Abstract
Quantitative analysis of dynamic processes in living cells using time-lapse microscopy requires not only accurate tracking of every particle in the images, but also reliable extraction of biologically relevant parameters from the resulting trajectories. Whereas many methods exist to perform the tracking task, there is still a lack of robust solutions for subsequent parameter extraction and analysis. Here a novel method is presented to address this need. It uses for the first time a deep learning approach to segment single particle trajectories into consistent tracklets (trajectory segments that exhibit one type of motion) and then performs moment scaling spectrum analysis of the tracklets to estimate the number of mobility classes and their associated parameters, providing rich fundamental knowledge about the behavior of the particles under study. Experiments on in-house datasets as well as publicly available particle tracking data for a wide range of proteins with different dynamic behavior demonstrate the broad applicability of the method.
Similar content being viewed by others
Introduction
Single particle tracking in live cell fluorescence microscopy imaging data serves as a powerful tool to study the dynamics of a wide range of different particles. Here, “particle” is a generic term that can, amongst others, refer to small fluorophores, single molecules, macromolecular complexes, viruses, organelles or microspheres1,2. Consequently, single particle tracking (SPT) can be broadly applied in microrheology3,4,5,6 as well as in studying dynamic processes in live cells. Examples of such processes are microtubule assembly and disassembly7, cell migration governed by focal adhesions8, membrane dynamics9, intracellular transport10, chromatin assembly and gene transcription11, genome maintenance12,13, and virus trafficking14. Since manual tracking is subjective and becomes quite cumbersome for large datasets, automated tracking is preferred15,16. Many different software tools are available for SPT and new methods are still being developed17. SPT results in a series of coordinates over time for every single particle (also called “trajectories”), but by itself does not provide direct insights into the dynamic process of interest.
In order to relate trajectories of individual particles to the behavior of the population, mobility patterns must be analyzed in an automated, unbiased and statistically relevant way. As molecular behavior is commonly linked to function and structure, mobility analysis is connected to a deeper understanding of the associated biological process. The goal is to quantify behavior by determining physical properties of the particle of interest, such as velocity, processivity, confinement or spatial distribution18,19. Additional biological insights into the dynamic behavior of populations with mixed mobility can be provided by determining the relative fractions of particles in different functional states under varying conditions.
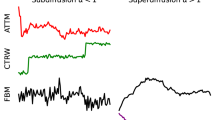
There are several approaches to this type of analysis, each with its own drawbacks. Methods based on single time steps, such as hidden Markov modeling (HMM)20,21,22,23 and probability density function (PDF) or cumulative distribution function (CDF) fitting, are problematic for detecting motion types that exhibit patterns over longer time-scales (Supplementary Note 1). There are also methods that use rolling windows of multiple time-points for classification. The main methods in this category are based on machine learning24,25 and, most commonly used in biological research, mean square displacement (MSD) analysis9,26,27,28 (Supplementary Note 2). A drawback of these methods is that a set window size introduces a trade-off between sensitivity and accuracy. Moreover, MSD-based methods are mostly limited to quantitative analysis of particles that exhibit pure diffusion, while in practice confined (subdiffusive) motion and highly correlated (superdiffusive) motion are quite common.
A critical limitation is that most of these methods are not able to detect switching from one behavior type to another within single trajectories, while change in behavior is the core of biological function. One approach to capturing this transient behavior uses image segmentation to distinguish between free motion and trapping of molecules29. In this method, the trapping state is characterized by the accumulation of trajectory segments, leading to a denser cloud in the image. However, in applications such as ours, this type of approach leads to inaccuracies when the times spent in each state become relatively short, or when trajectories are not long enough or do not form compact and well defined regions where particles are trapped.
More recently, a method was developed that uses divide-and-conquer classification (where trajectories first get an initial segmentation that is refined in subsequent steps) in combination with the moment scaling spectrum (MSS)30, an advanced measure for random motion characterization that has also been used in a variety of other motion studies31,32,33. By uncoupling segmentation and further motion analysis, this method allows to detect different types of motion as well as mobility switches. However, the number and the location of switching points are not always determined accurately, segmentation takes multiple steps, and there is a higher probability of misclassification for shorter trajectories30.
In this paper, a novel general method is presented to robustly analyze particle trajectories, providing information about the type of motion, associated parameters, and switching behavior. Here, particle trajectories are analyzed using state-of-the-art deep learning techniques in combination with advanced post-processing. The proposed method consists of two components and will henceforth be referred to as DL-MSS (Deep Learning followed by Moment Scaling Spectrum analysis).
Firstly, a deep learning (DL) neural network is trained with simulated data containing trajectories that switch between different types of mobility. This self-contained deep learning approach does not require any specific modelling or manual parameter tuning. Even though one could also use manually annotated real data, if available, the approach of using simulated trajectories to estimate the dynamics of real systems has been shown to be fruitful previously34. The trained network is applied to real microscopy imaging data to segment trajectories into segments, referred to as “tracklets”, that exhibit the same type of motion.
