri, V. P. Matrix-metalloproteinase-9 is cleaved and activated by cathepsin K. BMC Res. Notes 8, 322 (2015).
Dong, Z. et al. Matrix metalloproteinase activity and osteoclasts in experimental prostate cancer bone metastasis tissue. Am. J. Pathol. 166, 1173–1186 (2005).
Jain, A. & Bahuguna, R. Role of matrix metalloproteinases in dental caries, pulp and periapical inflammation: an overview. J. Oral Biol. Craniofac. Res. 5, 212–218 (2015).
Linsuwanont, B., Takagi, Y., Ohya, K. & Shimokawa, H. Expression of matrix metalloproteinase-9 mRNA and protein during deciduous tooth resorption in bovine odontoclasts. Bone 31, 472–478 (2002).
Tezuka, K. et al. Identification of matrix metalloproteinase 9 in rabbit osteoclasts. J. Biol. Chem. 269, 15006–15009 (1994).
Reponen, P., Sahlberg, C., Munaut, C., Thesleff, I. & Tryggvason, K. High expression of 92-kD type IV collagenase (gelatinase B) in the osteoclast lineage during mouse development. J. Cell. Biol. 124, 1091–1102 (1994).
Zhang, H. et al. MMP9 protects against LPS-induced inflammation in osteoblasts. Innate Immun. 26, 259–269 (2020).
Akkoc, A., Inan, S. & Sonmez, G. Matrix metalloproteinase (MMP-2 and MMP-9) and steroid receptor expressions in feline mammary tumors. Biotech. Histochem. 87, 312–319 (2012).
Orriss, I. R., Key, M. L., Hajjawi, M. O. R. & Arnett, T. R. Extracellular ATP released by osteoblasts is a key local inhibitor of bone mineralisation. PLoS ONE 8, e69057 (2013).
Yeung, D., Kharidia, R., Brown, S. C. & Gorecki, D. C. Enhanced expression of the P2X4 receptor in Duchenne muscular dystrophy correlates with macrophage invasion. Neurobiol. Dis. 15, 212–220 (2004).
Zippel, N. et al. Purinergic receptors influence the differentiation of human mesenchymal stem cells. Stem Cells Dev. 21, 884–900 (2012).
Paic, F. et al. Identification of differentially expressed genes between osteoblasts and osteocytes. Bone 45, 682–692 (2009).
Dougall, W. C. Molecular pathways: osteoclast-dependent and osteoclast-independent roles of the RANKL/RANK/OPG pathway in tumorigenesis and metastasis. Clin. Cancer Res. 18, 326–335 (2012).
Halleen, J. M., Tiitinen, S. L., Ylipahkala, H., Fagerlund, K. M. & Väänänen, H. K. Tartrate-resistant acid phosphatase 5b (TRACP 5b) as a marker of bone resorption. Clin. Lab. 52, 499–509 (2006).
Grant, M., Wilson, J., Rock, P. & Chapple, I. Induction of cytokines, MMP9, TIMPs, RANKL and OPG during orthodontic tooth movement. Eur. J. Orthod. 35, 644–651 (2013).
Verma, R. P. Hydroxamic acids as matrix metalloproteinase inhibitors. EXS 103, 137–176 (2012).
Franco, C., Patricia, H. R., Timo, S., Claudia, B. & Marcela, H. Matrix metalloproteinases as regulators of periodontal inflammation. Int. J. Mol. Sci. 18, 440 (2017).
Tsuchiya, M. et al. Comparison of expression patterns of cathepsin K and MMP-9 in odontoclasts and osteoclasts in physiological root resorption in the rat molar. Arch. Histol. Cytol. 71, 89–100 (2008).
Acknowledgements
This work was supported by the Biotechnology and Biological Sciences Research Council, Fiona and Ian Russell Seed Corn Grant Fund, The University of Edinburgh and The Sym Charitable Trust and MSD Animal Health.
Author information
Authors and Affiliations
Contributions
S.L. designed the study, performed the experiments, interpreted the results, and drafted the original manuscript. S.J.B analysed and interpreted RNA-seq data. S.T phenotyped and classified TR. N.M performed qPCR and immunohistochemistry. C.F and G.T.B designed the study and supervised the experiments. S.J.B, C.F and G.T.B reviewed manuscript and conducted critical revision. All authors read and commented on the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Lee, S., Bush, S.J., Thorne, S. et al. Transcriptomic profiling of feline teeth highlights the role of matrix metalloproteinase 9 (MMP9) in tooth resorption. Sci Rep 10, 18958 (2020). https://doi.org/10.1038/s41598-020-75998-3
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-020-75998-3
This article is cited by
-
CX3CL1 promotes M1 macrophage polarization and osteoclast differentiation through NF-κB signaling pathway in ankylosing spondylitis in vitro
Journal of Translational Medicine (2023)
-
Gingiva-derived Stromal Cells Isolated from Cats Affected with Tooth Resorption Exhibit Increased Apoptosis, Inflammation, and Oxidative Stress while Experiencing Deteriorated Expansion and Anti-Oxidative Defense
Stem Cell Reviews and Reports (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.