Abstract
MicroRNAs (miRNAs) are small non-coding RNAs that function in RNA silencing and post-transcriptional regulation of gene expression in most organisms. The water flea, Daphnia magna is a key model to study phenotypic, physiological and genomic responses to environmental cues and miRNAs can potentially mediate these responses. By using deep sequencing, genome mapping and manual curations, we have characterised the miRNAome of D. magna. We identified 66 conserved miRNAs and 13 novel miRNAs; all of these were found in the three studied life stages of D. magna (juveniles, subadults, adults), but with variation in expression levels between stages. Forty-one of the miRNAs were clustered into 13 genome clusters also present in the D. pulex genome. Most miRNAs contained sequence variants (isomiRs). The highest expressed isomiRs were 3′ template variants with one nucleotide deletion or 3′ non-template variants with addition of A or U at the 3′ end. We also identified offset RNAs (moRs) and loop RNAs (loRs). Our work extends the base for further work on all species (miRNA, isomiRs, moRNAs, loRNAs) of the miRNAome of Daphnia as biomarkers in response to chemical substances and environment cues, and underline age dependency.
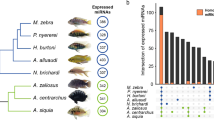
Similar content being viewed by others
Introduction
Non-coding, regulatory small RNAs constitute one of the epigenetic mechanisms of gene regulation. These regulatory RNA molecules are evolutionary conserved across phyla, and elements are found in a vast majority of organisms from bacteria to animals and even viruses1,2,3,4. The main types of small RNAs in animals are microRNA (miRNA), piwi-interacting RNAs (piRNAs) and small interfering RNAs (siRNAs)5. Of these, miRNAs have been the most studied group and are found to regulate a massive number of biological processes by interacting with mRNAs and non-coding RNAs. The canonical miRNA genes are transcribed mainly by RNA polymerase II into a primary transcript called pri-miRNA. Pri-miRNAs are processed by the DROSHA/Pasha (Pasha is known as DGCR8 in mammals) microprocessor complex to precursor miRNAs (pre-miRNAs) (~70 nucleotides) which are then transported to cytoplasm by Exportin 5/Ran-GTP and further processed by Dicer to ~22 bp duplex sequence with 2 nucleotides overhang at the 3′ end. Usually either the 5′- or the 3′-end (5p or 3p arm) of the pre-miRNA is chosen as mature guide miRNA when associated with Argonaute (Ago) proteins in a functional RNA-induced silencing complex (RISC) that is the regulatory unit. The other strand (passenger strand or miRNA*) may be discarded5,6. However, arm switching may occur where the dominant guide miRNA shifts from one arm to the other of the pre-miRNA across different tissues, developmental stages and between animal species7,8. The pri-miRNA and pre-miRNA transcripts may undergo alternative cleavage by Drosha and Dicer and thereby produce sequence variants with different start and end positions compared to the mature miRNAs (3′ - and 5′ template variants). Sequence variants may also be produced by nucleotide additions (3′- and 5′-end) and single nucleotide substitutions (SNPs) (non-template variants). All these variants of mature miRNA are called isomiRs and seem to be functional and have regulatory properties8,9,10. The nucleotides of position 2–7 from the 5′ end of the mature guide miRNA form the seed sequence that recognize and bind to the target sequences. The binding may also involve nucleotide eight and to a lesser extent nucleotide at position 13–1611. Adding or deleting nucleotides at the 5′ end and SNPs in the seed sequence of the mature miRNA will change the seed sequence and may give different targeting properties12. Previous deep sequencing projects have also revealed reads from miRNA offset RNAs (called moRs) of the pri-miRNA transcript adjacent to the 5′ end of mature miRNA of the 5p arm and to the 3′ end of mature miRNAs of the 3p arm13,14. Reads have also been identified from the loop of pre-miRNAs15,16 and these loop-derived sequences are named loRs17. The functions of moRs and loRs are still unclear.
MiRNAs are prevalent in insects18 and are involved in regulation of several biological processes19, as for example insect physiology20 like metamorphosis21 or immune response22 and even targeting viruses23.
Daphnia magna and D. pulex are the aquatic key invertebrate model organisms in ecotoxicology, ecology and evolution24. Responses to different triggers and stressors, including environmental pollutants, pesticides and GMOs, have been studied on its life-history and physiology as well as on a genetic and epigenetic levels25,26,27,28,29. Recently, some attention has been given to describe miRNAs in the genus Daphnia. In D. pulex, 229 microRNAs were identified in silico and verified by qPCR30 and microarrays31 while in D. magna 205 putative mature miRNAs were identified based on small RNA sequencing and mapping against databases32. Eighteen of these were confirmed by secondary structure analysis of miRNA hairpins. Recently, Hearn et al.33 reported 33 miRNAs in D. magna by deep sequencing and genome mapping. The authors also investigated whether transgenerational effect could be mediated via miRNAs and showed that miRNAs played a role in maternal provisioning rather than longer-term transgenerational responses. Indeed, the expression profiles of miRNA are a promising step to further examine organism responses to stressors, in particular on specific genes, genetic pathways or networks34. Thus, miRNAs may extend their use as important biomarkers31,33,35. However, it has recently been reported that annotating miRNAs without mapping sequenced reads to the same species genome may largely overestimate the presence of miRNAs36. For instance, Yalle et al.36 found that insects contained 65 conserved families with very low variation.
In this study, we characterize the miRNAome from three different life stages (juveniles, subadults, adults) in D. magna by a four step process: 1) deep sequencing of small RNAs, 2) alignment of the trimmed, clean reads to the D. magna genome, 3) identification of putative miRNA by aligning the mapped reads to miRBase and 4) manual inspection of sequence and reads in the genome at the position of the putative miRNA. With this approach we aimed at improving the coverage and verification of authentic presence of this important class of gene regulating nucleic acids, building on previous miRNA studies in D. magna32,33. We identify conserved and novel miRNAs, miRNA clusters, isomiRs, offset RNAs (moRs) and loop RNAs (loRs), and compare these data between juveniles, subadults and adults. Our work presents novel miRNA discoveries and characterisation on age-specific miRNAs in Daphnia which can be used for further verification and investigation.
Results and Discussion
Analyses of the small RNA sequencing data
The raw reads from deep sequencing of three small RNA libraries, one each from the three life stages of pooled D. magna juveniles, subadults and adults, were processed into trimmed clean reads of 17–30 nucleotides (Table 1).
The trimmed clean reads were mapped to the genome of D. magna and about 96% of the clean reads of 17–30 nucleotides from all three libraries aligned to the genome sequence (Table 1).
