Abstract
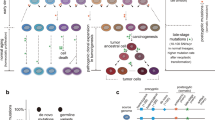
Somatic mutations are the cause of cancer and have been implicated in other, noncancerous diseases and aging. While clonally expanded mutations can be studied by deep sequencing of bulk DNA, very few somatic mutations expand clonally, and most are unique to each cell. We describe a detailed protocol for single-cell whole-genome sequencing to discover and analyze somatic mutations in tissues and organs. The protocol comprises single-cell multiple displacement amplification (SCMDA), which ensures efficiency and high fidelity in amplification, and the SCcaller software tool to call single-nucleotide variations and small insertions and deletions from the sequencing data by filtering out amplification artifacts. With SCMDA and SCcaller at its core, this protocol describes a complete procedure for the comprehensive analysis of somatic mutations in a single cell, covering (1) single-cell or nucleus isolation, (2) single-cell or nucleus whole-genome amplification, (3) library preparation and sequencing, and (4) computational analyses, including alignment, variant calling, and mutation burden estimation. Methods are also provided for mutation annotation, hotspot discovery and signature analysis. The protocol takes 12–15 h from single-cell isolation to library preparation and 3–7 d of data processing. Compared with other single-cell amplification methods or single-molecular sequencing, it provides high genomic coverage, high accuracy in single-nucleotide variation and small insertions and deletion calling from the same single-cell genome, and fewer processing steps. SCMDA and SCcaller require basic experience in molecular biology and bioinformatics. The protocol can be utilized for studying mutagenesis and genome mosaicism in normal and diseased human and animal tissues under various conditions.
Key points
-
Protocol for single-cell whole-genome sequencing to discover and analyze somatic mutations in tissues and organs, using single-cell multiple displacement amplification alongside the SCcaller software.
-
Compared with bulk sequencing approaches, single-cell whole-genome sequencing allows discovery of most, if not all, mutations in the same single-cell genome. This enables quantification of the mutation burden per cell, discovery of mutational hotspots and establishment of cell lineages.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout








Similar content being viewed by others
Code availability
The source code for the data analysis pipeline and its documentations is freely available on GitHub via ref. 24 and at https://github.com/XiaoDongLab/SCcaller-pipeline.
References
Failla, G. The aging process and cancerogenesis. Ann. N. Y. Acad. Sci. 71, 1124–1140 (1958).
Szilard, L. On the nature of the aging process. Proc. Natl Acad. Sci. USA 45, 30–45 (1959).
Vijg, J. & Dong, X. Pathogenic mechanisms of somatic mutation and genome mosaicism in aging. Cell 182, 12–23 (2020).
Nowell, P. C. The clonal evolution of tumor cell populations. Science 194, 23–28 (1976).
Alexandrov, L. B. et al. The repertoire of mutational signatures in human cancer. Nature 578, 94–101 (2020).
The ICGC/TCGA Pan-Cancer Analysis of Whole Genomes Consortium. Pan-cancer analysis of whole genomes. Nature 578, 82–93 (2020).
Jaiswal, S. et al. Age-related clonal hematopoiesis associated with adverse outcomes. N. Engl. J. Med. 371, 2488–2498 (2014).
Kessler, M. D. et al. Common and rare variant associations with clonal haematopoiesis phenotypes. Nature 612, 301–309 (2022).
Erickson, R. P. Somatic gene mutation and human disease other than cancer. Mutat. Res. 543, 125–136 (2003).
Erickson, R. P. Somatic gene mutation and human disease other than cancer: an update. Mutat. Res. 705, 96–106 (2010).
Bae, T. et al. Analysis of somatic mutations in 131 human brains reveals aging-associated hypermutability. Science 377, 511–517 (2022).
Bianconi, E. et al. An estimation of the number of cells in the human body. Ann. Hum. Biol. 40, 463–471 (2013).
