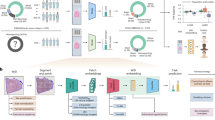
Abstract
Chromosomes are a principal target of clinical cytogenetic studies. While chromosomal analysis is an integral part of prenatal care, the conventional manual identification of chromosomes in images is time-consuming and costly. This study developed a chromosome detector that uses deep learning and that achieved an accuracy of 98.88% in chromosomal identification. Specifically, we compiled and made available a large and publicly accessible database containing chromosome images and annotations for training chromosome detectors. The database contains five thousand 24 chromosome class annotations and 2,000 single chromosome annotations. This database also contains examples of chromosome variations. Our database provides a reference for researchers in this field and may help expedite the development of clinical applications.
Similar content being viewed by others
Background & Summary
The human cell has one pair of sex chromosomes and 22 pairs of other chromosomes. Abnormalities in the total number or structure of chromosomes are referred to as chromosomal aberrations and are the leading cause of genetic disorders1. The conventional sampling method is amniocentesis, during which amniotic fluid from the uterus is aspirated under sonographic guidance. Approximately one in 150 babies have chromosomal abberations2. Common chromosomal aberrations occur on chromosomal pair 13 (trisomy 13), which is associated with Patau syndrome, pair 18 (trisomy 18), which is associated with Edwards syndrome, and pair 21 (trisomy 21), which is associated with Down syndrome. According to the National Center for Biotechnology Information, these chromosomal aberrations cause 50%–60% of early miscarriages. Karyotyping is clinically important in prenatal genetic diagnosis3.
Karyotyping is a diagnostic method in which characteristic dark and light bandings of chromosomes are visualised on images for examination by physicians or senior technicians. Abnormality is determined according to the number and structure of abnormal chromosomes and sex-related chromosomes. The procedure typically takes approximately 20 min for an experienced examiner. The examiner needs to sort, cut, orient, and rearrange the mapping of a raw chromosomal cell, and at least four chromosomal images need to be processed for an individual subject to ensure a correct diagnosis. Chromosomal analysis is labour intensive and is an urgent issue because of increasing shortages of medical manpower. Automated chromosome classification systems are scarce. Most current systems are based on artificial intelligence (AI) approaches involving machine learning and deep learning4,5,6,7. Earlier studies on chromosome classification were based on segmenting overlaps and adherent chromosomes and employed conventional methods like border detection8,9, the watershed method10, and straightening of bent chromosomes11,12. These methods depended heavily on image preprocessing, resulting in distorted chromosome features that could result in misdiagnoses. Recent research in this field has a growing preference for chromosome prototypes over preprocessing.
Chromosomes are classified by basically one of two approaches. The first approach involves the analysis of single chromosomes. This requires a human examiner, takes substantial time and effort, and is often complemented by background image segmentation and noise suppression13. Convolutional neural networks (CNNs)14,15 may be used for classifying images; however, the accuracy is unsatisfactory due to low data volume. This approach, due to its repetitiveness and the variability of chromosome features, has limited clinical application. The second approach involves the analysis of original images by using deep learning–based object detection models16,17,18,19,20 to identify and classify chromosomes. For example, DeepACEv221 requires no manual preprocessing and uses object detection as the backbone to frame and classify individual chromosomes, and this is followed by final confirmation and manual editing by a human examiner. This approach is clinically more applicable. In the literature, chromosome images are relatively easy to identify and classify from chromosome images. Despite the simplicity of these images, an examiner must spend substantial time and effort to identify the chromosomes. An automatic chromosome recognition system is essential for handling more difficult images for better clinical application.
The application of AI models for medical imaging is constrained by the complexity of medical images. In a clinical setting, an incomplete AI model would not be practically useful and may even decrease staff productivity. In the event of an incomplete database, the trust of experts and patients cannot be gained22. Many examples of AI in medical research require a large database to improve the credibility and stability of the AI model23,24,25,26,27.
