Abstract
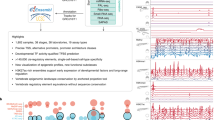
The zebrafish has recently emerged as a model system for investigating the developmental roles of glucocorticoid signaling and the mechanisms underlying glucocorticoid-induced developmental programming. To assess the role of the Glucocorticoid Receptor (GR) in such programming, we used CRISPR-Cas9 to produce a new frameshift mutation, GR369-, which eliminates all potential in-frame initiation codons upstream of the DNA binding domain. Using RNA-seq to ask how this mutation affects the larval transcriptome under both normal conditions and with chronic cortisol treatment, we find that GR mediates most of the effects of the treatment, and paradoxically, that the transcriptome of cortisol-treated larvae is more like that of larvae lacking a GR than that of larvae with a GR, suggesting that the cortisol-treated larvae develop GR resistance. The one transcriptional regulator that was both underexpressed in GR369- larvae and consistently overexpressed in cortisol-treated larvae was klf9. We therefore used CRISPR-Cas9-mediated mutation of klf9 and RNA-seq to assess Klf9-dependent gene expression in both normal and cortisol-treated larvae. Our results indicate that Klf9 contributes significantly to the transcriptomic response to chronic cortisol exposure, mediating the upregulation of proinflammatory genes that we reported previously.
Similar content being viewed by others
Introduction
The vertebrate hypothalamus–pituitary–adrenal (HPA) axis orchestrates physiological, behavioral, and metabolic adjustments required for homeostasis, by dynamically regulating production and secretion of adrenal steroids known as glucocorticoids. In humans the primary glucocorticoid is cortisol, the biological activity of which is mediated by two regulatory proteins in the nuclear receptor family, the ubiquitous glucocorticoid receptor (GR) and the more tissue-restricted mineralocorticoid receptor (MR). The GR binds cortisol less avidly than the MR and is thus more dynamically regulated over the normal physiological range of cortisol fluctuations1,2. The GR and MR function both as transcription factors and as non-nuclear signaling proteins, including in the central nervous system where both proteins are highly expressed1,2,3,4,5. Given that the GR is more widely expressed and more dynamically regulated by cortisol, it is generally thought to be the principal mediator of cortisol-induced genomic responses to circadian rhythms and acute stress5. An important question for understanding GR function is what downstream transcriptional regulatory genes does it regulate, and to what end? Answering this question is not only important for understanding the physiological function and regulation of the GR, but also for deciphering the gene regulatory networks that orchestrate adaptive developmental programming in response to chronic glucocorticoid exposure such as occurs with chronic early life stress6.
The zebrafish has recently emerged as a model system well-suited to investigating the developmental functions of glucocorticoid signaling and mechanisms underlying stress-induced developmental programming7,8,9,10,11. As in humans, the endogenous glucocorticoid in zebrafish is cortisol, which is produced by the interrenal gland, the functional equivalent of the mammalian adrenal. Thus, the zebrafish homolog of the HPA axis is the Hypothalamus-Pituitary-Interrenal (HPI) axis. Eight different zebrafish GR mutants have been described in the literature to date12,13,14,15. The first of these, grs357, is a missense mutation that substitutes a cysteine for an arginine in the DNA binding domain, abolishing GR DNA binding activity12. The other mutations, all produced by targeted mutagenesis using CRISPR-Cas9 or TALEN technology, consist of frameshift indels in exons 2 and 513,14,15, which respectively encode domains N- and C-terminal to the DNA binding domain (see Fig. 1). All of the zebrafish GR mutants manifest behavioral defects but are viable as homozygotes.
A new CRISPR-Cas9-induced mutation of the zebrafish GR. (A) Schematic of the nr3c1 gene and encoded GR showing location of the targeted sequence and resulting premature stop codon with respect to the protein domains: N-terminal domain (NTD), DNA binding domain (DBD), and ligand binding domain (LBD). (B) Nucleotide sequence of nr3c1 exon 3, showing sequence targeted by the gRNA (underlined), the 20 base deletion resulting from injection of that gRNA and Cas9 mRNA (gray), and the resulting premature stop codon (bold italic). (C) Predicted amino acid sequence of the full-length zebrafish GR, plus the extra amino acids introduced by the frameshift shown in (B) (red font). The sequence eliminated by the premature stop is shown in gray, with the DBD underlined. Methionines corresponding to potential alternative initiation sites are shown in green. The positions of all reported frameshift mutations (including that reported here, E369) are boxed. The arginine that is changed to a cysteine in the grs357 mutation is shown in orange. (D) Schematic of the GR showing locations of the two previously reported exon 2 frameshift mutations resulting in truncations at amino acids 178 and 310, and the one in exon 3 reported here truncating at amino acid 369, and residues included in N310 and N369 mRNA constructs produced for microinjection. (E) Relative expression of fkbp5 in 6 hpf homozyogous GR369- embryos, either uninjected or injected with mRNA encoding full-length GR (GR-FL), N-terminally truncated GR isoforms (GR-N310 and GR-N369), or a nonspecific control mRNA encoding Xenopus elongation factor 1α (Xeno EF).
We showed previously that zebrafish larvae treated chronically with 1 µM cortisol upregulate proinflammatory genes and give rise to adults that maintain chronically elevated cortisol and dysregulated expression of those same genes9. Thus in zebrafish, chronic cortisol exposure during early development results in persistent, long-term dysregulation of the HPI axis and gene expression downstream thereof, a ‘developmental programming’ effect similar to what has been observed with early life stress in mammals9,11. To ask whether and to what extent that developmental programming depends on the GR, we used CRISPR-Cas9 technology to mutate the GR-encoding gene nr3c1. Here we report a new loss-of-function mutation, consisting of a deletion in nr3c1 exon 3 that results in a frameshift and premature stop codon immediately upstream of the DNA binding domain which eliminates GR activity as a transcriptional activator. Using this mutant line, we performed an RNA-seq experiment to identify genes regulated by the GR and to parse GR-dependent and GR-independent effects of chronic cortisol treatment on larval gene expression. We found that most of the transcriptomic effects of the chronic cortisol treatment are mediated by the GR and identified klf9, which encodes a ubiquitously expressed16 member of the krüppel-like family of zinc finger transcription factors, as the one regulatory gene consistently upregulated by the GR under both normal conditions and in response to chronic cortisol treatment. We therefore used CRISPR-Cas9 to mutate klf9 and RNA-seq to interrogate its role with respect to the transcriptomic response to chronic cortisol treatment. Our results indicate that klf9 mediates the cortisol-induced upregulation of pro-inflammatory genes that we reported previously9, and suggest that klf9 is a core feedforward regulator of the transcriptional response to glucocorticoid signaling.
Results
A frameshift deletion introduced into exon 3 of zebrafish nr3c1 eliminates GR transcriptional function
To mutate the GR, we employed CRISPR-Cas9 using a guide RNA (gRNA) that targets nr3c1 exon 3 (the second coding exon; Fig. 1A). This resulted in a 20 base deletion and frameshift that introduces sixteen new amino acids after E369, followed by a premature translational stop immediately upstream of the DNA binding domain (Fig. 1B,C), a mutation hereafter referred to as GR369-. F0 males with GR369- in the germline were identified and outbred to wild-type (AB) females, and siblings of that cross were mated to generate F2s, from which homozygous males and females were identified. Homozygous GR369- males crossed with wild-type females produced viable embryos, whereas most homozygous GR369- females crossed with either mutant or wild-type males produced embryos that all died within 24 h of fertilization. However, one produced 10 viable offspring that were raised to adulthood, suggesting that maternal GR is not essential for early development. However, earlier studies using morpholino antisense-mediated knockdown of the GR did suggest that maternal GR is essential for early development17,18, so it is also possible that the mutants but not the morphants are able to compensate for loss of maternal GR as has been documented for other genes19. When inbred these F3 fish produced no viable offspring. However, females homozygous for GR369- following another generation of outcrossing produced variable numbers of viable offspring. These results suggest that loss of GR function may compromise egg quality, albeit in a context-dependent way.