Subsequently, these tracklets are further analyzed using the moment scaling spectrum (MSS) and clustered according to their diffusion constant and type of motion to determine parameters associated with each class of mobility. The concept of the MSS is not new in more theoretical fields, but its practical application is overshadowed by the simpler MSD analysis, which can provide only a limited understanding of the underlying random behavior. MSS is a very robust tool to analyze and understand what modes of motion are present in a dataset, and implicitly contains other frequently used methods such as MSD and correlation between subsequent displacements35,36.
DL-MSS is able to perform segmentation in a single step without being limited to any trajectory length and returns a number of mobility classes with their associated parameters, providing fundamental knowledge about the behavior of the particle in question. Since this method can separate different populations in a dataset, DL-MSS makes it possible to compare the collective mobility of a specific type of molecule at different conditions.
There are many potential applications of single particle tracking (SPT) and trajectory analysis using DL-MSS. In this article, the focus is on mobility patterns of different nuclear proteins that exhibit multiple types of random walk-type behavior. DL-MSS was inspired by observable switches in motion for breast cancer susceptibility protein 2 (BRCA2). This large, multifunctional protein is most well-known for its role in the repair of double strand breaks (DSBs) in DNA37,38. Since DSBs can be introduced in live cells artificially through ionizing radiation39, BRCA2 mobility pattern analysis provides a nice showcase to detect behavioral changes upon DNA damage induction. As control datasets, histone protein H2B and nuclear localization signal (NLS) were used, as these molecules are characterized as mostly stuck or mostly free, respectively. The immobile H2B dataset can simultaneously be used to confirm that global movements of the cell are negligible compared to the local movement of single molecules12,40,41. Moreover, four publicly available datasets (generously provided by the authors of42) were used that contain trajectories of several proteins that are expected to exhibit different types of mobility, ranging from immobile to freely diffusing. DL-MSS was used successfully to classify and analyze all these different datasets in accordance with expected results.
Results
Deep learning neural network
DL-MSS consists of two elements: a deep learning part followed by a post-processing part (Fig. 1). A long short term memory (LSTM) deep learning recurrent neural network was used for trajectory segmentation. This type of model was chosen because LSTM networks are known to be flexible to input size (which in this study depends on the trajectory length) and to be able to retain information over longer timescales43,44,45 (Supplementary Note 3). This network was trained with simulated trajectories that switch between three mobility classes: one fast diffusing state (diffusion constant 1.0 μm2/s), one slow diffusing state (diffusion constant 1.0 μm2/s) and one immobile state. The immobile state is chosen to reflect the situation where a molecule is “stuck”, e.g. where it is impossible to distinguish between motion of the molecule, the movement of the cell41 and the localization error of the molecules. These classes were chosen to reflect the mobility patterns of fluorescently labeled BRCA2, which served as the incentive to develop DL-MSS and showed at least two mobility classes, namely immobile and diffusing12. One extra class was added to increase the flexibility of the model without introducing overfitting to extra clusters that do not provide useful information. As the proposed deep learning network will not inadvertently detect motion types that are not actually present (Supplementary Note 4) and potential additional mobility classes can be detected later on in MSS analysis (Supplementary Note 5), this three-state model provides a simple yet flexible basis for classification. On simulated three-state mobility data, the trained network achieved an accuracy of 0.94 on the training set and an accuracy of 0.92 on the testing set (Online Methods), out of a maximum accuracy of 1. The trained network can be applied to unseen simulated data as well as trajectories extracted from real microscopy data and classifies these trajectories per time step for any length of trajectory (Supplementary Note 6). The same trained network was used for classification of all datasets mentioned in this paper.
Overview of the DL-MSS method. Automatic tracking software is used to obtain single molecule trajectories from fluorescence microscopy data. A trained deep learning (DL) neural network is applied to these trajectories to segment them into “tracklets” of consecutive track points that were classified to have the same type of mobility. Tracklets are further analyzed using the moment scaling spectrum (MSS) to acquire the properties associated with each class.
Moment scaling spectrum analysis
Segmented trajectories produce so-called “tracklets”, which are segments that are classified to one of the three states. These tracklets are further analyzed using the moment scaling spectrum (MSS). As opposed to classical methods such as MSD-based analysis, which makes use of only the second moment (x2 ∼ τ with x position and τ time step), MSS utilizes higher order moments27,35:
where N is the number of trajectories, Tn is the duration of trajectory n, τ is the time step, xn(t) is the position of the nth particle at time t, and p is the moment order. This means that xp ∼ τγp, where the plot of γp versus p gives the MSS. The slope of the MSS, denoted SMSS, indicates the motion type of the tracklet. In this spectrum, SMSS = 0.5 represents pure diffusion, \(0 < {S}_{MSS} < 0.5\) represents restricted motion, and \(0.5 < {S}_{MSS} < 1.0\) represents more directed motion30 (Supplementary Note 7). The SMSS can be calculated along with the diffusion constant D (to distinguish between “faster” and “slower” motion, Supplementary Note 8) for every tracklet in order to obtain a scatterplot of all tracklets together in SMSS-D space46. This procedure yields clusters of tracklets with the same kind of mobility, showing the properties of the different classes of tracklets. Because MSS analysis is less reliable for shorter tracklets, only tracklets of more than ten time frames are used for clustering (Supplementary Note 9). However, as MSS analysis is used only to determine the properties for clusters of tracklets with the same classification label, these properties can still be assigned to shorter tracklets as well, because they were classified by the deep learning neural network to have the same type of mobility as the longer tracks. Note that this method does not deny there can be more than three classes of mobility in a given dataset. Clusters can be subdivided into multiple classes recursively. DL-MSS aims to find the major clusters of motion so the proportions of tracklets in those clusters can be compared between different datasets.