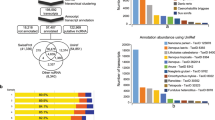
As a first step to identify miRNAs, reads mapping to the D. magna genome were aligned to miRNAs in miRBase release 2137 and dpu-miRNAs from D. pulex30. The percentage of mapped reads that aligned to miRNA sequences was similar for juveniles (35.4%) and adults (36.7%) but only 15.2% for subadults (Table 1). The mapped read counts were normalized and distributed on read lengths (Fig. 1). For all three life-stages, reads of 22 nucleotides were most abundant, but the number of reads of 22 nucleotides from subadults was less than one third of the reads of 22 nucleotides from juveniles and adults. The next highest counts were found for reads of 18 nucleotides (juveniles, adults) and 19 nucleotides (subadults). Overall, juveniles and adults had more read counts of lengths below 24 nucleotides, while subadults had more reads with lengths between 25–30 nucleotides (Fig. 1).
Identification of conserved miRNAs in D. magna
The reads that mapped to the D. magna genome aligned to several hundred putative miRNA species in miR Base and dpu-miRNAs. To verify that the putative miRNAs identified in miRBase and dpu-miRNAs were real D. magna miRNAs, we mapped the mature miRNA sequence of each putative miRNAs back to the D. magna genome. The positions in the genome, where each of these mature miRNA sequences aligned, were then manually scanned for sequence and mapped reads in the same regions. Through these analyses we identified the presence of reads that could be annotated to 66 conserved microRNAs in miRBase and dpu-miRNAs, representing 51 miRNA-families (Table 2). This number of miRNA families is similar to a recent re-evaluation of the number of miRNAs in insects, which identified 65 conserved miRNA families36. Hearn et al.33 identified 72 precursor miRNAs in D. magna of which all the 33 annotated and nine non-annotated miRNAs were present among our 66 miRNAs. Of the 205 mature sequences representing 188 miRNAs and the 18 miRNA hairpin sequences identified by Ünlü et al.32 we found 53 miRNA and 17 hairpin sequences in our data. (Fig. 2, Supplementary2 Table S1). Overall, about 52% (34 of 66) of the miRNAs reported by Hearn et al.33, Ünlü et al.32 and us overlap (Fig. 2, see Supplementary2 Table S1 for details on the overlap of miRNAs in previous studies32,33 and this study). In addition 37 miRNAs of the 229 found in D. pulex30,31 were identified in D. magna in this work (Supplementary2 Table S1).
The number of normalized reads aligned to the miRNA hairpins (pre-miRNA + adjacent sequences) ranged from a few reads (miR-153) to more than 3 million (miR-1) (Table 2). When comparing the miRNA expressions in the three life stages, the adults had highest expression of 28 miRNA hairpins, followed by juveniles with highest expression of 24. Subadults had highest expression of only 14 miRNA hairpins (Table 2).
Notably, the sequenced reads from the 66 miRNA hairpins were from whole animals, thus lacking further details on how different body parts, organs, etc., differ in expression. However, the three age groups expressed all the same 66 species of miRNA, but with some proportional differences between the three different life stages (Fig. 3, Table 2). The heat map (Fig. 3) also depicts some larger groups of miRNAs where the expression pattern of miRNAs in each group was similar across the three life stages. Cluster analysis of the total miRNA read counts of each life stages indicated that juveniles and subadults were the two most similar whereas the adults deviated from these two (Fig. 3).
Heat map and clustering of conserved microRNA hairpins from juvenile, subadult and adult D. magna, based on normalized read counts. Clustering of the three life stages is shown at the top of the heat map, while clustering of the miRNAs is shown to the left. The clustering was performed by using Pearson distance measurement and average linkage methods.
Most of the conserved pre-miRNAs were between 57–100 nucleotides in length. However, four miRNAs differed with longer lengths: pre-let-7 (168 nucleotides), pre-miR-263a (231 nucleotides), pre-miR375 (241 nucleotides) and pre-miR-750 (152 nucleotides) (Fig. 4). Using the mature miRNA sequences from D. magna to map the D. pulex genome, we found similar length for dpu-pre-let-7 (179 nucleotides), dpu-pre-miR-375 (282 nucleotides) and dpu-pre-mir-750 (153 nucleotides), while dpu-pre-mir-263a was shorter (92 nucleotides). An interesting observation is that the D. magna pre-mir-750 contains a putative novel miRNA inside the mature miR-750-5p and -3p (see below). In the pre-miR-750 of D. pulex a putative novel pre-miR hairpin with a rather strong folding free energy37 of ∆G < −20 kcal/mol could be detected, but with a different sequence than in D. magna.
Conserved mature miRNAs
Mature guiding strands of miRNAs can be expressed from the 5p arm, 3p arm or both in the pre-miRNAs by changing from one to the other arm in different tissue in the same organism7,38. Usually the pre-miRNA arm containing the mature miRNA sequence with the highest read counts is treated as the mature guide miRNA, leaving the sequence of the other strand as passenger (or star) miRNA5,39. Sometimes miRNA sequences form both pre-miRNA arms are expressed as mature guide miRNAs and may co-mature due to different expression in different cells (so called arm switching)38. In this study, with the primary aim to discover new, and strengthen the description and verification of previously described miRNA in D. magna, we did a single sequencing of small RNAs from each of three life stages (relatively large, pooled batches of whole-body juveniles, subadults and adults).
Based on the counts of read sequences and folding of pre-miRNAs, we could identify the mature miRNA sequences from both arms of the vast majority of the conserved miRNA hairpins (Supplementary1 Table S2, Supplementary1 Fig. S1). It should be noted that for the identical mature miRNAs, the reads were equally divided between miR-2a-1-3p and miR-2a-2-3p, and between miR-87-1-3p and miR-87-2-3p due to the impossibility of knowing which miRNA the reads mapped to.
The majority of the mature guide miRNA sequences in D. magna were expressed from the 3p arm (44 miRNA) of the pre-miRNA (Supplementary1 Table S2). Almost all of the mature miRNA sequences had more than 50% of the total read counts mapped to its arm of the miRNA hairpin (Supplementary1 Table S3), but with variation from 0% (miR-279b-5p) to 94.6% (miR-2944-5p) (Supplementary2 Table S1). In addition to the read counts of each mature miRNA, additional read counts to each arm were mainly due to miRNA isoforms (isomiRs) (Supplementary2 Table S1. See also below). Although for most of the miRNAs the mature guide miRNA had much more reads than the passenger miRNA, several of the passengers also exhibited rather high read counts (Supplementary1 Table S2). This could indicate that some of these passenger miRNAs may be active regulatory species. However, for a few miRNAs the read counts of mature miRNA sequences from both 5p and 3p arms were similar. To indicate co-expression of mature miRNA from both arms, we used the ratio of <2 between the read counts of the most expressed arm divided by the number of reads from the less expressed arm36. We found that some of the mature miRNAs displayed co-expression of both arms in different combinations of juveniles, subadults and adults as shown in Table 3. Furthermore, some mature miRNAs had undergone arm-switching of the most expressed miRNA from one arm to the other in the different life stages of D magna (Table 3, Supplementary1 Table S4). The co-expression and arm-switching for miR-2a-1 and miR-2a-2 are highly uncertain since the expression levels of mature miRNAs were only estimated to be equal (see above). We also found that some miRNAs displayed shift in expression between the mature miRNA and its isomiR in the three life stages (see below).