The zebrafish nr3c1 transcript has multiple AUG codons that could potentially initiate translation to allow production of alternative N-terminal isoforms from a single mRNA (Fig. 1C,D; Fig. S1), as occurs with human nr3c120 and some other transcription factors (e.g.21). One potential in-frame initiation codon upstream of the DNA binding domain lies downstream of all previously reported frameshift mutations in exon 213,14,15 (Fig. 1C,D), suggesting that nr3c1 mRNA transcribed in those mutants might still allow translation of an N-terminally truncated GR isoform with an intact DNA binding domain. To assess the transcriptional activity of different N-terminal isoforms we injected homozygous GR369- zygotes with mRNA encoding full-length GR (FL) or GRs lacking the first 310 or 369 amino acids of the full-length GR (N310 and N369, Fig. 1D). The N310 mRNA corresponds to the shortest translation initiation variant with a DNA binding domain potentially available to the previously described exon 2 mutants13,14,15, whereas the N369 corresponds to what is available in GR369- mutants. The embryos were collected at 6 h post fertilization (hpf) and quantitative reverse transcription and polymerase chain reaction (qRT-PCR) was used to measure the expression of the GR-target gene fkbp5. Unlike uninjected embryos or embryos injected with N369 or a Xenopus control mRNA, embryos injected with N310 mRNA upregulated fkbp5, indicating that N310 retains activity as a transcription factor (Fig. 1E). Zygotes injected with mRNA encoding full length GR showed ~ 50-fold stronger upregulation of fkbp5 than those injected with N310 (Fig. 1E). We conclude that the previously described nr3c1 exon 2 frameshift mutations13,14,15 strongly abrogate but do not abolish the potential to translate a GR with transcriptional function (although for one of those mutants it was shown that nr3c1 transcript levels were also significantly reduced, likely through nonsense-mediated decay13) , whereas GR369- eliminates potential for translating a transcriptionally active GR.
RNA-seq shows that the transcriptomic response to chronic cortisol treatment is largely GR-dependent but converges with that caused by loss of GR function
To identify GR-dependent zygotic gene expression and assess the extent to which the GR contributes to the transcriptional effects of chronic cortisol exposure9, we used a visual background adaptation (VBA) assay to identify homozygous GR369- larvae from a heterozygous cross. This assay makes use of the fact that larval melanophores mount a camouflaging response to changes in background, dispersing when larvae are transferred from the dark to a light background12. This response is mediated by the GR, so larvae lacking a functional GR can be identified as those in which melanophores fail to respond, remaining clustered in a dark patch after transfer to a light background. At four days post fertilization (dpf) we used the VBA screen to separate homozygous GR369- (VBA-) larvae from their heterozygous and wild type (VBA +) siblings developed under normal (vehicle-treated) conditions or in medium supplemented with 1 µM cortisol (Fig. 2A). In support of the screen’s validity, the number of VBA- larvae was ~ 1/4 of the total number of larvae, the expected Mendelian ratio for homozygous mutants. Moreover, in a pilot experiments high resolution melt analysis (HRMA) showed that the screen effectively segregated homozygous mutants from mixed heterozygotes and wild-type larvae at 4 dpf (data not shown), and qRT-PCR showed that the VBA- larvae underexpress nr3c1 (Fig. S2A), likely owing to nonsense-mediated decay. The larvae developed for another day under the same conditions (vehicle- or cortisol-treated), at which time four biological replicates of each sample were collected for RNA-seq (Fig. 2A).
RNA-seq identifies GR-dependent and GR-independent genes and effects of chronic cortisol treatment. (A) Diagram of experimental design using Visual Background Adaptation (VBA) selection. (B) Principal component plot of the RNA-seq data, showing the location of each replicate sample with respect to the first two principal components. Cortisol-treated samples are shown in red, vehicle-treated controls in black. VBA+ samples (containing at least one functional GR allele) are filled shapes, VBA- samples are empty shapes. (C) Venn diagram showing numbers of genes upregulated in each of the indicated comparisons (keyed as in (A)): VBA+ vs. VBA- (vehicle-treated controls); cortisol-treated vs. vehicle-treated VBA+; and cortisol-treated vs. vehicle treated VBA-. Upregulated genes in common respectively between the first two and the last two comparisons are listed on the right. (D) Box plots of Z-transformed expression levels of klf9 and npas4a obtained from the RNA-seq data. Sample numbers correspond to those in (B). Asterisks indicate significant differential expression (adjusted p values): ***< .0001; *< .05.
A principal component analysis (PCA) of the RNA-seq data indicated that 60% of the variance in gene expression among samples is captured in the first two PCs, which respectively correlate with the two treatments (chronic cortisol and absence of a GR, respectively accounting for 46% and 14% of the variance; Fig. 2B). The VBA- samples again underexpressed nr3c1, indicating that the VBA screen was effective in identifying homozygous mutants (Fig. S2B). The eight VBA+ (i.e. mixed wild type and heterozygous mutant) samples cluster according to whether they were treated with cortisol, with the two clusters (cortisol-treated and vehicle-treated controls) segregating toward opposite poles of PC1. This is not the case in the VBA- (i.e. homozygous GR369-) fish, the four cortisol-treated replicates of which are widely dispersed along PC1. This indicates that the global effect of the cortisol treatment captured in PC1 is GR-dependent and suggests as well that gene expression is less constrained overall in larvae lacking a GR. PC2 correlates with the presence and absence of a GR (VBA+ vs. VBA− respectively). Interestingly, along PC2 chronic cortisol treatment displaces the VBA+ transcriptome toward the VBA- pole, suggesting that the cortisol-treated fish adapt to the exposure by developing GR resistance.
The regulatory roles of the GR were further assessed by analyzing differential gene expression (DGE) between pairs of treatments, using an adjusted p value (false-discovery) threshold of 0.05 as the criterion for differential expression. Comparison of VBA+ and VBA- larvae identified 405 genes affected by loss of the GR in 5-dpf larvae, 103 of which are underexpressed (Fig. 2C) and 302 of which are overexpressed in VBA- larvae compared to their VBA+ counterparts (Fig. S3A). Gene ontology (GO) analysis shows that the underexpressed genes (i.e. genes normally upregulated by the GR) are involved in sugar metabolism and response to heat, whereas the overexpressed genes (i.e. genes that are normally downregulated via the GR) are involved in basement membrane organization, epidermis development, cell adhesion, locomotion, and growth (Figs. S3 and S4; Table S1).
A DGE analysis comparing cortisol-treated VBA+ fish to their vehicle-treated VBA+ counterparts showed that in cortisol-treated larvae with a functional GR, 4,298 genes were differentially expressed (Fig. S3B), 2,177 of which were upregulated (Fig. 2C) and 2,121 of which were downregulated. GO enrichment analysis of the upregulated genes identified biological processes related to nervous system development and function as well as cell adhesion, locomotion, and growth, whereas the downregulated genes were largely involved in protein synthesis and metabolism (Figs. S5, S6; Table S2). Interestingly, the transcriptome of cortisol-treated VBA+ larvae overlapped more with that of vehicle-treated VBA- larvae than with that of vehicle treated VBA+ larvae (Fig. S3C, Table S3), and accordingly, many of the biological processes affected by the absence of GR function in VBA- larvae were similarly affected by the chronic cortisol-treatment in VBA+ larvae (Fig. S8). This again suggests that the latter larvae develop a GR resistant phenotype. This resistance is probably not associated with any effect on levels of the GR or MR in the cortisol-treated larvae, as neither nr3c1 nor nr3c2 displays significant differential expression in response to the cortisol treatment.