Showcase 1: BRCA2 behavioral change upon treatment with ionizing radiation
The first showcase for the application of DL-MSS is the response of BRCA2 protein mobility upon treatment of the cells with ionizing radiation (IR), which introduces DSBs into DNA. BRCA2 plays an important role in the repair of DSBs, and is known to accumulate at nuclear sites of DNA damage39,47. Consequently, the corresponding hypothesis is that more BRCA2 molecules should become immobile upon IR treatment compared to untreated cells12.
From the SMSS versus D scatterplot of wildtype BRCA2 without any treatment (Fig. 2a), it is clear that the three-state mobility model fits the data well; the clusters of data points with common mobility characteristics are well defined, well sorted by class (each cluster contains only one color), and well separated. Moreover, no extra clusters are visible, indicating there are no additional mobility classes. The location of each cluster mean (indicated with “+” in Fig. 2a) in SMSS-D space specifies the properties of the corresponding class of tracklets. For BRCA2 without IR, DL-MSS yields three motion types: the first is very slow and immobile, the second is slow and close to free diffusion, and the third is fast and close to free diffusion as well. The immobile cluster presumably reflects protein localized to perform its repair function. The possibility to detect multiple mobile states (in this case slow and fast diffusion) is important because these different states can be biologically relevant, as proteins can be modified and can also interact with other molecules and structures in the cell.
SMSS versus D plots for the BRCA2 protein without and with ionizing radiation (IR). (a,b) scatterplot for BRCA2 –IR/BRCA2 + IR where red, blue and grey color coding corresponds to fast, slow and immobile tracklets, respectively. Histograms on the sides show the distributions of the tracklets in different clusters relative to each other for the different axes. Cluster means are indicated by the + symbol. (c,d) kernel density estimation plot for BRCA2 –IR/BRCA2 + IR, color intensity indicates density (see colorbar).
The same type of scatterplot for BRCA2 tracklets from cells treated with IR (Fig. 2b) shows that even though there is very little variation in the location of the data clusters in SMSS-D space, their relative fractions change upon IR-treatment. This means that particle mobility characteristics do not change but the portion of particles in the different classes does change, which becomes even more clear when comparing the kernel density estimation (KDE) maps (Fig. 2c,d). These density maps show a shift from the diffusive states (mainly the fast diffusive state) to the immobile state after inducing DSBs, corresponding to the idea that more damage sites require more BRCA2 molecules to become “stuck” in order to perform their task. DL-MSS serves as a tool to successfully detect this behavioral change in a unique way, as it provides information about how fast the molecules move around as well as the specific type of motion. What is striking about this type of analysis is that it reveals how the relative intensities of the data clusters change rather than the cluster locations.
Showcase 2: Unimodal mobility of histone protein H2B-HaloTag and nuclear localization signal (HaloTag-NLS)
Of course, not every molecule exhibits three types of mobility. In order to test whether or not DL-MSS is prone to overfitting, trajectories were analyzed for two molecules for which the behavior is known to be very simple. HaloTag labelled histone protein H2B and a nuclear localization signal (HaloTag-NLS) were chosen for this purpose because they are known to be predominantly immobile and fast diffusing, respectively48,49. DL-MSS finds only an immobile cluster for H2B (Fig. 3a,c) and almost exclusively finds fast diffusing tracklets for NLS (Fig. 3b,d), where the corresponding diffusion constant is considerably higher than that of the fast diffusive population of BRCA2 (Fig. 2). This result was obtained using the same network as for the first showcase, trained on the same simulated three-state mobility data. These results not only show that DL-MSS identifies the expected clusters for these control datasets, but also that this method does not find mobility classes that are not present in the data and that clusters are not necessarily bound to specific locations.
SMSS versus D plots for the H2B protein and NLS. HaloTag was used for tracking. (a,b) scatterplot for H2B and NLS where red, blue and grey color coding corresponds to fast, slow and immobile tracklets, respectively. Histograms on the sides show the distribution of tracklets in the clusters relative to each other for the different axes. Cluster means are indicated by the + symbol. (c,d) kernel density estimation plot for H2B and NLS, color intensity indicates density (see colorbar).
Showcase 3: Publicly available datasets for H2B, CTCF, Sox2 and 3 × NLS
Finally, DL-MSS was applied to four datasets that were made publicly available by the authors of42, which were imaged and tracked in a different way than the datasets analyzed above (Online Methods). This was done in order to demonstrate that the applicability of DL-MSS is not limited to our own type of imaging data, particle dynamics, or tracking algorithm. The four datasets contain trajectories of histone protein H2B, transcription factors CTCF and Sox2 and a protein consisting of three tandem repeats of nuclear localization signal (3 x NLS), all fused to a HaloTag. What makes these datasets interesting for DL-MSS analysis is that they range from being mostly stuck to being mostly free, in the order H2B – CTCF – Sox2 – 3 x NLS (see Fig. 4G,H in42). This spectrum of different types of behavior should become visible after DL-MSS analysis through a shift from the immobile state to the free state. The kernel density estimation plots for the four datasets (Fig. 4) clearly illustrate that DL-MSS indeed picks up the shift in SMSS-D space from immobile to free. The difference in D found for this publicly available dataset of H2B compared to the in-house H2B dataset from the previous showcase can be explained by the difference in frame rate for data acquisition (5 vs 30 ms interval, respectively). A higher frame rate means that interframe displacements can be smaller, while the detection error remains the same. This means that the detection error gets larger relative to the displacements between frames, leading to an overestimation of the associated diffusion constant.