The mature guide miRNA sequences of D. magna were identical or similar to most of the conserved mature guide miRNAs in miR Base, release 2137. The mature guide miRNAs (highest expression) are shown in Supplementary1 Table S2, and their putative differential expression between the three developmental stages is displayed in the heat map in Supplementary1 Fig. S2. The clustering of the mature miRNA profiles (Supplementary1 Fig. S2) indicate a closer relationship between juveniles and subadults, as was seen for the miRNA hairpin profiles (Fig. 3). Again, we observed some groups of miRNAs where the pattern of expression in each group was similar across juveniles, subadults and adults. It is interesting to observe that miR-1, let-7, miR-184, miR-275 and miR-276 which are among the highest expressed miRNAs in D. magna, are also among the highest expressed miRNAs in the closely related, white shrimp Litopenaeus vannamei40. In contrast, only miR-1 and miR-276 (and then miR-276-5p and not miR-276-3p which is the highest in our result) were among the five highest expressed D. magna miRNAs analysed by Hearn et al.33. Many of the conserved miRNAs found in D. magna have been revealed to have diverse and important biological functions in insects18,19,21,36. For instance, let-7 is the most known miRNA that regulates developmental timing in animals. Mir-1, predominantly expressed in muscle tissue, in insects, is responsible for maintaining muscles integrity and mir-184 is involved in regulation of oogenesis19.
Identification of novel miRNAs
Many miRNAs are clustered in the genome due to the formation of new miRNAs close to pre-existing ones41,42. In mammalian species, most of the miRNA genes (about 2/3) are clustered within 50 kb43. Based on this knowledge we first performed a query for putative novel miRNAs in D. magna by manual scanning of scaffolds and contigs that contained the conserved miRNAs. Using the criteria for annotation of novel miRNA37 described in the Methods section we identified 13 putative novel miRNAs (Supplementary1 Table S5) of which six were identical with non-annotated precursor miRNAs in D.magna reported by Hearn et al.33 (Supplementary2 Table S1). Five (miR-nov-1, miR-nov-3, miR-nov-4, miR-nov-10 and miR-nov-12) fulfilled the criteria of novel miRNAs in D. magna (Table 4). The secondary structures of these five miRNA hairpins with their folding free energies (∆G) are shown in Fig. 5.
In addition, eight other pre-miRNAs fulfilled several of the criteria for the annotation as novel miRNAs37. However, for most of them mature miRNA reads were only detected from the putative miRNA sequence on one of the hairpin arms (Supplementary1 Table S5, Supplementary1 Fig. S3).
Comparisons of the mature sequences of miR-nov-7-3p, miR-nov-8-3p and miR-nov-9-3p shows that the seed sequence (nucleotide 2-7) is identical in all three (Supplementary1 Table S5). Thus, these miRNAs are of the same family. Several of the novel miRNAs showed a lower expression level in the juveniles compared to subadults and adults, i.e. miR-nov-3-3p, mR-nov-4-3p, miR-nov-12-3p, miR-nov-2-3p, miR-nov-7-3p, miR-nov-9-3p and miR-nov-11-5p) (Table 4, Supplementary1 Table S5). We also observed a tendency of increased expression of some novel miRNAs from juvenile to subadult and to adult stages (e.g. miR-nov-3-3p, miR-nov-4-3p, miR-nov-2-3p). These differently expressed novel miRNAs in different age-classes indicate important roles in the ontogenetic development of D. magna, but further studies need to investigate this specifically.
Mapping the D. pulex genome with the mature sequence of the novel D. magna miRNAs showed that several of these miRNA sequences were also present in D. pulex (Supplementary1 Table S6).
Ikeda et al.44 reported that 21 of the novel miRNAs in Triops cacriformis (tadpole shrimp) showed more than 80% sequence similarities with the genomic sequence of these miRNAs in D. pulex. In our data we found only four of the T. cancriformis novel miRNAs as expressed miRNAs in D. magna. It is interesting that the three novel D. magna miRNAs: miR-nov-7-3p, miR-nov-8-3p and miR-nov-9-3p have sequences identical to tca-miR-n504-3p, tca-mir-n512-3p and tca-miR-n503-3p from T. cancriformis. Moreover, miR-nov-13 and tca-miR-3477 have identical seed sequences, but with five mismatches outside the seed region.
Genomic clustering of miRNAs
MiRNAs are often found as clusters in metazoan genomes. In flies and worms about 30% of all miRNAs are clustered as two or more miRNAs within 10 kb, while ~45% of conserved miRNAs are found in clusters in these species (and mammals)42. Moreover, many of these miRNAs clustered within 10 kb, and especially when clustered within 1 kb, are co-expressed45,46,47,48. In D. melanogaster, miRNAs separated by less than 1 kb seem to be highly co-expressed in different tissues49.
Identification of clustered miRNAs (both conserved and novel) within 10 kb of D. magna genome and comparison with clustering of the same miRNAs in the D. pulex genome, is shown in Supplementary1 Table S7. Both species contain the same 13 clusters with the same order of the miRNAs in each cluster. Of the 66 conserved miRNAs in D. magna, 31 (47%) are grouped into 11 clusters. Furthermore, miR-nov-7, miR-8, and miR-nov-9, which are clustered in D. magna, have their identical miRNAs tca-miR-n504, tca-miR-n512 and tca-miR-504 clustered in the same orientation in T. cancriformis44. We also note that miR-283, miR-nov-13, miR-12 build a cluster in D. pulex, but these miRNAs are on different scaffolds in D. magna. However, scaffold sc1551 (miRNA-283) may be connected to scaffold sc2703 (miR-nov-13, miR-12) since the Thyroid receptor-interacting protein (http://arthropods.eugenes.org:8091/gbrowse/cgi-bin/gbrowse/daphnia_magna7/) is encoded across these two scaffolds. Thus, these three miRNAs seem to be within a 10 kb distance also in D. magna. Another support for this cluster is that similar clusters are documented in other organisms. For example, the tca-miR-12/miR-3477 (same seed as miR-nov-13)/miR-283 cluster is within 1000 bp in T. cancriformis44, the miR12/miR-1889/miR-283 in Anopheles gambiae50, and similar clusters in several other arthropods (see miRBase).