We reasoned that some of the transcriptomic effects of the cortisol treatment might stem from GR-induced upregulation of a GR target gene that functions as a feedforward transcriptional regulator of GR signaling. To the extent that basal expression of such a gene depends on the presence of a GR it might be expected to be underexpressed in VBA- larvae (which lack GR function) but upregulated in VBA+ larvae in response to chronic cortisol (i.e., opposite of the predominant trend noted above). Of the 2177 genes upregulated in cortisol-treated VBA+ larvae only four were basally underexpressed in VBA- larvae (Fig. 2C), two of which encode transcription factors: klf9 and per1a. Of these, only klf9 was also found to be significantly upregulated in our previous RNA-seq analysis of the effects of chronic cortisol treatment in wild-type fish (Fig. S3C), being one of the most highly upregulated transcription factors9. A plot comparing klf9 expression in each of the conditions reveals that the GR contributes to both its normal developmental expression and its upregulation in response to the cortisol treatment (Fig. 2D), which was confirmed by qRT-PCR in another experiment (Fig. S2C). However, the plot also suggests that cortisol affects klf9 in a GR-independent fashion, albeit more variably, as indicated by the range of expression levels in the cortisol-treated VBA- samples shown in Fig. 2D, which correlate with the spread of the cortisol-treated VBA- samples along PC1 shown in Fig. 2B.
In contrast to the situation in VBA+ larvae, only 8 genes were differentially expressed in cortisol-treated VBA- larvae compared to their vehicle-treated VBA- siblings, all of them upregulated (Fig. 2C, Fig. S3D). The genes included the immediate early genes (IEGs) npas4a, egr1, egr4, fosab, and ier2b (Fig. 2C, Fig. S9). The GR-independence of their cortisol-induced upregulation is clearly seen in a plot of the expression levels of the most highly upregulated gene of this set, npas4a (Fig. 2D), a neuronal IEG that along with the other IEGs was also found to be upregulated in our previous RNA-seq analysis of cortisol-treated larvae (Fig. S3D)9. This indicates that the GR mediates nearly all the transcriptomic effects of chronically elevated cortisol, except for a small subset that appears to relate to neuronal activity.
A frameshift deletion introduced into exon 1 of zebrafish klf9 eliminates the DNA binding domain and significantly reduces expression of the mature transcript
The fact that klf9 was the transcriptional regulatory gene most consistently found to be upregulated by chronic cortisol exposure in a GR-dependent way suggested that it may contribute to the transcriptomic effects of the exposure. To test this, we mutated klf9 using CRISPR-Cas9 with a gRNA that targets exon 1 (Fig. 3A,B). This resulted in a frameshift mutation upstream of the DNA binding domain (Fig. 3A,B), producing a transcript encoding a truncated protein predicted to lack function as a transcription factor. Klf9 loss-of-function mutations are viable in mice22 and similarly, the klf9 mutant fish were viable and fertile when bred to homozygosity, although mutant embryos survive at a lower rate than wild type (data not shown).
CRISPR-Cas9-mediated disruption of zebrafish klf9. (A) Schematic of the klf9 locus and encoded protein showing location of the gRNA target in exon 1, which introduces a premature stop codon upstream of the DNA binding domain. (B) Sequence of klf9 exon 1, with 5′ untranslated region in small case and coding sequence in upper case. The location of the 2 base pair deletion generated by CRISPR is shown in gray font; the gRNA target sequence is underlined; and the premature stop codon introduced by frameshift is noted with an asterisk. (C) Effect of chronic cortisol treatment and the frameshift mutation on relative levels of klf9 mRNA and pre-mRNA, measured by qRT-PCR. The results show the averages and SEM of three biological replicates; significance values were obtained by ANOVA.
To ask how the mutation affects klf9 expression we used qRT-PCR to compare klf9 transcript levels in wild type and klf9 homozygous mutant (hereafter referred to as klf9-/-) larvae under both normal conditions and in response to chronic cortisol treatment. This provided further confirmation that the cortisol treatment leads to upregulation of klf9 and revealed that klf9 mRNA levels are significantly reduced in the klf9-/- larvae, probably due to nonsense-mediated decay triggered by the premature stop codon (Fig. 3C). In support of this possibility, there was no significant effect on klf9 pre-mRNA levels, measured by qRT-PCR of the intron (Fig. 3C). We conclude from these experiments that the frameshift mutation introduced into klf9 exon 1 abrogates Klf9 function.
RNA-seq shows that klf9 is required for the pro-inflammatory transcriptomic effects of chronic cortisol treatment
To identify Klf9 target genes and ask whether Klf9 contributes to the transcriptomic response to cortisol treatment we used RNA-seq to query gene expression in 5-dpf wild type and klf9-/- mutant larvae from sibling parents, developed both normally and in the presence of 1 µM cortisol. Samples were collected at the same time on day 5 post-fertilization as in the previous GR knockout experiment and processed similarly. However, PCA revealed the largest source of variance in this experiment was not due to genotype or treatment, but rather the order in which the samples were collected and processed (Fig. S10A). GO analysis of genes correlated with the first principal component showed that the later samples (replicates 3 and 4) had increased synaptic signaling and decreased translation (Table S4), suggestive of a physiological stress response (e.g. to the stress of capture). However, a further confound is that PC1 also correlates with preparation of the RNA in two batches on separate days, suggesting that it may also reflect technical variance in sample preparation (see Materials and Methods). Reassuringly, after normalizing for this variation the samples segregate along two principal components representing genotype and treatment (Fig. S10B). For subsequent DGE analysis we therefore treated the batch effect as a categorical covariate.
DGE analysis using a false-discovery rate of 0.05 identified 239 genes affected by loss of klf9 function in vehicle-treated larvae, 100 of which were upregulated and 139 of which were downregulated. Gene ontology term enrichment analysis shows that the upregulated genes are largely involved in complement activation (e.g. c3b, c3c, c4, cfb ), glucose and carbohydrate metabolism (e.g. pgm1, tpi1b, and pfkmb), and nucleosome positioning (e.g., h1fx, h1f0, and smarca5; Fig. S11, Table S5). Genes downregulated in response to loss of klf9 function are largely involved in sterol metabolism (e.g. faxdc2, sqlea, tm7sf2, sc5d, cyp51, lss, and msmo1; Fig. S12, Table S5). Interestingly, several of the processes associated with carbohydrate metabolism that we identified as being positively regulated by the GR are negatively regulated by klf9 (Fig. S13).
To ask how loss of kf9 function affects the transcriptomic response to cortisol treatment, we compared the response in wild-type embryos to that in klf9-/- larvae (Fig. 4A). Strikingly, with a false discovery threshold of 0.05 the major difference between the two responses was that ~ 70% (408) of the genes upregulated in by cortisol treatment in wild type embryos were not upregulated by the treatment in klf9-/- embryos, which instead upregulated a mostly different set of genes, albeit less strongly (Fig. 4A,B). This indicates that Klf9 is a key regulator of the transcriptomic response to cortisol. Of the 408 genes upregulated by the cortisol-treatment in a klf9-dependent way (Fig. 4A,B), about a quarter (91) were also identified in our previous study9 as being significantly upregulated by chronic cortisol exposure (Table S6). Examples of the latter include irg1l and marco (Fig. 4C and Fig. S14), as well as irg1, irf1a, ifi35, mpeg1.1, mpeg1.2, mxc, socs1a, socs3b, stat1b, and stat4 (Table S6). Gene ontology term enrichment analysis of the 408 genes upregulated by chronic cortisol treatment in wild-type but not klf9-/- embryos revealed significant enrichment for genes involved in defense and immunity (Fig. 4D, Table S7), the same biological processes that we previously found to be the most strongly affected by the chronic cortisol treatment9, whereas these processes were not associated with either the 176 genes upregulated in both wild-type and klf9-/- embryos, or the 903 genes upregulated in klf9-/- embryos but not wild-type (Table S7). Notably the set of genes upregulated by cortisol in wild-type but not klf9-/- embryos included four interferon regulatory factors (irf1a, irf8, irf9, and irf10), two interleukins (il1b and il34), and four interleukin receptors (il4r.1, il6r, il13ra1, and il20ra). These results indicate that Klf9 contributes in a significant way to the pro-inflammatory gene expression induced by chronic cortisol exposure.