Discussion
DL-MSS is a general method to analyze single particle trajectories through single time step classification and clustering of segmented tracklets in SMSS-D space. This procedure yields specific mobility classes, which was exemplified by the showcases presented in this paper. As opposed to traditional analysis methods, DL-MSS defines mobility clusters based on the diffusion constant as well as the type of mobility. Additionally, this method makes use of state-of-the-art deep learning techniques for classification, which not only makes it possible to accurately segment trajectories into tracklets before calculating any biological parameters, but also allows classification of tracklets that would ordinarily be too short for mobility analysis. Moreover, DL-MSS is flexible, meaning that mobility classification is not restricted to either the number of classes the network was trained with, or the parameters (D, SMSS) that were assigned to the training data. Finally, this method is user-friendly; results can be obtained by running one single script, while still allowing the user to supervise all intermediate steps in classification and further analysis.
All these properties of DL-MSS can facilitate new insights into biological problems. In the example of BRCA2, it was already shown that this protein has multiple states of mobility12. However, instead of fitting a certain number of diffusion classes, DL-MSS yields new information about the types of motion in BRCA2 behavior (one fast diffusive class, one slow subdiffusive class and one immobile class). Furthermore, it was shown that the mobility classes of BRCA2 do not change with regard to their location in SMSS-D space when DNA damage is introduced into the cells. Rather, it is the relative density of the three clusters that changes. Additionally, the flexibility of DL-MSS was illustrated by the showcases of H2B, NLS and Spot-On datasets. These results clearly showed that this method is not restricted to the classification of mobility patterns that exhibit behavior similar to the simulated data the network was trained with, irrespective of the methods that were used to obtain the trajectory data.
Altogether, DL-MSS is a very versatile method that can be used for a wide range of applications. Moreover, DL-MSS is not only useful to analyze different molecules separately, but also to compare mobility patterns between different types of molecules. By analyzing mobility classes from datasets obtained through different experiments, the meaning and function of those classes can be elucidated. This is especially interesting when certain molecules are suspected to interact with each other, when there are different variants of the same molecule, or when multiple datasets are available of the same molecule but within different environments or with different treatments. Comparing different datasets to one another can easily be done using DL-MSS, as different datasets can be fully analyzed in parallel in only a few minutes up to a few hours, depending on the size of the dataset and computing power. All datasets mentioned in this paper were analyzed within 1 hour on a normal laptop (1.8–2.4 GHz Intel i7 CPU with 8 GB RAM) with the exception of the larger Spot-On H2B dataset, which took 2.5 hours (Supplementary Note 10). Of course, this method requires training, which takes 2–3 hours on the GPU used in this study (Nvidia GTX 980), or 6–7 hours on the CPU used. However, the training has to be done only once, after which the model can be saved and applied as many times as needed.
The DL-MSS software is not limited to producing the type of results shown in this paper. Depending on the application and the needs of the user, the software can be used to extract additional useful parameters for the dataset and there is a large variety of visualization options (Supplementary Note 11). For example, classification results can be used to determine the switching probabilities from one state to another, the dwell times per state, and the fraction of time points spent in each state. In terms of visualization, molecule trajectories can be plotted inside the cell (nucleus) with different colors per state in order to see if there are certain patterns. In the example of BRCA2, this type of figure could be useful to see if there are regions inside the cell nucleus where more BRCA2 proteins are immobile, possibly indicating the presence of DNA damage in these regions. Additionally, DL-MSS can be used to detect inconsistencies in tracking, which manifest themselves as clusters at unexpected locations (Supplementary Note 12).
Since there are no clearly defined rules for deep learning, there are many ways to adapt and elaborate the network that is part of the DL-MSS method proposed in this paper to increase accuracy on simulated training data, where the ground truth is available. Theoretically it should even be possible to train a network that outputs the diffusion constant D and moment scaling spectrum slope SMSS at once. However, this would likely lead to an increase in computational demand. The relatively straightforward network presented here offers a nice balance between simplicity, performance and flexibility. Moreover, the uncoupled MSS analysis gives the opportunity to monitor and control the output of the network. Overall, DL-MSS provides a new, robust and very flexible tool for particle mobility analysis.