Several miRNA clusters are conserved among animals42. In mammals, clustered miRNAs may function as a unit in regulation of biological processes51. Many of the D. magna (and D. pulex) clusters are found as similar or identical clusters in other arthropods (miRBase release 2137). Particularly many of the D. magna clusters are conserved in T. cancriformis (10 clusters)44, Tribolium castaneum (9 clusters)52 and Apis mellifera (9 clusters) (miRBase release 2137). Among the D. magna clusters with the highest expressed miRNAs are the miR-100/let-7/miR-125 cluster which have been found conserved in all metazoans53 and is involved in regulating the larvae-to-adult transition as well as neural development54. The miR-2/miR-13/miR-71 cluster is conserved in many invertebrates55 and was recently reported to be involved in resistance to the insecticide deltamethrin in mosquitos56.
miRNA isoforms (isomiRs) in D. magna
Deep sequencing of small RNAs has shown that most miRNAs are expressed both as mature sequences and as variants (isomiRs)8,9,57. We have analysed the read sequences that mapped with more than 10 read counts to miRNA hairpins. The read sequences have been divided into seven groups: i) mature miRNAs; ii) 3′ template isomiRs containing reads with the mature 5′ end and deletions or additions at the 3′ end complementary to the genome sequence; iii) 3′ non-template isomiRs containing reads with mature 5′ end and 3′ additions not complementary to the genome sequence; iv) isomiR reads not part of groups ii) or iii) and with the mature 5′ end but with single nucleotide polymorphisms (SNPs) not altering the mature seed sequence; v) 5′ template isomiRs with reads containing the mature 3′ end and deletions or additions at the 5′ end complementary to the genome sequence; vi) 5′ non-template isomiRs containing reads with intact mature 3′ end and additions at the 5′ end not complementary to the genome sequence; and vii) isomiR reads not part of group v) or vi) and with SNPs altering the seed sequence10. Although the majority of D. magna read counts were mature miRNA sequences (conserved and novel), expressed in all the three developmental stages (juveniles, subadults, adults), all miRNA hairpin arms (5p or 3p) mapped with more than 10 reads, contained isomiRs (Supplementary2 Table S1). For each miRNA calculation of the ratio of read counts for each of the seven isomiR groups, divided on the total read count for the miRNA arm, showed large variation both for each miRNA and each isomiR group (Supplementary2 Table S1). Moreover, determining the mean ratio for each isomiR group (across all miRNAs) revealed large variation also in the sizes of each isomiR group. Generally, the highest expressed isomiRs were the 3′-templated variants (mean 20.7% of the total read counts for 5p+3p arms). The mean level of expression of the other variants was as following: 3′ non-template isomiRs, 3.7%; 5′ template isomiRs, 4.2%; isomiR reads with SNPs changing seed sequence, 3.5%; isomiR reads with SNPs not altering seed, 2.2% and 5′ non-template isomiRs, 0.08%. (Fig. 6, Supplementary1 Table S8). Supplementary1 Figure S4a & b) shows mean level of expression for each arm. Although some of these variants displaying one or a few reads may be sequencing errors, most isomiR sequences have several read counts and are present in all three independent sequencing samples (juvenile, subadult, adult stages) supporting that they are real isomiR sequences.
Interestingly, the proportional relationship between the different isomiR variants in D. magna is similar to that reported for the feline (cat family) miRNAome58, suggesting that the roles and functions of mature miRNAs and their isomiRs may be conserved by evolution, even across distantly related organisms. A recent review suggests that isomiRs may function as an additional repression tool beside the mature miRNA59. For some miRNAs we found 3′ template variants that had a similar level of expression as their mature miRNAs: miR-1175-5p (two variants), miR-12-5p (mainly one variant), miR-279b-3p (juveniles, mainly one variant), miR-71-5p (mainly three variants), miR-993-5p (mainly three variants), miR-998-3p (juveniles and subadults, one variant; adults, two variants), miR-iab-8-5p (juveniles and adults, two variants; subadults, three variants), miR-9b-5p (two variants), miR-31-5p (one variant), miR-34-5p (mainly three variants), miR-210-5p (one variant), miR-285-3p (mainly one variant), miR-2944-3p (one variant), miR-317-3p (mainly two variants), miR-92b-3p (one variant), miR-981-3p (one variant) and 5′ template variants: miR-210-3p, miR-305-3p, miR-9a-3p (summarised in Table 5). In some cases the level of expression between mature miRNA and one of its 3′ template or 5′ template isomiR variants were noticeably different between the three developmental stages, e.g miR-12-5p, miR-210, miR-285-3p, miR-31-5p miR-998-3p, miR-305-3p, miR-9a-3p (Table 5). MiR-71-5p was special in this regard since the 3′ template variants had from 55% to 120% more read counts than the mature sequence (Supplementary2 Table S1, Table 5). However, the 3′ template isomiRs reads are here mainly divided on three different sequences with one to three deletions at the end (Table 5).
Although 3′ template isomiR variants usually are the most common, very little is known about their functions. In a recent report Yu et al.60 found that 3′ template isomiRs of miR-222 could play functional roles in human cell lines. We found that the highest expressed 3′ template variants were usually those with one deletion at the end (Supplementary 2 Table S1). However, some exceptions were also found where the 3′ end additions were non-template A or U (see below).
Non-templated 3′ variants generated by post-transcriptional uridylation or adenylation have been found to change stability, affect expression and target selection43,61,62. In our data the highest read counts were 3′ non-template variants with A or U additions at the end and some of these had expression values similar to 3′ template variants, e.g. isomiRs from let-7-5p, miR-1-3p, miR-100-5p, miR-275-3p, miR-8-3p, miR-965-3p. In a few cases the most highly expressed isomiR sequence were a 3′ non-templated one, e.g. let-7-5p, miR-100-5p, miR-13-3p, miR-184-3p, miR-277-3p, miR-34-3p, miR-8-3p (Supplementary2 Table S1).