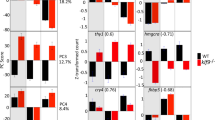
RNA-seq identifies klf9-dependent transcriptomic effects of chronic cortisol exposure. (A) Scatter plot comparing differential gene expression in response to chronic cortisol treatment in wild type (WT) and klf9-/- larvae. (B) Venn diagram comparing numbers of genes upregulated by cortisol treatment in wild type and klf9-/- larvae, and overlap. (C) Examples of two of the genes upregulated by chronic cortisol in a klf9-dependent fashion that were also identified in our previous analysis. (D) Treemap generated by REVIGO57 (https://revigo.irb.hr/) of gene ontology biological process terms found by GOrilla34 to be enriched in the set of 408 genes upregulated by chronic cortisol in a klf9-dependent way.
Genes found to be consistently upregulated by chronic cortisol treatment in multiple RNA-seq experiments depend on klf9 for that upregulation
As a final analysis we assessed the overlap between the transcriptomic effects of chronic cortisol treatment across all our RNA-seq experiments with wild-type and VBA+ (mixed wild-type and heterozygous GR369-) larvae, including the experiment published previously9, in order to identify a set of high-confidence genes that consistently respond the cortisol treatment and then ask how loss of klf9 function affects that response. To eliminate any technical artifacts emanating from the use of different parameters in the different analyses the sequence reads from all the experiments were reanalyzed from scratch using a common pipeline (see Materials and Methods). A PCA of the variance across all experiments produced several interesting observations. First, it showed that the effect of chronic cortisol treatment across all experiments was subtle, found only in PCs 4, 5, and 6 accounting for 14.3% of the total variance. PC5 captures a cortisol-treatment effect common to all three experiments (accounting for 2.77% of the total variance) and when plotted against PC4 clearly shows segregation of the cortisol-treated and control (vehicle-treated) samples (Fig. 5A). Gratifyingly, GO analysis of a single gene list ranked by upregulation along PC5 showed the same biological response to chronic cortisol as that which we reported previously9, i.e. upregulation of processes related to defense, inflammation and immunity (Fig. 5B, Table S8). The fourth principal component, accounting for 8.95% of the total variance, shows a cortisol treatment effect in the two experiments reported here, but not in that which we reported previously9 (Fig. 5A and Fig. S15). This suggests that PC4 represents cortisol-treatment effects that are dependent on the circadian light–dark cycle under which embryos in this study developed, which was absent in our previous study in which embryos developed in the dark23 (see below, Discussion). The sixth PC (2.57%) is the complement of PC4, suggesting cortisol-induced effects that are only apparent in the larvae that were developed in the dark (Fig. S15).
Combined analysis of three different RNA-seq datasets examining the transcriptomic effects of chronic cortisol exposure. (A) Principal component plot showing the location of each wild-type or VBA+ sample from all three RNA-seq experiments with respect to PCs 4 and 5. (B) Treemap generated by REVIGO57 (https://revigo.irb.hr/) of GO biological process terms found by GOrilla34 to be associated with PC5 ranked by upregulation in response to chronic cortisol treatment.
The spread of the cortisol treatment effects across three principal components likely reflects differing biological responses to the treatment among different experiments. That the effects are somewhat different under total dark compared with light–dark cycles is not surprising given what is known about the interplay of GC signaling and the circadian clock23,24,25,26. Indeed GO analysis of PC2, which segregates our previously published experiment9 from the two reported here (accounting for 18.5% of the variance), clearly shows that effects on circadian and light-responsive gene expression (Table S9). Additional sources of variance are less clear. The first PC, which segregates the wild-type samples in the last (klf9-/- vs. wild-type) experiment from the wild-type samples in our previous experiment as well as the VBA + samples described here (accounting for 37.6% of the variance, Fig. S15), is heavily loaded with genes involved in synaptic signaling and neurogenesis (Table S10) suggestive of differences in neurodevelopment and/or responsiveness to stress in those samples. The third PC (14.2% of the variance) segregates replicates 1 and 2 of the Klf9 experiment from all other samples, possibly reflecting the batch effect in sample preparation noted above.
Unsurprisingly given the large amount of gene expression variance across experimental samples unrelated to the cortisol treatment (i.e. noise), only 12 genes were found to be consistently upregulated by the cortisol treatment in all the experiments with statistical significance (adjusted p < 0.05), whereas none were consistently downregulated in all experiments (Fig. 6A). The consistently upregulated set included klf9, the only gene in that set that encodes a transcription factor. We reasoned that many more genes are affected by the treatment, albeit not consistently with statistical significance owing to the abovementioned noise (see Discussion). This was borne out by plotting the estimated fold-change of the 149 genes that were upregulated by cortisol-treatment in at least two of the three experiments (Fig. 6B, Table S11), which also revealed that the upregulation of those genes is klf9-dependent. Furthermore, this plot shows that the cortisol treatment effect is stronger in the wild-type larvae than in the VBA+ (1:2 wild-type:heterozygous GR+/-369-), suggesting haploinsufficiency of the GR for some of the effect. Query of this set for enrichment of GO biological process terms indicated chronic cortisol exposure upregulates genes associated with response to organic substance (including response to stress, defense response, response to external biotic stimulus, and inflammatory response to wounding), similar to what we reported previously9, and gluconeogenesis (Fig. 6C, Table S12).
Identification and analysis of a common set of genes upregulated by chronic cortisol treatment in multiple RNA-seq experiments. (A) Venn diagrams showing numbers of upregulated and downregulated genes following reanalysis of the data from all three experiments and their overlap. Only 8 of the 12 genes in common between all experiments were annotated with gene names (listed). (B) Violin and box plots of estimated fold change of the 149 genes that are upregulated in at least 2 of the three experiments shown in (A); the differences between each experiment are statistically significant (Table S14). (C) Treemap generated by REVIGO57 (https://revigo.irb.hr/) of GO biological process terms found by GOrilla34 to be associated with the 149 genes upregulated by chronic cortisol treatment in at least two out of the three experiments.
Finally, we used HOMER motif enrichment analysis27 to ask what transcription factor binding sites are enriched in flanking regions of the set of 149 genes upregulated by cortisol-treatment in at least two of the three experiments. The resulting set of motifs included sites for the various krüppel-like factors as well as the GR (Table S13). The most significantly enriched motif, the Klf14 binding motif RGKGGGCGKGGC, matches the Klf9 consensus motif in the JASPAR database28,29 and would be expected to bind Klf9, which is in the same KLF subfamily as Klf1430,31. In contrast, HOMER analysis of the 408 genes identified as klf9-dependent in Fig. 4 recovered a somewhat different list of motifs that included binding sites for several immunoregulatory transcription factors (Table S13; note that there are 65 genes in common between the 408 identified in Fig. 4 and the 149 identified in Fig. 6), suggesting that the larger set includes more indirect targets of feedforward regulation downstream of Klf9. Consistent with this, the latter set of motifs includes sites for two immunoregulatory genes, irf1 and stat4 that are both consistently upregulated by chronic cortisol treatment (i.e. in the common list of 149 genes identified in Fig. 6) and dependent on klf9 for that upregulation (i.e. in the list of 408 genes identified in Fig. 4). Altogether these results underscore the conclusion that klf9 is a feedforward regulator of GR signaling that mediates the pro-inflammatory transcriptomic response to chronic cortisol exposure, likely involving both direct engagement of Klf9 with its transcriptional regulatory targets and downstream effects that those targets in turn have on the genes that they regulate.