Online Methods
Simulation of trajectories for training
The lengths of the simulated trajectories were randomly sampled according to \({L}_{{\rm{track}}} \sim Exp(\lambda )=\lambda {e}^{-\lambda x}\) with rate parameter λ50. Every track was randomly assigned an initial type of mobility and labeled correspondingly. The switching probability was modeled using a Markov model51 with state transition probability matrix
where pii is the probability of remaining in state i and pij is the probability of switching from state i to state j. With such a problem setup, the number of steps Si that a particle will remain in a certain state i can be sampled using a geometric distribution, where \(\Pr ({S}_{i}=k)={{p}_{ii}}^{k-1}(1-{p}_{ii})\) gives the probability that the kth step is followed by a switching event. To create training sets, it should be possible to generate any type of process, diffusive as well as anomalous. Pure diffusion (Brownian motion) can easily be simulated in 1D from the normal distribution \({\mathscr{N}}(\mu ,\,{\sigma }^{2})\) with μ = 0 and \(\sigma =\sqrt{2D\tau }\) (diffusion constant D and time step τ, where τ can be chosen but does not have to match the real data). For 2D or 3D cases, the simulation of displacements is done independently for each coordinate. Anomalous diffusion was modeled using fractional Brownian motion (fBm)52, where the type of motion depends on the Hurst component H, which is equal to 0.5 for pure diffusion, lower than 0.5 for subdiffusion and higher than 0.5 for superdiffusion. fBm can be simulated53 using
where \(\Delta x(t)\) is the displacement in x for one time step, n is the number of intervals that every time step is divided into, H is the Hurst component, \(\varGamma \) is the gamma function, t is the integer time, M is the range that can be covered in time t and \(\xi \) are independent and identically distributed samples from a normal (Gaussian) distribution with zero mean and unit variance. \(\Delta y(t)\) was simulated in the same way as \(\Delta x(t)\) to create any type of anomalous diffusion as well as pure diffusion (Supplementary Note 13). The data was subsequently scaled to appear at specific SMSS versus D using scaling factor η. For the model used in this paper, training was done with three-state simulated data with three corresponding labels: “\(0\)” for diffusion with \(D\,=\,1.0\,{\rm{\mu }}{{\rm{m}}}^{2}/{\rm{s}}\), “\(1\)” for diffusion with \(D\,=\,0.1\,{\rm{\mu }}{{\rm{m}}}^{2}/{\rm{s}}\) and “\(2\)” for the immobile state with Hurst coefficient \(H\,=\,0.1\) and scaling factor \(\eta \,=\,0.3\). The transition probability matrix is given by:
Deep learning using an LSTM recurrent neural network
As mobility state prediction requires sequential analysis as well as the ability to learn long-term dependencies, the model of choice for DL-MSS was a bidirectional Long Short Term Memory (LSTM) network45,54. Using a bidirectional network increased the performance of both ends of the window as well as in the middle compared to only forward and only reverse networks (Supplementary Note 14). For each time step, the distance travelled by the molecule was fed into the network, along with the x- and y- coordinates of the two points flanking this distance and some higher order average distances (Supplementary Note 15). The number of LSTM units corresponds to the number of time steps in a trajectory and every unit outputs a class label by passing the resulting hidden state (containing \(200\) hidden units, Supplementary Note 16) through a fully connected layer before passing the hidden state on to the next unit. The model was implemented in Keras (with TensorFlow as backend) and optimized using categorical cross entropy as the loss function and Adam as the optimization method55,56. EarlyStopping, an algorithm that stops the training process when the validation error considerably exceeds the training error, was used as a generalization method57.
The simulated trajectories were split into time windows of \(25\) frames (Supplementary Note 16) and used for training (\(10,000\) windows), validation (\(5,000\) windows), and testing (\(5,000\) windows) with a batch size of \(\,256\). Ten “splits” were performed to get a reliable estimate for the accuracy of prediction, meaning that new training, validation and test sets were picked ten times from the total pool of available time windows to repeat the training procedure. The trained network can be applied to trajectories of any size.
MSS analysis
For moment scaling spectrum (MSS) analysis, only tracklets that have a length of ten or more frames and do not have a negative \(D\) or \({S}_{MSS}\) due to unstable linear regression were selected to get a reliable result. For kernel density estimation (KDE) on the data points in SMSS-D space, a Gaussian kernel was used with a bandwidth (\({\rm{bw}}\)) corresponding to Scott’s rule (\({\rm{bw}}=\,{n}^{-1/(d+4)}\)58, where n is the number of data points and d is the number of dimensions).
DL-MSS software
The DL-MSS method was implemented in the Python programming language with Keras and using the TensorFlow library as the deep-learning backend. The software and other scripts used in the presented experiments are publicly available at GitHub: https://github.com/ismal/DL-MSS.
Single molecule tracking experiments for BRCA2, H2B-HaloTag and HaloTag-NLS
IB10 mouse embryonic stem cells (mESCs) were cultured on gelatinized plates (0.1% porcine gelatin (Sigma)) in 50% DMEM (High-Glucose, Ultraglutamine, Lonza), 40% BRL conditioned medium and 10% FCS supplemented with non-essential amino acids, 0.1 mM β-mercaptoethanol, pen/strep and 1,000 U/ml leukemia inhibitory factor.