Sender, R., Fuchs, S. & Milo, R. Revised estimates for the number of human and bacteria cells in the body. PLoS Biol. 14, e1002533 (2016).
Brunner, S. F. et al. Somatic mutations and clonal dynamics in healthy and cirrhotic human liver. Nature 574, 538–542 (2019).
Blokzijl, F. et al. Tissue-specific mutation accumulation in human adult stem cells during life. Nature 538, 260–264 (2016).
Martincorena, I. & Campbell, P. J. Somatic mutation in cancer and normal cells. Science 349, 1483–1489 (2015).
Dong, X. et al. Accurate identification of single-nucleotide variants in whole-genome-amplified single cells. Nat. Methods 14, 491–493 (2017).
Zhang, L. et al. Single-cell whole-genome sequencing reveals the functional landscape of somatic mutations in B lymphocytes across the human lifespan. Proc. Natl Acad. Sci. USA 116, 9014–9019 (2019).
Brazhnik, K. et al. Single-cell analysis reveals different age-related somatic mutation profiles between stem and differentiated cells in human liver. Sci. Adv. 6, eaax2659 (2020).
Huang, Z. et al. Single-cell analysis of somatic mutations in human bronchial epithelial cells in relation to aging and smoking. Nat. Genet. 54, 492–498 (2022).
Milholland, B. et al. Differences between germline and somatic mutation rates in humans and mice. Nat. Commun. 8, 15183 (2017).
Zhang, L. et al. Maintenance of genome sequence integrity in long- and short-lived rodent species. Sci. Adv. 7, eabj3284 (2021).
Sun, S. et al. Single-cell analysis of somatic mutation burden in mammary epithelial cells of pathogenic BRCA1/2 mutation carriers. J. Clin. Invest. 132 https://doi.org/10.1172/jci148113 (2022).
Zhang, L. et al. Single-cell whole-genome sequencing for discovering somatic mutations. GitHub https://doi.org/10.5281/zenodo.7826180 (2023).
Luquette, L. J., Bohrson, C. L., Sherman, M. A. & Park, P. J. Identification of somatic mutations in single cell DNA-seq using a spatial model of allelic imbalance. Nat. Commun. 10, 3908 (2019).
Bohrson, C. L. et al. Linked-read analysis identifies mutations in single-cell DNA-sequencing data. Nat. Genet. 51, 749–754 (2019).
Min, S. et al. Absence of coding somatic single nucleotide variants within well-known candidate genes in late-onset sporadic Alzheimer’s disease based on the analysis of multi-omics data. Neurobiol. Aging 108, 207–209 (2021).
Zhang, L. et al. Analysis of somatic mutations in senescent cells using single-cell whole-genome sequencing. AgingBio 1, 1–6, https://doi.org/10.59368/agingbio.20230005 (2023).
Schmitt, M. W. et al. Detection of ultra-rare mutations by next-generation sequencing. Proc. Natl Acad. Sci. USA 109, 14508–14513 (2012).
Abascal, F. et al. Somatic mutation landscapes at single-molecule resolution. Nature 593, 405–410 (2021).
Maslov, A. Y. et al. Single-molecule, quantitative detection of low-abundance somatic mutations by high-throughput sequencing. Sci. Adv. 8, eabm3259 (2022).
Zafar, H., Navin, N., Chen, K. & Nakhleh, L. SiCloneFit: Bayesian inference of population structure, genotype, and phylogeny of tumor clones from single-cell genome sequencing data. Genome Res. 29, 1847–1859 (2019).
Wang, F. et al. MEDALT: single-cell copy number lineage tracing enabling gene discovery. Genome Biol. 22, 70 (2021).
Telenius, H. et al. Degenerate oligonucleotide-primed PCR: general amplification of target DNA by a single degenerate primer. Genomics 13, 718–725 (1992).
Dean, F. B., Nelson, J. R., Giesler, T. L. & Lasken, R. S. Rapid amplification of plasmid and phage DNA using Phi 29 DNA polymerase and multiply-primed rolling circle amplification. Genome Res. 11, 1095–1099 (2001).