5′ variants of isomiRs had a mean expression of 4.2% for the 5′ template and only 0.08% for the 5′ non-template variants, respectively (Fig. 6, Supplementary1 Table S8). IsomiRs of the 5′ template type were the highest expressed isomiRs of miR-1175-3p, miR-125-3p, miR-210-3p, miR-252a-3p, miR-275-5p, miR-279c-5p, miR-305-3p, miR-71-3p and miR-9a-3p. However, it is interesting to notice that miR-9a-3p, miR-210-3p, and miR-305-3p displayed 5′ template read counts at a similar level as the potential mature miRNA, and that the most highly expressed sequence of mature miRNA and its 5′ template isomiRs may differ between the three developmental stages (Table 5, Supplementary2 Table S1). Especially the 5′ template variant of miR-1175-3p showed a different pattern of expression (percentage ratio of total read counts) which was low in juveniles (12%) and adults (5%) but high in subadults (41%) (Suplementary2 Table S9a). The same sequence variants of the 5′ template of miR-1175-3p are present in all three life stages although the read counts of each sequence vary between the stages (Suplementary2 Table S9a). Of the isomiRs that displayed seed shifting, the 5′ template isomiRs were generally higher expressed than the 5′ non-template sequences. Since 5′ variants shift the seed sequence they have been found to regulate other RNA sequences than the mature miRNA12,63. However, recently it was reported that 5′ variants also may be able to regulate the same mRNA as the mature miRNA64.
We observed a mean expression of 3.5% of reads with SNPs that alter the seed sequence and a mean expression of 2.2% of reads with SNPs that did not affect the seed sequence (Fig. 6, Supplementary 1 Table S8). Although some of these SNP variants may be sequencing errors, we find that several variants have the same mismatch in identical sequences with several read counts across the three independent sequencing samples (juvenile, subadult, adult) supporting that these are real isomiRs (Suplementary2 Table S1). The SNP variants with altered seed sequences were the highest expressed isomiRs of miR-190-3p, miR-279c-5p, and miR-750-5p (Suplementary2 Table S9b). Editing of miRNA may result in SNPs both altering the seed sequence and the sequence outside the seed region65,66.
Taken together our sequencing results show expression of a variety of isomiRs of miRNAs in D. magna. The observed proportions for all three life stages shows that the 3′ variants and variants with SNPs that do not alter the seed, have a mean percentage of 26.5% while the 5′ variants and variants with SNPs changing the seed have a mean percentage of 7.8% (Supplementary1 Table S8). The 3′ variants show the highest expression for most of the miRNAs while the 5′ variants are expressed to a lower degree. IsomiRs have been suggested as promising, additional biomarkers in diagnostic assays based on mature miRNA for studying illness such as for example cancers67,68. In D. magna isomiRs may be developed into important biomarkers in addition to the role of mature miRNAs for analysing biological changes.
Offset- and loop miRNAs
Analysis of reads adjacent and in the loop of pre-miRNAs revealed one or more reads which were recognised as moRs and/or loRs (Supplementary2 Table S10). Nine miRNAs display moRs and/or loRS with more than 10 reads in at least one the three life stages of D. magna (Table 6). MiR-7 is interesting since it contains 5′ moR reads that add up to relatively high percentages of the mature miR-7 reads (juveniles: 27%, subadults: 14% and adults: 96%) (Supplementary2 Table S10). As reported previously17, we also found that moRs at the 5′ site (5p) of mature miRNAs are more frequent and with higher read counts than on the 3′ site (3p) (Supplementary2 Table S10). In the three investigated life stages the total expressions of loRs were at similar level as for 5′ moRs. However, the expression level of the sum of 5′moR + 3′moR was somewhat higher than the expression of loRs (Supplementary2 Table S10). Both moRs and loRs may play functional roles15,69.
Conclusions
Daphnia magna is an important model organism for ecotoxicology, ecology and evolution. Its remarkable ability to respond to changing environmental conditions, i.e. its plasticity, could potentially be mediated via age-specific miRNA expression. As such, miRNAs could constitute an important epigenetic mechanism in clonal organisms. Age or life-stage in an animal represent an important organismal context and needs to be considered for future experimental designs whenever miRNA is of interest. This is also in agreement with previous work33.
The main purpose of this study was to characterise the miRNAome and thereby identify and validate the number of miRNAs in D. magna by deep sequencing, mapping of reads to its genome combined with manual curation of putative miRNAs. By this procedure we identified and further characterized 66 conserved and 13 novel miRNAs from three life stages of D. magna (the pre-miRNA sequences are shown in Supplementary3 Table 11). Of the 66 miRNAs we found, 42 were already reported by Hearn et al.33 and 17 miRNA hairpins and 53 miRNA were already identified by Ünlü et al.31 All the miRNAs we found were expressed in all the three life stages and displayed similar miRNA profiles. However, clustering of the three miRNA profiles indicated a closer relationship between juveniles and subadults. The mature guide miRNAs were generally expressed with the highest read counts (>50% of total read counts). The additional reads were sequence variants (isomiRs) which were expressed by almost all miRNAs. We identified isomiRs which were 3′-, 5′- and SNP- variants. Several miRNAs contained 3′ template- or 5′ template variants of isomiRs that had a similar expression levels as their mature miRNAs. Furthermore, we found that miRNAs with the highest expression levels could shift between the mature miRNA and one of its 3′ template -or 5′ template isomiR variants over the three life stages. Several miRNAs displayed reads adjacent (offset RNAs (moRs)) and in the loop (loRs) of the pre-miRNAs.
Our work extends the background for further work on using both mature miRNAs and their sequence variants (isomiRs) as biomarkers of stress in Daphnia and should also help to exclude age-related confounding in follow-up studies. For future studies we recommend: i) independent validation of miRNAs in other laboratories, where at least 20 million reads is necessary (since we see some differences in our data to previous reports), ii) linked analyses of mRNA and miRNA expression to reveal more of the functional roles of miRNA in general, iii) as complete and detailed as possible reporting of the atlas of miRNA expression in different organs and sub-systems in the organism, also at different stages of development, and iv) test individual miRNAs for their ability and sensitivity to function as biomarkers of different types.
Materials and Methods
Experimental setup
All individuals of D. magna used in the experiments were from the same clonal population. Animals were kept in M7 medium70, fed Desmodesmus subspicatus, ad libitum and never crowded. The juveniles (n = 120) that were sequenced were born within 24 h from the third clutch of a set of n = 25 mothers. The subadults (n = 40) were also born within 24 hours from a set of n = 25 mothers and reared for 6 days. We checked for visible eggs in the brood chamber of each individual and confirmed that eggs were not present. For the adults, we selected animals with visible eggs at day 9 (n = 30). During the course of the experiment the subadult and adult groups were fed Desmodesmus subspicatus green algae with 0.15 mg Carbon per animal per day.