BRCA2 in these mESCs was tagged with HaloTag at the C-terminus by modification of the endogenous locus of BRCA2 using CRISPR/Cas9. A detailed description of the exact methods can be found in59. In short, cells were electroporated with 15 µg of both the px459 Cas9/gRNA plasmid (gRNA: gctgttgagtcttagcctcc) as well as the donor plasmid consisting of homology arms and HaloTag-F2A-neo cassette12. After antibiotic selection, clones were picked and validated for homozygous integration of the cassette by PCR genotyping and western blotting. H2B-HaloTag and HaloTag-NLS were cloned into a PiggyBac vector60 containing a CAG promoter and PGK-puro selection cassette, and stable cell lines were generated by Lipofectamine 3000 transfection followed by puromycin selection.
For imaging, cells were seeded in µ-Slide 8 Well Glass Bottom (Ibidi) coated with 25 ug/ml Laminin (Roche) the day before the experiment. For ionizing radiation (IR) cells were treated with \(5\) Gy of X-rays. Cells were labeled with fluorescent 5 nM JF549-HaloTag61 ligand (500 pM for HaloTag-H2B and HaloTag-NLS) for 15 minutes in Fluorobrite medium (ThermoFisher), complemented with 10%FCS, non-essential amino acids, 0.1 mM β-mercaptoethanol, pen/strep and 1,000 U/ml leukemia inhibitory factor. To remove free HaloTag ligand from the cells, twice Fluorobrite medium was exchanged with 15-minute interval. Experiments were done around 2 hours after irradiation. Imaging was performed using HiLo illumination on an Elyra PS1 system with 100 × 1.49NA α Plan Apochromat DIC (Zeiss) TIRF objective and Tokai Hit stage and objective heating (37 °C and 5% CO2). For excitation of JF549 a 100 mW 561 nm laser was used with a 570–650 nm bandpass filter. Signal was detected on an Andor iXon DU897 with 256 × 256 pixel region at 32 ms interval with an EMCCD gain of 300. In total 2000 frames were recorded per cell. JF549-HaloTag ligand was a kind gift from Luke Lavis.
A software tool (plug-in for ImageJ, which is publically available at http://smal.ws/wp/software/sosplugin/), to extract the protein trajectories from microscopy data for this application already existed in our group62,63. This algorithm finds the bright spots that represent single molecules in every time frame, fits a 2D Gaussian-like intensity profile to these spots and then constructs tracks by connecting spots from different time frames together through nearest-neighbor linking.
Single molecule tracking experiments for Spot-On datasets (H2B, CTCF, Sox2 and 3xNLS)
The Spot-On publicly available datasets were acquired from HaloTag-Sox2 knock-in mESCs and HaloTag-3xNLS, H2B-HaloTag-SNAP and C32 HaloTag-CTCF knock-in human U2OS osteosarcoma cells (H2B, CTCF and 3 × NLS). Cells were labelled with PA-JF646 dye and imaged in phenol red-free medium at \(37\) °C and \(5 \% \) CO2 using TIRF microscopy. The datasets chosen for this paper were imaged at a frame rate of \(201\) Hz (time step of \(5\) ms and pixel size of \(0.16\) µm) and are available at https://zenodo.org/record/834781#.XC-YHlVKjX6. Molecules in these datasets were tracked using a custom-written Matlab implementation of the MTT-algorithm64. Additional information on data acquisition and single molecule tracking for the Spot-On datasets (H2B, CTCF, Sox2 and 3xNLS) can be found in42.
References
Chenouard, N. et al. Objective comparison of particle tracking methods. Nature Methods 11, 281–289 (2014).
Saxton, M. J. Single-particle tracking: connecting the dots. Nature Methods 5, 671–672 (2008).
Valentine, M. et al. Colloid surface chemistry critically affects multiple particle tracking measurements of biomaterials. Biophysical Journal 86, 4004–4014 (2004).
Valentine, M. T. et al. Investigating the microenvironments of inhomogeneous soft materials with multiple particle tracking. Physical Review E 64, 061506 (2001).
Mason, T., Ganesan, K., Van Zanten, J., Wirtz, D. & Kuo, S. C. Particle tracking microrheology of complex fluids. Physical Review Letters 79, 3282 (1997).
Josephson, L. L., Furst, E. M. & Galush, W. J. Particle tracking microrheology of protein solutions. Journal of Rheology 60, 531–540 (2016).
Akhmanova, A. & Steinmetz, M. O. Tracking the ends: a dynamic protein network controls the fate of microtubule tips. Nature Reviews Molecular Cell Biology 9, 309–322 (2008).
Berginski, M. E., Vitriol, E. A., Hahn, K. M. & Gomez, S. M. High-resolution quantification of focal adhesion spatiotemporal dynamics in living cells. PloS One 6, e22025 (2011).
Saxton, M. J. & Jacobson, K. Single-particle tracking: applications to membrane dynamics. Annual Review of Biophysics and Biomolecular Structure 26, 373–399 (1997).
Jandt, U. & Zeng, A.-P. In Genomics and Systems Biology of Mammalian Cell Culture 221–249 (Springer, 2011).
Sinha, B. et al. Dynamic organization of chromatin assembly and transcription factories in living cells. Methods in Cell Biology 98, 57–78 (2010).
Reuter, M. et al. BRCA2 diffuses as oligomeric clusters with RAD51 and changes mobility after DNA damage in live cells. The Journal of Cell Biology 207, 599–613 (2014).