We have developed here a detector called the ‘Automated Chromosome Detector Based on Metaphase Cell Images Using Deep Learning’ that is capable of locating and classifying chromosomes in images. The images used in this study have more chromosome overlaps and adherences than those used in other studies. Chromosome overlaps and adherences can be confusing for specialists. A probabilistic two-stage algorithm was adopted to improve chromosome detection accuracy. The method was trained and validated using data from 5,000 chromosomal images of fetuses. High accuracy (98.88%) was achieved—higher than that achieved by experienced specialists.
The chromosomal images and annotations used to train the detector have been provided in this study. This is the first publicly available large database of chromosome annotations. The database contains 2,000 annotations for single chromosomes and 5,000 annotations for 24 chromosomes [Fig. 1b,c, respectively]. We also provide criteria for defining difficult images and notes from our experts on classifying chromosomes as a series of common points in the clinical recognition of difficult images. What we provide is a good benchmark dataset for researchers in this field that can expedite technical development in this application area. For example, using 5,000 annotations for 24 chromosomes, better accuracy can be achieved. These images can also help develop algorithms and expert recommendations for those images that are difficult to examine. Finally, single chromosome segmentation data can help segment chromosome overlap and adherence or to standardise the orientation of the short arms of chromosomes for examination by clinicians.
Methods
Data overview
Our dataset contains three collections, 5,000 annotations of 24 chromosome categories, 2,000 annotations of single chromosome categories, totalling 229,852 chromosomes. The data set was compiled from the data of 1,598 fetuses of pregnant women undergoing prenatal chromosomal studies between 2014 and 2021 at the Cytogenetic Laboratory, Department of Women’s Medicine, Taichung Veterans General Hospital. These data collections were approved by the Internal Review Board of Taichung Veterans General Hospital (IRB no. CE20369B). We informed all subjects and obtained their consent to use their data in relevant research.
Each collection contains a file of images and a file of annotations. Content includes the file name, image size, file path, category, and object box coordinates or segmentation coordinates for each chromosomal image. All chromosomal annotations represent markers made by an assistant trained for 3 months by specialist technicians within the department, and the results were acceptable. When the chromosomal images were collected, they were visually inspected and contained no personal information that could be linked to the subjects.
Annotation methods
We used the Image Labeler apps in Matlab software (version: 2022a) to annotate the chromosomal images. Once finished, all annotations, image addresses, categories, and annotations were stored separately in the Matlab default storage format, gTruth. When another computer language is needed for annotation, the gTruth format is converted to the xml file format. For easy reference, an image is saved as a separate xml file with the same header name (e.g. 104011.jpg is saved as 104011.xml).
In addition to the basic chromosome information, each original chromosome image is accompanied by a karyotype (Fig. 2). First, each chromosome is identified by its characteristics and checked against the karyotype for accuracy. At the end of each chromosome image, the number of chromosomes is checked for accuracy. Finally, each chromosome annotation is checked by another marker to ensure that the correct total number of chromosomes is recorded.
Definition of difficult image for recognition
The 5,000 chromosomal notes contain simple and difficult images for identification. A difficult image combines the senior technicians’ perception and identification results of our model. The classification in this study is intended to help other users to develop models that are more compatible with clinical applications. Three kinds of difficult image features are illustrated in Fig. 3a–c. The three definitions of difficult features adopted by the cytogenetic technician are described as follows:
-
a.
Multiple chromosomal overlaps: overlaps of two chromosomes can be easily handled by the examiner and by the model, whereas an overlap of more than two chromosomes makes identification difficult.
-
b.
Suboptimal banding: in the event of dull colour and unclear features, the examiner needs to adjust the microscope preference to turn images darker; also with poor staining, the dark and light bands appear dull.
-
c.
The chromosome is too elongated: for example, if point c often occurs at the same place as point b, the elongated chromosome stretches the band feature and is more prone to overlap.