Every third day, we transferred each animal to a new glass with fresh M7 medium. Temperature was held constant at 22 + /− 1.5 degrees C. Light regime was 16 hours of light and 8 hours of darkness. Throughout the experiment, the average pH of the medium was 7.8, oxygen saturation always >95% and average conductivity 575 µs cm−1.
Total RNA extraction
D. magna animals were filtered from the growth cultures. For each life stage (juveniles, subadults and adults), 100 mg of animals were added to 1000 µl of Trizol (Invitrogen) and homogenized by bead milling with ceramic beads (Roche Applied Science, Basel, Switzerland) in a Precellys homogenizer (Bertin instruments). The Trizol extraction of total RNA was according to the manufacturer’s protocol with modifications involving prolonged precipitation and centrifugation steps to ensure recovery of the small RNA fraction. Purified total RNAs were dissolved in nuclease free water and stored at −70 °C. RNA quantity was determined with a NanoDrop spectrophotometer (NanoDrop Technologies). The quality of RNA was analysed with an Agilent 2100 Bioanalyzer using an Agilent RNA 600 Nano kit (Agilent Technologies).
Construction of small RNA library and Illumina sequencing
Total RNAs were shipped on dry ice to the Norwegian Sequencing Center (NSC), Ullevål University Hospital, Oslo (http://www.sequencing.uio.no/) for small RNA library construction and Illumina sequencing. The small RNA libraries were prepared as described in the “Small RNA Sample Preparation Guide (Illumina, Cat # FC 930-1004, Part # 1004239 Rev B, August 2009). After PCR amplification gel-purified small cDNA libraries (fragment size about 100 bp) were sequenced on an Illumina Genome Analyzer.
Analyses of sequenced reads
Three datasets containing de-multiplexed, clean reads (reads with adaptor clipped off and removed non-clipped reads, adaptor-only reads, N reads, reads < 10 nucleotides) were received from NSC as fastaq files. These clean reads were analysed on a CLC Genomics Workbench (Qiagen) using the following pipeline: 1) The clean reads were trimmed to 17–30 nucleotides (“Trimmed Clean reads”). 2) The trimmed reads were mapped to the D. magna genome (dmagna-v2.4-20100422-assembly (http://wfleabase.org/)71 using “Map Reads to Reference” module and following mapping parameters: mismatch costs 2, insertion cost and deletion cost each 3, length fraction 0.85, similarity fraction 0.8, map randomly. Reads mapped to D. magna genome were then aligned to miRNAs in miRBase release 21 (http://www.mirbase.org)37 and in-silico determined dpu-miRNAs30 using the various “Mature length variants” and “Alignment settings” parameters. The DNA sequence of potential D. magna miRNAs recognised from miRBase and dpu-miRNAs alignments (matches) were mapped to the D. magna genome and manually inspection of sequence and reads in the genome at the position of the putative miRNA was used to identify and annotate miRNAs. Pre-miRNA sequences were manually identified and secondary structures and their folding free energy (ΔG) of miRNA hairpins were determined by using the “Predict Secondary Structure RNA” parameter in the CLC software.
For examination of the presence of D. magna miRNAs in D. pulex, miRNA sequences were mapped to the D. pulex genome (http://www.ftp://ftp.ensemblgenomes.org/pub/release-34/metazoa/fasta/daphnia_pulex/dna/)72
Annotating novel D. magna miRNAs was performed mainly by the criteria described previously17,37,73 except that we also included putative novel miRNA with reads aligning only to one of the pre-miRNA arms.
MoRNAs were defined as reads that were immediately adjacent either to the 5′ end of mature miRNA of the 5′ arm and/or adjacent to the 3′ end of the mature miRNA of the 3′ arm14. LoRNAs were annotated as reads between mature miRNAs of the 5′ and 3′ arm15. For both moR and loR we also included reads that started outside the mature miRNA and ended inside the miRNA and which could not be regarded as isomiRs.
We used DESeq2 package (R package) to normalise the raw read counts as described previously74. Shortly, as input the DESeq2 package expects raw count data obtained from sequencing of miRNA. The DESeq2 model internally corrects for differences in library size (estimateSizeFactor function) and delivers normalized values. Normalized counts were accessed by “counts” function on recalculated counts object. We did not perform statistics to look for significant differences in miRNA expression between the groups due to our experimental design (one independent library per age class). Hierarchical clustering, presented as heat maps, were generated by Heatmapper75 using average linkage and Pearson’s distant methods.
Data availability
The data sets supporting the results of this article are included within the article and its supplementary information files. Fastaq files of clean raw reads are uploaded to ENA under our Bioproject accession number PRJEB29358.
References
Daugaard, I. & Hansen, T. B. Biogenesis and Function of Ago-Associated RNAs. Trends Genet. 33, 208–219 (2017).
Ghildiyal, M. & Zamore, P. D. Small silencing RNAs: an expanding universe. Nat. Rev. Genet. 10, 94–108 (2009).
Morris, K. V. & Mattick, J. S. The rise of regulatory RNA. Nat. Rev. Genet. 15, 423–437 (2014).
Wagner, E. G. H. & Romby, P. Small RNAs in Bacteria and Archaea. Who They Are, What They Do, and How They Do It. Advances in Genetics 90, 133–208 (2015).
Ha, M. & Kim, V. N. Regulation of microRNA biogenesis. Nat. Rev. Mol. Cell Biol. 15, 509–524 (2014).
Bartel, D. P. Metazoan MicroRNAs. Cell 173, 20–51 (2018).
Griffiths-Jones, S., Hui, J. H. L., Marco, A. & Ronshaugen, M. MicroRNA evolution by arm switching. EMBO Rep. 12, 172–177 (2011).
Marco, A., Ninova, M. & Griffiths-Jones, S. Multiple products from microRNA transcripts. Biochem. Soc. Trans. 41, 850–854 (2013).
Guo, L. & Chen, F. A challenge for miRNA: multiple isomiRs in miRNAomics. Gene 544, 1–7 (2014).
Neilsen, C. T., Goodall, G. J. & Bracken, C. P. IsomiRs – the overlooked repertoire in the dynamic microRNAome. Trends Genet. 28, 544–549 (2012).
Klein, M., Chandradoss, S. D., Depken, M. & Joo, C. Why Argonaute is needed to make microRNA target search fast and reliable. Sem. Cell Dev. Biol. 65, 20–28 (2017).
Xia, J. & Zhang, W. A meta-analysis revealed insights into the sources, conservation and impact of microRNA 5′-isoforms in four model species. Nucleic Acids Res. 42, 1427–1441 (2014).
Bortoluzzi, S., Biasiolo, M. & Bisognin, A. MicroRNA–offset RNAs (moRNAs): by-product spectators or functional players? Trends Mol. Med. 17, 473–474 (2011).