Discussion
Several previously published studies have characterized loss-of-function mutations of the zebrafish GR, including four frameshifting indels that introduce a premature stop codon in exon 213,14,15 (Fig. 1C). Here we report a new frameshift deletion (GR369-) within exon 3 (Fig. 1A,B), which unlike the previous mutations removes all possible in-frame initiation codons upstream of the DNA binding domain (Fig. 1C), eliminating the transcriptional function of the mutant gene (Fig. 1E). We used RNA-seq to determine how this loss of GR function affects the larval transcriptome, both in larvae developed under normal conditions and in larvae treated chronically with cortisol. To overcome the low survival of embryos from homozygous mutant females and avoid nonspecific maternal effects that might be associated with poor egg quality, we took advantage of a visual background adaptation (VBA) screen to identify homozygous GR369- progeny of a heterozygous cross, as those larvae lack a VBA response (VBA- larvae). Based on the observation that VBA− larvae comprised ~ ¼ of the population, as expected for a recessive Mendelian trait, larvae that successfully mount a VBA response (VBA+ larvae) are predicted to consist of a 1:2 mixture of homozygous wild-type and heterozygous GR369- mutants, and hence to contain at least one intact nr3c1 allele encoding a functional GR. A principal component analysis of the RNA-seq data revealed that absence of a functional GR has a profound effect on gene expression in both normal and cortisol-treated larvae, such that transcriptomes from larvae lacking a GR are clearly distinguished from those that have one, and that cortisol treatment produces a coherent effect only in larvae with a functional GR (Fig. 2A). A pair-wise comparison of gene expression between VBA+ and VBA− larvae (accounting for PC2) identified with statistical significance about four hundred genes that are regulated by the GR in 5-dpf larvae at the time they were collected (midmorning). GO term enrichment analysis indicated that genes upregulated by the GR at that time are involved in metabolism and stress response, as would be expected, while genes downregulated by the GR are involved in epidermis development, cell adhesion, growth, and basement membrane formation, suggesting that the GR may function as a switch to downregulate those morphogenetic processes in late development, or to temporally segregate them to a certain time of day given the circadian dynamic of glucocorticoid signaling. Interestingly, numerous biological processes and individual genes affected by loss of GR function were similarly affected by chronic cortisol treatment in larvae with a GR. This suggests that one effect of the chronic treatment is to promote development of GR resistance, and moreover, that it does so via the GR. Such resistance might be construed as an adaptive response to the chronic exposure.
Comparing gene expression in VBA+ larvae treated with cortisol versus vehicle (accounting for PC1 in that experiment) identified over four thousand genes that are differentially expressed in response to chronic cortisol treatment. This latter number is substantially larger than the 555 differentially expressed genes identified in our previous analysis9 of the effects of chronic cortisol treatment and yielded a somewhat different result when subjected to GO term enrichment analysis (Figs. S5, S6). One major difference between the analyses reported here and that reported previously9 is that in the latter the embryos and larvae were cultured in the dark, whereas in the present study we cultured them from fertilization in a diurnal light–dark cycle. In zebrafish larvae the circadian clock is not synchronized until the fish are exposed to a light–dark cycle32,33, so our previous results may have had circadian asynchrony as a confounding variable. Indeed, the impact of this difference on the transcriptome is clearly seen in the PCA of the combined analysis of all three RNA-seq experiments (Fig. S15), accounting for nearly 20% of the variance. Despite this, the RNA-seq results reported here assessing effects of chronic cortisol exposure in wild-type and klf9-/- larvae (Fig. 4) were from embryos developed with light–dark cycles, and in the wild type larvae produced an effect on proinflammatory gene expression similar to that of our earlier study, demonstrating that that effect was not an artifact of circadian asynchrony. Another difference between both studies using wild-type larvae and that depicted in Fig. 2 was that the latter measured transcriptomic effects of chronic cortisol treatment in a 1:2 mixture wild-type and heterozygous mutant larvae (VBA +); thus the results in VBA + larvae would be expected to be less sensitive to any effects of the chronic exposure for which the GR is haploinsufficient, a possibility supported by the comparison shown in Fig. 6B. Further work is required to more fully assess the effects of GR gene dosage on the transcriptome under both normal conditions and in response to chronic cortisol exposure.
The meta-analysis comparing all our RNA-seq experiments examining the transcriptomic effects of chronic cortisol treatment in wild-type or VBA + larvae (Figs. 5 and 6 and Fig. S15) provides some important insights that are broadly relevant to RNA-seq data interpretation. One is that different experiments that examine the effects of a single variable under somewhat different conditions and in a limited number of biological replicates will often produce different lists of differentially expressed genes passing an arbitrary threshold of statistical significance (e.g. adjusted p < 0.05). The reason is clear enough: biological systems are highly responsive to genetic and environmental factors that vary between experiments and affect gene expression, sometimes stochastically and/or in ways that are difficult to control and measure, especially with a limited number of biological replicates. Nevertheless, GO analyses of the lists obtained from different experiments can detect consistent biological effects even if the gene lists differ in the individual genes that they include, particularly if methods such as GOrilla34 are used to test for statistically significant enrichment of GO terms toward one end or the other of a single list of genes ranked by some measurable criterion (e.g. a principal component that accounts for a specific condition as in Fig. 5). This underscores the important but often unappreciated point that statistical significance does not equate to biological significance, and generally is not a good sole criterion to assess the effects of a given condition on the expression of a given gene using high throughput methods such as RNA-seq. By comparing across the three experiments, we were able to identify both a small but “very high confidence” set of 12 genes as well as a larger "high confidence” set of 149 genes consistently affected by the cortisol treatment that would not have been discernible without our integrated meta-analysis (Fig. 6). Moreover, use of unbiased approaches such as PCA to parse the variance in the data can help identify robust condition-specific effects and provide insight into the biology underlying those effects when combined with GO term enrichment analysis. For the experiments reported here this approach validated our earlier finding that chronic cortisol exposure leads to upregulation of pro-inflammatory gene expression and extended that result by showing that the upregulation depends on the GR target gene klf9.
The set of “very high confidence” genes identified in the meta-analysis included klf9, one of only four genes showing GR-dependent expression in normal 5-dpf larvae that were also upregulated in VBA+ larvae in response to chronic cortisol, and only one of two that encode transcription factors, the other being per1a (Fig. 2B). Both klf9 and per1a are involved in circadian regulation, and both have been shown to be GR targets in other vertebrate models16,35,36,37. Interestingly, klf9 was also found to be the most commonly upregulated transcription factor gene in a recent meta-analysis of glucocorticoid-induced gene expression in the mammalian brain38. We have found by ATAC-seq that the promoter region of klf9 is one of the most differentially open regions of chromatin in blood cells of adults derived from cortisol-treated embryos (Hartig et al., submitted). In mice klf9 was recently shown to mediate glucocorticoid-induced metabolic dysregulation in liver39. Among other things Klf9 functions as a transcriptional repressor40,41, and in mouse macrophages as an incoherent feedforward regulator of the GR target klf242, which functions to control inflammation43. Further work is needed to determine how klf9 contributes to pro-inflammatory gene expression in response to chronic cortisol exposure, which could either be directly as a feedforward activator (possibly via effects on metabolism), indirectly as a feedforward repressor of an anti-inflammatory regulator like Klf2, or both. GO analysis also showed that genes involved in sterol biosynthesis are downregulated by loss of klf9, and more experiments are required to determine if klf9 regulates the metabolism of cortisol and other steroid hormones. Our motif enrichment analysis of flanking sequences from the set of 149 genes upregulated by chronic cortisol in at least 2 of our 3 RNA-seq experiments indicated enrichment for KLF binding sites (Table S13); further work involving chromatin immunoprecipitation is needed to determine whether any of those sites are bound by Klf9 or Klf2. Additional studies are also required to determine whether loss of Klf9 alters the function of immune cells or the inflammatory response to injury or infection.
Perhaps unsurprisingly, our results (Fig. 2 and Fig. S3) indicate that the GR is required for nearly all the transcriptomic effects of chronically elevated cortisol. However, eight genes upregulated by the treatment were found in the RNA-seq analysis to be upregulated in both VBA+ and VBA− larvae (Fig. 2B), indicating that the GR is not required for their upregulation. Interestingly most of these genes are known IEGs, and include the neuronal activity-dependent gene npas4a, the mammalian homologue of which is directly repressed by the GR44, as well as egr1 which has been shown to differentially regulate GR in rat hippocampus depending on level of maternal care during development45. One possible explanation is that the genes are upregulated by increased MR activity, which was recently shown to contribute to stress axis regulation in zebrafish larvae14. Further work in MR mutant fish14 will be needed to test this.