Stracy, M. et al. Single-molecule imaging of UvrA and UvrB recruitment to DNA lesions in living Escherichia coli. Nature Communications 7, 12568 (2016).
Brandenburg, B. & Zhuang, X. Virus trafficking–learning from single-virus tracking. Nature Reviews Microbiology 5, 197–208 (2007).
Dorn, J. F., Danuser, G. & Yang, G. Computational processing and analysis of dynamic fluorescence image data. Methods in Cell Biology 85, 497–538 (2008).
Huth, J. et al. Significantly improved precision of cell migration analysis in time-lapse video microscopy through use of a fully automated tracking system. BMC Cell Biology 11, 24 (2010).
Manzo, C. & Garcia-Parajo, M. F. A review of progress in single particle tracking: from methods to biophysical insights. Reports on Progress in Physics 78, 124601 (2015).
Holcman, D., Hoze, N. & Schuss, Z. Analysis and interpretation of superresolution single-particle trajectories. Biophysical Journal 109, 1761–1771 (2015).
Pécot, T., Zengzhen, L., Boulanger, J., Salamero, J. & Kervrann, C. A quantitative approach for analyzing the spatio-temporal distribution of 3D intracellular events in fluorescence microscopy. eLife 7, e32311 (2018).
Das, R., Cairo, C. W. & Coombs, D. A hidden Markov model for single particle tracks quantifies dynamic interactions between LFA-1 and the actin cytoskeleton. PLoS Computational Biology 5, e1000556 (2009).
Monnier, N. et al. Inferring transient particle transport dynamics in live cells. Nature Methods 12, 838–840 (2015).
Persson, F., Lindén, M., Unoson, C. & Elf, J. Extracting intracellular diffusive states and transition rates from single-molecule tracking data. Nature Methods 10, 265–269 (2013).
Schuster‐Böckler, B. & Bateman, A. An introduction to hidden Markov models. Current Protocols in Bioinformatics 18, A.3A.1–A.3A.9 (2007).
Helmuth, J. A., Burckhardt, C. J., Koumoutsakos, P., Greber, U. F. & Sbalzarini, I. F. A novel supervised trajectory segmentation algorithm identifies distinct types of human adenovirus motion in host cells. Journal of Structural Biology 159, 347–358 (2007).
Kinder, M. & Brauer, W. Classification of trajectories-Extracting invariants with a neural network. Neural Networks 6, 1011–1017 (1993).
Michalet, X. Mean square displacement analysis of single-particle trajectories with localization error: Brownian motion in an isotropic medium. Physical Review E 82, 041914 (2010).
Qian, H., Sheetz, M. P. & Elson, E. L. Single particle tracking. Analysis of diffusion and flow in two-dimensional systems. Biophysical Journal 60, 910–921 (1991).
Gal, N., Lechtman-Goldstein, D. & Weihs, D. Particle tracking in living cells: a review of the mean square displacement method and beyond. Rheologica Acta 52, 425–443 (2013).
Weihs, D., Gilad, D., Seon, M. & Cohen, I. Image-based algorithm for analysis of transient trapping in single-particle trajectories. Microfluidics and Nanofluidics 12, 337–344 (2012).
Vega, A. R., Freeman, S. A., Grinstein, S. & Jaqaman, K. Multistep track segmentation and motion classification for transient mobility analysis. Biophysical Journal 114, 1018–1025 (2018).
Sbalzarini, I. F. & Koumoutsakos, P. Feature point tracking and trajectory analysis for video imaging in cell biology. Journal of Structural Biology 151, 182–195 (2005).
Zambrano, H. A., Walther, J. H., Koumoutsakos, P. & Sbalzarini, I. F. Thermophoretic motion of water nanodroplets confined inside carbon nanotubes. Nano Letters 9, 66–71 (2008).
Siebrasse, J. P. et al. Trajectories and single-particle tracking data of intracellular vesicles loaded with either SNAP-Crb3A or SNAP-Crb3B. Data in Brief 7, 1665–1669 (2016).
Weihs, D., Teitell, M. A. & Mason, T. G. Simulations of complex particle transport in heterogeneous active liquids. Microfluidics and Nanofluidics 3, 227–237 (2007).
Ferrari, R., Manfroi, A. & Young, W. Strongly and weakly self-similar diffusion. Physica D: Nonlinear Phenomena 154, 111–137 (2001).
Izeddin, I. et al. Single-molecule tracking in live cells reveals distinct target-search strategies of transcription factors in the nucleus. eLife 3, e02230 (2014).
Holloman, W. K. Unraveling the mechanism of BRCA2 in homologous recombination. Nature Structural & Molecular Biology 18, 748–754 (2011).
Liu, J., Doty, T., Gibson, B. & Heyer, W.-D. Human BRCA2 protein promotes RAD51 filament formation on RPA-covered single-stranded DNA. Nature Structural & Molecular Biology 17, 1260–1262 (2010).
Yuan, S.-S. F. et al. BRCA2 is required for ionizing radiation-induced assembly of Rad51 complex in vivo. Cancer Research 59, 3547–3551 (1999).
Essers, J. et al. Dynamics of relative chromosome position during the cell cycle. Molecular Biology of the Cell 16, 769–775 (2005).