Data Records
Images and their associated annotations are publicly available on CELL IMAGE LIBRARY, which is a well-known website with a diverse library of cellular images (data set link: https://doi.org/10.7295/W9CIL54816)28. We stored images and annotations in folders with the structure shown in BOX 1. The xml format is shown in BOX 2. All 24 chromosome annotations are stored in the file ‘24_chromosomes_object’. Table 1 shows the number of each chromosome type in the data set. Annotations are stored in the annotation files, and images are stored as JPEG files. We categorised the data set into simple and difficult images according to our assessment criteria. The number and proportion of simple and difficult images in the training and test sets are shown in Table 2. The annotated files (xml) of the data sets are stored under the simple file and difficult file directories accordingly.
Single chromosomal annotations are stored in the ‘single_chromosomes_object’ files. Annotations and images are stored in the folders for annotation files and JPEG files, respectively. This data set only provides users with a distinction between chromosomes and backgrounds. Chromosomes are not classified; therefore, having the same number of annotations for the 24 chromosomes is not required.
Three additional csv files are provided. One records the image file names and related descriptions corresponding to the normal cases of chromosomes (file name: normal.csv), another records the cases of abnormal numbers (file name: number_abormalities.csv), and the other records the cases of structural abnormality (file name: structural_abormalities.csv). These relevant descriptions contain information on which pairs of chromosomes are abnormal in number or structure. This information will allow researchers in the field of chromosomal abnormalities to make better use of our database. Table 3 shows the total cases of normal, abnormal number, and structural abnormality in our data set.
Comparison with other data sets
Table 4 compares our data set with other chromosome data sets7,10,21,29,30. The total number of chromosomes and images in our data set is much higher than in the other data sets. Except for our data set and the SRAS-net data set29, the data sets used in the other studies are not publicly available. Although chromosome painting (e.g. spectral karyotyping) is a different method that allows the identification of both numerical and structural chromosomal aberrations and the chromosomes from which the fragments originate, the most common method used at present for chromosome analysis is the G-banding technique. Moreover, only our data set and DeepACEv221 contain difficult images. Difficult images can facilitate the development of a more clinically applicable system.
Technical Validation
The objective of chromosomal identification and classification is to address the drawbacks of existing automated chromosomal identification software, to reduce manual involvement, and to improve expert efficiency. To verify the validity of the chromosomal database, two classification models were trained using the database to test its validity and clinical applicability. The first model was used to identify chromosomes and backgrounds. The results of the model were truncated and exported for image classification and semantic segmentation. In this test, an accuracy of 98.91% was achieved. The output of the model can be applied to semantic segmentation to further obtain single chromosomes without the background. In this test, the results, classified as general, simple, and difficult, reached accuracies of 98.88%, 99.15%, and 98.78%, respectively. Results were validated by experts. Images of detected chromosomes are shown in Fig. 4.
Two images containing detected chromosomes. (a) Simple and (b) difficult images. Detection accuracy was 100% with a simple image. Multiple overlapping and adherent chromosomes make detection more difficult. Chromosomes not captured correctly were those that fell between three overlapping chromosomes.
Table 5 shows the detection accuracy for each of the 24 chromosome categories. Detection accuracy was higher than 98.5% for all categories except for pair G22, chromosome X, and chromosome Y, for which the detection accuracy was still higher than 97.4%. The reason for the lower accuracy is that their body size is relatively short compared to the other chromosomes and they are easily covered or overlapped by other chromosomes and not easily detected.
Figure 5 shows the curve of number of images and model accuracy (%). We started recording with 81.79% accuracy by using 800 images. After increasing the number of images, the accuracy increased dramatically and reached 98.87% by using 2,000 images and 98.91% by using 5,000 images. This result shows that if the number of images in the data set is not large enough, the model cannot achieve good accuracy. Moreover, the images in our proposed data set were all obtained by using the G-banding method. This method uses an AI model that can determine cases with abnormal chromosome numbers or structural variations. In other words, our data set allows researchers to develop highly accurate and clinically practical assisted chromosome detection systems without using chromosome painting images that require expensive and special image analysis systems. This also explains the necessity and value of our data set.