Shi, W., Hendrix, D., Levine, M. & Haley, B. A distinct class of small RNAs arises from pre-miRNA–proximal regions in a simple chordate. Nat. Struct. Mol. Biol. 16, 183–189 (2009).
Okamura, K., Ladewig, E., Zhou, L. & Lai, E. C. Functional small RNAs are generated from select miRNA hairpin loops in flies and mammals. Genes Dev. 27, 778–792 (2013).
Winter, J. et al. Loop-miRs: active microRNAs generated from single-stranded loop regions. Nucleic Acids Res. 41, 5503–5512 (2013).
Desvignes, T. et al. miRNA Nomenclature: A View Incorporating Genetic Origins, Biosynthetic Pathways, and Sequence Variants. Trends Genet. 31, 613–626 (2015).
Lucas, K. & Raikhel, A. S. Insect MicroRNAs: Biogenesis, expression profiling and biological functions. Insect Biochem. Mol. Biol. 43, 24–38 (2013).
Asgari, S. MicroRNA functions in insects. Insect Biochem. Mol. Biol. 43, 388–397 (2013).
Lucas, K. J., Zhao, B., Liu, S. & Raikhel, A. S. Regulation of physiological processes by microRNAs in insects. Curr. Opin. Insect Sci. 11, 1–7 (2015).
Belles, X. MicroRNAs and the Evolution of Insect Metamorphosis. Annu. Rev. Entomol. 62, 111–125 (2017).
Chen, H. et al. An invertebrate-specific and immune-responsive microRNA augments oyster haemocyte phagocytosis by targeting CgIκB2. Sci. Rep. 6, 29591 (2016).
Pedersen, I. M. et al. Interferon modulation of cellular microRNAs as an antiviral mechanism. Nature 449, 919–922 (2007).
Lampert, W. Daphnia: Development of a Model Organism in Ecology and Evolution (International Ecology Institute) (2011).
Bøhn, T., Primicerio, R., Hessen, D. O. & Traavik, T. Reduced fitness of Daphnia magna fed a Bt-transgenic maize variety. Arch. Environ. Toxicol. 55, 584–592, https://doi.org/10.1007/s00244-008-9150-5 (2008).
Bøhn, T., Traavik, T. & Primicerio, R. Demographic responses of Daphnia magna fed transgenic Bt-maize. Ecotoxicology 19, 419-430 https://doi.org/10.1007/s10646-009-0427-x. Open Access. (2010).
Bøhn, T., Rover, C. M. & Semenchuk, P. Daphnia magna negatively affected by chronic exposure to purified Cry-toxins. Food Chem. Toxicol. 91, 130–140, https://doi.org/10.1016/j.fct.2016.03.009 (2016).
Kusari, F., O’Doherty, A. M., Hodges, N. J. & Wojewodzic, M. W. Bi-directional effects of vitamin B12 and methotrexate on Daphnia magna fitness and genomic methylation. Sci. Rep. 7, 11872, https://doi.org/10.1038/s41598-017-12148-2 (2017).
Nys, C. et al. Mixture toxicity of nickel and zinc to Daphnia magna is noninteractive at low effect sizes but becomes synergistic at high effect sizes: Interactive effects in a binary Ni-Zn mixture. Environ. Toxicol. Chem. 34, 1091–1102 (2015).
Chen, S., McKinney, G. J., Nichols, K. M. & Sepúlveda, M. S. In Silico Prediction and In Vivo Validation of Daphnia pulex Micrornas. PLoS One 9, e83708 (2014).
Chen, S., McKinney, G. J., Nichols, K. M., Colbourne, J. K. & Sepúlveda, M. S. Novel Cadmium Responsive MicroRNAs in Daphnia pulex. Environ. Sci. Technol. 49, 14605–14613 (2015).
Ünlü, E. S., Gordon, D. M. & Telli, M. Small RNA Sequencing Based Identification of MiRNAs in Daphnia magna. PLoS One 10, e0137617 (2015).
Hearn, J. et al. Daphnia magna microRNAs respond to nutritional stress and ageing but are not transgenerational. Mol. Ecol. 27, 1402–1412 (2018).
Wojewodzic, M. W. & Beaton, M. J. The Future of Environmental Epigenetics: Insights Using the Clonal Water Flea Model in Advances in Insect Physiology (ed. Verlinden, H.) 287–312 (Elsevier 2017).
He, Y., Ju, C. & Zhang, X. Roles of small RNAs in the immune defence mechanisms of crustaceans. Mol. Immunol. 68, 399–403 (2015).
Ylla, G., Fromm, B., Piulachs, M.-D. & Belles, X. The microRNA toolkit of insects. Sci. Rep. 6, 37736 (2016).
Kozomara, A. & Griffiths-Jones, S. miRBase: annotating high confidence microRNAs using deep sequencing data. Nucleic Acids Res. 42, D68–D73 (2014).
Pundhir, S. & Gorodkin, J. Differential and coherent processing patterns from small RNAs. Sci. Rep. 5, 12062 (2015).
Czech, B. & Hannon, G. J. Small RNA sorting: matchmaking for Argonautes. Nat. Rev Genet. 12, 19–31 (2011).
Xi, Q.-Y. et al. Genome-wide discovery of novel and conserved microRNAs in white shrimp (Litopenaeus vannamei). Mol. Biol. Rep. 42, 61–69 (2015).
Mohammed, J., Siepel, A. & Lai, E. C. Diverse modes of evolutionary emergence and flux of conserved microRNA clusters. RNA 20, 1850–1863 (2014).
Wang, Y., Luo, J., Zhang, H. & Lu, J. microRNAs in the Same Clusters Evolve to Coordinately Regulate Functionally Related Genes. Mol. Biol. Evol. 33, 2232–2247 (2016).
Chiang, H. R. et al. Mammalian microRNAs: experimental evaluation of novel and previously annotated genes. Genes Dev. 24, 992–1009 (2010).
Ikeda, K. et al. Identification, expression, and molecular evolution of microRNAs in the “living fossil” Triops cancriformis (tadpole shrimp). RNA 21, 230–242 (2015).
Baskerville, S. & Bartel, D. P. Microarray profiling of microRNAs reveals frequent coexpression with neighboring miRNAs and host genes. RNA 11, 241–247 (2005).
Chaulk, S. G., Ebhardt, H. A. & Fahlman, R. P. Correlations of microRNA:microRNA expression patterns reveal insights into microRNA clusters and global microRNA expression patterns. Mol BioSyst 12, 110–119 (2016).