Finally, gene ontology analysis of genes upregulated by chronic cortisol treatment in VBA+ progeny of the GR+/369- cross indicated a strong effect on biological processes associated with nervous system development and function. This is consistent with a recent report that injection of cortisol into eggs leads to increased neurogenesis in the larval brain10. In this regard it is interesting that klf9 is a stress-responsive gene that regulates neural differentiation and plasticity35,46. The long-term dysregulation of the HPA/I axis caused by early life exposure to chronic stress and/or chronically elevated cortisol suggests that the exposure perturbs brain development and activity. Given its role in regulating plasticity in brain regions relevant to neuroendocrine function, it will be interesting to determine whether klf9 contributes to those effects.
Materials and methods
Zebrafish strains, husbandry, and embryo treatments
The AB wild-type strain was used for all genetic modifications. Husbandry and procedures were as described previously9. All animal procedures were approved by the Institutional Animal Care and Use Committee (IACUC) of the MDI Biological Laboratory, and all methods were performed in accordance with the relevant guidelines and regulations. Embryo culture and cortisol treatments were performed as previously described9, with one difference: embryos were cultured in a diurnal light–dark cycle (14 h light–10 h dark). Briefly, fertilized eggs were collected in the morning, disinfected and at ~ 4 h post fertilization placed in dishes with either 1uM cortisol or vehicle (DMSO) added to embryo media. Embryos developed in a 28.5° C incubator with a 14/10 light/dark cycle synchronized with the core fish room. Media was changed daily.
Construction of nr3c1 and klf9 mutant lines
To mutate nr3c1 and klf9 we used CRISPR-Cas947, injecting zygotes with multiple guide RNAs for each gene and mRNA encoding Cas9. Guide RNAs were designed using the CHOP-CHOP algorithm48,49.
To generate the GR369- mutant line fertilized wild-type AB embryos were injected at the 1 cell stage with 1–2 nL of a gRNA cocktail targeting nr3c1 exons 2 and 3 (final concentration 40 ng/µL for each gRNA, 230 ng/µL Cas9 mRNA, 0.1 M KCl, and 0.01% phenol red indicator dye). Individual whole injected larvae were screened for mutations of the targeted regions by high resolution melt analysis (HRMA) of PCR amplicons containing those regions50. Detected mutations were then verified by Sanger sequencing of a PCR amplicon containing the targeted region. F0 adults bearing mutations were identified by HRMA of DNA extracted from tailfin clips, and germline mutations were then identified by PCR and HRMA/sequencing of sperm. F0 males with germline mutations were outcrossed to AB females, and heterozygous progeny were screened via sequencing from tailfin clips. The GR369- mutation was identified and selected for by breeding over 2 additional generations to yield F3 homozygous progeny. Subsequent generations were maintained as heterozygotes for health and breeding purposes.
To generate the klf9-/- mutant line fertilized wild-type AB embryos were injected at the 1-cell stage with < 20% cell volume of injection mix consisting of 200 ng/ul Cas9 mRNA, 100 ng/ul guide RNA, 0.05% phenol red dye, and 0.2 M KCl. Individual F0 larvae were screened with HRMA to confirm CRISPR efficacy. Larvae were placed into system and raised. Young adult fish were genotyped via HRMA using DNA extracted from fin clips, and mutations were confirmed by Sanger sequencing. F0 fish positive for mutation were outcrossed to wild-type (AB) fish. F1 offspring of this cross were screened by HRMA to confirm germline transmission, and Sanger sequencing identified the 2 bp frame-shift mutation in one female founder. This female was out crossed with WT AB males. The resulting F2 fish were screened as young adults via fin clip and HRMA, and males positive for mutation were back crossed to the F1 founder female. Resulting F3 generation fish were screened via fin clip HRMA and sequenced to identify homozygous mutants as well as homozygous wild-type siblings. F3 generation was Mendelian 1:2:1 wild-type:heterozygote:homozygous-mutant ratio.
In vitro mRNA transcription and injection
Total RNA was extracted from 5dpf wild-type larvae using Trizol. RNA was treated with DNase I (NEB), and full-length nr3c1 cDNA was reverse transcribed using a specific primer (ggtcaaggttagtttaatgaattagtctgac) and Primescript RT kit (TaKara). Template DNA for transcripts was amplified from this cDNA using Q5 high fidelity polymerase (NEB), full-length (agtaatgcaaaatggatcaaggagg), truncated 310- (ctctttgggaacagctcgcc) or 369- (gggccagtttatgcttttcca) forward primers with upstream T7 promoters, and a common reverse primer (catcgtgtcctgctgttggg) downstream of the stop codon. Template DNA was run through an agarose gel to verify size, extracted and purified using E.Z.N.A. kit from Omega Biotek. Transcription reactions were run using mMessage mMachine Ultra T7 kit from Invitrogen, and yield quantified by spectrophotometry. Xenopus elongation factor 1a transcript from pTRI-Xef included in the mMessage mMachine kit was used as a control. Homozygous GR369- mutant embryos were injected at the one-cell stage with a mix containing 200 ng/ul mRNA and 0.05% phenol red dye, with an injection volume of ~ 20% cell volume. Injected embryos were snap-frozen at 6 h post fertilization for RNA extraction and qRT-PCR.
Visual background adaptation (VBA) screen
To identify larvae lacking a functional GR a VBA screen was performed on 4-day old larvae as described12. Briefly, larvae were incubated for 20 min in a dark incubator then transferred to a white background and immediately examined under a stereomicroscope with brightfield optics. Larvae that failed to mount a VBA response were identified by the failure of melanophores to disperse, remaining clustered in a dark patch on the dorsal surface12. Larvae were segregated as VBA+ and VBA- cohorts and returned to culture for an additional day before they were collected for RNA-seq.
RNA-seq and data analysis
At 3 h zeitgeber time (post lights-on) on day 5 post-fertilization four biological replicates of cortisol-treated and control embryos were collected as follows. For the first RNA-seq experiment, one by one, a dish of larvae corresponding to a single condition was removed from the incubator, and 8 larvae per replicate (4 replicates) were collected in a 1.5 mL tube with minimal water and immediately snap frozen in liquid nitrogen. All replicates from one condition were collected sequentially before moving on to the next condition. Total RNA was extracted using the Qiagen RNA-Easy Plus mini kit (Qiagen). For the second RNA-seq experiment, four replicates of n = 10 larvae were collected from a single dish for each condition and immediately snap frozen in liquid nitrogen. Collection of 16 samples occurred over 22 min. RNA was prepared as described above on two different days. On the first day (experimental replicates 1 and 2) the lysis buffer was added to all 8 frozen samples before homogenization, while on the second day the lysis buffer was added to each sample which was then immediately homogenized. This difference in sample preparation likely accounts for the large variance between samples prepared on day 1 and day 2. RNA was sent to the Oklahoma State Genomics Facility for Illumina library preparation and single-end sequencing.
RNA-seq libraries were generated with Illumina-compatible KAPA libraries and sequenced on an Illumina NextSeq 500 High Output sequencer. klf9-/- and matched control samples were sequenced as single end 75-bp reads. VBA+ and VBA- samples were sequences as paired-end 75-bp samples.
Fastq formatted read files were preprocessed with Trimmomatic51 version 0.38 with default options, and then aligned to the Zebrafish genome version 11 as presented in Ensembl52 version 93, using the STAR aligner53 version 2.6.1b. The Ensembl transcriptome was preprocessed with a splice junction overhang of 100 nt. Following alignment, the resulting BAM files were processed with RSEM54 version 1.3.0 for isoform and gene-level expression estimates. The resulting gene-level expression values were merged into a single expression matrix with an in-house python script. EDASeq55 carried out in R version 3.6.1 was used to further normalize data for systematic effects, using gene-level length and GC-content as downloaded from Ensembl version 98 using EDAseq’s included scripts. “WithinLane” normalization with GC content was judged as superior to that based on length. Final normalized gene-level counts (which = “full”) were generated using GC-based WithinLaneNormalization followed by BetweenLaneNormalization. Subsequent differential expression analysis was carried out in R version 3.6.1 with the DESeq256 version 1.24.0, using either treatment (DMSO/cortisol) or genetics (VBA+/VBA− or klf9-/-/WT) as the comparison. The klf9-/- experiment analysis also included a two-level categorical batch covariate. DESeq2 was also used to generate a rlog-matrix which was subsequently Z-transformed to normalize each gene across all samples. The Z-transformed rlog matrix was then loaded into JMP version 15 and used for PCA (using the “Wide Method”) after thresholding to an average rlog expression value of 7.5.