Zong, C., Lu, S., Chapman, A. R. & Xie, X. S. Genome-wide detection of single-nucleotide and copy-number variations of a single human cell. Science 338, 1622–1626 (2012).
Chen, C. et al. Single-cell whole-genome analyses by linear amplification via transposon insertion (LIANTI). Science 356, 189–194 (2017).
Huang, L., Ma, F., Chapman, A., Lu, S. & Xie, X. S. Single-cell whole-genome amplification and sequencing: methodology and applications. Annu. Rev. Genomics Hum. Genet. 16, 79–102 (2015).
Spits, C. et al. Whole-genome multiple displacement amplification from single cells. Nat. Protoc. 1, 1965–1970 (2006).
Xing, D., Tan, L., Chang, C. H., Li, H. & Xie, X. S. Accurate SNV detection in single cells by transposon-based whole-genome amplification of complementary strands. Proc. Natl Acad. Sci. USA https://doi.org/10.1073/pnas.2013106118 (2021).
Sarangi, V. et al. SCELLECTOR: ranking amplification bias in single cells using shallow sequencing. BMC Bioinforma. 21, 521 (2020).
Motyer, A. et al. The mutational landscape of single neurons and oligodendrocytes reveals evidence of inflammation-associated DNA damage in multiple sclerosis. Preprint at bioRxiv https://doi.org/10.1101/2022.04.30.490132 (2022).
Bloom, J. C. & Schimenti, J. C. Sexually dimorphic DNA damage responses and mutation avoidance in the mouse germline. Genes Dev. 34, 1637–1649 (2020).
Gundry, M., Li, W., Maqbool, S. B. & Vijg, J. Direct, genome-wide assessment of DNA mutations in single cells. Nucleic Acids Res. 40, 2032–2040 (2012).
Gravina, S., Dong, X., Yu, B. & Vijg, J. Single-cell genome-wide bisulfite sequencing uncovers extensive heterogeneity in the mouse liver methylome. Genome Biol. 17, 150 (2016).
Ailshire, J. A., Beltran-Sanchez, H. & Crimmins, E. M. Becoming centenarians: disease and functioning trajectories of older US Adults as they survive to 100. J. Gerontol. Ser. A, Biol. Sci. Med. Sci. 70, 193–201 (2015).
Van der Auwera, G. A. & O'Connor, B. D. Genomics in the Cloud: Using Docker, GATK, and WDL in Terra 1st edn (O'Reilly Media, 2020).
Lodato, M. A. et al. Aging and neurodegeneration are associated with increased mutations in single human neurons. Science 359, 555–559 (2018).
Sherman, M. A. et al. PaSD-qc: quality control for single cell whole-genome sequencing data using power spectral density estimation. Nucleic Acids Res. 46, e20 (2018).
Wang, K., Li, M. & Hakonarson, H. ANNOVAR: functional annotation of genetic variants from high-throughput sequencing data. Nucleic Acids Res. 38, e164 (2010).
Chang, X. & Wang, K. wANNOVAR: annotating genetic variants for personal genomes via the web. J. Med. Genet. 49, 433–436 (2012).
Yang, H. & Wang, K. Genomic variant annotation and prioritization with ANNOVAR and wANNOVAR. Nat. Protoc. 10, 1556–1566 (2015).
Consortium, E. P. An integrated encyclopedia of DNA elements in the human genome. Nature 489, 57–74 (2012).
Mejia-Ramirez, E. & Florian, M. C. Understanding intrinsic hematopoietic stem cell aging. Haematologica 105, 22–37 (2020).
Wang, J. et al. Evidence for mutation showers. Proc. Natl Acad. Sci. USA 104, 8403–8408 (2007).
Nik-Zainal, S. et al. Mutational processes molding the genomes of 21 breast cancers. Cell 149, 979–993 (2012).