Highly accurate AI models are useful in clinical settings; however, they must be trained with large data sets containing images of sufficient complexity. No public chromosome image database with a large number of complex images is available. Furthermore, validating a model is difficult; even if a model can achieve 100% accuracy in a small, private data set, its clinical practicability may not be high. However, our data set exceeds that of existing public data sets in terms of quantity and the complexity of images it contains. Such a data set can be used to verify whether a developed model is actually clinically usable. In addition, considering current clinical regulations, AI cannot completely replace doctors or medical examiners. Most AI is still employed in decision-making assistance roles. Therefore, a model that can feasibly be applied in a clinical setting may be able to increase confidence to a level at which experts finally decide to trust it.
In a previous study, among fetuses with chromosome aberrations, 144 (69.56%) had trisomy 13, trisomy 18, trisomy 21, or sex chromosome disorder, and 63 (30.44%) had balanced translocation, unbalanced abnormality, inversion, or marker chromosome31. Because cases of chromosomal structural abnormalities are relatively rare in the general population, few images with chromosomal structural abnormalities are present in our data set. To allow researchers to develop AI models capable of detecting abnormalities in chromosome structure in the future, we will continue to increase the data on abnormalities in chromosome structure in our data set. To expand our data set, we have organised another four high-quality cytogenetic laboratories in Taiwan and will work together to build a better data set that can be used in clinical applications.
Usage Notes
The 24 chromosome class annotations and single chromosome annotations data sets are provided in the file ‘xml2coco_ob.py’. The code can convert the box coordinates to diagonal coordinates. Results are saved in the coco format. Similarly, for the single chromosome segments data set, xml2coco_seg.py is provided in the code to convert the mask to a polygon, and the results are saved in the coco format.
Code availability
We provide two model weights using Pytorch as a deep learning framework to detect the 24 chromosome categories for both object detection and single chromosome object detection. Our neural network model is based on YOLOv4. We recommend using argusswift’s code (https://github.com/argusswift/YOLOv4-pytorch) and provide a py file that converts xml files to coco format (xml2coco.py).
References
Wapner, R. J. et al. Chromosomal microarray versus karyotyping for prenatal diagnosis. New England Journal of Medicine 367, 2175–2184 (2012).
Carlson, L. M. & Vora, N. L. Prenatal diagnosis: screening and diagnostic tools. Obstetrics and Gynecology Clinics 44, 245–256 (2017).
Theisen, A. & Shaffer, L. G. Disorders caused by chromosome abnormalities. The application of clinical genetics 3, 159 (2010).
Jindal, S., Gupta, G., Yadav, M., Sharma, M. & Vig, L. in Proceedings of the IEEE international conference on computer vision workshops. 72–81.
Karvelis, P. S., Fotiadis, D. I., Georgiou, I. & Syrrou, M. in 2006 International Conference of the IEEE Engineering in Medicine and Biology Society. 3009–3012 (IEEE).
Litjens, G. et al. A survey on deep learning in medical image analysis. Medical image analysis 42, 60–88 (2017).
Qin, Y. et al. Varifocal-net: A chromosome classification approach using deep convolutional networks. IEEE transactions on medical imaging 38, 2569–2581 (2019).
Nimitha, N. et al. in IOP Conference Series: Materials Science and Engineering. 012194 (IOP Publishing).
Yan, W. & Shen, S. in 2008 2nd International Conference on Bioinformatics and Biomedical Engineering. 2390–2392 (IEEE).
Lin, C. et al. Cir-net: Automatic classification of human chromosome based on inception-resnet architecture. IEEE/ACM Transactions on Computational Biology and Bioinformatics (2020).
Ding, W., Chang, L., Gu, C. & Wu, K. in 2019 12th International Congress on Image and Signal Processing, BioMedical Engineering and Informatics (CISP-BMEI). 1–5 (IEEE).
Gu, J. et al. Recent advances in convolutional neural networks. Pattern recognition 77, 354–377 (2018).
Krizhevsky, A., Sutskever, I. & Hinton, G. E. Imagenet classification with deep convolutional neural networks. Communications of the ACM 60, 84–90 (2017).
Hu, X. et al. Classification of metaphase chromosomes using deep convolutional neural network. Journal of Computational Biology 26, 473–484 (2019).