Both RNA-seq dataset have been deposited in the NCBI Gene Expression Omnibus database, under accession numbers GSE144884 (GR+/− experiment) and GSE144885 (Klf9+/− experiment).
The previously published RNA-seq data set was reprocessed as just described, and an overall matrix of only wild-type or VBA+ samples was generated and jointly normalized with EDASeq and then subjected to PCA as described in the previous paragraph.
DESeq2 output tables were filtered with the following restrictions for the identification of statistically significant differentially expressed genes: BaseMean ≥ 75 (average of 5 counts per sample), padj ≤ 0.05, |log2foldchange|≥ 0.5, estimated standard error of the log2foldchange < 1.0.
The GOrilla algorithm34 (https://cbl-gorilla.cs.technion.ac.il/) was used for Gene Ontology term enrichment analysis, and the data were visualized using REVIGO57 (https://revigo.irb.hr/). Venn diagrams were generated using Venny 2.1 (https://bioinfogp.cnb.csic.es/tools/venny/). HOMER motif enrichment analysis27 was used to compare incidence of known vertebrate motifs in a list of promoters of interest with incidence in a background list of all zebrafish promoters by running the findMotifs program (https://homer.ucsd.edu/homer/microarray/index.html) using default settings except that sequence from − 1500 to + 500 bp relative to the transcription start site was searched for motifs from 10 to 18 bp in length.
Quantitative reverse transcription and polymerase chain reaction (qRT-PCR)
Total RNA purified from snap-frozen larvae using the Trizol method and used as template to synthesize random-primed cDNA using the Primescript cDNA synthesis kit (TaKaRa). Relative gene expression levels were measured by qRT-PCR, using the delta-delta Ct method as described previously9, and eif5a as a reference gene. Examination of the results of multiple RNA-seq data sets indicated that eif5a activity was highly stable across treatments and genotypes. In many experiments beta-actin was also used as a reference gene, and this did not substantially change the results.
Graphics
RNA-seq results (PCA plots, box plots, scatter plots, and violin plot) were graphed using JMP version 15 from SAS. qRT-PCR results were graphed using Microsoft Excel. All figures were drafted using Adobe Illustrator CS4. Some of the graphs (Figs. 2B,D, 4C, and 5A) were redrawn in Illustrator exactly without modifying the depiction of the data.
References
Kadmiel, M. & Cidlowski, J. A. Glucocorticoid receptor signaling in health and disease. Trends Pharmacol. Sci. 34, 518–530 (2013).
Gomez-Sanchez, E. & Gomez-Sanchez, C. E. The multifaceted mineralocorticoid receptor. Comp. Physiol. 4, 965–994 (2014).
McEwen, B. S. Influences of adrenocortical hormones on pituitary and brain function. Monogr. Endocrinol. 12, 467–492 (1979).
Karst, H. et al. Mineralocorticoid receptors are indispensable for nongenomic modulation of hippocampal glutamate transmission by corticosterone. Proc. Natl. Acad. Sci. USA 102, 19204–19207 (2005).
Spencer, R. L., Chun, L. E., Hartsock, M. J. & Woodruff, E. R. Glucocorticoid hormones are both a major circadian signal and major stress signal: How this shared signal contributes to a dynamic relationship between the circadian and stress systems. Front. Neuroendocrinol. 49, 52–71 (2018).
Coffman, J. A. Chronic stress, physiological adaptation, and developmental programming of the neuroendocrine stress system. Future Neurol. https://doi.org/10.2217/fnl-2019-0014 (2020).
Nesan, D. & Vijayan, M. M. The transcriptomics of glucocorticoid receptor signaling in developing zebrafish. PLoS ONE 8, e80726 (2013).
Nesan, D. & Vijayan, M. M. Role of glucocorticoid in developmental programming: evidence from zebrafish. Gen. Comp. Endocrinol. 181, 35–44 (2013).
Hartig, E. I., Zhu, S., King, B. L. & Coffman, J. A. Cortisol-treated zebrafish embryos develop into pro-inflammatory adults with aberrant immune gene regulation. Biol. Open 5, 1134–1141 (2016).
Best, C., Kurrasch, D. M. & Vijayan, M. M. Maternal cortisol stimulates neurogenesis and affects larval behaviour in zebrafish. Sci. Rep. 7, 40905 (2017).
Wilson, K. S. et al. Early-life glucocorticoids programme behaviour and metabolism in adulthood in zebrafish. J. Endocrinol. 230, 125–142 (2016).
Griffiths, B. B. et al. A zebrafish model of glucocorticoid resistance shows serotonergic modulation of the stress response. Front. Behav. Neurosci. 6, 68 (2012).
Facchinello, N. et al. nr3c1 null mutant zebrafish are viable and reveal DNA-binding-independent activities of the glucocorticoid receptor. Sci. Rep. 7, 4371 (2017).
Faught, E. & Vijayan, M. M. The mineralocorticoid receptor is essential for stress axis regulation in zebrafish larvae. Sci. Rep. 8, 18081 (2018).
Lee, H. B. et al. Novel zebrafish behavioral assay to identify modifiers of the rapid, nongenomic stress response. Genes Brain Behav. 18, e12549 (2019).
Shewade, L. H., Schneider, K. A., Brown, A. C. & Buchholz, D. R. In-vivo regulation of Kruppel-like factor 9 by corticosteroids and their receptors across tissues in tadpoles of Xenopus tropicalis. Gen. Comp. Endocrinol. 248, 79–86 (2017).
Nesan, D. et al. Glucocorticoid receptor signaling is essential for mesoderm formation and muscle development in zebrafish. Endocrinology 153, 1288–1300. https://doi.org/10.1210/en.2011-1559 (2012).
Pikulkaew, S. et al. The knockdown of maternal glucocorticoid receptor mRNA alters embryo development in zebrafish. Dev. Dyn. 240, 874–889. https://doi.org/10.1002/dvdy.22586 (2011).
Rossi, A. et al. Genetic compensation induced by deleterious mutations but not gene knockdowns. Nature 524, 230–233. https://doi.org/10.1038/nature14580 (2015).
Lu, N. Z. & Cidlowski, J. A. Translational regulatory mechanisms generate N-terminal glucocorticoid receptor isoforms with unique transcriptional target genes. Mol. Cell 18, 331–342 (2005).
Zeller, R. W., Coffman, J. A., Harrington, M. G., Britten, R. J. & Davidson, E. H. SpGCF1, a sea urchin embryo DNA-binding protein, exists as five nested variants encoded by a single mRNA. Dev. Biol. 169, 713–727 (1995).
Morita, M. et al. Functional analysis of basic transcription element binding protein by gene targeting technology. Mol. Cell Biol. 23, 2489–2500. https://doi.org/10.1128/mcb.23.7.2489-2500.2003 (2003).
Dickmeis, T. Glucocorticoids and the circadian clock. J. Endocrinol. 200, 3–22 (2009).
Dickmeis, T. et al. Glucocorticoids play a key role in circadian cell cycle rhythms. PLoS Biol. 5, e78 (2007).
Dickmeis, T., Weger, B. D. & Weger, M. The circadian clock and glucocorticoids–interactions across many time scales. Mol. Cell Endocrinol. 380, 2–15 (2013).
Weger, B. D. et al. Extensive regulation of diurnal transcription and metabolism by glucocorticoids. PLoS Genet. 12, e1006512 (2016).
Heinz, S. et al. Simple combinations of lineage-determining transcription factors prime cis-regulatory elements required for macrophage and B cell identities. Mol. Cell 38, 576–589 (2010).
Fornes, O. et al. JASPAR 2020: update of the open-access database of transcription factor binding profiles. Nucl. Acids Res https://doi.org/10.1093/nar/gkz1001 (2020).