Lora, D. Location and visualization of clustered somatic mutations. v1.0.1. CRAN-R Project https://cran.r-project.org/web/packages/ClusteredMutations/ClusteredMutations.pdf (2016).
Bhagat, T. D. et al. miR-21 mediates hematopoietic suppression in MDS by activating TGF-beta signaling. Blood 121, 2875–2881 (2013).
Kent, W. J. et al. The human genome browser at UCSC. Genome Res. 12, 996–1006 (2002).
Lee, B. T. et al. The UCSC Genome Browser database: 2022 update. Nucleic Acids Res. 50, D1115–d1122 (2022).
Alexandrov, L. B. et al. Mutational signatures associated with tobacco smoking in human cancer. Science 354, 618–622 (2016).
Alexandrov, L. B., Nik-Zainal, S., Wedge, D. C., Campbell, P. J. & Stratton, M. R. Deciphering signatures of mutational processes operative in human cancer. Cell Rep. 3, 246–259 (2013).
Alexandrov, L. B. et al. Signatures of mutational processes in human cancer. Nature 500, 415–421 (2013).
Sun, S., Wang, Y., Maslov, A. Y., Dong, X. & Vijg, J. SomaMutDB: a database of somatic mutations in normal human tissues. Nucleic Acids Res. 50, D1100–D1108 (2022).
Denkinger, M. D., Leins, H., Schirmbeck, R., Florian, M. C. & Geiger, H. HSC aging and senescent immune remodeling. Trends Immunol. 36, 815–824 (2015).
Osorio, F. G. et al. Somatic mutations reveal lineage relationships and age-related mutagenesis in human hematopoiesis. Cell Rep. 25, 2308–2316.e2304 (2018).
Mitchell, E. et al. Clonal dynamics of haematopoiesis across the human lifespan. Nature 606, 343–350 (2021).
Rodrigues-Moreira, S. et al. Low-dose irradiation promotes persistent oxidative stress and decreases self-renewal in hematopoietic stem cells. Cell Rep. 20, 3199–3211 (2017).
Acknowledgements
L.Z. is supported by the American Federation for Aging Research (the Sagol Network GerOmic Award for Junior Faculty). J.V. is supported by the US National Institutes of Health awards (P01 AG017242, P01 AG047200, P30 AG038072, U01 ES029519, U01 HL145560 and U19 AG056278). X.D. is supported by the NIH awards (R00 AG056656, P01 HL160476, U54 AG076041 and U54 AG079754).
Author information
Authors and Affiliations
Contributions
L.Z. and J.V. designed the wet-laboratory protocol. L.Z., M.L. and A.Y.M. validated the wet-laboratory protocol. X.D. designed and validated the dry-laboratory protocol. L.Z., C.M., J.V. and X.D. wrote the manuscripts with input from M.L. and A.Y.M.
Corresponding authors
Ethics declarations
Competing interests
L.Z., M.L., A.Y.M., J.V. and X.D. are co-founders and shareholders of SingulOmics Corp. C.M. declares no conflict of interest.
Peer review
Peer review information
Nature Protocols thanks Isidro Cortes-Ciriano, Thomas Mitchell and Jie Qiao for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Related links
Key references using this protocol
Dong, X. et al. Nat. Methods 14, 491–493 (2017): https://doi.org/10.1038/nmeth.4227
Zhang, L. et al. Proc. Natl Acad. Sci. USA 116, 9014–9019 (2019): https://doi.org/10.1073/pnas.1902510116
Zhang, L. et al. Sci. Adv. 7, eabj3284 (2021): https://doi.org/10.1126/sciadv.abj3284
Supplementary information
Supplementary Tables 1–5
Supplementary Tables 1–5.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Zhang, L., Lee, M., Maslov, A.Y. et al. Analyzing somatic mutations by single-cell whole-genome sequencing. Nat Protoc 19, 487–516 (2024). https://doi.org/10.1038/s41596-023-00914-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41596-023-00914-8
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.