Bochkovskiy, A., Wang, C.-Y. & Liao, H.-Y. M. Yolov4: Optimal speed and accuracy of object detection. arXiv preprint arXiv:2004.10934 (2020).
Kong, T. et al. Foveabox: Beyound anchor-based object detection. IEEE Transactions on Image Processing 29, 7389–7398 (2020).
Law, H. & Deng, J. in Proceedings of the European conference on computer vision (ECCV). 734–750.
Lin, T.-Y., Goyal, P., Girshick, R., He, K. & Dollár, P. in Proceedings of the IEEE international conference on computer vision. 2980–2988.
Redmon, J., Divvala, S., Girshick, R. & Farhadi, A. in Proceedings of the IEEE conference on computer vision and pattern recognition. 779–788.
Tan, M., Pang, R. & Le, Q. V. in Proceedings of the IEEE/CVF conference on computer vision and pattern recognition. 10781–10790.
Xiao, L. et al. DeepACEv2: Automated chromosome enumeration in metaphase cell images using deep convolutional neural networks. IEEE Transactions on Medical Imaging 39, 3920–3932 (2020).
Pesapane, F., Codari, M. & Sardanelli, F. Artificial intelligence in medical imaging: threat or opportunity? Radiologists again at the forefront of innovation in medicine. European radiology experimental 2, 1–10 (2018).
Hou, L. et al. Dataset of segmented nuclei in hematoxylin and eosin stained histopathology images of ten cancer types. Scientific data 7, 1–12 (2020).
Khalil, Y. A. et al. Multi-scanner and multi-modal lumbar vertebral body and intervertebral disc segmentation database. Scientific Data 9, 1–11 (2022).
Kromp, F. et al. An annotated fluorescence image dataset for training nuclear segmentation methods. Scientific Data 7, 1–8 (2020).
Liebl, H. et al. A computed tomography vertebral segmentation dataset with anatomical variations and multi-vendor scanner data. Scientific data 8, 1–7 (2021).
Shapey, J. et al. Segmentation of vestibular schwannoma from MRI, an open annotated dataset and baseline algorithm. Scientific Data 8, 1–6 (2021).
Lu, C.-H., Kuo, C.-E. & Tseng, J.-J. CIL:54816. Cell Image Library https://doi.org/10.7295/W9CIL54816 (2022).
Liu, X., Fu, L., Chun‐Wei Lin, J. & Liu, S. SRAS‐net: Low‐resolution chromosome image classification based on deep learning. IET Systems Biology (2022).
Xie, N., Li, X., Li, K., Yang, Y. & Shen, H. T. Statistical karyotype analysis using CNN and geometric optimization. IEEE Access 7, 179445–179453 (2019).
Tseng, J.-J. et al. Detection of chromosome aberrations in the second trimester using genetic amniocentesis: experience during 1995-2004. Taiwanese Journal of Obstetrics and Gynecology 45, 39–41 (2006).
Acknowledgements
This work was supported (in part) by the Taichung Veterans General Hospital (TCVGH-AI-10904 and TCVGH-AI-11003) and the National Science and Technology Council, Taiwan. (109-2221-E-005 -080 -MY3, 111-2221-E-075A-004 -MY2, and 111-2634-F-005-001 - project Smart Sustainable New Agriculture Research Center (SMARTer)).
Author information
Authors and Affiliations
Contributions
Jenn-Jhy Tseng: Resources, Data Curation, Writing - Original Draft. Chien-Hsing Lu: Resources, Project administration, Funding acquisition. Jun-Zhou Li: Methodology, Software, Data Curation, Writing - Original Draft. Hui-Yu Lai: Resources, Data Curation. Min-Hu Chen: Resources, Data Curation. Fu-Yuan Cheng: Resources, Data Curation. Chih-En Kuo: Conceptualization, Methodology, Investigation, Supervision, Project administration, Writing - original draft & editing.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Tseng, JJ., Lu, CH., Li, JZ. et al. An Open Dataset of Annotated Metaphase Cell Images for Chromosome Identification. Sci Data 10, 104 (2023). https://doi.org/10.1038/s41597-023-02003-7
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41597-023-02003-7