Sandelin, A., Alkema, W., Engstrom, P., Wasserman, W. W. & Lenhard, B. JASPAR: an open-access database for eukaryotic transcription factor binding profiles. Nucl. Acids Res 32, D91-94. https://doi.org/10.1093/nar/gkh012 (2004).
de Assuncao, T. M. et al. New role for Kruppel-like factor 14 as a transcriptional activator involved in the generation of signaling lipids. J. Biol. Chem. 289, 15798–15809. https://doi.org/10.1074/jbc.M113.544346 (2014).
Pearson, R., Fleetwood, J., Eaton, S., Crossley, M. & Bao, S. Kruppel-like transcription factors: a functional family. Int. J. Biochem. Cell Biol. 40, 1996–2001. https://doi.org/10.1016/j.biocel.2007.07.018 (2008).
Kaneko, M. & Cahill, G. M. Light-dependent development of circadian gene expression in transgenic zebrafish. PLoS Biol. 3, e34 (2005).
Dekens, M. P. & Whitmore, D. Autonomous onset of the circadian clock in the zebrafish embryo. Embo J. 27, 2757–2765 (2008).
Eden, E., Navon, R., Steinfeld, I., Lipson, D. & Yakhini, Z. GOrilla: a tool for discovery and visualization of enriched GO terms in ranked gene lists. BMC Bioinform. 10, 48 (2009).
Bonett, R. M., Hu, F., Bagamasbad, P. & Denver, R. J. Stressor and glucocorticoid-dependent induction of the immediate early gene kruppel-like factor 9: implications for neural development and plasticity. Endocrinology 150, 1757–1765 (2009).
Sporl, F. et al. Kruppel-like factor 9 is a circadian transcription factor in human epidermis that controls proliferation of keratinocytes. Proc. Natl. Acad. Sci. USA 109, 10903–10908 (2012).
Reddy, T. E., Gertz, J., Crawford, G. E., Garabedian, M. J. & Myers, R. M. The hypersensitive glucocorticoid response specifically regulates period 1 and expression of circadian genes. Mol. Cell Biol. 32, 3756–3767 (2012).
Juszczak, G. R. & Stankiewicz, A. M. Glucocorticoids, genes and brain function. Prog. Neuropsychopharmacol. Biol. Psychiatry 82, 136–168 (2018).
Cui, A. et al. Dexamethasone-induced Kruppel-like factor 9 expression promotes hepatic gluconeogenesis and hyperglycemia. J. Clin. Invest. 130, 2266–2278 (2019).
Knoedler, J. R., Subramani, A. & Denver, R. J. The Kruppel-like factor 9 cistrome in mouse hippocampal neurons reveals predominant transcriptional repression via proximal promoter binding. BMC Genom. 18, 299 (2017).
Ying, M. et al. Kruppel-like factor-9 (KLF9) inhibits glioblastoma stemness through global transcription repression and integrin alpha6 inhibition. J. Biol. Chem. 289, 32742–32756 (2014).
Chinenov, Y., Coppo, M., Gupte, R., Sacta, M. A. & Rogatsky, I. Glucocorticoid receptor coordinates transcription factor-dominated regulatory network in macrophages. BMC Genom. 15, 656 (2014).
Nayak, L. et al. Kruppel-like factor 2 is a transcriptional regulator of chronic and acute inflammation. Am. J. Pathol. 182, 1696–1704 (2013).
Furukawa-Hibi, Y., Yun, J., Nagai, T. & Yamada, K. Transcriptional suppression of the neuronal PAS domain 4 (Npas4) gene by stress via the binding of agonist-bound glucocorticoid receptor to its promoter. J. Neurochem. 123, 866–875 (2012).
Weaver, I. C. et al. Epigenetic programming by maternal behavior. Nat. Neurosci. 7, 847–854 (2004).
Besnard, A. et al. Targeting Kruppel-like factor 9 in excitatory neurons protects against chronic stress-induced impairments in dendritic spines and fear responses. Cell Rep. 23, 3183–3196 (2018).
Hwang, W. Y. et al. Efficient genome editing in zebrafish using a CRISPR-Cas system. Nat. Biotechnol. 31, 227–229. https://doi.org/10.1038/nbt.2501 (2013).
Labun, K., Montague, T. G., Gagnon, J. A., Thyme, S. B. & Valen, E. CHOPCHOP v2: a web tool for the next generation of CRISPR genome engineering. Nucleic Acids Res. 44, W272-276. https://doi.org/10.1093/nar/gkw398 (2016).
Montague, T. G., Cruz, J. M., Gagnon, J. A., Church, G. M. & Valen, E. CHOPCHOP: a CRISPR/Cas9 and TALEN web tool for genome editing. Nucleic Acids Res. 42, W401-407. https://doi.org/10.1093/nar/gku410 (2014).
Parant, J. M., George, S. A., Pryor, R., Wittwer, C. T. & Yost, H. J. A rapid and efficient method of genotyping zebrafish mutants. Dev. Dyn. 238, 3168–3174. https://doi.org/10.1002/dvdy.22143 (2009).
Bolger, A. M., Lohse, M. & Usadel, B. Trimmomatic: a flexible trimmer for Illumina sequence data. Bioinformatics 30, 2114–2120 (2014).
Zerbino, D. R. et al. Ensembl 2018. Nucl. Acids Res. 46, D754–D761. https://doi.org/10.1093/nar/gkx1098 (2018).
Dobin, A. et al. STAR: ultrafast universal RNA-seq aligner. Bioinformatics 29, 15–21. https://doi.org/10.1093/bioinformatics/bts635 (2013).
Li, B. & Dewey, C. N. RSEM: accurate transcript quantification from RNA-Seq data with or without a reference genome. BMC Bioinform. 12, 323 (2011).
Risso, D., Schwartz, K., Sherlock, G. & Dudoit, S. GC-content normalization for RNA-Seq data. BMC Bioinform. 12, 480. https://doi.org/10.1186/1471-2105-12-480 (2011).
Love, M. I., Huber, W. & Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 15, 550. https://doi.org/10.1186/s13059-014-0550-8 (2014).
Supek, F., Bosnjak, M., Skunca, N. & Smuc, T. REVIGO summarizes and visualizes long lists of gene ontology terms. PLoS ONE 6, e21800 (2011).
Acknowledgements
This work was supported by Grants from the National Institutes of Health (R03- HD099468, P20-GM104318, and P20-GM103423), by a Morris Scientific Discovery Award to JAC, and by a gift from The Linde Packman Lab for Biosciences Innovation to Colby College.
Author information
Authors and Affiliations
Contributions
IG and EIH performed most of the experiments and did the matings, microinjections and husbandry to establish the klf9 and nr3c1 mutant zebrafish strains; SZ designed the gRNAs and performed the initial PCR and sequencing analyses to identify fish containing the desired nr3c1 mutations; IG performed the mRNA injection experiments, qRT-PCR, RNA-seq data analysis, and HOMER motif enrichment analysis; ART contributed to the RNA-seq experimental design and data analysis; LNH wrote the code used in the RNA-seq data analysis; NJM performed the initial analysis of the RNA-seq data; JHG directed the computational analyses and performed the PCA and DGE analyses; JAC conceived the study, contributed to analysis of the data and drafted the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Gans, I., Hartig, E.I., Zhu, S. et al. Klf9 is a key feedforward regulator of the transcriptomic response to glucocorticoid receptor activity. Sci Rep 10, 11415 (2020). https://doi.org/10.1038/s41598-020-68040-z
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-020-68040-z
This article is cited by
-
Krüppel-like factor 9 (KLF9) links hormone dysregulation and circadian disruption to breast cancer pathogenesis
Cancer Cell International (2023)
-
Loss of Krüppel-like factor 9 deregulates both physiological gene expression and development
Scientific Reports (2023)
-
Molecular signature of excessive female aggression: study of stressed mice with genetic inactivation of neuronal serotonin synthesis
Journal of Neural Transmission (2023)
-
Sex-specific differences in zebrafish brains
Biology of Sex Differences (2022)
-
Identification and characterization of relapse-initiating cells in MLL-rearranged infant ALL by single-cell transcriptomics
Leukemia (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.