Dion, V. & Gasser, S. M. Chromatin movement in the maintenance of genome stability. Cell 152, 1355–1364 (2013).
Hansen, A. S. et al. Robust model-based analysis of single-particle tracking experiments with Spot-On. eLife 7, e33125 (2018).
Bengio, Y., Simard, P. & Frasconi, P. Learning long-term dependencies with gradient descent is difficult. IEEE Transactions on Neural Networks 5, 157–166 (1994).
Chung, J., Gulcehre, C., Cho, K. & Bengio, Y. Empirical evaluation of gated recurrent neural networks on sequence modeling. arXiv: 1412.3555 [cs.NE] (2014).
Hochreiter, S. & Schmidhuber, J. Long short-term memory. Neural Computation 9, 1735–1780 (1997).
Ewers, H. et al. Single-particle tracking of murine polyoma virus-like particles on live cells and artificial membranes. Proceedings of the National Academy of Sciences 102, 15110–15115 (2005).
Roy, R., Chun, J. & Powell, S. N. BRCA1 and BRCA2: different roles in a common pathway of genome protection. Nature Reviews Cancer 12, 68–78 (2012).
Hansen, A. S., Pustova, I., Cattoglio, C., Tjian, R. & Darzacq, X. CTCF and cohesin regulate chromatin loop stability with distinct dynamics. eLife 6, e25776 (2017).
Kimura, H. & Cook, P. R. Kinetics of core histones in living human cells: little exchange of H3 and H4 and some rapid exchange of H2B. The Journal of Cell Biology 153, 1341–1354 (2001).
Manhart, M., Kion-Crosby, W. & Morozov, A. V. Path statistics, memory, and coarse-graining of continuous-time random walks on networks. The Journal of Chemical Physics 143, 214106 (2015).
Rabiner, L. R. A tutorial on hidden Markov models and selected applications in speech recognition. Proceedings of the IEEE 77, 257–286 (1989).
Gmachowski, L. Fractal model of anomalous diffusion. European Biophysics Journal 44, 613–621 (2015).
Feder, J. Random walks and fractals. in Fractals, 163–183 (Springer, 1988).
Schuster, M. & Paliwal, K. K. Bidirectional recurrent neural networks. IEEE Transactions on Signal Processing 45, 2673–2681 (1997).
Goodfellow, I., Bengio, Y. & Courville, A. Deep Learning. (MIT press, 2016).
Kingma, D. P. & Ba, J. Adam: A method for stochastic optimization. arXiv:1412.6980 [cs.LG] (2014).
Yao, Y., Rosasco, L. & Caponnetto, A. On early stopping in gradient descent learning. Constructive Approximation 26, 289–315 (2007).
Scott, D. W. Multivariate density estimation: theory, practice, and visualization. (John Wiley & Sons, 2015).
Paul, M. W., Zelensky, A. N., Wyman, C. & Kanaar, R. Single-molecule dynamics and localization of DNA repair proteins in cells. Methods in Enzymology 600, 375–406 (2018).
Zelensky, A. N., Schimmel, J., Kool, H., Kanaar, R. & Tijsterman, M. Inactivation of Pol θ and C-NHEJ eliminates off-target integration of exogenous DNA. Nature Communications 8, 66 (2017).
Grimm, J. B. et al. A general method to improve fluorophores for live-cell and single-molecule microscopy. Nature Methods 12, 244–250 (2015).
Abràmoff, M. D., Magalhães, P. J. & Ram, S. J. Image processing with ImageJ. Biophotonics International 11, 36–42 (2004).
Meijering, E., Dzyubachyk, O. & Smal, I. Methods for cell and particle tracking. Methods in Enzymology 504, 183–200 (2012).
Sergé, A., Bertaux, N., Rigneault, H. & Marguet, D. Dynamic multiple-target tracing to probe spatiotemporal cartography of cell membranes. Nature Methods 5, 687–694 (2008).
Acknowledgements
The authors are grateful to the authors of25 for providing the Spot-On datasets used in part of the presented experiments. They also acknowledge financial support from Erasmus University Medical Center (I.S. and E.M.) and NWO (ECHO.15.CL1.069) and Oncode Institute Erasmus MC (C.W. and M.P.).
Author information
Authors and Affiliations
Contributions
All authors conceived of and planned the project. M.A. and I.S. designed the presented methods, implemented the software, performed the experiments, and analyzed the results. M.P. acquired and prepared the datasets used in the experiments and helped with the analysis. M.A. drafted the manuscript. C.W. and E.M. oversaw the execution of the project and contributed to the writing of the manuscript. We thank the optical imaging centre (OIC) at Erasmus MC for support with microscopes.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Arts, M., Smal, I., Paul, M.W. et al. Particle Mobility Analysis Using Deep Learning and the Moment Scaling Spectrum. Sci Rep 9, 17160 (2019). https://doi.org/10.1038/s41598-019-53663-8
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-019-53663-8
This article is cited by
-
Objective comparison of methods to decode anomalous diffusion
Nature Communications (2021)
-
Accelerating the experimental responses on cell behaviors: a long-term prediction of cell trajectories using Social Generative Adversarial Network
Scientific Reports (2020)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.