Lee, Y., Jeon, K., Lee, J.-T., Kim, S. & Kim, V. N. MicroRNA maturation: stepwise processing and subcellular localization. EMBO J. 21, 4663–4670 (2002).
Ventura, A. et al. Targeted Deletion Reveals Essential and Overlapping Functions of the miR-17∼92 Family of miRNA Clusters. Cell 132, 875–886 (2008).
Ryazansky, S. S., Gvozdev, V. A. & Berezikov, E. Evidence for post-transcriptional regulation of clustered microRNAs in Drosophila. BMC Genomics 12, 371 (2011).
Bryant, W. B., Mills, M. K., Olson, B. J. & Michel, K. Small RNA-Seq analysis reveals miRNA expression dynamics across tissues in the malaria vector. Anopheles gambiae. G3: Genes | Genomes | Genetics 9, 1507–1517 (2019).
Kabekkodu, S. P. et al. Clustered miRNAs and their role in biological functions and diseases. Biol. Rev. 93, 1955–1986, https://doi.org/10.1111/brv.12428 (2018).
Ninova, M., Ronshaugen, M. & Griffiths-Jones, S. MicroRNA evolution, expression, and function during short germband development in Tribolium castaneum. Genome Res. 26, 85–96 (2016).
Marco, A., Ninova, M., Ronshaugen, M. & Griffiths-Jones, S. Clusters of microRNAs emerge by new hairpins in existing transcripts. Nucleic Acids Res. 41, 7745–7752 (2013).
Sokol, N. S., Xu, P., Jan, Y.-N. & Ambros, V. Drosophila let-7 microRNA is required for remodeling of the neuromusculature during metamorphosis. Genes Dev. 22, 1591–1596 (2008).
Marco, A., Hooks, K. & Griffiths-Jones, S. Evolution and function of the extended miR-2 microRNA family. RNA Biol. 9, 242–248 (2012).
Guo, Q. et al. The role of miR-2∼13∼71 cluster in resistance to deltamethrin in Culex pipiens pallens. Insect Biochem. Mol. Biol. 84, 15–22 (2017).
Cammaerts, S., Strazisar, M., De Rijk, P. & Del Favero, J. Genetic variants in microRNA genes: impact on microRNA expression, function, and disease. Front. Genet. 6, 186 (2015).
Laganà, A. et al. Discovery and characterization of the feline miRNAome. Sci. Rep. 7, 9263 (2017).
Bodfill-De Ros, X., Yang, A. & Gu, S. IsomiRs: expanding the miRNA repression tools beyond the seed, BBA – Gene regulatory mechanisms, 10.1016/j.bbagram.2019.03.005
Yu, F. et al. Naturally existing isoforms of miR-222 have distinct functions. Nucleic Acids Res. 45, 11371–11385 (2017).
Burroughs, A. M. et al. A comprehensive survey of 3′ animal miRNA modification events and a possible role for 3′ adenylation in modulating miRNA targeting effectiveness. Genome Res. 20, 1398–1410 (2010).
Heo, I. et al. Mono-Uridylation of Pre-MicroRNA as a Key Step in the Biogenesis of Group II let-7 MicroRNAs. Cell 151, 521–532 (2012).
Azuma-Mukai, A. et al. Characterization of endogenous human Argonautes and their miRNA partners in RNA silencing. Proc. Natl. Acad. Sci. USA 105, 7964–7969 (2008).
Mercey, O. et al. Characterizing isomiR variants within the microRNA-34/449 family. FEBS Lett. 591, 693–705 (2017).
Nishikura, K. A-to-I editing of coding and non-coding RNAs by ADARs. Nat. Rev. Mol. Cell Biol. 17, 83–96 (2016).
Tomaselli, S. et al. ADAR Enzyme and miRNA Story: A Nucleotide that Can Make the Difference. Int. J. Mol. Sci. 14, 22796–22816 (2013).
Telonis, A. G. et al. Knowledge about the presence or absence of miRNA isoforms (isomiRs) can successfully discriminate amongst 32 TCGA cancer types. Nucleic Acids Res. 45, 2975–2985 (2017).
Wu, C. W. et al. A Comprehensive Approach to Sequence-oriented IsomiR annotation (CASMIR): demonstration with IsomiR profiling in colorectal neoplasia. BMC Genomics 19, 401 (2018).
Zhao, J. et al. MicroRNA-Offset RNA Alters Gene Expression and Cell Proliferation. PLoS One 11, e0156772 (2016).
Samel, A., Ziegenfuss, M., Goulden, C. E., Banks, S. & Baer, K. N. Culturing and bioassay testing of Daphnia magna using Elendt M4, Elendt M7, and COMBO media. Ecotoxicol. Environ. Saf. 43, 103–110 (1999).
Colbourne, J. K., Singan, V. R. & Gilbert, D. G. wFleaBase: the Daphnia genome database. BMC Bioinformatics 6, 45 (2005).
Colbourne, J. K. et al. The Ecoresponsive Genome of Daphnia pulex. Science 331, 555–561 (2011).
Fromm, B. et al. A Uniform System for the Annotation of Vertebrate microRNA Genes and the Evolution of the Human microRNAome. Annu. Rev. Genet. 49, 213–242 (2015).
Love, M. I., Huber, W. & Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 15, 550, https://doi.org/10.1186/s13059-014-0550-8 (2014).
Babicki, S. et al. Heatmapper: web-enabled heat mapping for all. Nucleic Acids Res. 44, W147–W153 (2016).
Acknowledgements
Thanks to Anita Ursvik and Tareq Hassan Khan for technical assistance during the isolation of total RNA. We thank Dag Hessen, University of Oslo for used Daphnia magna clone, originating from Germany. MW received funding from the European Union’s Intra- European Fellowships for Career Development (FP7-MC-IEF), Marie Skłodowska-Curie grant agreement No. 629892 and from the Norwegian Research Council, IS-TOPP No. 238820.
Author information
Authors and Affiliations
Contributions
D.H.C. and T.B. conceived and designed the study. D.H.C. was involved in the laboratory work, performed most of the data analyses and interpretation of the results, and drafted the manuscript. M.W.W. contributed to data analyse and comparison the results to previously published miRNA sets. D.H.C., T.B. & M.W.W. revised the manuscript and read and approved the final manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Coucheron, D.H., Wojewodzic, M.W. & Bøhn, T. MicroRNAs in Daphnia magna identified and characterized by deep sequencing, genome mapping and manual curation. Sci Rep 9, 15945 (2019). https://doi.org/10.1038/s41598-019-52387-z
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-019-52387-z
This article is cited by
-
Daphnia magna egg piRNA cluster expression profiles change as mothers age
BMC Genomics (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.